 American Journal of Plant Sciences, 2011, 2, 619-628 doi:10.4236/ajps.2011.24073 Published Online October 2011 (http://www.SciRP.org/journal/ajps) Copyright © 2011 SciRes. AJPS 619 Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis in Artemisia annua L., as Reported by a Promoter-GUS Fusion Hongzhen Wang, Linda Olofsson, Anneli Lundgren, Peter E. Brodelius School of Natural Sciences, Linnaeus University, Kalmar, Sweden. Email: peter.brodelius@lnu.se Received August 16th, 2011; revised September 7th, 2011; accepted September 20th, 2011. ABSTRACT Artemisia annua L. produces small amounts of the sesquiterpenoid artemisinin, which is used for treatment of malaria. A worldwide shortage of the drug has led to intense research to increase the yield of artemisinin in the plant. In o rder to study the regulation of expression of a key enzyme of artemisinin biosynthesis, the promoter region of the key enzyme amorpha-4,11-diene synthase (ADS) was cloned and fused with the β-glucuronidase (GUS) reporter gene. Transgenic plants of A. annua expressing this fusion were g enerated and studied . Transgenic plants expressing the GUS gene were used to establish the activity of the cloned promoter by a GUS activity staining procedure. GUS under the control of the ADS promoter showed specific expression in glandular trichomes. The activity of the ADS promoter varies temporally and in old tissues essentially no GUS staining could be observed. The expression pattern of GUS and ADS in aerial parts of the transgenic plant was essentially the same indicating that the cis-elements controlling glandular trichome specific expression are included in the cloned promoter. However, some cis-element(s) that control expression in root and old leaf appears to be missing in the cloned promoter. Furthermore, qPCR was used to compare the ac tivity of the wild-type ADS promoter with that of the cloned ADS promoter. The latter promoter showed a considerably lower activ- ity than the wild-type promoter as judged from the levels of GUS and ADS transcripts, respectively, which may be due to the removal of an enha ncing cis-element from the ADS promoter. The ADS gene is specifically expressed in stalk and secretory cells of glandular trichomes of A. annua. Keywords: Agrobacterium Tumefaciens, Amorpha-4,11-Diene Synthase, Artemisia annua, Artemisi nin Biosynthesi s, β-Glucuronidase, Gene Regulation, Promoter Activity, Stable Transformation 1. Introduction Artemisinin is an effective anti-malarial drug, which has become an important component of artemisinin-based combination therapies (ACTs) [1]. The content of ar- temisinin in Artemisia annua is, however, very low and ranges between 0.1% and 0.8% of dry weight [2]. Selec- tion of high-producing varieties of A. annua has met lim- ited success but hybrids producing increased amounts of artemisinin have been produced [3]. However, there is still a shortage in the supply o f artemisinin [4]. Different methods have been tried to improve the artemisinin pro- duction such as treatment with abscisic acid [5], gibbere- lic acid [6-8] and elicitors [9-11]. Various transgenic A. annua plants have been produced to increase the yield of artemisinin. These attempts include down-regulation of squalene synthase [12,13] or up-regulation of farnesyl diphosphate synthase (FDS) [14]. Cyclization of farnesyl diphosphate (FDP) to amor- pha-4,11-diene by amorpha-4,11-diene synthase (ADS) is the initial step of the artemisinin biosynthetic pathway [15] and amorpha-4,11-diene is the committed precursor [16]. Tissue specificity of ADS expression has been shown by GUS expression in Arabidopsis thaliana using a fusion of the ADS promoter and the reporter gene [17]. In the following step of the artemisinin biosynthesis, amorpha-4,11-diene is hydroxylated to yield artemisinic alcohol. This reaction is catalyzed by a cytochrome P450 dependent amorpha-4,11-diene 12-hydroxylase (CYP71AV1)  Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis 620 in Artemisia annua L., as Reported by a Promoter-GUS Fusion [18]. This enzyme is multifunctional and can also oxidize the alcohol to artemisinic aldehyde and then further on to artemisinic acid [19]. However, artemisinic acid is a di- rect precursor of arteannuin B biosynthesis. For artemisi- nin biosynthesis the artemisinic aldehyde must be re- duced to dihydroartemisinic aldehyde, which subse- quently is oxidized to dihydroartemisinic acid and then converted to artemisinin [20]. These two steps are cata- lyzed by artemisinic aldehyde 11 (13) reductase [21] and aldehyde dehydrogenase 1 [22], respectively. Glan- dular trichome specific expression of ADS and CYP71- AV1 [17,18] strongly supported the notion that artemisi- nin is sequestered and localized to glandular trich omes of A. annua [23,24]. In order to understand the mechanisms behind the in- crease in artemisinin biosynthesis after treatment with hormones [5-8] or biotic elicitors [9-11], a detailed knowledge about th e regulation of expr ession of differ ent enzymes involved in artemisinin biosynthesis will be essential. To our knowledge, there