Journal of Biophysical Chemistry
Vol.3 No.2(2012), Article ID:19444,8 pages DOI:10.4236/jbpc.2012.32020
Build mutant and build homology protein structure predictions for indonesian avian influenza neuraminidase
![]()
Departemen Fisika, Fakultas MIPA, Universitas Indonesia, Depok, Indonesia; *Corresponding Author: sigit.jaya.herlambang@gmail.com
Received 1 March 2012; revised 22 March 2012; accepted 11 April 2012
Keywords: Protein; Modeling; Mutation; Build Model Protocols
ABSTRACT
Protein structure modeling using a homologous template is one of many routines that accompany the molecular dynamics simulation for biological material. There are currently two protocols of protein modeling available in Accelrys Discovery Studio 2.1, Build Mutants and Build Homology Modeling protocols. Both are template-based modeling, but with a different process. In this study, two different templates, 3CKZ and 274Y, have been used to see how much the differences will be made by those two protocols if the templates has significant percentage of identity. Evaluation of structure models has been performed using DOPE score, 3D-profile, and PROCHECK. The results indicated that Build Mutants Protocols produces more stable structures but has a low reliability values and low quality of stereochemistry when using a template that has a lower percentage of identity. The results also yield more stable, reliable, and higher percentage of residues in most favoured and additionally allowed region for the usage of Build Homology Modeling Protocol on both templates. These observations suggest that Build Homology Modeling protocol is recommended for protein modeling.
1. INTRODUCTION
The early generation of protein molecular structure using x-ray crystallography performed in the late 1956’s by Perutz and Kendrew [1], thenceforth more than 62293 x-ray crystal structures of proteins have been generated [2]. Adhering to that, the knowledge of proteins structural-function has increased dramatically. There is no doubt that the x-ray structure crystallography takes much time and involves difficult methods which requires significant amount of expertise and resources to generate only one protein structure. To deal with this problem, many alternative methods have been developed. Among them, there are two protocols in Accelrys Discovery Studio 2.1 which adopted MODELLER program [3], Build Homology Modeling and Build Mutants protocols, to generate structure models based on their templates.
It has been generally known that the medium identity percentage of the sequences resulted only slight differences in their 3D structure [4]. Although the results of homology models still need an assessment, the performance of Build Homology Modeling protocol has been well understood and documented [5,6]. These protein structure prediction protocol could build backbone, sidechains, loop (if needed), and any necessary cofactors and ligand.
In contrast to the Build Homology Modeling protocol, Build Mutants protocol uses the template without changing backbone, side-chains, loop and any necessary cofactors and ligand. This protocol mutates one or more residues of wild-type protein and optimizes the local structure in the specified range while the outside radius is made fix. Therefore, the local energy minimization that occurs will greatly affect if the mutated residues are quite a lot.
Both Build Homology Modeling and Build Mutants protocols were used to generate H5N1 avian influenza virus (AIV) neuraminidase (NA) structure models. Avian influenza NA has a tetramer quaternary structure. The proper assembly of its monomer is critical for a functional protein. The residue in the codon 274 is one of several functional residues that provide direct interactions with neuraminidase substrate and inhibitors. The mutation in this codon could help the NA to avoid the interaction with inhibitors and able to bind its native substrate, sialic acid (SA). Because of it, the modeling of NA should be taken gently.
In this study, we try to theoretically investigate to what extent the neuraminidase models will vary if both Build Homology Modeling and Build Mutants protocols were utilized to generate models using different identity percentage templates. Structural and quantitative analysis was performed on all models using models root mean square deviation (RMSD), three protein assessment methods (DOPE, 3D-profile, and PROCHECK) and visualization comparison. Evaluation of models generated from these two protocols is important as a reference for selecting modeling methods to be used, particularly for the sequence with low percentage of identity.
2. MATERIALS AND METHODS
2.1. Sequences and Templates Collection
Amino acid sequence of the Indonesian H274Y mutant NA has been taken from National Center of Biotechnology Information (NCBI) [7]. The accession code of these NA is ABW06159 and has been extracted from (A/Indonesia/560H/2006(H5N1)) AIV. The sequence file is labeled as 274Y and manually has been modified by changing tyrosine, in codon 274, into histidine. This modification was based on the sequence from (A/Indonesia/ CDC625/2006(H5N1)) AIV neuraminidase (found to have an accession code ABI36369). The sequence is then labeled 274H and has been aligned with 274Y and 3CKZ template sequences. The 274Y template has been generated and selected by three protein assessment in our previous study [8] from the 3CKZ template. The second template, the 3CKZ structure, was obtained at the RCSB [9] by the recommendation of BLASTP 2.2.16 [10] and InterProScan [11] results provided by Swiss Expasy [12-16]. All atoms that are not needed have been removed from both templates to optimize the structure prediction process. After refinement, the CHARMm force field was applied to the templates to add the missing hydrogen atoms [17,18].
