Journal of Software Engineering and Applications
Vol.07 No.05(2014), Article ID:46055,10 pages
10.4236/jsea.2014.75039
Bleeding and Ulcer Detection Using Wireless Capsule Endoscopy Images
Jinn-Yi Yeh1, Tai-Hsi Wu2, Wei-Jun Tsai1
1Department of Management Information Systems, National Chiayi University, Taiwan
2Department of Business Administration, National Taipei University, Taiwan
Email: jyeh@mail.ncyu.edu.tw, taiwu@mail.ntpu.edu.tw
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 16 April 2014; revised 10 May 2014; accepted 16 May 2014
ABSTRACT
Wireless capsule endoscopes (WCEs) have been used widely to detect abnormalities inside regions of the small intestine that are not accessible when using traditional endoscopy techniques. However, an experienced clinician must spend an average of 2 hours to view and analyze the approximately 60,000 images produced during one examination. Therefore, developing a computer- aided system for processing WCE images is crucial. This paper proposes a novel method used for detecting bleeding and ulcers in WCE images. This approach involves using color features to determine the status of the small intestine. The experimental results revealed that the proposed scheme is promising in detecting bleeding and ulcer regions.
Keywords:
Wireless capsule endoscope (WCE), Computer-aided diagnosis, Bleeding, Ulcer

1. Introduction
Traditional endoscopy enables a physician to view both ends of a patient’s digestive tract, including the eso- phagus, stomach, duodenum, colon, and terminal ileum. However, examining tissues in the small intestine with- out performing a surgical operation was a difficult task until recently. The solution to this problem was the wireless capsule endoscope (WCE), which was first invented by Given Imaging in 2000 and involves using wireless transmission to send images from inside the intestine to the outside environment [1] . The WCE primarily consists of a complementary metal oxide semiconductor image sensor, a lens, white light-emitting diodes, and an application-specific integrated circuit transmitter, which transmits images captured by the image sensor wirelessly. Figure 1(a) illustrates the appearance of the WCE. The WCE is 11 mm in diameter and 26 mm in length. Figure 1(b) shows the components of the WCE, which is a pill-shaped device that contains miniaturized

Figure 1. Given imaging capsule [4] . (a) Capsule endoscope, (b) Components diagram of WCE.
elements [2] . The WCE was approved by the US Food and Drug Administration in 2001, and this new technology has been reported to be valuable in evaluating gastrointestinal bleeding, Crohn’s disease, ulcers, and other diseases that develop in the digestive tract [3] .
After a WCE is swallowed by a patient, who must not eat any food but can drink water for approximately 12 hours before swallowing the device, the small device is propelled through peristalsis and begins to capture images while moving along the digestive tract. Simultaneously, the images are sent wirelessly to a recorder attached to the waist. The process continues until the battery is depleted (approximately 8 hours). Finally, all of the images are downloaded to a computer, and physicians can review the images and analyze the potential sources of various diseases in the gastrointestinal tract [2] . Approximately 60,000 images are captured per examination of one patient, and an experienced clinician spends an average of 2 hours viewing and analyzing all of the frames. Therefore, a computer-aided diagnosis system for processing WCE images must be developed. In this paper, a new method used for detecting bleeding and ulcers in WCE images is proposed. The new approach involves using color features to determine the status of the small intestine. Gastrointestinal bleeding refers to mucosa damage in the esophagus, stomach, small intestine, or large intestine. When patients repeatedly experience gastrointestinal bleeding, physicians usually use upper gastrointestinal endoscopy (gastroscopy) or lower gastrointestinal endoscopy (colonoscopy) to detect the bleeding region. If physicians cannot locate the bleeding point, then it is highly likely that the small intestine lesions are caused by unknown gastrointestinal bleeding. A WCE can be used to view the entire small intestine directly, without pain, sedation, or air insufflation; therefore, it can be used to detect gastrointestinal bleeding.
