Journal of Cancer Therapy
Vol.3 No.2(2012), Article ID:18678,4 pages DOI:10.4236/jct.2012.32024
Thymidylate Synthase 5’ UTR Polymorphic Allele Distribution in South Florida Population
![]()
1Department of Biology, Miller School of Medicine, University of Miami, Miami, USA; 2Department of Medicine, Division of Hematology and Oncology, Miller School of Medicine, University of Miami, Miami, USA.
Email: *spochi@med.miami.edu
Received February 16th, 2012; revised March 22nd, 2012; accepted April 12th, 2012
Keywords: TYMS enhancer region; polymorphic repeats; translation regulation; 5-FU resistance; colorectal cancer
ABSTRACT
Thymidylate synthase (TS; TYMS) is a pivotal enzyme in the DNA synthesis pathway. The 5’UTR of TYMS gene has a polymorphic 28 bp segment. Presence of two or three repeats of this unique 28 bp sequence is common. A distinct population distribution pattern for this polymorphic trait among different racial groups had been reported. We analyzed TYMS genotype in the peripheral blood mononucleocytes (PBMC) of 74 individuals in the South Florida region of the United States of America. The number of 28 bp repeats in the 5’ UTR was determined by PCR followed by agarose gel electrophoresis. The distribution of the three different genotypes was found to be 35.1% for 2R/2R, 39.2% for 2R/3R and 24.3% for 3R/3R. One individual was detected with 3R/4R genotype. Functional analyses associated homozygous for three repeats (3R/3R) to higher TYMS expression and therefore poor prognosis to chemotherapy. The other possible genotypes viz 2R/2R or 2R/3R is proposed to have better prognosis. However, there are reports that challenge this observation.
1. Introduction
Colorectal cancer (CRC) is the third most common cause of cancer-related death in the US (American Cancer Society, Cancer Facts & Figures 2012). Predicting tumor response and toxicity may help tailor individual therapy. It is known that individual genotype influence the response to chemotherapeutic drugs. Therefore, understanding the relationship between genotype and treatment outcome may help in this field of personal medicine [1,2].
Thymidylate synthase (TS; TYMS) is the only enzyme that methylates deoxyuridylate (dUMP) to deoxythymidylate (dTMP). In its absence no dTMP is formed. Therefore, TYMS is a pivotal enzyme for DNA synthesis. The uracil analog 5-fluorouracil (5-FU) serves as the building block of 5-fluorodeoxyuridylate (5-FdUMP). 5-FdUMP forms an inactive ternary complex composed with TYMS, and tetrahydrofolate. Thus 5-FdUMP serves as an irreversible inhibitor of thymidylate synthase [3].
The 5’ untranslated region (UTR) of TYMS gene has a 28 bp tandemly repeated sequence. The number of repeats normally varies between two and three [4]. More than three repeats had been observed in a few cases [5]. This 28 bp repeats is proposed to regulate TYMS translation. Triple homozygotes (3R/3R) are associated with a 2 - 4 fold higher TYMS activity compared to double repeat homozygotes (2R/2R) or heterozygotes (2R/3R) [6]. Various reports link high levels of TYMS with poor response to 5-FU treatment [7]. As a consequence of its effect on TYMS level, the 28 bp repeat can influence the treatment outcome with 5-FU [8]. Consequently, there is widespread interest in studying population distribution, influence of treatment outcome and the mechanism of regulation by this polymorphism. We investigated the distribution of TYMS 5’ UTR polymorphism in the patient population that receives treatment at Sylvester Comprehensive Cancer Center, University of Miami Miller School of Medicine, Miami, FL. The results of this study are reported here.
2. Materials and Methods
2.1. Patients and Methods
Individuals over 18 years of age, who are willing to give informed consent, were eligible to participate in this study. This basic science research protocol (SCCC 2003073) was approved by Institutional Review Board. Between January 2004 and December 2005, 74 volunteers consented to participate in this genotyping survey. Participants’ demographic details are listed in Table 1.