2.2. Sequences Alignment
The sequences alignment of 274H, 274Y, and 3CKZ has been performed with Align Two Profiles alignment type. The 274H sequence has been aligned in two steps. First, with the 274Y sequence, and then, the result is aligned with the 3CKZ sequence. Both alignments have been carried out using Align123 based on ClustalW program [19] which employ progressive pairwise alignment algorithm. Sequence alignment was executed with the main parameters such as fast pairwise alignment, BLOSUM scoring matrix [20], and Kabsch-Sander secondary structures information [21]. The result is then used as the input for both Build Homology Modeling and Build Mutants protocols.
2.3. Structures Modeling
The models have been generated by the two methods with each method uses two templates. For each template, we construct five 274H NA models. In the 274Y template, only one residue that will be mutated while in 3CKZ template there are ten residues. The generation of 274H NA structures by Build Homology Modeling protocols, from both templates, has been executed with identical parameters, such as high optimization level and cut overhangs to remove the terminal-unaligned residues in model sequence. The second method, Build Mutants protocol, has been carried out using high optimization level, high resolution DOPE method, and 4.5 Å cut radius parameters (the atoms in larger specified radius from the changed residues will be defined as fix atoms). The disulfide bridges, Cis-Prolines, and additional restraints were not added to simplify the models building.
2.4. Models Assessments
All models have been evaluated with Discrete Optimized Protein Energy (DOPE) method [22] using MODELER and 3D-profiles (verify score) [23,24]. DOPE method has been conducted with with high resolution whereas in 3Dprofiles the parameters such as smooth windows sized 10 and Kabsch-Sander algorithm for secondary structure method [25] has been given. All models are also sent to Swiss Expasy to be analyzed with PROCHECK [26-28] to get insight the stereo-chemical perspective of our models. All the generated models were also super-imposed to analyze root mean square deviation (RMSD) in the structure.
3. RESULTS
The evaluation of protein structure models have been taken to elucidate the structures reliability, stability, and stereo-chemistry as the assessment to choose a proper model for simulation. Each assessment is represented by the value of 3D-profile, DOPE score, and PROCHECK, respectively. The combination of these three assessments is able to produce a significant result in the model selection instead using only one protein assessment.
The aligned sequences file was used as the input for NA’s modeling. Investigation on the aligned sequences file shows that the 274H sequence have an identity percentage of 99.7% with 274Y whereas with 3CKZ is only 97.4%. Further comparison on those three sequences, we have found that there are nine different residues between 274H and 3CKZ whereas between 274H and 274Y is only one residue (see Figure 1). The usage of two different identity percentage templates will cause different results in the generation of models, particularly if we also use different modeling methods.
There are a total of 20 models have been generated from the template 274Y and 3CKZ using a variation of Build Mutants and Build Homology Modeling protocols.
Figure 1. The multiple alignment sequences result.
The models are then classified into four separate groups to clear the course of the discussion. The groups are the homology modeling models from 3CKZ template group (HM3CKZ group), the homology modeling models from 274Y template group (HM274Y group), the build mutants from 3CKZ template group (BM3CKZ group), and the build mutants from 274Y template group (BM274Y group). The results of the structure models evaluation have been listed in the Table 1, and the structural visualization results are displayed in the Figures 2 and 3. All models RMSD with the reference used is the closest native structure of Indonesian H274Y NA, 3CKZ structure, have been calculated and shown in the Table 2.
The reliability of models can be seen from the models 3D-profile score (contained in the 3D-profile column in Table 1). The 3D-profile score values were summarized from all amino acid profile scores, which have been calculated from the string of residue environments and the pre-calculated scoring matrix. The pre-calculated scoring matrix have a correlation with the probabilities of finding each amino acids in each environment classes as observed in a database of known structures and related sequences. Larger value indicates the fittest protein structure model.
