Journal of Biosciences and Medicines
Vol.04 No.10(2016), Article ID:71635,12 pages
10.4236/jbm.2016.410009
Association between Toll-Like Receptor 4 (Thr399Ile and Asp299Gly), Toll-Like Receptor 2 196-174 Deletion Polymorphisms with cryptosporidiosis
Asmita Dey1, Ujjala Ghoshal1*, Vikas Agarwal2, Uday Chand Ghoshal3
1Departments of Microbiology, Sanjay Gandhi Postgraduate Institute of Medical Sciences, Lucknow, India
2Departments of Immunology, Sanjay Gandhi Postgraduate Institute of Medical Sciences, Lucknow, India
3Departments of Gastroenterology, Sanjay Gandhi Postgraduate Institute of Medical Sciences, Lucknow, India

Copyright © 2016 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY 4.0).
http://creativecommons.org/licenses/by/4.0/



Received: September 8, 2016; Accepted: October 28, 2016; Published: October 31, 2016
ABSTRACT
Toll-like receptors (TLRs) 2 and 4 specifically recognize lipopolysaccharide motifs on surfaces of microorganisms. Polymorphims in the TLR genes result in structural variations in the proteins, abnormal host-pathogen interactions and hence, altered immune response. Opportunistic parasite Cryptosporidium spp. infects immuno- compromised patients who present with heterogeneous clinical manifestations ranging from mild infection resolving in a few weeks, to severe form characterized by voluminous diarrhoea, prolonged symptoms and recurrences. The present study investigates the role of TLR2 196_174del, TLR4 Asp299Gly and Thr399Ile polymorphisms in pathogenesis of cryptosporidiosis. Stool samples were collected from 210 immuno-compromised (renal transplant recipients and patients with Human immunodeficiency virus infection) both with and without cryptosporidiosis, and 200 healthy subjects for detection of Cryptosporidium spp. Blood samples were also collected from the patients and healthy subjects for genotyping of Δ22 (TLR2 196_174del) polymorphism, A896G (TLR4 Asp299Gly) and C1196T (Thr399Ile) single nucleotide polymorphisms (SNPs) by polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP). Cryptosporidium spp. was detected in 70 immunocompromised patients, while it was absent in all the healthy controls. No significant difference was observed in Δ22, A896G and C1196T alleles and genotypes (P > 0.05), between cases and controls. Thus, TLR2 196_174del, TLR4 Asp299Gly and Thr399Ile polymorphisms are not significantly associated with the pathogenesis or progression of cryptosporidiosis, in the Indian population included in the study.
Keywords:
Cryptosporidium, HIV, Renal Transplant, Immuno-Compromised, TLR, Immune Response

1. Introduction
Cryptosporidium spp. is a Coccidian enteric parasite causing severe diarrhea in immuno-compromised patients, such as human immunodeficiency virus (HIV) infected patients and organ transplant recipients [1] - [5] . This obligate intracellular parasite infects the small intestine. Severity increases as larger parts of the gastrointestinal mucosa get infested [6] . Clinical manifestations range from mild infection resolving in a few weeks, to severe forms characterized by voluminous diarrhea, prolonged symptoms and recurrences [6] - [8] . Such heterogeneity in clinical presentations might depend on genetic variations in the host leading to impaired immune mechanisms. The innate immune response is the first line of defense against invading pathogens. Therefore, genes encoding for the innate immune system components might be important candidates for studying the varying degrees of clinical manifestations or host immune response against the parasite. Also, association between genetic variations and infection might serve as risk factors.