is no data available on the regulation of expression of biosynthetic genes in A. annua using promoter-reporter gene fusions in transgenic plants of A. annua. We have initiated studies on the regulation of terpene metabolism in different tissues of A. annua [25] and here we report on results from transgenic A. annua expressing a fusion of the reporter gene (GUS) and the promoter of the key enzyme ADS. 2. Materials and Methods 2.1. Plant Materials Plants of A. annua were grown under 16 h days and 8 h nights at 22˚C to a height of approximately 1 m followed by induction of flowering at 8 h days and 16 h nights at 22˚C. Flower buds, young leaves, old leaves, stems and roots were collected for GUS staining. Samples of these tissues were also frozen in liquid nitrogen, ground and used for RNA extraction. 2.2. Promoter Cloning Genomic DNA was extracted from fresh young leaves of A. annua using the CTAB method [26]. The promoter sequence of ADS was available in the GenBank (acces- sion number AY528931). The sequence was used to de- sign primers for PCR amplification (primers 1 and 2 in Table 1) of a 1929 bp fragment of the ADS promoter using Pfu polymerase (Fermentas). Primers 1 and 2 car- ried Xm a I and NcoI restriction site, respectiv ely, used for cloning the fragment into the plant transformation vector. 2.3. Construction of Transformation Vector The pCAMBIA 1381Z vector (CambiaLabs, Brisbane, Table 1. Nucleotide sequence of primers used. Restriction sites are underlined; f = forward; r = reverse. PrimerNamePrimer Sequence 1 pADSfTCCCCCGGGAAAGTTAATTGCACATAT 2 pADSrCATGCCATGGATTTTCAAAACTTTGAATAT 3 GUSf AACCGTTCTACTTTACTGGCTTTGG 4 GUSr GCATCTCTTCAGCGTAAGGGTAAT 5 ADSf GGGAGATCAGTTTCTCATCTATGAA 6 ADSr CTTTTAGTAGTTGCCGCACTTCTT 7 β-actinfCCAGGCTGTTCAGTCTCTGTAT 8 β-actinrCGCTCGGTAAGGATCTTCATCA Australia) carrying the GUS-gene was used for the trans- formation of A. annua. However, the vector was modi- fied in that the plant resistance gene was changed from hygromycin to kanamycin. An XhoI fragment (0.88 kb) carrying the NPTII gene was cut out from pCAMBIA 2301 and ligated into pCAMBIA 1381Z digested with XhoI. Restriction analysis was used to determine that the XhoI fragment had been ligated in the right direction. The ADS promoter was double-digested with NcoI/ XmaI and inserted into the modified pCAMBIA 1381Z vector digested with the same restriction enzymes. The plant transformation vector obtained (pCAMBIA1381Z- pADS-GUS) was introduced into E. coli BL21, which was grown on LB medium containing kanamycin (50 mg/L). The vector was purified using the GeneJet Plas- mid Miniprep Kit (Fermentas) and used for transforma- tion of Agrobacetrium tumefaciens EHA105. The plant transformation vector was introduced into A. tumefaciens EHA105 by the freeze and thaw method. A. tumefaciens EHA105 carrying the plant transformation vector was grown on YEP medium containing kanamy- cin (100 mg/L), rifampicin (40 mg/L) and streptomycin (25 mg/L) at 28˚C to an OD600 = 0.8 – 1. The cells were collected by centrifugation and resuspended in MSMO liquid medium to an OD600 = 0.3 – 0.5 . 2.4. Plant Transformation A. annua seeds (variety Chongqin) were immersed in 75% ethanol for 1 min, surfaced sterilized with 5% so- dium hypochlorite (NaOCl) for 20 min, washed 3 - 4 times with sterile distilled water. The sterilized seeds were germinated on solid MSMO medium (Sigma) (MSMO powder 4.4 g/L, sucrose 30 g/L, agar 8 - 9 g/L, pH = 5.8) at 28˚C for 20 - 25 days. The leaflets of 20 - 25 days old seedlings were used as explants for transforma- tion. The explants were immersed into a suspension of A. tumefaciens EHA105 carrying the pCAMBIA1381Z- pADS-GUS transformation vector. After 30 min the ex- plants were transferred to co-cultivation medium (MSMO Copyright © 2011 SciRes. AJPS  Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis 621 in Artemisia annua L., as Reported by a Promoter-GUS Fusion medium containing 6-BA (0.5 mg/L), NAA (0.05 mg/L)) and the A. tumefaciens EHA105. Co-cultivation was car- ried out in darkness at 28˚C for 48 h. Subsequently, the explants were transferred to selection and germination medium (MSMO medium containing 6-BA (0.5 mg/L), NAA (0.05 mg/L), kanamycin (100 mg/L) and carbeni- cillin (250 mg/L)). The explants were transferred to new medium every second week. Shoots started to grow out after around two weeks. The shoots were transferred to root inducing medium (half strength MSMO medium containing kanamycin (100 mg/L) and carbenicillin (100 mg/L)). After rooting the plantlets were left 4 weeks be- fore they were transferred to soil. 2.5. Scanning Electron Microscopy Scanning electron microscopy (SEM) was carried out on unfixed tissues from wild-type and transgenic A. annua plants using a relatively low energy beam (5 kV) on a LEO 435VP scanning electron microscope. 2.6. GUS Assay Leaf primordia at the apex and expanded leaves at dif- ferent nodes, stems, roots from shoots and flowering plants, and flower buds were sampled from transgenic A. annua plants to perform GUS analysis. GUS histo- chemical staining of the various plant tissues was carried out as previously described [27]. GUS stained tissues were studied under a microscope (Nikon ECLIPSE E400) and photographs were taken using a digital camera (Nikon DP11). 