The reliability of models was then investigated. The 3D-profile score of models in HM3CKZ and HM274Y groups are almost similar. Both groups also reveal the intensity of Build Homology Modeling protocol to predict the entire molecules. Even when a lower identity percentage template has been used, there are no signifycant differences in the reliability of the models. This is in contrast to the more subtle value differences between BM3CKZ and BM274Y groups. The highest 3D-profile score of each group are 200.04, 201.05, 186.59, and 191.38; for HM3CKZ, HM274Y, BM3CKZ, and BM274Y groups respectively.
The Dope score column filled with the energetic values which measures the relative stability of structures with respect to other structure of the same protein. The most negative value indicates the most stable. From the DOPE score column can be seen that BM3CKZ group models have a lower DOPE scores than the other groups. Following BM3CKZ, there are HM3CKZ, BM274Y, and HM274Y, respectively. Energetic difference among them may be caused by the differences of local conformation areas which have been minimized. The most negative value of DOPE score have been observed from HM3CKZ, HM274Y, BM3CKZ, and BM274Y groups are –38801.99, –35752.53, –39973.06, and –36804.14 kcal/mole.
The most favoured & additionally allowed residues percentages (collected from PROCHECK protein stereochemistry assessment) have been shown in the third column. These values have been extracted from ramachandran

Table 1. The model assessments of Na structure models using three different assessment methods.
 (a)
(a) (b)
(b) (c)
(c)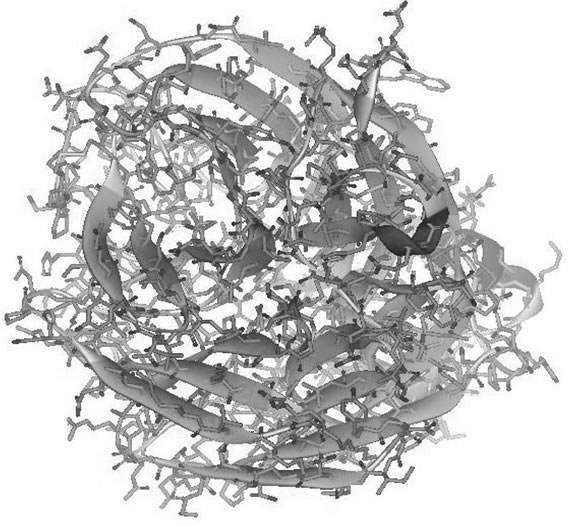 (d)
(d)
Figure 2. The best structure model of: (a) 274H best model structure generated from build homology modeling using 3CKZ template; (b) 274H best model structure generated from build homology modeling using 274Y template; (c) 274H best model structure generated from build mutants using 3CKZ template; (d) 274H best model structure generated from build mutants using 274Y template.
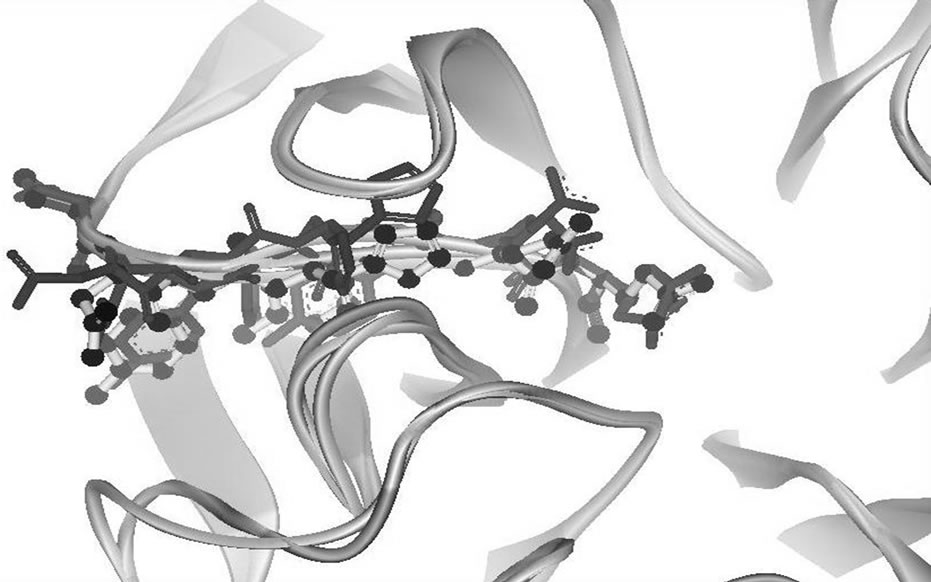
Figure 3. (a) 274H best model structure generated from build homology modeling using 3CKZ template (black); (b) 274H best model structure generated from build homology modeling using 274Y template (yellow); (c) 274H best model structure generated from build mutants using 3CKZ template (red); (d) 274H best model structure generated from build mutants using 274Y template (blue).