Toll-like receptors (TLRs), a class of pattern recognition receptor (PRRs) proteins expressed on cells of the innate immune system recognize motifs on pathogen surfaces (pathogen-associated molecular patterns or PAMPs), thus, triggering protective inflammatory response. TLR 2/4 specifically recognizes lipopolysaccharide motifs on surfaces of invading microorganisms and initiate series of signaling events leading to expression of pro-inflammatory genes such as cytokines, which might result in symptomatic disease. Polymorphism in the genes of TLRs results in abnormal host-pathogen interaction due to variation in TLR proteins and thus, anomalous immune response. The two most common types of genetic variations in TLR 4 are the single-nucleotide polymorphism (SNP) Asp/Gly at amino acid 299 due to A/G transition, (A896G) and Thr/Ile at amino acid 399 due to C/T transition (C1196T) and that in TLR 2 is 22 base pair insertion/deletion polymorphisms in the first un-translated exon at position 196- 174 (Δ22). Various TLR polymorphisms have been implicated in susceptibility and progression of diseases like neurocysticercosis, Hepatitis C virus, asthma, cervical cancer and asymptomatic filariasis [9] - [11] . A few previous studies have reported that intestinal and biliary invasion of Cryptosporidium spp. activates the TLR4/MyD88/NF-kB loop [12] [13] . Similarly, C. parvum infection of cultured human cholangiocytes induced recruitment of TLR2 and TLR4 at the infection sites [13] . However, there is no study describing the role of TLR polymorphisms in cryptosporidiosis. In fact, these receptors or host factor (TLR) might serve as promising targets for cryptosporidiosis risk assessment, thus preventing the infection by proper control measures or helping in early detection and management of the infection.
We undertook a prospective study in order to elucidate the association between these polymorphisms and pathogenesis of cryptosporidiosis.
2. Materials and Methods
2.1. Study Population and Sample Collection
Immuno-compromised subjects (210), with and without cryptosporidiosis were included in the study. These subjects were renal transplant recipients (RT) and patients with Human immunodeficiency virus (HIV) infection attending the Departments of Nephrology and Immunology respectively, of a tertiary care institute in India, during a period between December 2008 and October 2013. Patients negative for HIV infection and/or those without renal transplantation were excluded. The sample size was calculated by software QUANTO (Ver. 1.2.4). The calculated sample size of cases (immune-compromised subjects with cryptosporidiosis) was 62 and of diseased controls (immune-compromised subjects without cryptosporidiosis) was 124. We rounded off the figures to 70 cases and 140 controls. Two hundred healthy subjects were also included in the study as controls. Three consecutive stool samples and blood in EDTA vacutainer vials were collected from each patient and control subject. Stool samples were subjected to microscopy and bacterial culture as soon as possible. Aliquots of the stool and blood samples were stored at −80˚C for DNA extraction. Details of each subject were noted in a pre-designed proforma and informed consent was also obtained from them. The study protocol was approved by the Institutional ethical committee (Reference numbers: PGI/EP/EC/37/20/3/2007, PGI/PhD/IEC/57/21.10.2011).
2.2. Detection of Cryptosporidium Infection
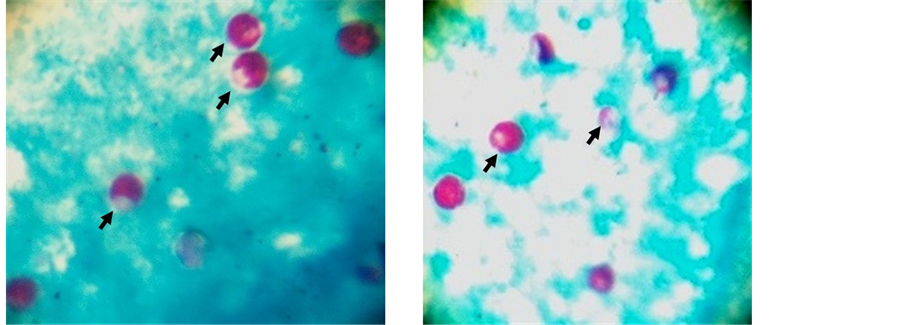
Microscopy for detection of Cryptosporidium spp.: Stool samples were subjected to normal saline and iodine wet mount for detection of oocysts and trophozoites of parasites and to exclude other parasitic infections. Subsequently, samples were concentrated by formol-ether sedimentation technique and subjected to modified Kinyoun’s staining, as per standard laboratory protocol [14] . Briefly, smears were methanol fixed and stained for 20 minutes in modified Kinyoun’s stain, decolorized with 10% acid alcohol for two minutes and counterstained with malachite green stain for one and half to two minutes. Stains were air dried and observed under light microscopy. Partially stained pink oocysts of Cryptosporidium spp. were visible against green background (Figure 1).