As illustrated in Figure 2(a), bleeding regions in WCE images exhibit different extents of variation in color texture compared with the surrounding regions. This specific property motivated us to investigate the color textural features of these images. Clinicians use the color texture information in WCE images as a primary indicator to diagnose various diseases [5] . Karargyris and Bourbakis [6] proposed using the expectation maximization clustering algorithm and Bayesian information criterion to detect bleeding regions automatically. Miaou et al. [7] applied fuzzy c-means cluster analysis in recognizing images with large abnormal areas to identify the suspected blood indicator. An ulcer is a discontinuity or break in a bodily membrane that impedes the organ of which that membrane is a part from performing its normal functions. Peptic ulcers commonly occur in the stomach (gastric ulcers) and the duodenum (duodenal ulcer), but can also occur in the small intestine or gastrointestinal tract. Peptic ulcers can cause severe gastrointestinal bleeding or gastrointestinal perforation. Ulcers appear as white spots in WCE images. The normal and ulcer regions in WCE images are differentiated using textural features, because texture information is a primary indicator analyzed by clinicians. As illustrated in Figure 2(b), abnormal regions in the WCE images of ulcers exhibit variations in texture that can be compared with the surrounding regions.
This characteristic and the aforementioned review encouraged us to investigate the textural features of WCE images. Li and Meng [5] proposed a new textural feature analysis algorithm that uses the curvelet transform and local binary patterns to detect ulcers. Using a multilayer perceptron, a neural network, and support vector ma- chines as the classifiers, these authors verified that the new method exhibited a more satisfactory performance in ulcer detection than several traditional algorithms did. Miaou et al. [7] used a back-propagation neural network to detect ulcers. Each bleeding and ulcer image exhibits 360 and 366 features, respectively. The presence of too many features degrades the performance of the classifier, because these features may be highly related or simi-

Figure 2. Some WCE images with bleeding and ulcer. (a) Bleeding image; (b) Ulcer images.
lar. Therefore, we conducted feature selection to reduce the number of features. In this study, we applied ReliefF, support vector machine recursive feature elimination (SVM-RFE), and One Rule (OneR) to implement feature selection based on a ranking method. The numbers of features used for the comparison were 20, 40, and 60. The classification methods used were the support vector machine (SVM), neural networks, and the decision tree. The performance of the proposed method was evaluated according to accuracy, sensitivity, specificity, and kappa.
2. Materials and Methods
2.1. Data Sets
The WCE images used in this study were obtained from Capsuleendoscopy.org and RAPID Atlas. CapsuleEndoscopy.org is an international gastroenterology professional care education website that is committed to the teaching, learning, and sharing of PillCam capsule endoscopy information. RAPID Atlas contains image data sets of Given Imaging capsule endoscopy. We obtained 220, 159, and 228 images of bleeding, ulcers, and nonbleeding/ulcers, respectively. All of the 607 images were captured in the small intestine by using the PillCam SB WCE. Ulcer samples were extracted from the candidate ulcer area. An image could contain multiple ulcers. We obtained a total of 190 ulcer samples and 258 nonulcer samples. The numbers of bleeding and non-bleeding samples were 220 and 228, respectively. The data distribution is shown in Table 1.
2.2. Color Features
WCE image analysis can be considered a color-related problem because many color spaces, such as red/ green/blue (RGB) and hue/saturation/value (HSV) spaces, are available. RGB representation is often employed because of its compatibility with additive color reproduction systems. The RGB model is represented as a cube using nonnegative values within a 0 - 1 range; black is assigned to the origin at the vertex (0, 0, 0), and the intensity of values along the 3 axes increases up to white at the vertex (1, 1, 1), diagonally opposite to black. An RGB triplet (r, g, b) represents the 3-dimensional coordinate of the location of the given color within the cube, on its faces, or along its edges. However, applying the grayscale algorithms directly to the RGB components of an image may result in color shifts caused by high correlations among RGB channels in natural images. Color shifts are especially undesired in medical images because color plays a crucial role in determining the statuses of tissues and organs.