Peripheral blood was collected in heparinized tubes. Genomic DNA was isolated as per the method of Subbarayan et al. [9], which is briefly summarized below: 300 μl of whole blood was mixed with 900 μl wash buffer (20 mM Tris-Cl; pH 7.6); incubated at room temperature for 10 minutes, then centrifuged for 30 sec at 10,000 X g at room temperature. The supernatant was discarded. Leucocyte enriched pellet was washed four times with wash buffer, and mechanically dispersed in 600 μl of ice cold rapid mammalian cell lysis buffer (10mM Tris-Cl pH 8.0, 1mM EDTA, and 0.1% SDS). To this suspension we added 3 μl of 4mg/ml DNase-free RNase A and incubated at 37˚C for 30 min, followed by the addition of 200 μl of 3 M sodium acetate, pH 5.2. This mixture was centrifuged at 10,000 g for 3 min at 4˚C. We added 600 μl of isopropanol to the supernatant and the precipitated genomic DNA was recovered by centrifuging at 10,000 g for 1 min at room temperature. The DNA pellet was washed once with 70% ethanol, vacuum dried then suspended in 100 μl of nuclease free water. DNA was quantitated by measuring absorbance at 260 and 280 nm.
The position of 28 bp repeat with reference to the ATG start codon, sequence variations etc are as described by Kaneda et al. [10,11]. The TYMS 5’ UTR region encoding the 28-bp repeat was synthesized using the forward primer (TS5’ sense) 5’-GTGGCTCCTGCGTTTCCCCC-3’ and reverse primer (TS5’ anti-sense) 5’- GCTCCGAGCCGGCCACAGGCATGGCGCGG-3’. The composition of the PCR reaction mixture, thermal cycler settings, expected product length and agarose gel analysis were as described in Villafranca et al. 2001 [11,12].
2.2. Statistical Analysis
We analyzed whether the sampled population reached Hardy-Weinberg equilibrium using the online statistical
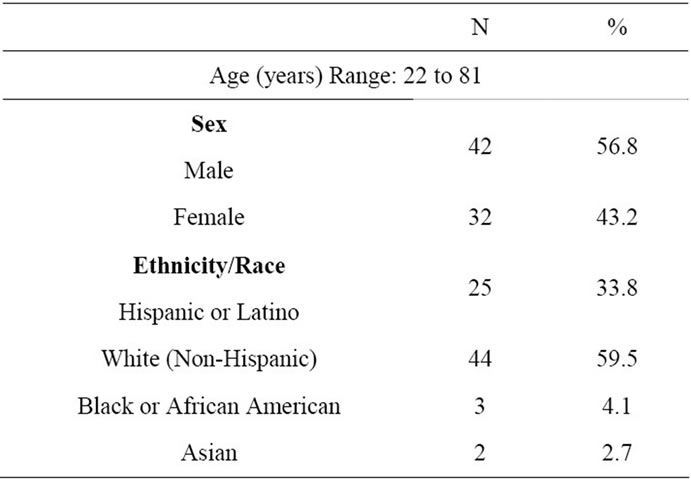
Table 1. Participants’ characteristics.
tool provided at http://krunch.med.yale.edu/hwsim. The data was subjected to 100,000 iterations. Deviations in the expected and observed values in the polymorphic distribution pattern above 0.05 (p < 0.05) will be considered significant.
3. Results
The results of the polymorphic analysis carried out in 74 individuals are summarized in Table 2. Near equal number of heterozygous (2R/3R) and homozygous for two repeats (2R/2R) was observed in this study population. These accounted for 39.2% & 35.1% respectively of the total. Eighteen individuals were homozygous for three repeats (3R/3R) constituting 24.3% of the sample population. Only one individual was detected to have four repeats (3R/4R; 1.4%).
The data was subjected to Hardy-Weinberg equilibrium analysis. The occurrence of four or more repeats is rare. In the sampled population, we encountered one 4R repeat (3R/4R). Therefore, we analyzed the data under two different scenarios. In first set, we combined 3R/4R with 3R/3R genotype. In the second we excluded 3R/4R genotype from the analysis. 100,000 iterations under simulated conditions, yielded c2 values of 3.2367 and 2.7993 respectively with 1 degree of freedom. The p values were not significant. The results reflect the changing population dynamics of South Florida, that is home to a large number of immigrants from all over the world.
4. Discussion
This study utilized the blood samples of individuals residing in South Florida that comprises Miami-Dade, Broward and Palm Beach counties. The year 2000 census data reveals this region to be home to a heterogeneous population consisting of Whites (72.3% - 80.4%), African American (14.9% - 22.2%), Asian and other ethnic groups (7.2% - 10.0%) (Profiles of General Demographic Characteristics: Florida. 2000 US Census Bureau). The percentages may add up to more than 100 because, individuals may report more than one race. As indicated in the Table 1, most of the enrolled volunteers were caucasians (Non-Hispanics and Hispanics; 69%). This number is similar to the census data.