Molecular analysis for detection of Cryptosporidium spp.: DNA extraction from stool: DNA was extracted from the fecal samples of all patients and controls by QIAamp DNA mini kit (Qiagen Inc., Valencia, CA, USA), according to the manufacturer’s protocol. Some modifications were made, such as, 200 mg of stool suspension was initially washed in 2% polyvinyl polypyrolidone (PVPP) and dissolved in 1× phosphate buffered saline (PBS) and, later, the heat incubation with stool lysis buffer
Figure 1. Oocysts of Cryptosporidium (partially pink) stained with modified Kinyoun’s staining method, counterstained with malachite green.
(Buffer ASL) was done at a higher temperature i.e. at 80˚C for 10 minutes, with occasional stirring in between. Further, manufacturer’s recommendations were followed for purification and elution of DNA.
Polymerase chain reaction (PCR) for detection of Cryptosporidium spp.: Polymerase chain reaction (PCR) of the small subunit rRNA (SSU rRNA) gene was performed for detection of Cryptosporidium spp. Primers SCL1 and CPB-DIAGR were used to amplify a 1016 bp region of the SSU rRNA gene according to the PCR conditions mentioned by Coupe et al. [15] . Amplicon of the first round of PCR was subjected to a second round of nested PCR, with SCL2 and SCR2 primers, in a total reaction volume of 50 µl, amplifying a 214-bp fragment. PCR conditions were same as mentioned previously (Coupe et al., 2005) except that the annealing temperature was 50˚C (Figure 2).
Genotyping of the TLR4/2 gene: for the TLR4/2 gene: DNA extraction from blood, PCR and RFLP: DNA was extracted from blood using QIAamp Blood DNA mini kit (Qiagen Inc, Valencia, CA, USA), according to the manufacturer’s protocol. Thereafter, the DNA was subjected to polymerase chain reaction-restriction fragment length polymorphism (PCR-RFLP) for TLR 4 Thr_399_Ile and TLR 4 Asp_299_Gly and TLR2 196 to 174 del polymorphisms, using previously described primers and conditions [12] [16] . Hinf1 and Nco1 restriction enzymes were used for TLR 4 Thr_399_Ile and TLR 4 Asp_299_Gly respectively. PCR and RFLP products were run on 3.5% - 4% agarose gel and checked for band patterns. In case of TLR2 196_174 del polymorphism, band of 286 bp size denoted wild type, 264 bp homozygous and both together heterozygous. Similarly in case of TLR 4 Thr_399_Ile and Asp_299_Gly polymorphisms, bands of 406 bp and 249 bp sizes denoted wild type, 377 bp and 233 bp were homozygous, and both together in each case denoted heterozygous, for each polymorphism.
2.3. Statistical Analysis
Genotype frequency among control population were determined and tested for consistency according to the Hardy-Weinberg equilibrium. Chi-squared and Fisher’s exact
Figure 2. First and second rounds of nested PCR results: (a) 1st PCR (1016 bp) of small subunit (SSU) rRNA gene of Cryptosporidium spp. (M = Molecular marker, 1-3―Cryptosporidium positive samples, 4 = Negative control, 5 = Positive control); (b) 2nd nested PCR (214 bp) of SSU rRNA gene of Cryptosporidium spp. (M = Molecular marker, 1-4―Cryptosporidium positive samples, 5 = Positive control).
tests, as applicable, were used to compare the differences in genotype distribution between cases and controls. Binary logistic regression was used to estimate odds ratio (OR) and 95% confidence interval (CI) to estimate the risk of cryptosporidiosis in relation to TLR polymorphisms. Probability values of less than 0.05 (p < 0.05) were considered statistically significant in all analyses. All the statistical analyses were done by using Statistical Package for the Social Sciences-(SPSS 15, Inc., Chicago, IL, USA). Haplotype analysis was done using Arlequin software (Version 3.5, The University of Berne, Swiss Institute of Bioinformatics, Switzerland).