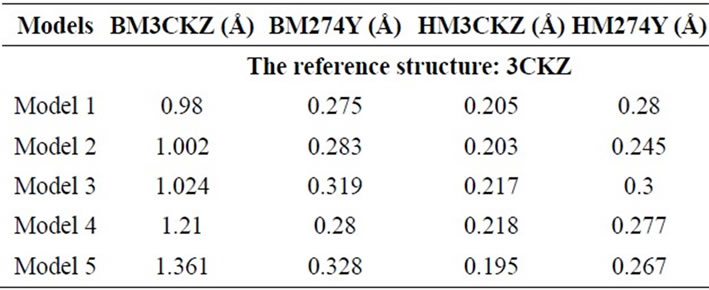
Table 2. The RMSD of all models compared with the native structure 3CKZ.
plots to simplify the evaluation process. The results of PROCHECK assessment are similar with 3D-profile in terms of the order from the highest to the lowest. The HM3CKZ and HM274Y groups show their similar order of the most favoured & additionally allowed residues percentages whereas in BM3CKZ and BM274Y groups showed a different result when the different identity percentage templates have been used.
Also from this column can be seen that Build Homology Modeling protocol should do better than Build Mutants protocol for NA’s modeling when using low identity percentage template. The highest most favoured & additionally allowed residues percentage for each HM3CKZ, HM274Y, BM3CKZ, and BM274Y groups are 99.4%, 99.1%, 93.2%, and 98.7%, respectively.
Overall, we have found that HM3CKZ models are preferable to be used in the simulation. They have better structure models reliability, stability, and stereo-chemistry compared with other groups. Further selection has been undertaken to choose the best model from each groups. For all groups, the model 4 was chosen since it is more reliable (can be seen from its 3D-profile score), stable (represented in the DOPE score), and higher stereo-chemistry value compared with other models. The compareson of each group best model showed that the protein models generated from Build Mutants protocol should not be taken if the template has low identity percentage. It can be seen from the BM3CKZ group that has decreased in all protein assessment scores. The evaluation of the best model selected from each group was not discussed deeply in here to keep the shape of the paper aim and goal.
Analysis of structure divergences (represented with their RMSD) between all models generated and 3CKZ showed that models of HM3CKZ group has a lower range of RMSD (0.195 - 0.218 Å) compared with other groups (see Table 2). In contrast with that, the models of BM3CKZ group showed significant deviation from 3CKZ structure (0.98 - 1.361 Å). Other groups, HM274Y and BM274Y show similar order of RMSD (around 0.275 - 0.328 Å). This indicates only BM3CKZ group showed a relatively high RMSD with 3CKZ structure.
The structures visualization of the best model selected of each group have been displayed in Figures 2(a), (b), (c), and (d). It can help to directly evaluate the models by harnessing the view of NA’s monomer quaternary structure. Visual structural comparison show that the best model from Figure 2(c) (BM3CKZ structure) is more cluttered than other groups best models. This supports the results of protein assessments and RMSD where the BM3CKZ group models has low quality. Observation in Figures 2(a) and (b) show a little difference in the surface regions. In Figure 2(d) is shown the BM274Y neuralminidase best model have several disoriented in structure backbone near the mutated residue. These results are consistent as the RMSD results also showed significant differences between BM3CKZ models and 3CKZ structure.
In the Figure 3, the residues adjacent to the changed residue were shown to compare four best models local conformation. The residues shown in this figure are from codon 271 to 277. The local conformation of the best model of HM3CKZ and HM274Y groups, seen as black ball-stick and yellow stick figures, has an overlapping local conformation. The local conformation of Build Mutants protocol best models, figured in blue (BM274Y group) and red (BM3CKZ group), has shown little disorientation as the results of the optimization structure after selected residues were mutated.
4. DISCUSSION
The aim of this study is to select an appropriate method to build the best model of avian influenza NA. In order to achieve that, the RMSD and three different protein assessments have been utilized to evaluate all models generated. The variation of templates (with different identity percentage) and model building protocols were combined. All models have been evaluated through the PROCHECK, 3D-profile, and DOPE score.
