Advances in Bioscience and Biotechnology
Vol.1 No.1(2010), Article ID:1590,5 pages DOI:10.4236/abb.2010.1109
A comparative investigation of interaction between metal ions with L-methionene and related compounds such as alanine, leucine, valine, and glycine in aqueous solution
![]()
Sharif University of Technology, Institute of Water & Energy, Tehran, Iran.
Email: sajadi@sharif.ac.ir
Received 7 February 2010; revised 25 February 2010; accepted 5 March 2010.
Keywords: Methionine; Metal Ion; Potentiometric Titration; Acidity and Stability Constants
ABSTRACT
The acidity and stability constants of M-L (M: M2+; L: Met, L-methionine) complexes, determined by potentiometric pH titrations, were used to make a comparative investigation. The stability constants of the 1:1 complexes formed between M2+ and L–, were determined by potentiometric pH titration in aqueous solution (I = 0.1 M, NaNO3, 25°C). The order of the stability constants was reported. It is shown that regarding to M ion – binding properties, vital differences on complex bilding were considered. It is demonstrated that in M-L complexes, M ion is coordinated to the carboxyl group, is also able to bild macrochelate over amine group. The aforementioned results demonstrate that for M (Met) complex, the stability constants are also largely determined by the affinity of Cu ion for amino group. It is indicated that this additional interaction with amino groups can influence the character of some amino acid complexes in biological systems.
1. INTRODUCTION
The α-amino and α-carboxyl groups of amino acids play prominent roles in metal ion binding. There are many examples of side chain functional groups that also interact with metal ions. Peptides interact with metal ions primarily through side chain functional groups, although there are many examples of peptide amide nitrogen which are functioning as donor atoms with certain metal ions [1]. Many physiologically important peptides function as metal complexes. Methionine is an essential amino acid, which is one of the two sulfur-containing amino acids (Figure 1). The side chain is quite hydrophobic and methionine is usually found buried within proteins. unlike cysteine, the sulfur of methionine is not highly nucleophilic, although it will react with some electrophilic centers. It is generally not a participant in the covalent chemistry that occurs in the active centers of enzymes. The thiol ether can be chemical linkage of the sulfur in methionine. We can compare this terminology with that of the oxygen containing ethers. The sulfur of methionine, as with that of cysteine, is prone to oxidation. The first step, yielding methionine sulfoxide, can be reversed by standard thiol containing reducing agents. The second step yields methionine sulfone, and is effectively irreversible. It is thought that oxidation of the sulfur in a specific methionine of the elastase inhibitor in human lung tissue by agents in cigarette smoke is one of the causes of smoking-induced emphysema [2]. Data on the complexation of essential metal ions and the bioactive ligands methionine and cysteine give insight into many physicochemical processes. The significance of these amino acids is enhanced by the fact that they display independent therapeutic activity [3]. Exposure of the HIV-2 protease to H2O2 resulted in conversion of the two methionine residues (Met-76 & Met-96) to methionine sulfoxide as determined by amino acid analytical and mass spectroscopy [4]. Based on above mentioned essential role of Met is interesting to study the interaction between other metal ions with Met [5-8]. The interesting question is also, is there any interaction between sulfur group and metal ions. In other words, is it possible to detect the last mentioned interaction in aqueous solution?