3. Results
3.1. Study Population
Seventy immunocompromised patients were found positive for Cryptosporidium spp. by both microscopy and PCR-RFLP (Figure 1 and Figure 2). Out of the 70 Cryptosporidium positive patients, 28/70 were HIV positive and 42/70 were renal transplant recipients. To simplify analysis, the subjects were divided into three groups: the Cryptosporidium positive patients under group I, 140 immunocompromised patients (80 renal transplant recipients and 60 HIV positive patients) without cryptosporidiosis were classified as disease controls (denoted as group II), while healthy controls constituted Group III. Demographic parameters of patients and controls and distribution of their TLR polymorphisms are presented in Table 1.
3.2. Allelic Frequencies and Genotype Distributions of TLR Variants
TLR2 and TLR4 gene polymorphisms were analyzed in all the three groups. The genotype and allele frequencies of the gene polymorphisms among cases and controls are shown in Table 2 and Table 3.
TLR2 (−196 to −174 del/ins) gene polymorphism:
There was no association between TLR2 196_174del homozygosity or heterozygosity (OR = 0.548, 95% CI (0.26 - 1.156), p = 0.114; OR = 2.576, 95% CI (0.159 - 41.8), p = 0.506) and occurrence of cryptosporidiosis when Cryptosporidium positive cases were compared with healthy controls. Similar results were observed when Cryptosporidium positive cases and immunocompromised patients without cryptosporidiosis, were compared (OR = 0.76, 95% CI (0.396 - 1.451), p = 0.403).
Table 1. Demography and distribution of TLR genotypes in different groups of subjects.
*HWE (p): Hardy Weinberg equilibrium (p value); SD: Standard deviation; n: Frequency; no.: number; ins: insertion; del: deletion.
Table 2. Association between TLR gene polymorphisms and cryptosporidiosis (Group I vs Group III).
CI: confidence interval; OR: Odds ratio, ratio of the lower to upper 95% confidence limit; n: number; ins: insertion; del: deletion.
TLR4_Asp_299_Gly gene polymorphism:
Logistic regression analysis revealed no association between TLR4_299 heterozygote and homozygote states and occurrence of cryptosporidiosis, when cases were compared with healthy controls and with immunocompromised patients without cryptosporidiosis (OR = 1.239 95% CI = (0.457 - 3.36), p = 0.674, and OR = 1.145, 95% CI = (0.45 - 2.94), p = 0.777, respectively).
TLR4_Thr_399_Ile gene polymorphism:
TLR4_399 gene homozygote and heterozygote states were not associated with occurrence of cryptosporidiosis when cases were compared with healthy controls and with immunocompromised patients without cryptosporidiosis (OR = 1.235, 95% CI = 0.31 - 4.91, p = 0.765, and OR = 1.31, 95% = 0.38 - 4.52, p = 0.669, respectively).
Table 3. Association between TLR gene polymorphisms and cryptosporidiosis (Group I vs Group II).
CI: confidence interval; OR: Odds ratio, ratio of the lower to upper 95% confidence limit; n: number; ins: insertion; del: deletion.
4. Discussion
In the present study, we investigated the association between TLR 2 (196_174 del) and TLR 4 Asp299Gly and Thr399Ile) gene polymorphisms and pathogenesis of cryptosporidiosis among immunocompromised patients in northern India. No significant association was observed between these three polymorphisms and cryptosporidiosis.
Previous studies on Cryptosporidium spp. have reported that TLR/Myd88 pathway help in defense against C. parvum infection of cholangiocyte cells [2] [14] . C. parvum infection of human cholangiocytes in culture induced recruitment of TLR2 and TLR4, to the infection sites, but other TLRs were not expressed. Also, C. parvum attachment to and invasion of cholangiocytes was found to be independent of TLRs, as mutation of TLR2 and TLR4 did not affect parasite attachment and invasion. However, TLR2 and TLR4 signals activated several downstream effectors like IRAK, enabled phosphorylation of p-38 and thus, activation of NF-kB signaling pathway triggering expression of human-defensin-2, cytokines/chemokines against C. parvum. The human immunodeficiency virus type 1 (HIV-1) Tat protein, inhibited expression of TLR4 protein on the surface of cholangiocytes through translational inhibition, which thus prevented the initiation of an innate immune response against C. parvum [6] .
