Advances in Microbiology
Vol.04 No.15(2014), Article ID:51673,9 pages
10.4236/aim.2014.415120
Dps Is a Stationary Phase-Specific Protein of Escherichia coli Nucleoid
Ali Azam Talukder1,2,3,4,5*, Akira Ishihama2,3
1Department of Microbiology, Jahangirnagar University, Savar, Bangladesh
2Department of Molecular Genetics, National Institute of Genetics, Mishima, Japan
3Micro-Nano Technology Research Center, Hosei University, Tokyo, Japan
4Japan Science and Technology Corporation, Kawaguchi, Japan
5Japan Society for the Promotion of Science, Tokyo, Japan
Email: *tunubj@yahoo.com, aat@juniv.edu
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 27 September 2014; revised 5 November 2014; accepted 20 November 2014
ABSTRACT
Bacterial genomic DNA is highly organized into one or few compacted bodies known as nucleoid, which is composed of DNA, RNA and several DNA-binding proteins. These DNA-binding proteins require essential alterations in their expression during stationary phase of growth in order to respond to stressful environmental conditions. Dps (DNA-binding protein from starved cells) is one of such DNA-binding proteins, which accumulates most when E. coli cells reach to the stationary phase. Here, we have characterized Dps protein under various growth phases. Immunofluorescent microscopic observation reveals that Dps plays a key role in final round of genome compaction during the stationary phase. Similar results are also obtained by Western immunoblot analysis, after quantification of Dps protein from the exponential phase and early stationary phase nucleoid bound fractions, separated by sucrose density gradient centrifugation. Our results support the conclusion that Dps occupies more than half of the stationary phase nucleoid in E. coli.
Keywords:
Dps, DNA-Binding Protein, Stationary Phase, E. coli, Nucleoid

1. Introduction
The chromosomes in the eukaryotic nucleus are folded with the nucleosome and its histone proteins as the basic unit [1] [2] . In bacteria there is no such nucleosome, but the DNA is nevertheless organized in a nucleoid body with group of proteins within the cells [3] [4] . Studies of the E. coli nucleoid over many years have revealed an irregularly shapedandcompact structure confined to less than half of the intracellular space [5] . This structure rapidly changes in stressful environmental conditions when gene expression is modulated [6] . One of such conditions is stationary phase. At this growth phase, supplies of various nutrition as well as other physical and chemical growth parameters are turned off and therefore cells have to survive without nutrition. This unfavorable condition alters E. coli cell physiology and metabolism, which help counter stressful conditions for long time, even years, with the synthesis or accumulation of several new components, including various nucleoid proteins, growth factors or hormones and enzymes, used to overcome stress conditions [7] . Several hundred of genes are expressed when cells reach the stationary phase [7] . These genes are controlled by stationary phase specific σfactorσS [8] .
One of such stress tolerant gene is Dps, which produces Dps protein at stationary phase, which was earlier identified as an abundant sequence non-specific DNA-binding protein in E. coli [9] [10] . We previously reported sequence recognition specificity and the DNA-binding affinity and intracellular localization E. coli based on the profiles of twelve species of the nucleoid-associated proteins including Dps [11] - [13] . Furthermore, several in vivo and in vitro studies have revealed that the Dps protein plays an important role in organizing genomic DNA into the nucleoid [14] . Here, we carried out an indirect immunofluorescence microscopic study and Western im- munoblot analysis to investigate the association of Dps protein in organization of E. coli genomic DNA into the nucleoid at various growth phases of life cycle.
2. Materials and Methods
2.1. Bacterial Strains and Growth Conditions
The bacterial strains for analysis here are the A-type lineage of E. coli W3110, which carries the intact forms of both σS and σF [6] . Strain ZK126 is a wild type Dps and ZK1056 is a Dps inactivated mutant strain [12] . Cells were grown at 37˚C under aeration in luria broth (LB). Growth was monitored by measuring the turbidity with a Klett-Summerson photometer. Samples were collected at different time interval where necessary, sieged the growth and then stored at around 4˚C until further use.
2.2. Viable Count and Determination of Total Number of Cells
The number of cells was counted with a Coulter Multisizer II equipped with a 30 μm orifice and a 100 μl volume. Gain and current settings of 2.5 and 5.5, respectively. The cell suspensions were diluted serially in LB broth and spread in triplicate on LB agar plates. After incubation at 37˚C overnight, the plates were scored for viable colonies and the surviving fraction was determined by comparing experimental with control cultures.