2. EXPERIMENTAL
2.1. Materials
The L-methionine (extra pure) was purchased from Merck, Darmstadt, Germany. The nitrate salt of Na+, Mn2+, Co2+, Cu2+, and Zn2+ (all pro analysi) were from Merck. Potassium hydrogen phthalate and standard solutions of sodium hydroxide (titrasol), nitric acid, EDTA and of the buffer solutions of pH 4.0, 7.0 and 9.0 were all from Merck. All solutions were prepared with de-ionized water. Water was purified by Milil-Q water purification
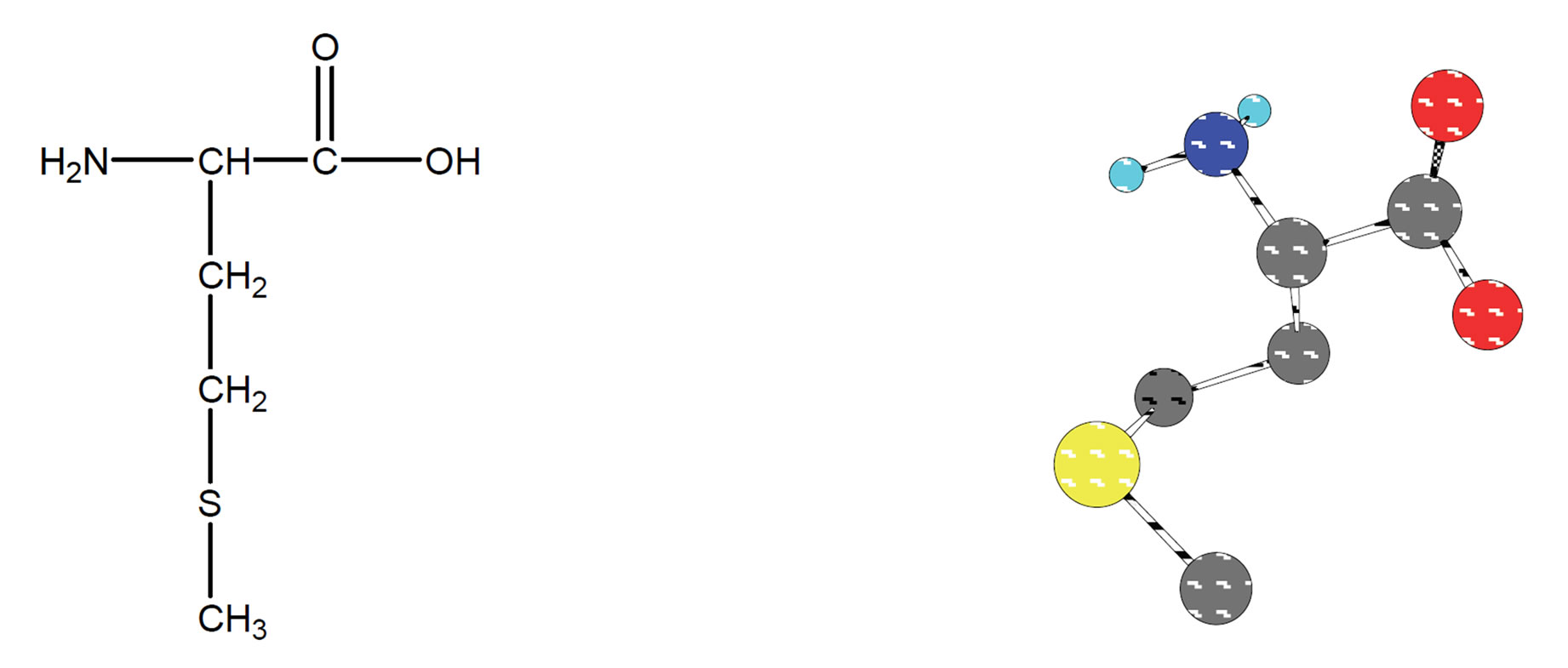
Figure 1. Chemical structure (2D & 3D) of L-methionine.
system, de-ionized and distillated.
2.2. pH Titrations
2.2.1. Reagents
Carbonate-free sodium hydroxide solution 0.03 M was prepared and standardized against sodium hydrogen phthalate and a standard solution of nitric acid 0.54 mM. M(II) nitrate solution (0.6 mM) was prepared by dissolving the above substance in water and was standardized with standard solution of EDTA 0.1 M (triplex).
2.2.2. Apparatus
All pH titrations were performed using a Metrohm 794 basic automatic titrator (Titrino), coupled with a thermostating bath Hero at 25℃ (± 0.1℃) and a Metrohm combined glass electrode (Ag/AgCl). The pH meter was calibrated with Merck standard buffer solutions (4.0, 7.0 and 9.0).
2.2.3. Procedure
For the determination of acid dissociation constants of the ligand Met, an aqueous solution (0.6 mM) of the protonated ligand was titrated with 0.03 M NaOH at 25℃ under nitrogen atmosphere and ionic strength of 0.1 M, NaNO3. For the determination of binary (a ligand and M2+) system, the ratios used were 1:1, M(II): Ligand and 1:1, M(II): Met, 0.6 mM. This solution was titrated with 0.03 M NaOH under the same conditions mentioned above. Each titration was repeated seven times in order to check the reproducibility of the data.