Our study results did not show association between TLR 4 Asp299Gly and Thr399Ile), TLR 2 (196_174 del) gene polymorphisms and cryptosporidiosis. This is in concordance with previous studies on other parasitic infections. In a previous study, frequencies of TLR4 Asp299Gly and Thr399Ile polymorphism did not show any significant differences in the distribution of alleles or genotypes between patients with chagasic cardiomyopathy and asymptomatic subjects [17] . In another study, TLR2 196-174 del heterozygosity was associated with protection from cerebral malaria (OR 0.34, 95% confidence intervals 0.16, 0.73) [18] . A study reported comparison of distribution of TLR4 Asp299Gly and Thr399Ile polymorphism in patients with visceral leishmaniasis and ethnically matched controls; it did not show any difference both in case of alleles and genotypes (P > 0.05) [19] . However, some infections have been associated with TLR polymorphisms. The TLR2 haplotype 196-174 del was found to be strongly associated with an increased risk of asymptomatic bancroftian filariasis [9] . In a study on neurocysticircosis (NCC), TLR4 Asp299Gly and Thr399Ile polymorphisms were associated with its occurrence (P < 0.001 for Asp299Gly; P = 0.003 for Thr399Ile) and progression to symptomatic NCC, compared with control subjects or asymptomatic NCC [20] . In another study, the frequency of the TLR2 196 - 174 del allele was higher in patients with HCV-associated hepatocellular carcinoma (22.5%) than in HCV-infected patients without hepatocellular carcinoma (15.6%, P = 0.016) and healthy controls (15.3%, P = 0.003) [21] . Thus, association between genetic polymorphisms of innate immunity components and outcome might depend on the type of infection or pathogen involved.
Oocysts of Cryptosporidium undergo excystation in the small intestine and the sporozoites multiply and disrupt the intestinal epithelium causing villous atrophy, crypt hyperplasia leading to malabsorption. The principle defense mechanism against protozoan parasites is innate immune response. Toll like receptors expressed on host cell surfaces recognize specific molecular motifs present on pathogen surfaces, which activate various intracellular signaling molecules, such as NF-κB, leading to the production of cytokines/chemokines involved in immune response. Polymorphisms in the genes of these TLRs result in abnormal host-pathogen interaction due to variation in TLR proteins and thus, abnormal immune response. Hence, TLR polymorphisms can lead to the occurrence of diseases. However, our study did not reveal association between TLR2 196-174 del, TLR4 Asp299Gly and Thr399Ile polymorphisms and cryptosporidiosis. There can be two possible explanations for these. There can be two possible explanations for our results. Firstly, the very low allele frequency of these polymorphisms in the studied population. According to previous reports there is absence of TLR4 Asp299Gly homozygotes in Indian population and very low frequency of Thr399Ile homozygotes (0.5%) [22] . Similar result was observed in northern Indian population where the prevalence of each polymorphism was found to be 0.7% [10] [20] [23] .
The second reason can be the low sample size, i.e. lower number of Cryptosporidium positive cases included in the study and thus, restricted statistical power to detect minor differences.
5. Conclusion
In conclusion, the present study explored the role of TLR 2_196-174 del, TLR4 Asp299Gly and Thr399Ile polymorphisms in Cryptosporidium infection. No significant differences in the genotype or allele distributions between case and control groups were observed. This study also confirmed the low frequency of the three polymorphisms in our population, as observed in other studies. Although, no significant association was found between these polymorphisms and occurrence of cryptosporidiosis, this may not infer a lack of involvement of the same. Studies involving very large population from different independent geographically located/ethnic groups may provide a clear and more comprehensive picture about the involvement of these polymorphisms with the infection. These may serve as risk factors in estimating the occurrence of cryptosporidiosis in immunocompromised patients, thus aiding in its timely control or proper management.
Acknowledgements
Ujjala Ghoshal acknowledges The Department of Biotechnology (DBT), New Delhi, India for providing financial assistance for conducting this study (Reference No. DBT SAN No.:102/IFD/SAN 1839/2008-09). Asmita Dey is also thankful to The University Grants Commission (UGC), New Delhi for granting her Ph.D. fellowship (F. No. 2-89/98 (SA-I).
Declaration of Interests Statement
The authors report no declarations of interest.