2.7. qPCR Different tissues of transgenic A. annua plants were sampled to analyze the relative expression pattern of pADS-GUS and pADS-ADS using β-actin as the reference gene. RNA was extracted using PurelinkTM Plant RNA Reagent (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instruction. Genomic DNA was re- moved by treatment with DNase I (Fermentas, St Leo- Roth, Germany). RNA (1 μg) was reverse transcribed using RevertAidTM H Minus-MuLV reverse transcriptase (Fermentas) primed with 0.5 μg oligo (dT)18 primer. RNA was removed from the cDNA obtained by treat- ment with RNas e H (Fermentas). The qPCR was performed on a 7500 qPCR thermocy- cler (Applied Biosystems, Foster City, USA) using prim- ers listed in Table 1 for GUS (primers 3 and 4), ADS (primers 5 and 6) or actin (primers 7 an d 8). First single- stranded cDNA was used as template in a 20 μl reaction mixture containing 10 μl Power SYBR® Green PCR Master Mix (Applied Biosystems) and 2 pmol of each primer. The qPCR thermal cycling was performed at 50˚C (2 min), 95˚C (10 min), 40 cycles at 95˚C (15 sec), 60˚C (1 min) and finally a dissociation state at 95˚C (15 sec), 60˚C (1 min) and 95˚C (15 sec). Triplet samples were run for each cDNA sample. 2.8. Statistical Methods Grubbs test (G = |Suspect value-xmean|/s) was used to test for outliers. The outliers were rejected if Gcalculated > Gcritical at P = 0.05. The critical value of G is 1.155 when the sample size is three [19]. 3. Results and Discussion 3.1. Cloning of Promoter of ADS Based on the sequence of the promoter region of ADS (GenBank accession number AY528931), a promoter region of 1929 bp upstream of the translation initial codon (ATG) as shown in Figure 1 was isolated from A. annua genomic DNA by PCR using high fidelity poly- merase and primers 1 and 2 as listed in Table 1. 3.2. Prediction of Transcription Start Site and Core Promoter Elements The transcription start site (TSS) of the cloned promoter was predicted using the TSSP software (http://linux1. softberry.com/berry.phtml). A putative TSS of ADS (la- beled +1 in Figure 1) was predicted 51 bp upstream of the translation initiation ATG-codon. The TATA-box is a cis-regulatory element found in the promoter region of many genes in eukaryotes with the core sequence or a variant, which is usually follo wed by three or more adenine bases. It is considered to be the core promoter sequence and the binding site of either general tran scription factors or histones (the binding of a transcription factor blocks the binding of a histone and vice versa). It is involved in the process of transcription by RNA polymerase and is usually located around 25 bp upstream of the TSS. A putative TATA-box was found at positions –29 to –24 (TATAAA) in the ADS promoter (Figure 1) upstream of the putative TSS. The CAAT-box signal is the binding site for the RNA transcription factor, and is typically accompanied by a conserved consensus sequence (GGCCAATCT). Genes that have this element seem to require it for the gene to be transcribed in sufficient quantities. This box along with the GC-box is known for binding general transcrip- tion factors. CAAT- and GC-boxes are primarily located in the region from 100 - 150 bp upstream from the TATA-box. Binding of specific protein is required for the CAAT-box activation. These proteins are known as CAAT-box binding proteins/CAAT-box binding factors. e could not find any CAAT-box with the consensus W Copyright © 2011 SciRes. AJPS 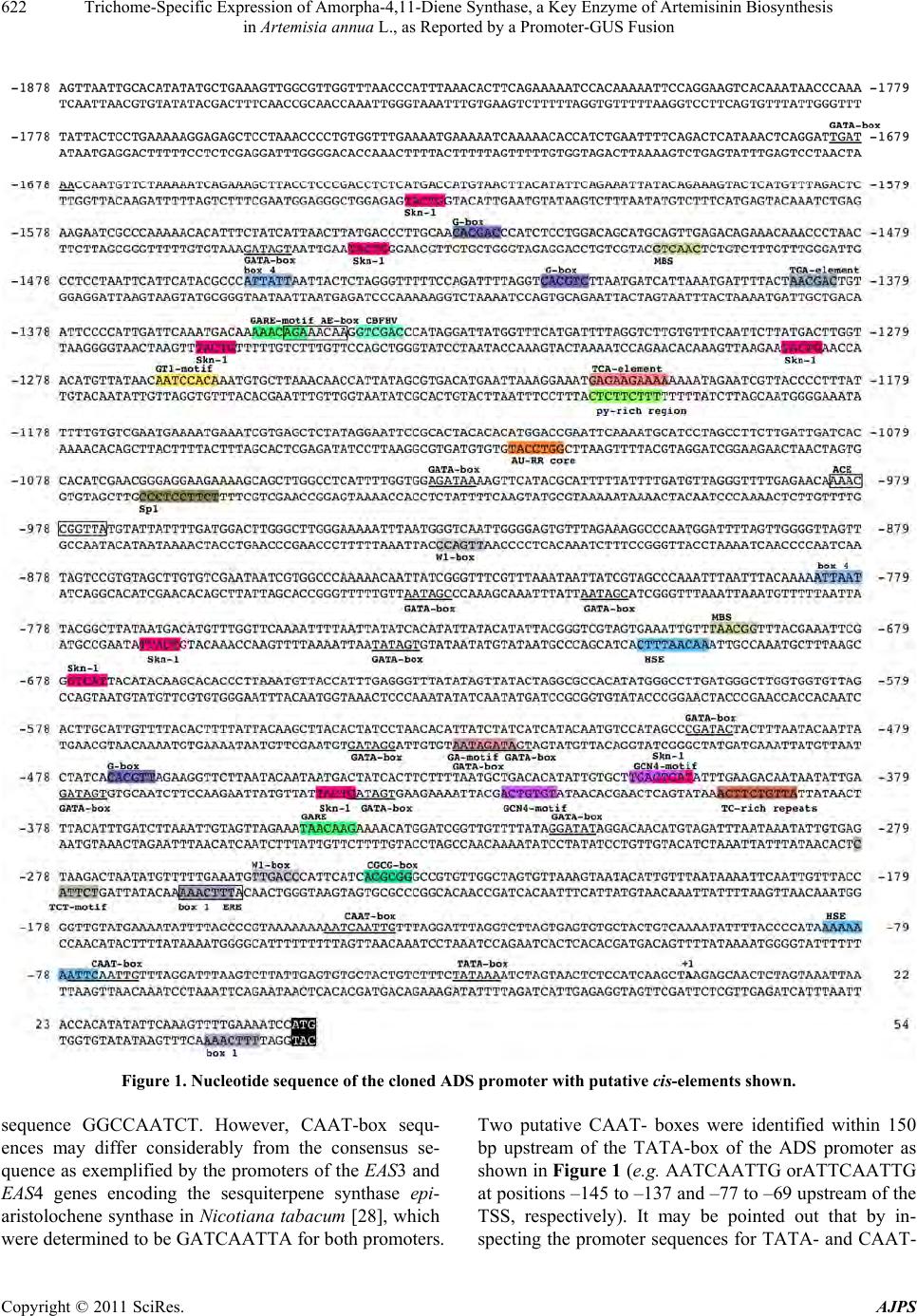 Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis in Artemisia annua L., as Reported by a Promoter-GUS Fusion Copyright © 2011 SciRes. AJPS 622 Figure 1. Nucleotide sequence of the cloned ADS promoter with putative cis-elements shown. sequence GGCCAATCT. However, CAAT-box sequ- ences may differ considerably from the consensus se- quence as exemplified by the promoters of the EAS3 and EAS4 genes encoding the sesquiterpene synthase epi- aristolochene synth ase in Nicotiana tabacum [28], which were determined to be GATCAATTA for both promoters. Two putative CAAT- boxes were identified within 150 bp upstream of the TATA-box of the ADS promoter as shown in Figure 1 (e.g. AATCAATTG orATTCAATTG at positions –145 to –137 and –77 to –69 upstream of the TSS, respectively). It may be pointed out that by in- specting the promoter sequences for TATA- and CAAT-  Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis 623 in Artemisia annua L., as Reported by a Promoter-GUS Fusion boxes a number of alternative putative TSSs may be found. 