2.2.4. Calculation
The acid dissociation constants, 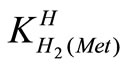 and
and 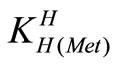 for H2(Met)+ were calculated by an algebraic method. The equilibria involved in the formation of 1:1 complex of Met and a divalent metal ion may be expressed as eqs.3 and 4.
for H2(Met)+ were calculated by an algebraic method. The equilibria involved in the formation of 1:1 complex of Met and a divalent metal ion may be expressed as eqs.3 and 4.
3. RESULTS AND DISCUSSION
The potentiometric pH-titrations (25℃, 0.1 M, NaNO3) were carried out to obtain the acidity and stability constants which are summarized in Tables 1 and 2.
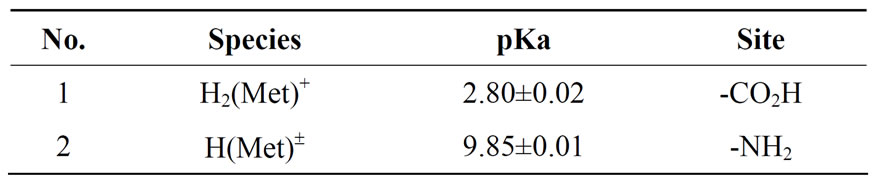
Table 1. Negative logarithm of the acidity constants of Met: L-methionine at 25°C, 0.1 M, NaNO3*, Eqs.1 and 2.
*The given errors are three times the standard error of the mean value or the sum of the propabable systematic errors.
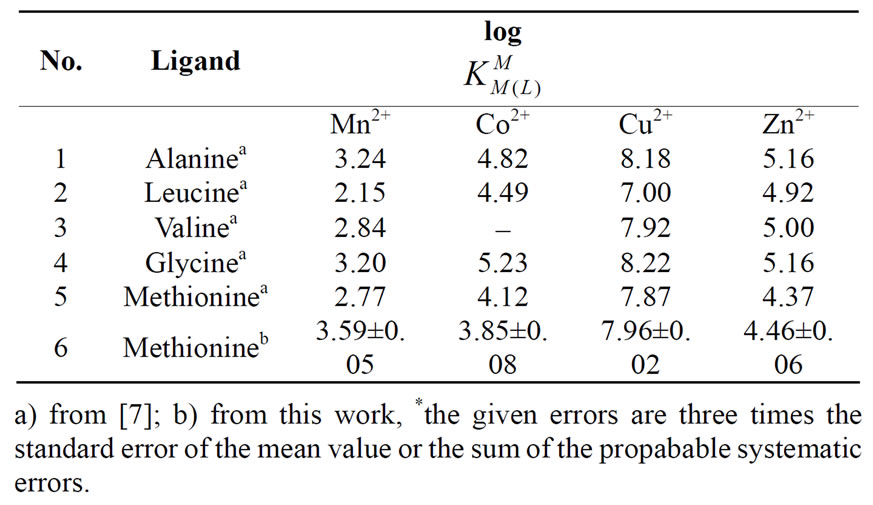
Table 2. Logarithm of the stability constants of binary complexes of M2+ at 25°C, 0.1 M, NaNO3*, eq.4. Met: methionine and related compounds such as Alanine, Leucine, Valine, and Glycine.
3.1. Acidity Constants
Methionate (Met), -O2CCH(NH2)CH2CH2SCH3, is a twobasic species, and thus it can accept two protons, given H2(Met)+, for which the following de-protonation equilibria are hold:
H2(Met)+  H+ + H(Met)± (1)
H+ + H(Met)± (1)
 = [H(Met)±][H+]/[H2(Met)+ (2)
= [H(Met)±][H+]/[H2(Met)+ (2)
H(Met)± H+ + Met- (3)
H+ + Met- (3)
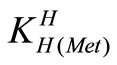 = [Met-][H+]/[H(Met)±] (4)
= [Met-][H+]/[H(Met)±] (4)
The two proton in H2(Met)+ are certainly bound at the terminal acetate and amino groups, i.e., it is released from -O2CCH(NH2)CH2CH2SCH3 according to equilibrium (1)
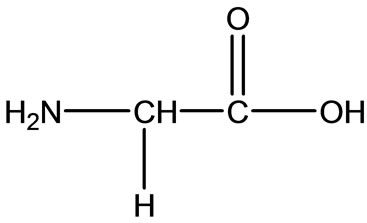 Gly
Gly 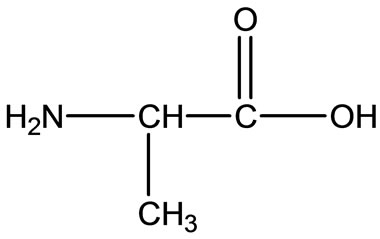 Ala
Ala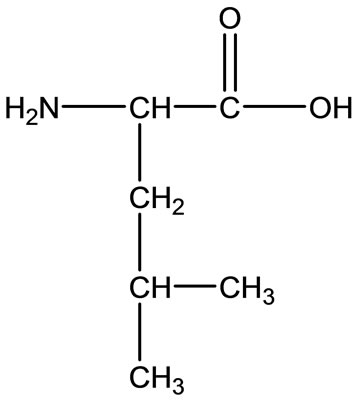 Leu
Leu  Val
Val
Figure 2. Chemical structure of Alanine (Ala), Glycine (Gly), Valine (Val), and Leucine (Leu).