The HSV color space is designed to be used intuitively in manipulating color and approximating the ways in which humans perceive and interpret color. Three properties of color, namely, hue, saturation, and value, are defined to differentiate the color components. Hue indicates the wavelength of a color by representing the color's name, such as green, red, or blue. Saturation denotes a measure of the purity of a color. Value measures the departure of a hue from black, the color of zero energy [8] . Luminance and color are separated using the HSV color space. Brightness is easily controlled because color is less susceptible to the influence of external light. Therefore, computing speed and color recognition rate can be improved. The saturation threshold is defined as the cutting point for distinguishing between bleeding and non-bleeding regions.
2.3. Color Coherence Vector
Color coherence is defined as the degree to which pixels of a color are members of large similarly colored re-
Table 1. Data distribution of WCE images.
gions, implying that coherent regions are critical for characterizing images. The classifier determines whether pixels are coherent or incoherent. Coherent pixels are a part of sizable contiguous regions, whereas incoherent pixels are not. A color coherence vector (CCV) is obtained by matching coherent pixels in one image to incoherent pixels in another image. This enables viewers to make fine distinctions that cannot be made using color histograms [9] .
The initial stage in computing a CCV is to blur the image slightly by replacing pixel values with the average value in a small local neighborhood. This eliminates small variations between neighboring pixels. The color space is then discretized such that there are only n distinct colors in the image. The next step is to classify the pixels within a given color bucket as either coherent or incoherent. A coherent pixel is part of a large group of pixels of the same color, whereas an incoherent pixel is not. Pixel groups are determined by computing the connected components. A connected component C is a maximal set of pixels grouped such that, for any 2 pixels p,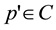 , a path in C travels between p and p'. Connected components are computed only within a given discretized color bucket. This process electively segments the image based on the discretized color space. When this process is complete, each pixel is allocated to exactly one connected component. Pixels are classified as either coherent or incoherent depending on the sizes of pixels of their connected component. A pixel is coherent if the size of its connected component exceeds a fixed value τ (coherent threshold); otherwise, the pixel is incoherent.
, a path in C travels between p and p'. Connected components are computed only within a given discretized color bucket. This process electively segments the image based on the discretized color space. When this process is complete, each pixel is allocated to exactly one connected component. Pixels are classified as either coherent or incoherent depending on the sizes of pixels of their connected component. A pixel is coherent if the size of its connected component exceeds a fixed value τ (coherent threshold); otherwise, the pixel is incoherent.
We next demonstrate the computation of a CCV. Suppose that after we slightly blur the input image, the resulting intensities are as Figure 3(a). Let us discretize the colorspace so that bucket 2 contains intensities 60 through 179, bucket 3 contains 180 through 299, and bucket 1 contains the others. Then after discretization the matrix is shown in Figure 3(b). The next step is to compute the connected components. Individual components will be labeled with letters (A, B,…) and we will need to keep a table which maintains the discretized color associated with each label, along with the number of pixels with that label. Of course, the same discretized color can be associated with different labels if multiple contiguous regions of the same color exist. The image may then be shown in Figure 3(c) and the connected components table will be listed in Table 2. The CCV for this image will be listed in Table 3 when the τ is defined as 4.
2.4. Textural Features
Texture describes the changes and distributions of the content in a grayscale image, such as directional lines or patterns. Image texture analysis can be divided into statistical and image conversion methods. A gray-level cooccurrence matrix is used for feature extraction and represents the spatial distribution and dependence of the gray levels within a local area. Each p(i, j) entry in the matrices represents the probability of going from one pixel with a gray level (i) to another pixel with a gray level (j) according to a predefined distance and angle. According to these matrices, several sets of statistical measures, or feature vectors, are computed to build various texture models. In this study, the WCE image was separated into windows of size L × L pixels with L/2 pixels overlapping. The cooccurrence matrix algorithm was then used to obtain information from the pixels in an image window. Four angles, namely 0˚, 45˚ 90˚, and 135˚, as well as a predefined distance of one pixel, were considered in the formation of the cooccurrence matrices. The L × L cooccurrence matrix and spatial features are defined as follows:
 (1)
(1)
 (2)
(2)

Figure 3. Computation of a CCV. (a) CCV example, (b) After discretized, (c) Compute connected components.