Earlier, Marsh et al. [5,13] reported Chinese population,
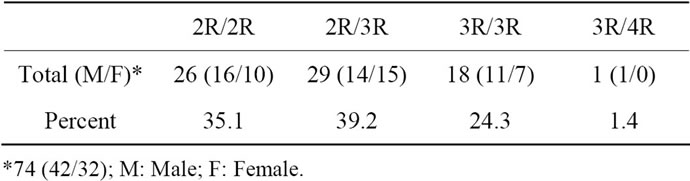
Table 2. Distribution of TYMS polymorphism in South Florida population.
to be predominantly homozygous for three repeats (3R/3R; 67%). On the other hand 38% of Caucasians, 40% of south west Asians and 24% of African Americans were reported to be homozygous for three repeats (3R/3R). The heterozygous genotype 2R/3R was found in 43%, 44% and 53% of the Caucasian, south west Asian and African American population respectively. In the above three groups, homozygous for two repeats (2R/2R) were 19%, 16% and 20% respectively. In contrast, hardly 2% of the sampled Chinese populations were homozygous for two repeats (2R/2R).
Besides the 5’ polymorphism, presence of a SNP in the 5’UTR and a six bp insertion at the 3’ UTR of TYMS had been reported. Studies have shown 5’ UTR to influence TYMS transcription [14] and the six bp insertion to increase the stability of TYMS mRNA [15]. Chu et al. [16] demonstrated that TYMS protein auto regulates its translation. Substrates like dUMP or FdUMP redirect the TYMS from its autoregulatory role towards the formation of thymidylate [16].
TYMS regulation can be classified as innate or induced. Polymorphism is innate whereas auto regulation is induced. As discussed earlier, the rate of TYMS transcription, translation and the transcript stability is regulated at many stages. Innately, if more TYMS is present as in the case of 3R/3R genotype, 5-FU based treatment may not be effective or may result in higher toxicity. For this particular reason, it would be preferable to screen patients for TYMS polymorphic status to formulate an effective treatment cocktail. In the present scenario, combination therapy with 5-FU and other agents is well established. The relationship among tumor tissue TYMS polymorphic status, expression and treatment outcome is debatable because only a subset of all cancers are TYMS dependent [14,17]. However, there still exists an opportunity to analyze the relationship among different polymorphic combinations to TYMS expression in general and patient populations. A conclusive difference in TYMS level in this populace may have cancer diagnostic value. Therefore, further well designed clinical studies are needed to understand the role of 5’ and 3’ polymorphism, SNP in the 5’UTR, TYMS expression in general and patient population.
5. Acknowledgements
We thank the participants who donated blood for the study. Technical help of undergraduate students Philip Lee and Alia Abdulla is appreciated. Research nurse Myra Lima coordinated blood collection. We also thank Dr. Isildinha Reis for critical reading of the manuscript.
REFERENCES
- F. Innocenti and M. J. Ratain, “Update on Pharmacogenetics in Cancer Chemotherapy,” European Journal of Cancer, Vol. 38, No. 5, 2002, pp. 639-644. doi:10.1016/S0959-8049(01)00434-8
- H. L. McLeod and W. E. Evans, “Pharmacogenomics: Unlocking the Human Genome for Better Drug Therapy,” Annual Review of Pharmacology and Toxicology, Vol. 41, No. 1, 2001, pp. 101-121. doi:10.1146/annurev.pharmtox.41.1.101
- Y. M. Rustum, “Thymidylate Synthase: A Critical Target in Cancer Therapy?” Frontiers in Bioscience, Vol. 9, 2004, pp. 2467-2473. doi:10.2741/1408
- S. Kaneda, K. Takeishi, D. Ayusawa, K. Shimizu, T. Seno and S. Altman, “Role in Translation of a Triple Tandemly Repeated Sequence in the 5’-Untranslated Region of Human Thymidylate Synthase mRNA,” Nucleic Acids Research, Vol. 15, No. 3, 1987, pp. 1259-1270. doi:10.1093/nar/15.3.1259.