2.3. Microscopic Observation of E. coli Cell, Nucleoid and Dps Protein
Various strains of E. coli were grown in LB medium and the culture conditions and sampling times were discussed in detail in the text where necessary. To observe clearly the shape and size of living and non-living cells and there nucleoids by fluorescent microscope, collected cells were fixed with 80% Methanol and stained with various dyes like DAPI (4’, 6-diamidino-2-phenylindole dihydrochloride, 5 μg/ml); Calcien and Propidium Idodide where necessary to observe nuceloid position on cell, living and dead cells, respectively, using the method developed previously [11] .
The procedure for indirect immunofluorescence microscopy was same as previously described [11] . In brief, various growth phase dependent samples were collected, fixed and stained with 500-fold-diluted first-step antibodies against the test Dps was placed on the sample, and left for 1 h at room temperature in a moisture chamber. After washing the sample with PBST buffer (140 mM NaCl, 2 mM KCl, 8 mM Na2HPO4, 1.5 mM KH2PO4 and 0.05% tween 20), the sample was treated with 500-fold diluted second-step goat anti-rabbit IgG antibodies conjugated with a fluorescence compound Cy3 at room temperature for 1 h in the moisture-controlled chamber. After washing with PBST, the immunostained sample was stained with 5 μl of DAPI for 1 min, and covered with one drop of mounting medium (1 mg/ml p-phenylenediamine, 90% glycerol in PBS [pH 9.0]). The immuno-stained samples were observed with a fluorescence and phase-contrast microscope (100× objective lens; Nikon, Japan) equipped with a chilled, color 3CCD camera, C5810-01 (Hamamatsu, Japan), connected with a computer. Filter cassettes corresponding to different fluorescence compound were used. The specificity of the observed immunostained fluorescent signal was verified by the absence of the signal in corresponding immuno- stained samples prepared using the null mutant cells (ZK1056). The images were transferred directly to a Power Macintosh and processed using Adobe Photoshop 6.0-j software.
2.4. Nucleoid Preparation and Isolation by Sucrose Density Gradient Centrifugation
Preparation of nucleoids and their sucrose density gradient centrifugation run were performed essentially according to the protocol devised by Kornberg et al. [15] , modified by Zimmerman [16] . In brief, an equal numbers of cells (each containing about 1.9 × 108 cells/ml) were collected from exponential to early-stationary phases, diluted where necessary, gently treated with lysozyme and then lysed with detergents in low salt media containing spermidine. These low-salt spermidine nucleoids have much higher protein and RNA contents than the high- salt nucleoids and contain DNA-binding proteins. In our cell lysate preparation, we have used 400 mg lysozyme/ml and chose to limit the exposure to lysozyme to 40 sec at 0˚C. The mild lysozyme concentration gave efficient detergent lysis under the conditions employed, as judged by recovery of cellular DNA in a nucleoid fraction. Linear gradients (12% - 60% sucrose in 10 mM Tris-HCl (pH 7.8 at 0˚C) containing 10 mM KCl, 1 mM EDTA, 0.2 mM DTT (dithiothreitol), 1 mM spermidine HCl) were formed in a cold room in 5 ml Ultraclear plastic centrifuge tubes (13 × 51 mm, Beckman) and were run at 10,000 rpm at 4˚C for 120 min in a Spinco SW50.1 rotor (the centrifugal force was reduced to 7000 g). Whole cell lysates from different growth phases were completely sedimented under these conditions. The relative positions of two independent nucleoids isolated from exponential and early stationary phases were reproducible at different speeds and for longer runs (data not shown). The speed and duration of the runs used in this study were chosen in order to maintain the fastest moving band (nucleoids from the 6.0 h cultures) within the bottom of the gradient. The tubes containing a single visible band are referred to as “nucleoid bound fraction”. Fractions in an average 0.45 ml (except nucleoid bound fractions in exponential phase was 0.6 ml and early stationary phase was 0.30 ml) were collected from the bottom of each gradient and mixed with 20 μl of 0.2 M EDTA, stored at −20˚C until further use.
2.5. Purification of DNA-Binding Proteins and Preparation of Antibodies
All eight DNA-binding proteins, CbpB (Rob), Dps, Fis, Hfq, H-NS, HU, IHF and StpA, were overexpressed with pMK19, pDPS1, pRJ1077, pHFQ607, pHOP11, pLhupAhupB, pSA5hiphimA and pT7stpA, respectively, and purified to apparent homogeneity as described previously [12] . Antibodies against each protein were produced in rabbits by injecting the purified proteins as described previously [12] .