The modeling of proteins subject remains a major bottleneck in preparing the material for simulations. The identity and similarity of protein structure template should be high enough to prevent errors in the model structure. Although Lesk et al. [4] had explained that the successful model building of an unknown protein structure depends on knowing the structure of a reasonably close relative, but a single change in the critical protein region, especially in the active/functional site of the protein, careful modeling method is still needed. This is because the mutation in such active site is able to trigger a different binding affinity of the complex molecule. From the RMSD results, we can conclude that the models still fit the Lesk et al. results. As explained in that publication that for proteins with sequence homologies of 50% or more the shifts are much smaller, lying in the range 0.3 - 1.5 Å. As commonly know, the RMSD is a method to represent the differences in structures appearance.
These indication and analysis are also supported by the Chotial et al. study on the relation between the divergence of sequence and structure in proteins [29]. In their publiccation has been explained that the RMSD in the positions of the main chain atoms is related to the fraction of mutated residues. Furthermore, in their publication were written that homologous proteins have regions which retain the same general fold and regions where the folds differ. For pairs of distantly related proteins (residue identity approximately 20%), the regions with the same fold may comprise less than half of each molecule. The regions with the same general fold differ in structure by amounts that increase as the amino acid sequences diverge. Moreover, other study shows the relationship between sequence and interaction divergence in proteins [30]. Aloy et al. found that the closest homologous (30% - 40%) almost interact the same way. Based on that, we could conclude that all templates used in here are good enough to get the same interaction for dynamics simulation. But, since proper method is needed for the generation of models, we still need to search for a better method.
The usage of Build Mutants protocol is not as popular as Build Homology Modeling protocol. The utilization of Build Mutants protocol is known published in a few studies and short report such as in generation of δ-crystallin [31], γC-crystallin [32], TLR2 protein [33], and GABA receptor [34]. All these studies explain that the usage of Build Mutants protocol was used only to mutate a single residue. Few of these studies use Build Homology Modeling protocol for the generation of wild-type model.
In contrast with Build Mutants protocol, the usage of Build Homology Modeling protocol is a widely-used method to generate the protein model. At this discussion, only studies that evaluate the Build Homology Modeling protocol models have been presented here. The models construction (using Build Homology Modeling protocol) and evaluation studies which have a similarity with our PROCHECK results are from Patel et al. [35], Khatri et al. [36], Satpathy et al. [37], and Singh et al. [38]. The amylase, ninjurin, lycopene, and H1N1 nucleocapsid models have been generated in those experiments. The percentages of the most favoured region are 87.3%, 87.3%, 88.2%, and in range 76.3% - 87.4%, respectively.
Better models have been generated from Build Homology Modeling protocol compared with Build Mutants protocol models. It is obvious from the results that show the more reliable and preferable stereo-chemistry from HM3CKZ and HM274Y groups. Although BM3CKZ groups has better stability that have been predicted in the DOPE score. But still, Build Mutants protocol is suggested not to be taken if the template has low identity percentage. The local structure optimization in Build Mutants protocol will cause serious problem in the tertiary and quaternary structure and may devastating the function of an important sites of enzymes. This measured structural divergence may be caused by the process of energy minimization produce the replacement of adjacent atoms until the targeted energy gradient is reached. If there are too many residues have been changed, the molecule is possible to be broken or damaged.
5. CONCLUSION
Build Homology Modeling and Build Mutants are two protocols which can be used for generating protein models using homologous template. We have made three suggestions related with the results of this study. First, Build Homology Modeling and Build Mutants protocols are good for models generation if the sequence possess a high identity percentage with the template. Second, the Build Mutants should not be used if the only template structure available has low identity percentage. Third, Build Homology Modeling protocol is preferable for either high or low identity percentage. Further research on the threshold is needed to determine the exact percentage of identity and a proper way of model generation should be taken with the best approach.
6. ACKNOWLEDGEMENTS
We would like to express gratitude towards Ding Ming Chee of Accelrys Singapore for the Accelrys Discovery Studio 2.1 trial sent to us.