- S. Marsh, M. M. Ameyaw, J. Githanga, A. Indalo, D. Ofori-Adjei and H. L. McLeod, “Novel Thymidylate Synthase Enhancer Region Alleles in African Populations,” Human Mutation, Vol. 16, No. 6, 2000, p. 528. doi:10.1002/1098-1004(200012)16:6<528::AID-HUMU11>3.0.CO;2-W
- N. Horie, H. Aiba, K. Oguro, H. Hojo and K. Takeishi, “Functional Analysis and DNA Polymorphism of the Tandemly Repeated Sequences in the 5’-Terminal Regulatory Region of the Human Gene for Thymidylate Synthase,” Cell Structure and Function, Vol. 20, No. 3, 1995, pp. 191-197. doi:10.1247/csf.20.191
- S. Popat, A. Matakidou and R. S. Houlston, “Thymidylate Synthase Expression and Prognosis in Colorectal Cancer: A Systematic Review and Meta-Analysis,” American Society of Clinical Oncology, Vol. 23, No. 9, 2004, pp. 529-536. doi:10.1200/JCO.2004.05.064.PMid:14752076
- B. Iacopetta, F. Grieu, D. Joseph and H. Elsaleh, “A PolyMorphism in the Enhancer Region of the Thymidylate Synthase Promoter Influences the Survival of Colorectal Cancer Patients Treated with 5-Fluorouracil,” British Journal of Cancer, Vol. 85, No. 6, 2001, pp. 827-830. doi:10.1054/bjoc.2001.2007.
- P. R. Subbarayan, M. Sarkar and B. Ardalan, “Isolation of Genomic DNA from Human Whole Blood,” BioTechniques, Vol. 33, No. 6, 2002, pp. 1231-1234.
- S. Kaneda, J. Nalbantoglu, K. Takeishi, K. Shimizu, O. Gotoh, T. Seno, et al., “Structural and Functional Analysis of the Human Thymidylate Synthase Gene,” The Journal of Biological Chemistry, Vol. 265, No. 33, 1990, pp. 20277-20284.
- H. Hammad, M. Sarkar, N. Gupta, B. Ardalan and P. R. Subbarayan, “The Presence of Three Repeats in the 5’UTR Region of Thymidylate Synthase (TS) Is Associated with Increased TS mRNA Expression in Cultured Human Cancer Cell Lines in Vitro,” Oncology Reports, Vol. 27, No. 1, 2012, pp. 246-249.
- E. Villafranca, Y. Okruzhnov, M. A. Dominguez, J. Garcia-Foncillas, I. Azinovic, E. Martinez, et al., “Polymorphisms of the Repeated Sequences in the Enhancer Region of the Thymidylate Synthase Gene Promoter May Predict Downstaging after Preoperative Chemoradiation in Rectal Cancer,” Journal of Clinical Oncology, Vol. 19, No. 6, 2001, pp. 1779-1786.
- S. Marsh, J. A. McKay, J. Cassidy and H. L. McLeod, “Polymorphism in the Thymidylate Synthase Promoter Enhancer Region in Colorectal Cancer,” International Journal of Oncology, Vol. 19, No. 2, 2001, pp. 383-386.
- M. Gusella and R. Padrini, “G>C SNP of Thymidylate Synthase with Respect to Colorectal Cancer,” Pharmacogenomics, Vol. 8, No. 8, 2007, pp. 985-996. doi:10.2217/14622416.8.8.985. PMid:17716232
- C. M. Ulrich, J. Bigler, C. M. Velicer, E. A. Greene, F. M. Farin and J. D. Potter, “Searching Expressed Sequence Tag Databases: Discovery and Confirmation of a Common Polymorphism in the Thymidylate Synthase Gene,” Cancer Epidemiology, Biomarkers & Prevention, Vol. 9, No. 12, 2000, pp. 1381-1385.
- E. Chu, D. M. Koeller, J. L. Casey, J. C. Drake, B. A. Chabner, P. C. Elwood, et al., “Autoregulation of Human Thymidylate Synthase Messenger RNA Translation by Thymidylate Synthase,” Proceedings of the National Academy of Sciences, Vol. 88, No. 20, 1991, pp. 8977-8981. doi:10.1073/pnas.88.20.8977
- R. Mauritz, E. Giovannetti, I. J. Beumer, K. Smid, C. J. van Groeningen, H. M. Pinedo, et al., “Polymorphisms in the Enhancer Region of the Thymidylate Synthase Gene Are Associated with Thymidylate Synthase Levels in Normal Tissues But Not in Malignant Tissues of Patients with Colorectal Cancer,” Clinical Colorectal Cancer, Vol. 8, No. 3, 2009, pp. 146-154. doi:10.3816/CCC.2009.n.024.
NOTES
*Corresponding author.