3.3. Prediction of Cis-Elements of Promoter Region of ADS Putative cis-elements of the promoter were predicted by using the PLANTCARE software (http://bioinformatics. psb.ugent.be/webtools/plantcare/html/). The py-rich sequence TTTCTTCTC is a cis-acting element conferring high transcription levels. Such an element was found at position –1204 to –1212 as shown in Figure 1. Yang and Poovaiah (2000) reported that the tobacco ethylene-responsive gene NtER1 encodes a calmodulin- binding protein [29] and demonstrated later that there is one NtER1 homolog as well as five related genes in Arabidopsis [30]. These six Arabidopsis genes are rap- idly and differentially induced by environmental signals such as extreme temperatures, UV-B, salt, wounding, hormones such as ethylene and abscisic acid (ABA), and signal molecules such as methyl jasmonate (MeJA), H2O2, and salicylic acid (SA). Hence, they were design- nated as AtSR1 to 6 (Arabidopsis thaliana signal-re- sponsive genes). AtSR1 targets the nucleus and specifi- cally recognizes a 6-bp CGCG-box (A/C/G)CGCG(G/T/ C). Multiple CGCG cis-elements are found in promoters of genes involved in ethylene, ABA and SA signaling, and light signal perception. A putative CGCG-box at positions –240 to –235 was found in the ADS promoter (Figure 1). The TCA-element with the seq u ence GAG AA GAATA is involved in responsiveness of genes to SA [31]. One such element with the sequence GAGAAGAAAA (–1212 to –1203) was foun d in the ADS promoter (Figure 1). A putative ethylene responsiveness ERE-element (ATTTC- AAA) was localized to –256 to –268. Furthermore, a putative gibberellin (GA)-responsive element (the GARE-motif AAACAGA) is present in the ADS pro- moter at position –1354 to –1348 and a modified GARE- motif (TAACAAG) at position –348 to –342. A TC-rich repeat, which is involved in defense and stress response, was localized to position –387 to –396 (ATTGTCTTCA). Finally, an AU-RR core sequence (GGTCCAT), which is involved in auxin responsiveness, was found at position –1116 to –1122. The elements described above may be involved in the response of A. annua to plant hormones (ABA, SA, eth- ylene, MeJA and GA). In fact, treatment of A. annua plants with ABA (10 μM) resulted in the induction of a number of genes encoding enzymes involved in artemis- inin biosynthesis [5]. The expression levels of 3-hy- droxy-3-methylglutaryl coenzyme A reductase (HMGR), farnesyl diphosphate synthase (FDS), CYP71AV1 and cytochrome P450 reductase (CPR) were significantly induced while ADS only showed a slight increase. The treatment with ABA resulted in a 65% increase in ar- temisinin content in A. annua plants [5]. Treatment of A. annua plants with 1 mM SA resulted in a gradual increase in the expression of the HMGR gene and a temporary peak in the expression of the ADS gene [32]. The expression of the FDS and CYP71AV1 genes showed little change. At 96 h after SA treatment, the concentration of artemisinin, artemisinic acid and dihydroartemisinic acid were 54%, 127% and 72% higher than th at of the control, respectively [32]. In a study on the production of artemisinin in cell sus- pension cultures of A. annua , acetyl-SA (20 mg/L), JA (5 mg/L) or GA (10 mg/L) were efficient elicitors and gave around 2-, 4- and 2.5-fold increased yield of artemisinin, respectively [10], indicating an increased expression of the key regulatory enzyme ADS. The beneficial effect of GA3 on artemisinin accumulation is also supported by the findings that treatment of A. annua plants with half- strength Hoagland’s solution containing 14 mM GA3 increased artemisinin content from 0.14% to 0.64% (w/w) when applied to 74-day-old plants [8]. No studies on the effects of GA on the expression level of enzymes in- volved in artemisinin biosynthesis have been reported. Two W1-boxes ((C/T)TGAC(C/T)), defined as elicit- tor-responsive elements, have been iden tified at positions –255 to –250 and –926 to –931 (Figure 1). The tran- scription factor WRKY binds to these elements trigger- ing the transcription of the gene. Transient expression of the A. annua transcription factor AaWRKY in agroinfil- trated leaves of A. annua resulted in increased levels of HMGR, ADS, CYP71AV1 and DBR2 indicating that several of the genes encoding enzymes involved in ar- temisinin biosynthesis are induced by binding of AaWRKY to the W-boxes [33]. In cotton, the GaWRKY1 is regulating the activity of cadinene synthase 1 (CAD1) involved in the biosynthesis of sesquiterpene phytoalex- ins [34]. The CAD1 promoter carries two W-boxes to which the GaWRKY 1 binds. A recen t study showed that foliar application of an elicitor (chitosan) resulted in in- crease of dihydroartemisinic acid and artemisinin by 72% and 53%, respectively, in A. annua [11]. Furthermore, semi-quantitative RT-PCR showed an increased level of ADS transcripts 2 h after application of chitosan. GATA-factors, involved in light-mediated regulation, are a class of transcriptional regulators present in plants that normally recognize the consensus sequence (T/A) GATA(G/A) [35]. Twelve putative GATA-boxes were found in the promoter r egion of ADS as shown in Figure 1. Other elements putatively involved in light-mediated Copyright © 2011 SciRes. AJPS 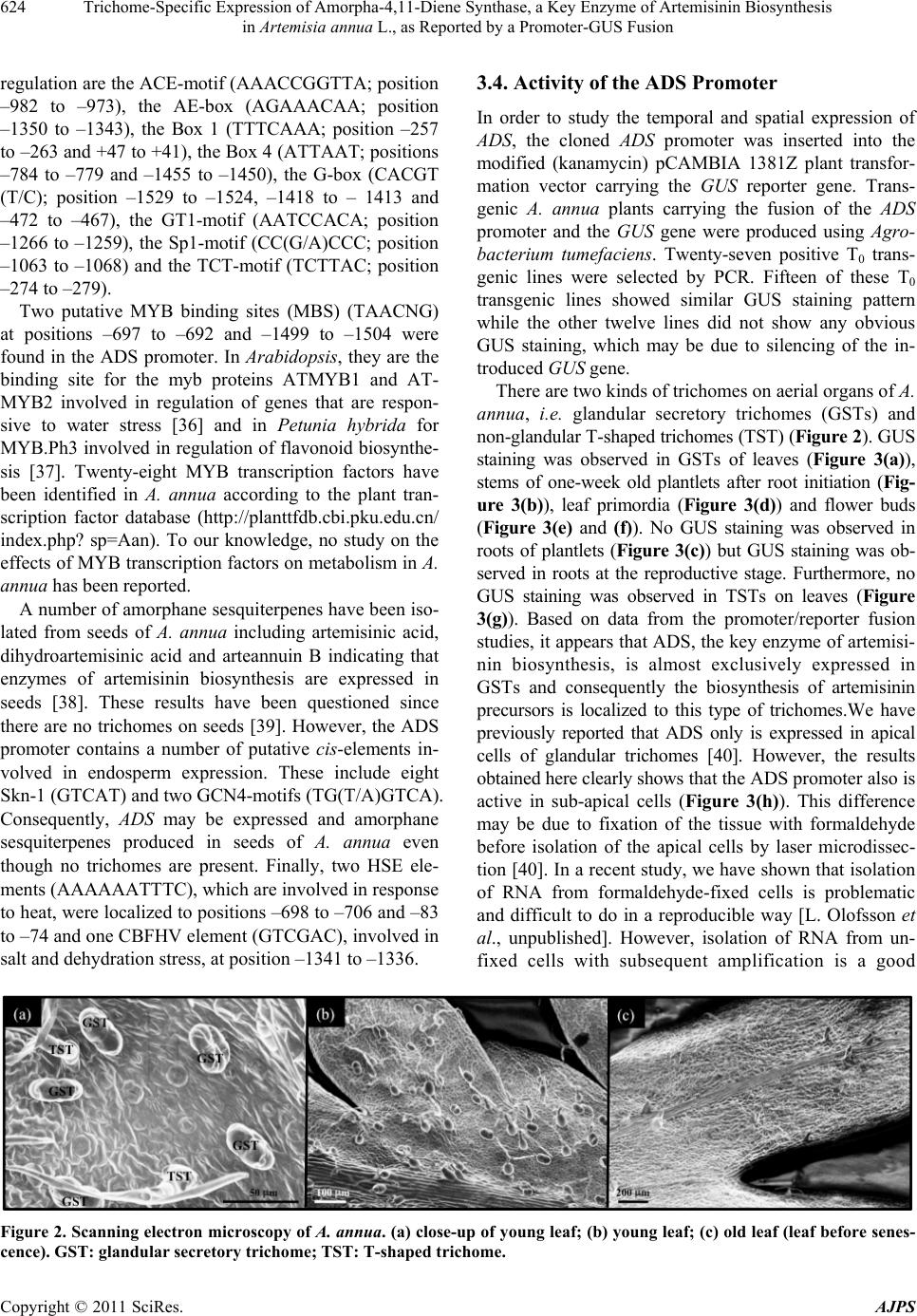 Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis in Artemisia annua L., as Reported by a Promoter-GUS Fusion Copyright © 2011 SciRes. AJPS 624 3.4. Activity of the ADS Promoter regulation