& (2). It is known as zwitter-ion. It is also closed to the de-protonation of acetate groups which occurs at the terminal acetate groups of aspartic acid [7,8].
3.2. Stability of Binary and Ternary Complexes
If we abbreviate for simplicity associating metal ions with M2+, then one may write the following two equlibria of (3) & (4):
M2+ + H(Met)±  M(H;Met)2+ (5)
M(H;Met)2+ (5)
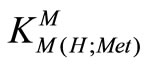 = [M(H;Met)2+]/[M2+][H(Met)±] (6)
= [M(H;Met)2+]/[M2+][H(Met)±] (6)
M2+ + (Met)-  M(Met)+ (7)
M(Met)+ (7)
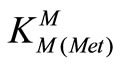 = [M(Met)+]/[M2+][Met-] (8)
= [M(Met)+]/[M2+][Met-] (8)
The experimental data of the potentiometric pH titrations may be completed by considering the above-mentioned equilibria (1) through (4), if the evaluation thereof is not carried into the pH range, where hyrdoxo complex formation occurs. No constant could be determined for the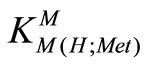 . The data were collected every 0.1 pH unit from the lowest pH which could be reached in the experiment to the beginning of the hydrolysis of M(aq)2+.
. The data were collected every 0.1 pH unit from the lowest pH which could be reached in the experiment to the beginning of the hydrolysis of M(aq)2+.
3.3. Potentiometric Analysis
The chemical structures of some related amino acid are shown in Figure 2. Also we noticed right away that all these amino acids consist of organic part R, carboxyland amino-groups. Just in Met we distinguish the thioether group, which can influence the character of Met or not, as we will consider as follows.
The results of all potentiometric pH-titration, i.e. acidity and stability constants, were summarized below in Tables 1 and 2. The de-protonated amino acid Met- can accept two protons, to give the acid H2(Met)+. The first
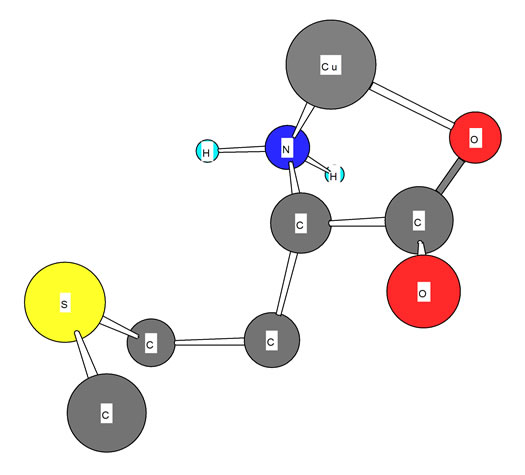
Figure 3. Schematic structures of the species with interactions according to equilibrium (5) for Cu-Met. The structure in the right part of the figure was drawn with the program CS Chem 3D, version 3.5, from Cambridge Software Corporation.
one of these two protons is released from carboxylate group; its pKa is shown in table 1. However, now Met± can release one more proton from neutral -NH3 group, which is the second acidity constant (table 1). The measured acidity constants in this work show good agreement with the same value received by other authors [7,9-12]. However, the carboxyl group is a far stronger acid than the amino group [13].