Cite this paper
Dey, A., Ghoshal, U., Agarwal, V. and Ghoshal, U.C. (2016) Association between Toll-Like Receptor 4 (Thr399Ile and Asp299Gly), Toll-Like Receptor 2 196-174 Deletion Polymorphisms with cryptosporidiosis. Journal of Biosciences and Medicines, 4, 85-96. http://dx.doi.org/10.4236/jbm.2016.410009
References
- 1. Bandin, F., Kwon, T., Linas, M.D., Guigonis, V., Valentin, A., Cassaing, S., Carol, A., Garnier, A., Baudouin, V. and Decramer, S. (2009) Cryptosporidiosis in Paediatric Renal Transplantation. Pediatric Nephrology, 24, 2245-2255.
http://dx.doi.org/10.1007/s00467-009-1274-y - 2. Botero, J.H., Castano, A., Montoya, M.N., Ocampo, N.E., Hurtado, M.I. and Lopera, M.M. (2003) A Preliminary Study of the Prevalence of Intestinal Parasites in Immunocompromised Patients with and without Gastrointestinal Manifestations. Revista do Instituto de Medicina Tropical de Sao Paulo, 45, 197-200.
http://dx.doi.org/10.1590/S0036-46652003000400004 - 3. Nahrevanian, H. and Assmar, M. (2008) Cryptosporidiosis in Immunocompromised Patients in the Islamic Republic of Iran. Journal of Microbiology, Immunology and Infection, 41, 74-77.
- 4. Ok, U.Z., Cirit, M., Uner, A., Ok, E., Akcicek, F., Basci, A. and Ozcel, M.A. (1997) Cryptosporidiosis and Blastocystosis in Renal Transplant Recipients. Nephron, 75, 171-174.
http://dx.doi.org/10.1159/000189527 - 5. Udgiri, N., Minz, M., Kashyap, R., Heer, M., Gupta, C.S., Mohandas, K., Minz, R.W. and Malla, N. (2004) Intestinal Cryptosporidiasis in Living Related Renal Transplant Recipients. Transplantation Proceedings, 36, 2128-2129.
http://dx.doi.org/10.1016/j.transproceed.2004.08.107 - 6. Current, W.L. and Garcia, L.S. (1991) Cryptosporidiosis. Clinical Microbiology Reviews, 4, 325-358.
http://dx.doi.org/10.1128/CMR.4.3.325 - 7. Arora, D.R. and Arora, B. (2009) AIDS-Associated Parasitic Diarrhoea. Indian Journal of Medical Microbiology, 27, 185-190.
http://dx.doi.org/10.4103/0255-0857.53199 - 8. Hunter, P.R. and Nichols, G. (2002) Epidemiology and Clinical Features of Cryptosporidium Infection in Immunocompromised Patients. Clinical Microbiology Reviews, 15, 145-154.
http://dx.doi.org/10.1128/CMR.15.1.145-154.2002 - 9. Junpee, A., Tencomnao, T., Sanprasert, V. and Nuchprayoon, S. (2010) Association between Toll-Like Receptor 2 (TLR2) Polymorphisms and Asymptomatic Bancroftian Filariasis. Parasitology Research, 107, 807-816.
http://dx.doi.org/10.1128/CMR.15.1.145-154.2002 - 10. Pandey, S., Mittal, R.D., Srivastava, M., Srivastava, K., Singh, S., Srivastava, S. and Mittal, B. (2009) Impact of Toll-Like Receptors [TLR] 2 (-196 to -174 del) and TLR 4 (Asp299Gly, Thr399Ile) in Cervical Cancer Susceptibility in North Indian Women. Gynecologic Oncology, 114, 501-505.
http://dx.doi.org/10.1016/j.ygyno.2009.05.032 - 11. Qian, F.H., Zhang, Q., Zhou, L.F., Jin, G.F., Bai, J.L. and Yin, K.S. (2010) Polymorphisms in the Toll-Like Receptor 2 Subfamily and Risk of Asthma: A Case-Control Analysis in a Chinese Population. Journal of Investigational Allergology and Clinical Immunology, 20, 340-346.