are the ACE-motif (AAACCGGTTA; position –982 to –973), the AE-box (AGAAACAA; position –1350 to –1343), the Box 1 (TTTCAAA; position –257 to –263 and +47 to +41), the Box 4 (ATTAAT; positio ns –784 to –779 and –1455 to –1450), the G-box (CACGT (T/C); position –1529 to –1524, –1418 to – 1413 and –472 to –467), the GT1-motif (AATCCACA; position –1266 to –1259), the Sp1-motif (CC(G/A)CCC; position –1063 to –1068) and the TCT-motif (TCTTAC; position –274 to –279). In order to study the temporal and spatial expression of ADS, the cloned ADS promoter was inserted into the modified (kanamycin) pCAMBIA 1381Z plant transfor- mation vector carrying the GUS reporter gene. Trans- genic A. annua plants carrying the fusion of the ADS promoter and the GUS gene were produced using Agro- bacterium tumefaciens. Twenty-seven positive T0 trans- genic lines were selected by PCR. Fifteen of these T0 transgenic lines showed similar GUS staining pattern while the other twelve lines did not show any obvious GUS staining, which may be due to silencing of the in- troduced GUS gene . Two putative MYB binding sites (MBS) (TAACNG) at positions –697 to –692 and –1499 to –1504 were found in the ADS promoter. In Arabidopsis, they are the binding site for the myb proteins ATMYB1 and AT- MYB2 involved in regulation of genes that are respon- sive to water stress [36] and in Petunia hybrida for MYB.Ph3 involved in regulation of flavonoid biosynthe- sis [37]. Twenty-eight MYB transcription factors have been identified in A. annua according to the plant tran- scription factor database (http://planttfdb.cbi.pku.edu.cn/ index.php? sp=Aan). To our knowledge, no study on the effects of MYB transcription factors on metabolism in A. annua has been reported. There are two kinds of trichomes on aerial organs of A. annua, i.e. glandular secretory trichomes (GSTs) and non-glandular T-shaped trichomes (TST) (Figure 2). GUS staining was observed in GSTs of leaves (Figure 3(a)), stems of one-week old plantlets after root initiation (Fig- ure 3(b)), leaf primordia (Figure 3(d)) and flower buds (Figure 3(e) and (f)). No GUS staining was observed in roots of plantlets (Figure 3(c)) but GUS staining was ob- served in roots at the reproductive stage. Furthermore, no GUS staining was observed in TSTs on leaves (Figure 3(g)). Based on data from the promoter/reporter fusion studies, it appears that ADS, the key enzyme of artemisi- nin biosynthesis, is almost exclusively expressed in GSTs and consequently the biosynthesis of artemisinin precursors is localized to this type of trichomes.We have previously reported that ADS only is expressed in apical cells of glandular trichomes [40]. However, the results obtained here clearly shows that the ADS pr o mo t e r also i s active in sub-apical cells (Figure 3(h)). This difference may be due to fixation of the tissue with formaldehyde before isolation of the apical cells by laser microdissec- tion [40]. In a recent study, we have shown that isolation of RNA from formaldehyde-fixed cells is problematic and difficult to do in a reproducible way [L. Olofsson et al., unpublished]. However, isolation of RNA from un- fixed cells with subsequent amplification is a good A number of amorphane sesquiterpenes have been iso- lated from seeds of A. annua including artemisinic acid, dihydroartemisinic acid and arteannuin B indicating that enzymes of artemisinin biosynthesis are expressed in seeds [38]. These results have been questioned since there are no trichomes on seeds [39]. However, the ADS promoter contains a number of putative cis-elements in- volved in endosperm expression. These include eight Skn-1 (GTCAT) and two GCN4-motifs (TG(T/A)GTCA). Consequently, ADS may be expressed and amorphane sesquiterpenes produced in seeds of A. annua even though no trichomes are present. Finally, two HSE ele- ments (AAAAAATTTC), which are involved in response to heat, were localized to positions –698 to –706 and –83 to –74 and one CBFHV element (GTCGAC), involved in salt and dehydration stress, at pos ition –1341 to –1336. Figure 2. Scanning electron microscopy of A. annua. (a) close-up of young leaf; (b) young leaf; (c) old leaf (leaf before senes- cence). GST: glandular secretory trichome; TST: T-shaped trichome. 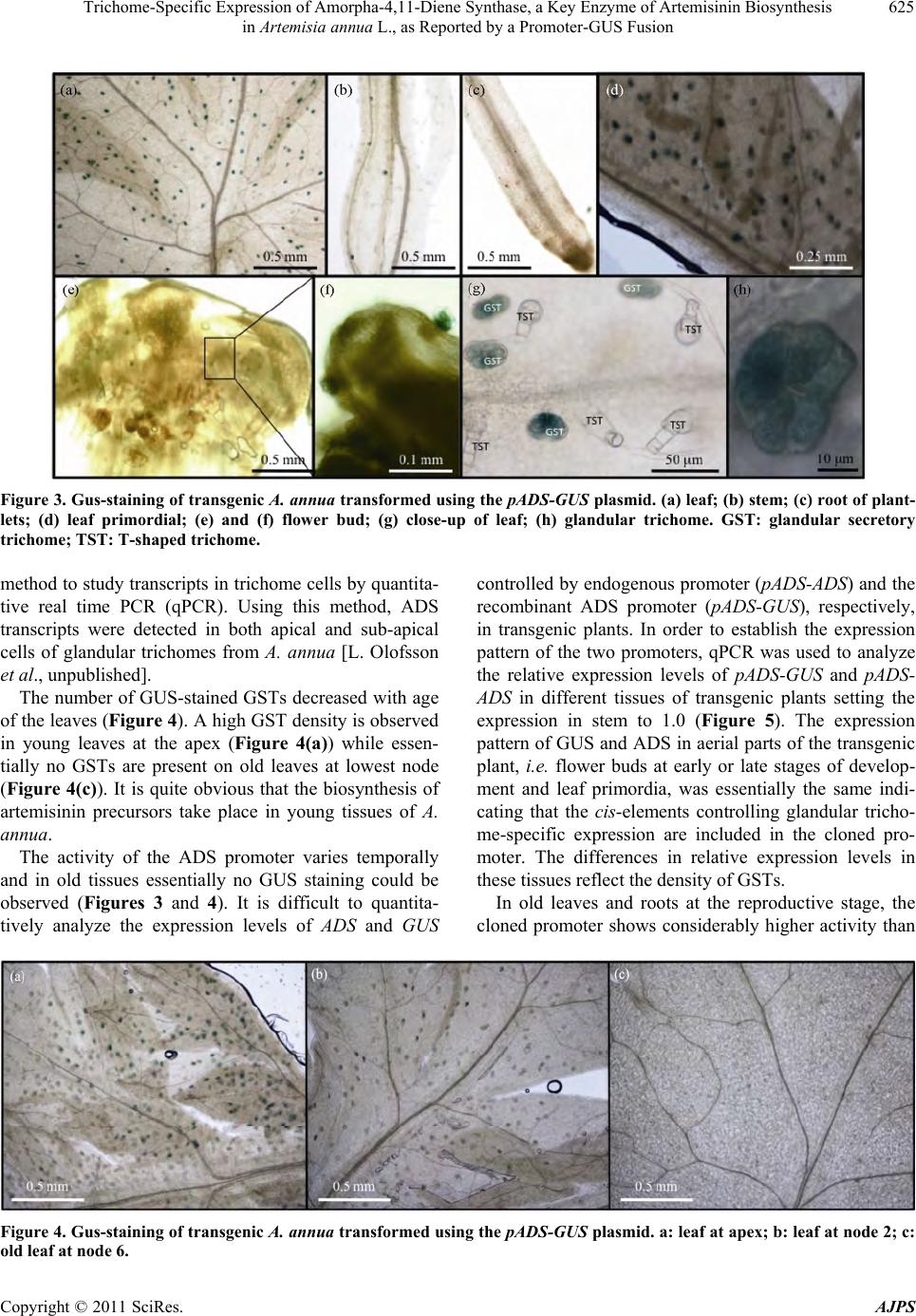 Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis 625 in Artemisia annua L., as Reported by a Promoter-GUS Fusion Figure 3. Gus-staining of transgenic A. annua transformed using the pADS-GUS plasmid. (a) leaf; (b) stem; (c) root of plant- lets; (d) leaf primordial; (e) and (f) flower bud; (g) close-up of leaf; (h) glandular trichome. GST: glandular secretory trichome; TST: T-shaped trichome. method to study transcripts in trichome cells by quantita- tive real time PCR (qPCR). Using this method, ADS transcripts were detected in both apical and sub-apical cells of glandular trichomes from A. annua [L. Olofsson et al., unpublished]. The number of GUS-stained GSTs decreased with age of the leaves (Figure 4). A high GST density is observed in young leaves at the apex (Figure 4(a)) while essen- tially no GSTs are