The stability constants of the binary complexes, such as M-Met were refined separately using the titration data of this system in a 1:1, ligand: M2+ ratio in the same conditions of temperature and ionic strength (according Eqs.3 and 4), as they were in good agreement with reported value [7,12]. We didn’t received reseanable results for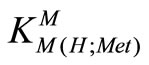 . All the stability constants of table 2 show the usual trend [14-17].
. All the stability constants of table 2 show the usual trend [14-17].
The stability constants of the binary complexes, such as Cu(Met) (figure 3) were refined separately using the titration data of this system in a 1:1, ligand:Cu2+ ratio in the same conditions of temperature and ionic strength
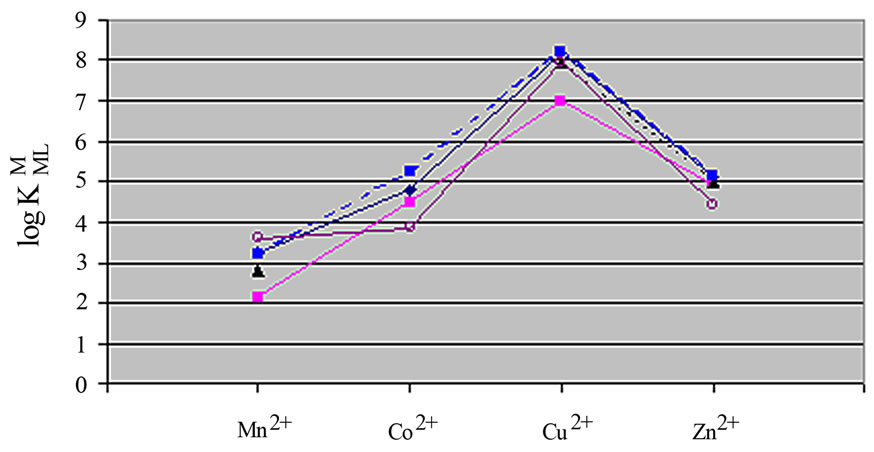
Figure 4. Irving-Williams sequence-type plot for the 1:1 complexes of Mn2+ to Zn2+ with some amino acids (see Table 2). ■: Leucine, ▲: Valine, ¨: Alanine, ●: Glycine, O: Methionine.
(according Eqs.3 and 4), as they were in good agreement with reported value [12,17].
The experimental results are summarized in the table 2. The sequence shows the same Irving-Williams sequence. Also in table 2 were listed the second stability constants of some related amino acids such as alanine, leucin, valine, and glycine.
If we compare these amino acids with each other, we see that they consist of three similar parts. They consist of an organic rest, a carboxyl-, and an amine group. The major difference is the presence of S-group in Met. When we compare the stability constant of these amino acids, we see that all of them show similar values. This means that stability of these amino acids is dependent on the interactions of metal ions with carboxyl-and aminogroups. In other words, it has nothing to do with interactions with the S group. These results are also shown in Figure 3 again. It is evident, that the stability constant shows the identical magnitude. If in M-Met would have seemed some additional interactions, would be observed an increase in stability. However, this was not observed, which means that there is no interaction between the metal ion to sulfur ether.
Also we noticed right away that these thermodynamic constants follow the Irving-Williams sequence (Figure 4). All the stability constants of table 2 show the usual trend. The obtained order is Ca2+ < Mg2+ < Mn2+ < Co2+ < Cu2+ > Zn2+. It is expected that all these metal ions build a chelate with amino acids over carboxyland amine groups, which can be seen in Figure 3. In Figure 3 is shown the structure of Met-complex with Cu2+. We see that Cu binds to carboxyl group on the one hand, and on other hand to amine group. Hard and borderline metal ions show a tendency for interaction with the two mentioned groups.