Table 2. Connected component table.
Table 3. CCV table.
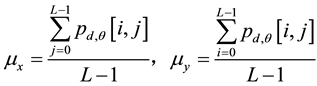 (3)
(3)







2.5. OneR
OneR generates a one-level decision tree expressed in the form of a set of rules that all test one particular feature. The rules are based on testing a single feature and branch accordingly. Each branch corresponds to a different value of the feature. The optimal classification to give each branch is the class that occurs most often in the training data. The error rate of the rules is then easily determined by counting the number of instances that do not contain the majority class. Each feature generates a different set of rules, and one rule is generated for each value of the feature. The error rate for each feature’s rule set is evaluated and the most favorable set is selected [10] .
2.6. ReliefF
The original Relief algorithm is employed to estimate the quality of attributes according to how well their values can be used to distinguish between instances that are near each other. Given a randomly selected instance R, the Relief algorithm searches for its 2 nearest neighbors: one from the same class, called nearest hit H, and the other from a different class, called nearest miss M. The algorithm then updates the quality estimation weights for all attributes A depending on their R, M, and H values. The process is repeated m times, where m is a user-defined parameter [11] .
2.7. Support Vector Machine and Support Vector Machine Recursive Feature Elimination
The SVM approach has been applied extensively in classification, regression, and density estimation. The SVM uses nonlinear mapping to transform the original training data into a higher dimension. In the new dimension, it searches for a linear optimal separating hyperplane. The SVM locates this hyperplane by using support vectors and margins. A hyperplane (w is the vector of hyperplane coefficients, b is a bias term) can be constructed as a quadratic optimization problem so that the margin between the hyperplane and the nearest point is maximized [12] .
The SVM-RFE method is used as a learning algorithm in recursive procedures to select a subset of features for classification. Nested subsets of features are selected in a sequential backward elimination process, which begins with all of the features and removes one feature each time. Thus, in the end, all of the feature variables are ranked. At each step, the coefficients of the weight vector w of a linear SVM are used as the feature ranking criteria [13] .
2.8. Decision Tree
A decision tree is a flow-chart-like tree structure, in which each internal node denotes a test on an attribute, each branch represents an outcome of the test, and each leaf node represents classes or class distributions. The topmost node in a tree is the root node. A test on an attribute is associated with a splitting criterion that is chosen to split the data sets into subsets that exhibit favorable class separability, thus minimizing occurrences of misclassification error. Once the tree is built from the training data, it is then heuristically pruned to avoid data overfitting, which tends to introduce classification errors in the test data [14] .
2.9. Neural Networks
Mimicking the processes found in biological neurons, artificial neural networks are used to learn and predict based on a given data set. Neural networks have been widely used in medical decision applications because they exhibit substantial predictive ability. In this study, we employed the traditional multilayer perception (MLP) neural network as the classifier used to conduct disease detection based on WCE images, because the MLP exhibits many advantages compared with other classifiers, such as greater generalization ability and more robust perfor- mance, and requires fewer training data. MLP is the most frequently used neural network architecture in medical image decision applications [14] .
2.10. Performance Evaluation
Ten fold cross-validation was used to split the samples into training and testing sets. The performance of the proposed method was determined using 4 well-known criteria, namely, accuracy, sensitivity, specificity, and kappa, which are defined as follows:




where TP represents the number of images that both the system and the expert considered to be abnormal; TN denotes the number of images that both the system and the expert considered to be normal; FP is the number of images identified as normal by the expert but classified as abnormal by the system; FN is the number of images identified as abnormal by the expert but classified as normal by the system; A is TP + FN; B is FP + TN; E is the number of total images; P0 is the relative observed agreement among raters; and Pc is the hypothetical probability that chance agreement occurs when using the observed data to calculate the probabilities of each observer randomly saying each category. The kappa value indicates the reliability of the system when it makes a binary decision as an expert does. A kappa value between 0.4 and 0.75 indicates that the system is reliable, and a kappa value between 0.75 and 1 implies that the system is highly reliable.