present on old leaves at lowest node (Figure 4(c)). It is quite obvious that the biosynthesis of artemisinin precursors take place in young tissues of A. annua. The activity of the ADS promoter varies temporally and in old tissues essentially no GUS staining could be observed (Figures 3 and 4). It is difficult to quantita- tively analyze the expression levels of ADS and GUS controlled by endog enous promoter (pADS-ADS) and the recombinant ADS promoter (pADS-GUS), respectively, in transgenic plants. In order to establish the expression pattern of the two promoters, qPCR was used to analyze the relative expression levels of pADS-GUS and pADS- ADS in different tissues of transgenic plants setting the expression in stem to 1.0 (Figure 5). The expression pattern of GUS and ADS in aerial parts of the transgen ic plant, i.e. flower buds at early or late stages of develop- ment and leaf primordia, was essentially the same indi- cating that the cis-elements controlling glandular tricho- me-specific expression are included in the cloned pro- moter. The differences in relative expression levels in these tissues reflect the density of GSTs. In old leaves and roots at the reproductive stage, the cloned promoter shows considerably higher activity than Figure 4. Gus-staining of transgenic A. annua transformed using the pADS-GUS plasmid. a: leaf at apex; b: leaf at node 2; c: ld leaf at node 6. o Copyright © 2011 SciRes. AJPS 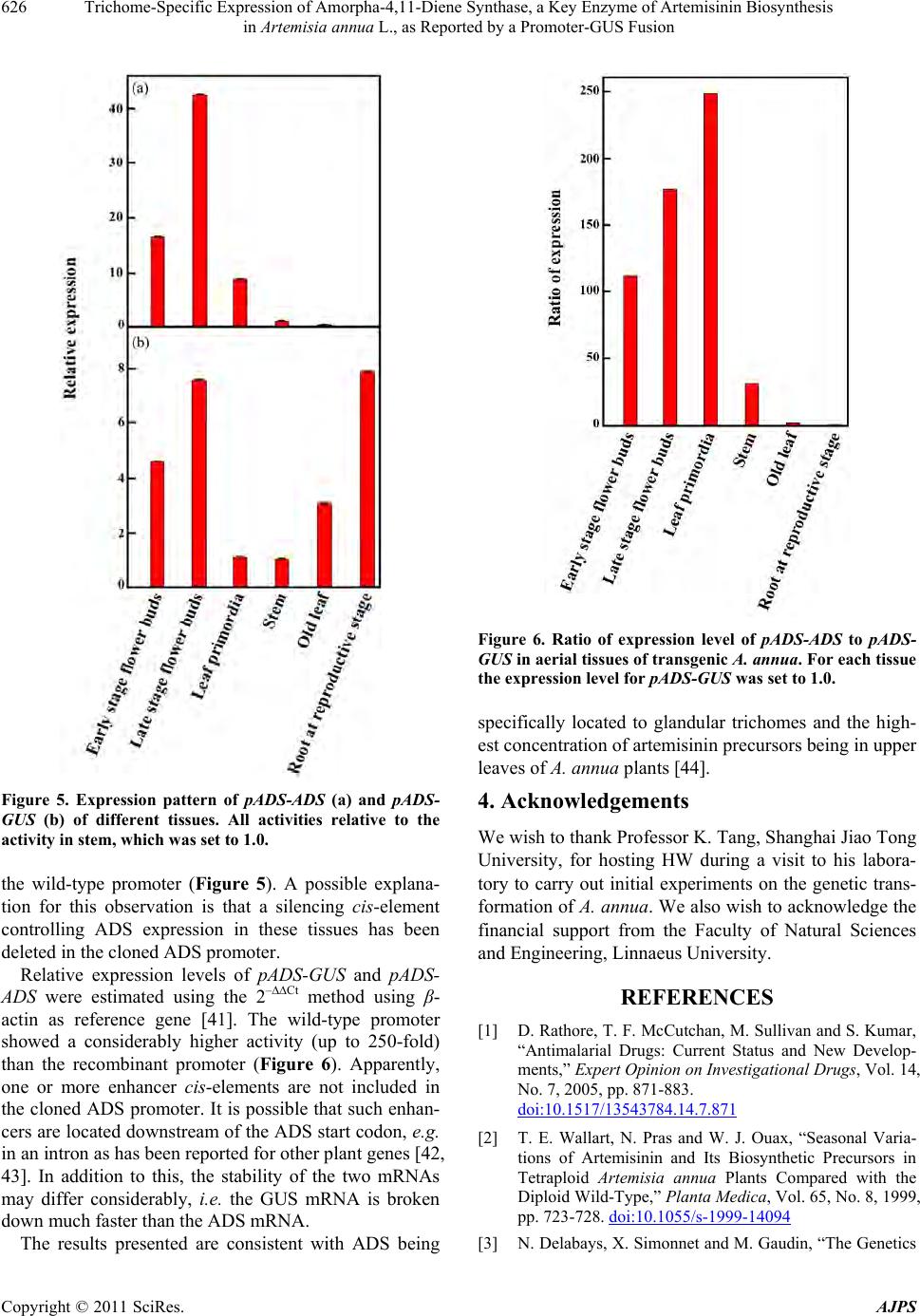 Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis 626 in Artemisia annua L., as Reported by a Promoter-GUS Fusion Figure 5. Expression pattern of pADS-ADS (a) and pADS- GUS (b) of different tissues. All activities relative to the activity in stem, which was set to 1. 0. the wild-type promoter (Figure 5). A possible explana- tion for this observation is that a silencing cis-element controlling ADS expression in these tissues has been deleted in the cloned ADS promoter. Relative expression levels of pADS-GUS and pADS- ADS were estimated using the 2–∆∆Ct method using β- actin as reference gene [41]. The wild-type promoter showed a considerably higher activity (up to 250-fold) than the recombinant promoter (Figure 6). Apparently, one or more enhancer cis-elements are not included in the cloned ADS promoter. It is possible that such enhan- cers are located downstream of the ADS start codon, e.g. in an intron as has been re po rt e d for ot her plant genes [42, 43]. In addition to this, the stability of the two mRNAs may differ considerably, i.e. the GUS mRNA is broken down much faster than the ADS mRNA. The results presented are consistent with ADS being Figure 6. Ratio of expression level of pADS-ADS to pADS- GUS in aerial tissues of transgenic A. annua. For each tissue the expression level for pADS-GUS was set to 1.0. specifically located to glandular trichomes and the high- est concentration of artemisinin precursors being in upper leaves of A. annua plants [44]. 4. Acknowledgements We wish to thank Professor K. Tang , Shanghai Jiao Tong University, for hosting HW during a visit to his labora- tory to carry out initial experiments on the genetic trans- formation of A. annua. We also wish to acknowledge the financial support from the Faculty of Natural Sciences and Engineering, Linnaeus University. REFERENCES [1] D. Rathore, T. F. McCutchan, M. Sullivan and S. Kumar, “Antimalarial Drugs: Current Status and New Develop- ments,” Expert Opinion on Investigational Drugs, Vol. 14, No. 7, 2005, pp. 871-883. doi:10.1517/13543784.14.7.871 [2] T. E. Wallart, N. Pras and W. J. Ouax, “Seasonal Varia- tions of Artemisinin and Its Biosynthetic Precursors in Tetraploid Artemisia annua Plants Compared with the Diploid Wild-Type,” Planta Medica, Vol. 65, No. 8, 1999, pp. 723-728. doi:10.1055/s-1999-14094 [3] N. Delabays, X. Simonnet and M. Gaudin, “The Genetics Copyright © 2011 SciRes. AJPS  Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis 627 in Artemisia annua L., as Reported by a Promoter-GUS Fusion of Artemisinin Content in Artemisia annua L. and the Breeding of High Yielding Cultivars,” Current Medicinal Chemistry, Vol. 8, No. 15, 2001, pp. 795-1801. [4] M. Hommel, “The Future of Artemisinins: Natural, Syn- thetic or Recombinant?” Journal of Biology, Vol. 7, No. 10, 2008, p. 38. doi:10.1186/jbiol101 [5] F. Jing, L. Zhang, M. Li, Y. Tang, Y. Wang, Y. Wang, Q. Wang, Q. Pan, G. Wang and K. Tang, “Abscisic Acid (ABA) Treatment Increases Artemisinin Content in Ar- temisia annua by Enhancing the Expression of Genes in Artemisinin Biosynthetic Pathway,” Biologia, Vol. 64, No. 2, 2009, pp. 319-323. doi:10.2478/s11756-009-0040-8 [6] T. C. Smith, P. J. Weathers and R. D. Cheetham, “Effects of Gibberellic Acid on Hairy Root Cultures of Artemisia annua: Growth and Artemisinin Production,” In Vitro Cellular & Developmental Biology-Plant, Vol. 33, No. 1, 1997, pp. 75-79. doi:10.1007/s11627-997-0044-4 [7] P. J. Weathers, G. Bunk and M. C. McCoy, “The Effect of Phytohormones on Growth and Artemisinin Production in Artemisia annua Hairy Root,” In Vitro Cellular & De- velopmental Biology-Plant, Vol. 