The ether sulfur is a very weak base. The protonation occurs only in strongly acidic solutions with a pK of -6.2, i.e. much weaker than oxygen ether and ethanol Sulfur atoms are soft bases, and the most favorable interaction with borderline or soft metal ions. This softness is the basis for the interaction of thioehers with metal ions. It is expected that the ether sulfur best with a soft metal ions interact, and for hard metal ions is expected, no significant interactions. Interactions with the ether in sulfure Met is even weaker. These results suggest that the thioether group of the complex does not contribute to the stability of the Met. Comparison of the coordination tendencies of tioeher group as part of bidetate and terdentate ligand offers us a greater tendency for thioether group as a donor atom of a bidentate ligand of lower stability constant than the third donor atom in a ligand terdentate clearly is greater stability constant. This tendency is for Cu, where the more favored tetragonal donor atoms occupy positions on the airwaves, and sulfur is increasingly forced to interact in an apical position, if at all. The thioehre group does not contribute significantly to the stability of hard or borderline metal ions with. Chelate formation of Cu in the glycinate and a weak interaction of the thioether group in the apical position of one of these observations. That the interaction is weak has been reported by the lack of absorption in 400 nm by complexation of a thioether group in the tetragonal plane supports Cu. It was a success for the crystallographic that crystals mainly reflect the structural requirements for structures in solution for labile metal ions. Thus, as expected, the thioether group is not coordinated with a borderline metal ions in complexes met. A trans Met-Pt complex with bidentate chelation in with S and N has not been characterized [18]. Crystal structure also show S-and N-bidentate chelate formation with Pd and Pt [19,20]. Both N and S are bound in terdendate Pt GlyMet-complex. These findings are in contrast to a complex of borderline terdendate GM-Cu, which were replaced a carboxylate O ether by S [21].
Metal ions may function to internally crosslink proteins. For example, at least ten motifs collectively known as zinc fingers have been described in nucleic acid– binding proteins. The M2+ allows relatively short stretches of polypeptide chain to fold into stable unites that can interact with nucleic acids. It is worth mention that a correct selection regarding to the kind of metal ions play a vital role in the biological system. This means the metal ions such as M2+, which coordinate strongly to Met and bild macrochelate is not suitable for right application of metal-ligand interactions in biological systems. For a successful design of identical metal-ligand complexes with similar or better function, such as new drugs, the right selection of both components is significant.
REFERENCES
- L. Stryer, (1995) Biochemistry, 4th Edition, W.H. Freeman Company, New York.
- Bill, G., Rick, H., Ken, W. and Thomas, O.B. (2003) The biology project. Department of Biochemistry and Molecular Biophysics, University of Arizona.
- Suzukin, O. and Navarin, M.S. (1965) Antibiotiki, 6, 562.
- Davis, D.A., et al. (2000) Biochemical Journal, 342(2), 305-311.
- IUPAC-IUBMB Joint Commission on Biochemical Nomenclature. Nomenclature and symbolism for amino acids and peptides. Recommendations on Organic & Biochemical Nomenclature, Symbols & Terminology etc., 17 May 2007.
- Nelson, D.L. and Cox, M.M. (2000) Lehninger, principles of biochemistry, 3rd Edition, Worth Publishing, New York.
- Martel, A.E. (2006) Critical stability constants of metal complexes, 26, Plenum Press, New York.
- Sajadi, S.A.A., Alamolhoda, A.A. and Alavi, A.N. (2009) Scientica Iranica, in Press.
- Handbook of Chemistry & Physics 55, D-129, 1975.
- Miranda, J.L. and Felcman, J. (2003) Polyhedron, 22(2), 225-233.
- Felcman, J. and Miranda, J.L. (1997) Journal of Brazilian Chemical Society, 8, 575.
- IUPAC Stability Conatants Database, Release 3, version 3.02, (1998) coplied by Pettit, L.D. and Powel, H.K.J. Academic Software Timble, UK.
- Sigel, H., Zuberbuehler, A.D. and Yamauchi, O. (1991) Analytica Chimica Acta, 255, 63.
- Ali, S., Sajadi, A., Song, B. and Sigel, H. (1998) Inorganica Chimica Acta, 283, 193-201.
- Sajadi, S.A.A., Song, B., Gregan, F. and Sigel, H. (1999) Inorganica Chimica, 38(3), 439-448.
- Sajadi, S.A.A., Song, B., Gregan, F. and Sigel, H. (1997) Bulletin of the Chemical Society of Ethiopia, 11(2), 121-130.
- Voet, D. (1997) Biochemistry, John Wiley, 560.
- Volshtein, L.M. Krylova, L.F. and Mogilevkina, M.F. (1967) Russian Journal of Inorganic Chemistry, 12, 832.
- Warren, R.C., McConnell, J.F. and Stephenson, N.C. (1970) Acta Crystallographica, B26, 1402.
- Freeman, H.C. and Golomb, M.L. (1970) Chemical Communications, 1523.
- Bear, C.A. and Freeman, H.C. (1976) Acta Crystallographica, B32, 2534.