3. Experimental Design and Analysis
3.1. Ulcer Candidate Extraction
Regions of ulcers in WCE images are usually small and not suitable for analyzing the entire image for classification. Therefore, using a small area of the subimage feature is more accurate than using the entire image. Li and Meng [2] divided an entire image into numerous small subimages that were then used to extract color features. The ulcer area and location are not fixed, causing subimage size setting to be a difficult task, because dividing images into subimages of a fixed size may not enable the ulcer area to be completely covered. Therefore, we first selected the candidate ulcer area, from which the texture and color features to be classified were selected.
The first step in ulcer detection is transforming the original image (Figure 4(a)) into a grayscale image (Figure 4(b)) that is binarized based on a predefined threshold. The ulcer area is usually white or pale yellow. In this study, the binarized ulcer area was rendered white, as shown in Figure 4(c). We applied a region growing method to segment the candidate regions from the binarized image. The regions were segmented from top to bottom and from left to right to obtain a value of 1 pixel in the image as a reference point. Based on the reference point, we continued outward expansion. If a pixel was adjacent to the reference point with a value of 1, then it was labeled as being in the same area. The procedure was repeated until the pixels in the surrounding regions achieved values of 0 and the area stopped growing. These steps were repeated until all of the pixels with a value of 1 were evaluated. Finally, all regions become ulcer candidate regions, as shown in Figure 4(d).
3.2. Parameter Setting
The experimental factors for parameter setting were the saturation threshold, coherent threshold, number of features, feature selection methods, and classifiers. The saturation threshold setting was based on the average saturation of the bleeding area. The coherent threshold settings were 5, 10, and 15. The numbers of features were 60, 40, and 20 for purposes of comparison. The feature selection methods used were the ReliefF, SVM-RFE, and OneR algorithms. The classification methods used were the SVM, neural network, and decision tree approaches. The factors and levels of the bleeding experiment are listed in Table 4, according to which 243 combinations of experimental parameters were tested. The parameters for the ulcer experiment were set in an identical manner.
3.3. Experimental Results and Analysis
3.3.1. Accuracy of bleeding detection
Table 5 lists the 5 highest accuracies observed in the bleeding classification experiments. Experiment 145 yielded the highest accuracy and kappa, and its sensitivity and specificity were 91.82% and 94.74%, respectively. In addition, the C4.5 decision tree was used as the classifier in all 5 experiments. Figure 5 shows the factor reaction figure of the bleeding classification accuracy according to 243 experimental results. Combining A1, B1, C3, D1, and E1 yielded the highest accuracy. The factors of classification accuracy were ranked from high to low according to reaction degree as follows E (classifier), A (saturation), B (coherent), D (feature selection), and C (number of features). The difference between A1 and the other 2 levels was large, possibly because the color saturation of the bleeding area was less than 70 and was thus excluded, resulting in a poor performance in classification. Figure 5 shows that C4.5 outperformed the other methods in accuracy.

Figure 4. Ulcer images. (a) Ulcer region, (b) Greyscale image, (c) Binary image, (d) Ulcer candidates.
Figure 5. Factor reaction figure for accuracy of bleeding classification..
Table 4. Parameter setting for bleeding experiment.
Table 5. The first 5 highest accuracies for experiments of bleeding classification.
3.3.2. Sensitivity of bleeding detection
A false negative criterion indicates when an abnormal image is misclassified as a normal image. Sensitivity is an indicator used to evaluate the performance of an abnormal image classification. Table 6 lists the 5 highest sensitivities observed in the experiments testing bleeding classification. Experiment 25 yielded the highest sen- sitivity, and its sensitivity and specificity were 92.86% and 93.64%, respectively. The feature selection method used was the ReliefF algorithm, and the classification method was C4.5. Figure 6 shows the factor reaction figure of the sensitivity of bleeding classification according to 243 experimental results. Similar to the accuracy, combining A1, B1, C3, D1, and E1 yielded the highest sensitivity. However, the factors of the sensitivity of bleeding classification were from high to low according to reaction degree as follows: E (classifier), A (saturation), C (number of features), D (feature selection), and B (coherent). The sensitivity of C3 was higher than the sensitivity of C1 and C2, indicating that feature selection is an effective method for reducing the feature dimension, thus enhancing the efficiency of classifier learning as well as the classification sensitivity. Figure 6 shows that C4.5 outperformed the other methods in sensitivity.