REFERENCES
- Kendrew, J. C., Bodo, G., Dintzis, H.M., Parrish, R.G., Wyckoff, H. and Phillips, D.C. (1958) A three-dimensional model of the myoglobin molecule obtained by x-ray analysis. Nature, 181, 662-666. doi:10.1038/181662a0
- http://www.rcsb.org/pdb/statistics/holdings.do
- Sali, A., Pottertone, L., Yuan, F., Van Vlijmen, H. and Karplus, M. (1995) Evaluation of comparative protein modeling by MODELLER. Proteins, 23, 318-326. doi:10.1002/prot.340230306
- Lesk, A.M. and Chothia, C. (1986) The response of protein structures to amino-acid sequence changes. Philosophical Transactions of the Royal Society of London. Series A. Mathematical and Physical Sciences, 317, 345-356. doi:10.1098/rsta.1986.0044
- Johnson, M.S., Srinivasan, N., Sowdhamini, R. and Blundell, T.L. (1994) Knowledge-based protein modeling. Critical Reviews in Biochemistry and Molecular Biology, 29, 1- 68. doi:10.3109/10409239409086797
- Congreve, M., Murray, C.W. and Blundell, T.L. (2005) Keynote review: Structural biology and drug discovery. Drug Discovery Today, 10, 895-907. doi:10.1016/S1359-6446(05)03484-7
- http://www.ncbi.nlm.nih.gov/genomes/FLU/Database/nph-select.cgi?go=database
- Herlambang, S. J. and Saleh, R. (2011) Indonesian avian influenza H274Y mutant neuraminidase homology models assessment. Journal of Biophysical Chemistry, 2, 345-352. doi:10.4236/jbpc.2011.23039
- http://www.pdb.org/pdb/home/home.do
- Altschul, S.F., Madden, T.L., Schaffer, A.A., Zhang, J., Zhang, Z., Miller, W. and Lipman, D.J. (1997) Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25, 3389- 3402. doi:10.1093/nar/25.17.3389
- Zdobnov, E.M. and Apweiler, R. (2001) InterProScan—An integration platform for the signature-recognition methods in InterPro. Bioinformatics, 17, 847-848. doi:10.1093/bioinformatics/17.9.847
- Arnold, K., Bordoli, L., Kopp, J. and Schwede, T. (2006) The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics, 22, 195-201. doi:10.1093/bioinformatics/bti770
- Kiefer, F., Arnold, K., Künzli, M., Bordoli, L. and Schwede, T. (2009) The SWISS-MODEL Repository and associated resources. Nucleic Acids Research, 37, 387-392. doi:10.1093/nar/gkn750
- Schwede, T., Kopp, J., Guex, N. and Peitsch, M.C. (2003) SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Research, 31, 3381-3385. doi:10.1093/nar/gkg520
- Guex, N. and Peitsch, M. C. (1997) SWISS-MODEL and the Swiss-Pdb Viewer: An environment for comparative protein modelling. Electrophoresis, 18, 2714-2723. doi:10.1002/elps.1150181505
- Peitsch, M.C. (1995) Protein modeling by E-mail (From amino acid sequence to protein structure: A free one-hour service). Nature Biotechnology, 13, 658-660. doi:10.1038/nbt0795-658
- Brooks, B.R., Bruccoleri, R.E., Olafson, B. D., States, D. J., Swaminathan, S. and Karplus, M. (1983) CHARMM: A program for macromolecular energy minimization and dynamics calculations. Journal of Computational Chemistry, 4, 187-217. doi:10.1002/jcc.540040211
- Brooks, B.R., Brooks III, C.L., MacKerell Jr., A.D., Nilsson, L., Petrella, R.J., Roux, B., Won, Y., Archontis, G., Bartels, C., Boresch, S., Caflisch, A., Caves, L.S.D., Cui, Q., Dinner, A.R., Feig, M., Fischer, S., Gao, J., Hodocek, M., Im, W., Kuczera, K., Lazaridis, T., Ma, J., Ovchinnikov, V., Paci, E., Pastor, R.W., Post, C.B., Pu, J.Z., Schaefer, M., Tidor, B., Venable, R.M., Woodcock III, H.L., Wu, X., Yang, W., York, D.M. and Karplus, M. (2009) CHARMM: The Biomolecular Simulation Program. Journal of Computational Chemistry, 30, 1545-1614.