41, No. 1, 2005, pp. 47-53. doi:10.1079/IVP2004604 [8] Y. S. Zhang, H. C. Ye, B. Y. Liu, H. Wang and G. F. Li, “Exogenous GA3 and Flowering Induce the Conversion of Artemisinic Acid to Artemisinin in Artemisia annua Plants,” Russian Journal of Plant Physiology, Vol. 52, No. 1, 2005, pp. 58-62. doi:10.1007/s11183-005-0009-6 [9] W. Putalun, W. Luealon, W. De-Eknamkul, H. Tanaka and Y. Shoyama, “Improvement of Artemisinin Produc- tion by Chitosan in Hairy Root Cultures of Artemisia an- nua L.,” Biotechnology Letters, Vol. 29. No. 7, 2007, pp. 1143-1146. doi:10.1007/s10529-007-9368-8 [10] A. Baldi and V. K. Dixit, “Yield Enhancement Strategies for Artemisinin Production by Suspension Cultures of Ar- temisia annua,” Bioresource Technology, Vol. 99, No. 11, 2008, pp. 4609-4614. doi:10.1016/j.biortech.2007.06.061 [11] C. Lei , D. Ma, G. Pu, X. Qiu, Z. Du, H. Wang, G. Li, H. Ye and B. Liu, “Foliar Application of Chitosan Activates Artemisinin Biosynthesis in Artemisia annua L.,” Indus- trial Crops and Products, Vol. 33, No. 1, 2011, pp. 176-182. doi:10.1016/j.indcrop.2010.10.001 [12] L. Zhang, F. Jing, F. Li, M. Li, Y. Wang, G. Wang, X. Sun and K. Tang, “Development of Transgenic Artemisia annua (Chinese Wormwood) Plants with an Enhanced Content of Artemisinin, an Effective Anti-Malarial Drug, by Hairpin-RNA-Mediated Gene Silencing,” Biote- chnology and Applied Biochemistry, Vol. 52, No. 3, 2009, pp. 199-207. doi:10.1042/BA20080068 [13] L. -L. Feng, R. -Y. Yang, X. -Q. Yang, X. -M. Zeng. W. -J. Lu and Q. -P. Zeng, “Synergistic Re-Channeling of Mevalonate Pathway for Enhanced Artemisinin Produc- tion in Transgenic Artemisia annua,” Plant Science, Vol. 177, No. 1, 2009, pp. 57-67. doi:10.1016/j.plantsci.2009.03.014 [14] J. -L. Han, B. -Y. Liu, H. -C. Ye, H. Wang, Z. -Q. Li and G. -F. Li, “Effects of Overexpression of the Endogenouse Farnesyl Diphosphate Synthase on the Artemisinin Con- tent in Artemisia annua L.,” Journal of Integrative Plant Biology, Vol. 48, No. 4, 2006, pp. 482-487. doi:10.1111/j.1744-7909.2006.00208.x [15] P. Mercke, M. Bengtsson, H. J. Bouwmeester, M. A. Posthumus and P. E. Brodelius, “Molecular Cloning, Ex- pression, and Characterization of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis in Artemisia annua L.,” Archives of Biochemistry and Bio- physics, Vol. 381, No. 2, 2000, pp. 173-180. doi:10.1006/abbi.2000.1962 [16] H. J. Bouwmeester, T. E. Wallaart, M. H. Janssen, B. va n Loo, B. J. Jansen, M. A. Posthumus, C. O. Schmidt, J. W. de Kraker, W. A. Konig and M. C. Franssen, “Amor- pha-4,11-Diene Synthase Catalyses the First Probable Step in Artemisinin Biosynthesis,” Phytochemistry, Vol. 52, No. 5, 1999, pp. 843-854. doi:10.1016/S0031-9422(99)00206-X [17] S. H. Kim, Y. J. Chang and S. U. Kim, “Tissue Specific- ity and Developmental Pattern of Amorpha-4,11-Diene Synthase (ADS) Proved by ADS Promoter-Driven GUS Expression in the Heterologous Plant, Arabidopsis thaliana,” Planta Medica, Vol. 74, No. 2, 2008, pp. 188-193. doi:10.1055/s-2008-1034276 [18] K. H. Teoh, D. R. Polichuk, D. W. Reed, G. Nowak and P. S. Covello, “Artemisia annua L. (Asteraceae) Trichome- Specific cDNAs Reveal CYP71AV1, a Cytochrome P450 with a Key Role in the Biosynthesis of the Antimalarial Sesquiterpene Lactone Artemisinin,” FEBS Letters, Vol. 580, No. 5, 2006, pp. 1411-1416. doi:10.1016/j.febslet.2006.01.065 [19] D. -K. Ro, E. M. Paradise, M. Ouel let, K. J. Fisher, K. L. Newman, J. M. Ndungu, K. A. Ho, R. A. Eachus, T. S. Ham, J. Kirby, M. C. Y. Chang, S. T. Withers, Y. Shiba, R. Sarpong and J. D. Keasling, “Production of the Anti- malarial Drug Precursor Artemisinic Acid in Engineered Yeast,” Nature, Vol. 440, No. 7, 2006, pp. 940-943. doi:10.1038/nature04640 [20] G. D. Brown and L. K. Sy, “In Vivo Transformations of Dihydroartemisinic Acid in Artemisia annua Plants,” Tetrahedron, Vol. 60, No. 5, 2004, pp. 1139-1159. doi:10.1016/j.tet.2003.11.070 [21] Y. Zhang, K. H. Teoh, D. W. Reed, L. Maes, A. Goossens, D. J. Olson, A. R. Ross and P. S. Covello, “The Molecular Cloning of Artemisinic Aldehy de 11(13) Reductase and Its Role in Glandular Trichome-Dependent Biosynthesis of Artemisinin in Artemisia annua,” The Journal of Biological Chemistry, Vol. 283, No. 31, 2008, pp. 21501-21508. doi:10.1074/jbc.M803090200 [22] K. H. Teoh, D. R. Polichuk, D. W. Reed and P. S. Covello, “Molecular Cloning of an Aldehyde Dehydro- genase Implicated in Artemisinin Biosynthesis in Ar- temisia annua,” Botany, Vol. 87, No. 6, 2009, pp. 635-642. doi:10.1139/B09-032 [23] M. V. Duke, R. N. Paul, H. N. Elsohly, G. Sturtz and S. O. Duke, “Localization of Artemisinin and Artemisitene in Copyright © 2011 SciRes. AJPS  Trichome-Specific Expression of Amorpha-4,11-Diene Synthase, a Key Enzyme of Artemisinin Biosynthesis in Artemisia annua L., as Reported by a Promoter-GUS Fusion Copyright © 2011 SciRes. AJPS 628 Foliar Tissues of Glanded and Glandless Biotypes of Ar- temisia annua L.,” International Journal of Plant Sci- ences, Vol. 155, No. 3, 1994, pp. 365-372. doi:10.1086/297173 [24] P. S. Covello, K. H. Teoh, R. Devin, D. R. Polichuk, “Function Genomic