- 12. Chen, X.M., O'Hara, S.P., Nelson, J.B., Splinter, P.L., Small, A.J., Tietz, P.S., Limper, A.H. and LaRusso, N.F. (2005) Multiple TLRs Are Expressed in Human Cholangiocytes and Mediate Host Epithelial Defense Responses to Cryptosporidium parvum via Activation of NF-kappaB. Journal of Immunology, 175, 7447-7456.
http://dx.doi.org/10.4049/jimmunol.175.11.7447 - 13. O'Hara, S.P., Small, A.J., Gajdos, G.B., Badley, A.D., Chen, X.M. and Larusso, N.F. (2009) HIV-1 Tat Protein Suppresses Cholangiocyte Toll-Like Receptor 4 Expression and Defense against Cryptosporidium parvum. The Journal of Infectious Diseases, 199, 1195-1204.
http://dx.doi.org/10.1086/597387 - 14. Garcia, L.S. (2001) Diagnostic Medical Parasitology. American Society for Microbiology, USA.
- 15. Coupe, S., Sarfati, C., Hamane, S. and Derouin, F. (2005) Detection of Cryptosporidium and Identification to the Species Level by Nested PCR and Restriction Fragment Length Polymorphism. Journal of Clinical Microbiology, 43, 1017-1023.
http://dx.doi.org/10.1128/JCM.43.3.1017-1023.2005 - 16. Lorenz, E., Frees, K.L. and Schwartz, D.A. (2001) Determination of the TLR4 Genotype Using Allele-Specific PCR. Biotechniques, 31, 22-24.
- 17. Zafra, G., Florez, O., Morillo, C.A., Echeverria, L.E., Martin, J. and Gonzalez, C.I. (2008) Polymorphisms of Toll-Like Receptor 2 and 4 Genes in Chagas Disease. Memórias do Instituto Oswaldo Cruz, 103, 27-30.
http://dx.doi.org/10.1590/S0074-02762008000100004 - 18. Greene, J.A., Sam-Agudu, N., John, C.C., Opoka, R.O., Zimmerman, P.A. and Kazura, J.W. (2012) Toll-Like Receptor Polymorphisms and Cerebral Malaria: TLR2 Delta22 Polymorphism Is Associated with Protection from Cerebral Malaria in a Case Control Study. Malaria Journal, 11, 47.
- 19. Rasouli, M., Keshavarz, M., Kalani, M., Moravej, A., Kiany, S. and Badiee, P. (2012) Toll-Like Receptor 4 (TLR4) Polymorphisms in Iranian Patients with Visceral Leishmaniasis. Molecular Biology Reports, 39, 10795-10802.
http://dx.doi.org/10.1007/s11033-012-1973-5 - 20. Verma, A., Prasad, K.N., Gupta, R.K., Singh, A.K., Nyati, K.K., Rizwan, A., Pandey, C.M. and Paliwal, V.K. (2010) Toll-Like Receptor 4 Polymorphism and Its Association with Symptomatic Neurocysticercosis. Journal of Infectious Diseases, 202, 1219-1225.
http://dx.doi.org/10.1086/656395 - 21. Nischalke, H.D., Coenen, M., Berger, C., Aldenhoff, K., Muller, T., Berg, T., Kramer, B., Korner, C., Odenthal, M., Schulze, F., et al. (2012) The Toll-Like Receptor 2 (TLR2)-196 to -174 Del/Ins Polymorphism Affects Viral Loads and Susceptibility to Hepatocellular Carcinoma in Chronic Hepatitis C. International Journal of Cancer, 130, 1470-1475.
http://dx.doi.org/10.1002/ijc.26143 - 22. Achyut, B.R., Ghoshal, U.C., Moorchung, N. and Mittal, B. (2007) Association of Toll-Like Receptor-4 (Asp299Gly and Thr399Ileu) Gene Polymorphisms with Gastritis and Precancerous Lesions. Human Immunology, 68, 901-907.
http://dx.doi.org/10.1016/j.humimm.2007.10.006 - 23. Tahara, T., Arisawa, T., Wang, F., Shibata, T., Nakamura, M., Sakata, M., Hirata, I. and Nakano, H. (2007) Toll-Like Receptor 2-196 to 174del Polymorphism Influences the Susceptibility of Japanese People to Gastric Cancer. Cancer Science, 98, 1790-1794.
http://dx.doi.org/10.1111/j.1349-7006.2007.00590.x