Table 6. The first 5 highest accuracies for experiments of bleeding classification.
Figure 6. Factor reaction figure for sensitivity of bleeding classification.
3.3.3. Accuracy of ulcer detection
Table 7 lists the 5 highest accuracies observed in the experiments testing ulcer classification. Experiment 88 yielded the highest accuracy and kappa, and its sensitivity and specificity were 87.37% and 89.15%, respectively. Figure 7 shows the factor reaction figure of the accuracy of ulcer classification. Combining A3, B2, C2, D2, and E1 yielded the highest accuracy. The factors of classification accuracy were ranked from high to low according to reaction degree as follows: E (classifier), A (saturation), C (number of features), D (feature selection), and B (coherent). The coherent threshold in ulcer classification exhibited a lower influence than the coherent threshold in bleeding classification did. Furthermore, the higher the saturation threshold was, the higher the accuracy, possibly because the average saturation of the ulcer regions was greater than 60. As observed in the bleeding experiment, C4.5 outperformed the other methods in accuracy according to Figure 7.
3.3.4. Sensitivity of ulcer detection
Table 8 lists the 5 highest sensitivities observed in the experiments testing ulcer classification. Experiment 124 yielded the highest sensitivity, and its accuracy, sensitivity, and specificity were 85.71 %, 87.90%, and 84.11%, respectively. The feature selection method was the ReliefF algorithm and the classification method was C4.5. Figure 8 shows the factor reaction figure of the sensitivity of ulcer classification according to 243 experimental results. Combining A2, B2, C2, D3, and E1 yielded the highest sensitivity. The factors of the sensitivity of ulcer classification were ranked from high to low according to the reaction degree as follows: E (classifier), A (saturation), C (number of features), D (feature selection), and B (coherent). In contrast to bleeding experiment, the degree of reaction increased when Factor A (saturation threshold) increased in the ulcer experiment, because the average saturation of the bleeding region was greater than the average saturation of the normal region. However, the average saturation of the ulcer was less than the average saturation of the normal region.
According to the analysis, C4.5 was the optimal classifier in both the bleeding and ulcer experiments. The classification performance was considerably improved after applying the color saturation threshold. The most favorable combination of parameters for automatically detecting bleeding and ulcers is listed in Table 9.
3.3.5. Performance of the color coherence vector
This study focused on the color features used for conducting WCE image analysis and, particularly, on using the CCV to improve the accuracy of classification. Table 10 lists the classification performance according to whe- ther CCV features were used. The experimental results revealed that the performance was substantially more favorable when using CCV than when CCV was not used for classification in both the bleeding and ulcer experiments. Thus, CCV features effectively improved the WCE image analysis.
Figure 7. Factor reaction figure for accuracy of ulcer classification.
Figure 8. Factor reaction figure for sensitivity of ulcer classification.
Table 7. The first 5 highest accuracies for experiments of ulcer classification.
Table 8. The first 5 highest sensitivity for experiments of ulcer classification.
Table 9. The best combination of parameters.
Table 10. The performance of classification depends on using and not using CCV features.
4. Conclusion
This paper proposes a novel method used for bleeding and ulcer detection in WCE images. The new approach involves using color features, including RGB, HSV, and CCV features, to determine the status of the small intestine. To identify an optimal classification solution, we implemented a series of experiments to test various parameter settings. The experimental results revealed that the saturation threshold was 70 when using the ReliefF method, the number of color features was reduced to 20. Using C4.5 effectively enhanced the accuracy and sensitivity of bleeding classification. Similar settings were applied to ulcer classification, except that the number of color features was reduced to 40. Furthermore, the accuracy and sensitivity of bleeding classification when using the CCV features were 92.86% and 93.64%, respectively, which were substantially higher than the accuracy and sensitivity of the bleeding classification observed when CCV features were not used (75.22% and 71.36%, respectively). The ulcer experiment revealed the same results. Therefore, the CCV features effectively improved the efficiency of WCE image analysis. Using the computer-aided system for WCE images, abnormal images were effectively and automatically detected, reducing the time required to analyze from 50,000 to 60,000 images.