- Thompson, J.D., Higgins, D.G. and Gibson, T.J. (1994) CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic acids research, 22, 4673-4680. doi:10.1093/nar/22.22.4673
- Henikoff, S. and Henikoff, J.G. (1992) Amino acid substitution matrices from protein blocks. Proceedings of the National Academy of Sciences of the USA, 89, 10915-10919. doi:10.1073/pnas.89.22.10915
- Kabsch, W. and Sander, C. (1983) Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometrical features. Biopolymers, 22, 2577-2637. doi:10.1002/bip.360221211
- Shen, M.-Y. and Sali, A. (2006) Statistical potential for assessment and prediction of protein structures. Protein Science, 15, 2507-2524.
- Lüthy, R., Bowie, J.U. and Eisenberg, D. (1992) Assessment of protein models with three-dimensional profiles. Nature, 356, 83-85. doi:10.1038/356083a0
- Ramachandran, G. N., Ramakrishnan, C. and Sasisekharan, V. (1963) Stereochemistry of polypeptide chain configuretions. Journal of Molecular Biology, 7, 95-99. doi:10.1016/S0022-2836(63)80023-6
- Kabsch, W. and Sander, C. (1983) Biopolymers Dictionary of protein secondary structure: Pattern recognition of hydrogen-bonded and geometrical features. Biopolymers, 22, 2577-2637. doi:10.1002/bip.360221211
- Fiser, R.K. and Sali, A. (2000) Modeling of loops in protein structures. Protein Science, 9, 1753-1773. doi:10.1110/ps.9.9.1753
- Laskowski, R.A., MacArthur, M.W., Moss, D.S. and Thornton, J.M. (1993) PROCHECK: A program to check the stereochemical quality of protein structures. Journal of Applied Crystallography, 26, 283-291. doi:10.1107/S0021889892009944
- Morris, A.L., MacArthur, M.W., Hutchinson, E.G. and Thornton, J.M. (1992) Proteins: Structure, Function, and Bioinformatics. Biochemistry and Molecular Biology, 12, 345-364.
- Chothia, C. and Lesk A.M. (1986) The relation between the divergence of sequence and structure in proteins. EMBO Journal, 5, 823-826.
- Aloy, P., Ceulemans, H., Stark, A. and Russel, R.B. (2003) The relationship between sequence and interaction divergence in proteins. Journal of Molecular Biology, 332, 989- 998. doi:10.1016/j.jmb.2003.07.006
- Huang, C.W., Chen, Y.H., Chen, Y.H., Tsai, Y.C. and Lee, H.J. (2009) The interacton of Glu294 at the subunit interface is important for the activity and stability of goose δ-crystallin. Molecular Vision, 15, 2358-2363.
- Kumar, M., Agarwal, T., Khokhar, S., Kumar, M., Kaur, P., Roy, T.S. and Dada, R. (2011) Mutation screening and genotype phenotype correlation of a-crystallin, g-crystallin and GJA8 gene in congenital cataract. Molecular Vision, 17, 693-707.
- Biswas, D., Gupta, S.K., Sindhwani and Patras, A. (2009) TLR2 polymorphisms, Arg753GLn and Arg677Trp, are not associated with increased burden of tuberculosis in Indian patients. BMC Research Notes, 2, 162-167.
- Zhang, J., Xue, F. and Chang, Y. (2008) Structural Determinants for Antagonist Pharmacology That Distinguish the ρ1GABAC Receptor from GABAA Receptor. Molecular Pharmacology, 74, 941-951. doi:10.1124/mol.108.048710
- Patel, A., Dewangan, R., Khatri, S., Choubey, J., Gupta, S.K. and Verma, M.K. (2009) Identification of insilico 3D structure of amylase (Drosophila melanogaster) and comparative computational studies. Journal of Engineering and Technology Research, 1, 39-45.
- Khatri, S., Patel, S., Choubey, J., Gupta, S.K. and Verma, M.K. (2010) Insilico 3D structure prediction of cell membrane associated protein ninjurin (homosapiens). Current Research Journal of Biological Sciences, 2, 1-5.
- Satpathy, R., Guru, R.K., Behera, R. and Priyadarshini, A. (2010) Homology modelling of lycopene cleavage oxygenase: The key enzyme of bixin production. Journal of Computer Science of System Biology, 3, 59-61.
- Singh, S., Kumar, A., Patel, A., Tripathi, A., Kumar, D. and Verma, D. (2010) In silico 3D structure prediction and comparison of nucleocapsid protein of H1N1. Journal of Modelling and Simulation of Systems, 1, 108-111.