and Biosynthesis of Artemisinin,” Phytochemistry, Vol. 68, No. 14, 2007, pp. 1864-1871. doi:10.1016/j.phytochem.2007.02.016 [25] L. Olofsson, A. Engström, A. Lundgren and P. E. Brode- lius, “Relative Expression of Genes of Terpene Metabo- lism in Different Tissues of Artemisia annua L.,” BMC Plant Biology, Vol. 11, No. 45, 2011, pp. 1-12. [26] J. Fütterer, A. Gisel, V. Iglesias, A. Klöti, B. Kost, O. Mittelsten Scheid, G. Neuhaus, G. Neuhaus-Url, M. Schrott, R. Shillito, G. Spangenberg and Z. Y. Wang, “Standard Molecular Techniques for the Analysis of Transgenic Plants,” In: I. Potrykus and G. Spangenberg, Eds., Gene Transfer to Plants, Springer Verlag, Berlin, 1995, pp. 215-263. [27] R. A. Jefferson, T. A. Kavanagh and M. W. Bevan, “GUS Fusions: Beta-Glucuronidase as a Sensitive and Versatile Gene Fusion Marker in Higher Plants,” The EMBO Journal, Vol. 6, No. 13, 1987, pp. 3901-3907. [28] S. Yin, L. Mei, J. Newman, K. Back and J. Chappell, “Regulation of Sesquiterpene Cyclase Gene Expression; Characterization of an Elicitor- and Pathogen-Inducible Promoter,” Plant Physiology, Vol. 115, No. 2, 1997, pp. 437-451. doi:10.1104/pp.115.2.437 [29] T. Yang and B. W. Poovaiah, “An Early Ethylene up-Regulated Gene Encoding a Calmodulin-Binding Pro- tein Involved in Plant Senescence and Death,” The Jour- nal of Biological Chemistry, Vol. 275, No. 49, 2000, pp. 38467-38473. doi:10.1074/jbc.M003566200 [30] T. Yang and B. W. Poovaiah, “A Calmodulin-Binding/ CGCG-Box DNA-Binding Protein Family Involved in Multiple Signaling Pathways in Plants,” The Journal of Biological Chemistry, Vol. 277, No. 47, 2002, pp. 45049-45058. doi:10.1074/jbc.M207941200 [31] L. D. Zhang, K. J. Zuo, F. Zhang, Y. F. Cao, J. Wang, Y. D. Zhang, X. F. Sun and K. X. Tang, “Conservation of Non-Coding Microsatellites in Plants: Implication for Gene Regulation,” BMC Genomics, Vol. 7, No. 323, 2006 , pp. 1-14. [32] G. -B. Pu, D. -M. Ma, J. -L. Chen, L. -Q. Ma, H. Wang, G. -F. Li, H. -C. Ye and B. -Y. Liu, “Salicylic Acid Acti- vates Artemisinin Biosynthesis in Artemisia annua L.,” Plant Cell Reports, Vol. 28, No. 7, 2009, pp. 1127-1135. doi:10.1007/s00299-009-0713-3 [33] D. Ma, G. Pu, C. Lei, L. Ma, H. Wang, Y. Guo, J. Chen, Z. Du, H. Wang, G. Li, H. Ye and B. Liu; “Isolation and Characterization of AaWRKY1, an Artemisia annua Transcription Factor that Regulates the Amorpha-4,11- Diene Synthase Gene, a Key Gene of Artemisinin Bio- synthesis,” Plant and Cell Physiology, Vol. 50, No. 12, 2009, pp. 2146-2161. doi:10.1093/pcp/pcp149 [34] Y. -H. Xu, J. -W. Wang, S. Wang, J. -Y. Wang and X. -Y. Chen, “Characterization of GaWRKY1, a Cotton Tran- scription Factor that Regulates the Sesquiterpene Syn- thase Gene (+)-δ-Cadinene Synthase-A,” Plant Physiol- ogy, Vol. 135, No. 1, 2004, pp. 507-515. doi:10.1104/pp.104.038612 [35] J. C. Reyes, M. I. Muro-Pastor and F. J. Florencio, “The GATA Family of Transcription Factors in Arabidopsis and Rice,” Plant Physiology, Vol. 134, No. 4, 2004, pp. 1718-1732. doi:10.1104/pp.103.037788 [36] T. Urao, K. Yamaguchi-Shinozaki, S. Urao and K. Shi- nozaki, “An Arabidopsis myb Homolog Is Induced by Dehydration Stress and Its Gene Product Binds to the Conserved MYB Recognition Sequence,” Plant Cell, Vol. 5, No. 11, 1993, pp. 1529-1539. [37] R. Solano, C. Nieto, J. Avila, L. Cañas, I. Diaz and J. Paz-Ares, “Dual DNA Binding Specificity of a Petal Epidermis-Specific MYB Transcription Factor (MYB. Ph3) from Petuni a hybrida,” The EMBO Journal, Vol. 14, No. 8, 1995, pp. 1773-1784. [38] G. D. Brown, G. -Y. Liang and L. -K. Sy, “Terpenoids from the Seeds of Artemisia annua,” Phytochemistry, Vol. 64, No. 1, 2003, pp. 303-323. doi:10.1016/S0031-9422(03)00294-2 [39] J. F. S. Ferre ira, J. E. Simon and J. Janick, “Developmen- tal Studies of Artemisia annua: Flowering and Artemisi- nin Production under Greenhouse and Field Conditions,” Planta Medica, Vol. 61, No. 2, 1995, pp. 167-170. doi:10.1055/s-2006-958040 [40] M. E. Olsson, L. M. Olofsson, A. L. Lindahl, A. Lundgren, M. Brodelius and P. E. Brodelius, “Localiza- tion of Enzymes of Artemisinin Biosynthesis to the Api- cal Cells of Glandular Secretory Trichomes of Artemisia annua L.,” Phytochemistry, Vol. 70, No. 9, 2009, pp. 1123-1128. doi:10.1016/j.phytochem.2009.07.009 [41] K. J. Livak and T. D. Schmittgen, “Analysis of Relative Gene Expression Data using Real-Time Quantitative PCR and the 2–CT Method,” Met h od s, Vol. 25, No. 4, 2001, pp. 402-408. doi:10.1006/meth.2001.1262 [42] D. Mascarenhas, I. J. Mettler, D. A. Pierce and H. W. Lowe, “Intron-Mediated Enhancement of Heterologous Gene Expression in Maize,” Plant Molecular Biology, Vol. 15, No. 6, 1990, pp. 913-920. doi:10.1007/BF00039430 [43] M. K. Deyholos and L. E. Sieburth, “Separable Whorl- Specific Expression and Negative Regulation by Enhan- cer Elements within the AGAMOUS Second Intron,” Plant Cell, Vol. 12, No. 10, 2000, pp. 1799-1810. [44] S. O. Duke and R. N. Paul, “Development an Fine Struc- ture of the Glandular Trichomes of Artemisia annua L.,” International Journal of Plant Sciences, Vol. 154, No. 1, 1993, pp. 107-118. doi:10.1086/297096
|