Acknowledgements
We are grateful for the grant provided by the National Science Council (NSC 101-2221-E-415-002).
References
- Iddan, G., Meron, G., Glukhovsky, A. and Swain, P. (2000) Wireless Capsule Endoscopy. Nature, 405, 725-729. http://dx.doi.org/10.1038/35013140
- Li, B.P. and Meng, M.Q.-H. (2009) Computer-Based Detection of Bleeding and Ulcer in Wireless Capsule Endoscopy Images by Chromaticity Moments. Computers in Biology and Medicine, 39, 141-147. http://dx.doi.org/10.1016/j.compbiomed.2008.11.007
- Adeler, D.G. and Gostout, C.J. (2003) Wireless Capsule Endoscopy. Hospital Physician, 39, 14-22.
- Kodogiannis, V.S., Boulougoura, M., Wadge, J.N. and Lygouras, J.N. (2007) The Usage of Soft-Computing Method- ologies in Interpreting Capsule Endoscopy. Engineering Applications of Artificial Intelligence, 20, 539-553. http://dx.doi.org/10.1016/j.engappai.2006.09.006
- Li, B.P. and Meng, M.Q.-H. (2009) Computer-Aided Detection of Bleeding Regions for Capsule Endoscopy Images. IEEE Transactions on Biomedical Engineering, 56, 1032-1039. http://dx.doi.org/10.1109/TBME.2008.2010526
- Karargyris, A. and Bourbakis, N. (2010) Wireless Capsule Endoscopy and Endoscopic Imaging: A Survey on Various Methodologies Presented. IEEE Engineering in Medicine and Biology Magazine, 29, 72-83. http://dx.doi.org/10.1109/MEMB.2009.935466
- Miaou, S.-G., Chang, F.-L., Timotius, I.K., Huang, H.-C., Su J.-L., Liao, R.-S. and Lin, T.-Y. (2009) A Multi-Stage Recognition System to Detect Different Types of Abnormality in Capsule Endoscope Images. Journal of Medical and Biological Engineering, 29, 114-121.
- Smith, A.R. (1978) Color Gamut Transform Pairs. Proceedings of the 5th Annual Conference on Computer Graphics and Interactive Techniques (SIGGRAPH '78), New York, 23-25 August 1978, 12-19.
- Pass, G., Zabih, R. and Miller, J. (1996) Comparing Images Using Color Coherence Vectors. Proceedings of the 4th ACM International Conference on Multimedia, Boston, 18-22 November 1996, 65-73. http://dx.doi.org/10.1145/244130.244148
- Witten, I.H. and Frank, E. (2011) Data Mining: Practical Machine Learning Tools and Techniques. 3rd Edition, Morgan Kaufmann, Burlington.
- Kononenko, I. (1994) Estimation Atributes: Analysis and Extensions of RELIEF. Proceedings of the 1994 European Conference on Machine Learning, Catania, 6-8 April 1994, 171-182.
- Ubeyli, E.D. (2007) Comparison of Different Classification Algorithms in Clinical Decision-Making. Expert Systems, 24, 17-31. http://dx.doi.org/10.1111/j.1468-0394.2007.00418.x
- Guyon, I., Weston, J., Barnhill S. and Vapnik, V. (2002) Gene Selection for Cancer Classification Using Support Vector Machines. Machine Learning, 46, 389-422. http://dx.doi.org/10.1023/A:1012487302797
- Han, J. and Kamber, M. (2011) Data Mining: Concepts and Techniques. 3rd Edition, Morgan Kaufmann, Burlington.