2.6. Quantitative Western Immunoblot Analysis
For the measurement of each DNA-binding protein in E. coli W3110 nucleoid bound and nuceloud unbound fractions, a quantitative Western immunoblot analysis was employed using the polyclonal anti-Dps and anti-Fis antibodies as described [12] . In brief, either nucleoid or cytosolic fractions were treated with a sodium dodecyl sulfate (SDS) sample buffer (50 mM Tris-HCl [pH 6.8], 2% SDS, 1% 2-mercaptoethanol, 10% glycerol, 0.025% bromophenol blue) and separated on SDS-15% polyacrylamide gels. Protein in the gel was directly electroblotted onto polyvinylidene difluoride (PVDF) membranes (Nippon Genetics, Japan). Blots were blocked overnight at 4˚C in 3% bovine serum albumin (BSA) in phosphate-buffered saline (PBS), probed with the specific antibodies against each protein, washed with 0.5% Tween 20 in PBS, and incubated with goat anti-rabbit immunoglobulin G conjugated with hydroxyl peroxidase (Cappel). The blots were developed with ECL Western blotting detection reagents (Amersham) and chemiluminescence was quantitated with a lumino-image analyzer LAS- 1000 (Fuji, Japan). For accurate measurement of the levels of each of the nucleoid proteins, we took care in the followings: (i) the protein range was determined for each protein, where the linear relationship existed between the protein concentration and the immunostaining intensity; (ii) the determination was carried out using several volumes of cell lysates, which contained the test proteins at the concentration range determined as above; (iii) the standard samples of known concentrations were always included in the assays; (iv) the determination was repeated at least three times for each DNA-binding protein.
3. Results and Discussion
3.1. Intact Cell Structure of E. coli Nucleoid
When E. coli cell reaches to stationary growth phase, various types of cell populations were observed, like viable and culturable cells, viable but non-culturable cells, dead cells etc [5] . Here, we studied whether these variations are reproducible in the laboratory condition, cells were grown for various time intervals and visualized by fluorescent microscope as described previously [16] . Collected cell were fixed, stained with DAPI (4’, 6-diami- dino-2-phenylindole dihydrochloride); Cal (Calcein) and PI (Propidium Iodide). Various cell populations were seen when E. coli cells stained with these dyes are shown in Figure 1. At least 3 types of cells were found at stationary phase; some are viable and culturable (green color), some are viable but non-culturable (in orange color) and some are dead cells (in red color). However, these types of cells were not observed at exponential phase (Figure 1, top panel). This result clearly demonstrates that the physico-chemical properties are altered at the stationary phase, which ensures survival of E. coli at stationary phase. The needed special protein like Dps or group of proteins/other components that play important roles during the formation of stationary phase nucleoid.
3.2. Intracellular Localization of E. coli Dps Protein
Overnight Dps wild type (ZK126) and mutant (ZK1056) cells were diluted 1000-fold into the fresh LB medium and the cultures were further incubated at 37˚C at least 7 days as shown in Figure 2. Samples were collected in different time intervals and their viable colony numbers were measured as shown in Figure 2(A). Viable colony numbers were more or less same until 5-days old cultures in both strains and thereafter decreased gradually in Dps inactivated colony (ZK1056) counts compare to wild type (ZK126) cells. This result clearly demonstrates that Dps protein is essential for stationary phase survival.
Figure 1. Growth phase dependent changes in the shape and size of E. coli nucleoids, visualized by fluorescent microscope. E. coli W3110 cell growth, sample collection and sample preparation for fluorescence microscopy were discussed in the Materials and Methods section. Samples were collected at various time intervals as indicated in the left from top to bottom, fixed, stained with various dyes and observed through microscope. In each of five planes from left to right abbreviated as: PC = Phase contrast, which represents the cell shape and size; DAPI = DNA specific dye which binds only in the nucleoid area of cell, which stains in blue color; PC + DAPI = Combined image of PC + DAPI, which indicates the shape/size of cell and the position of nucleoid onto the cell; Cal = Calcein which indicates living cell and stained in green in color at 260 nm and in orange in color, which indicates viable but non-culturable cell; PI = Propidium Idodide which could detects dead cells in red in color at 120 nm. Circle indicates dead cell, where nucleoid is damaged. Scale bar in the bottom left panel represents 5 μm.
Figure 2. Intracellular localization of Dps protein in E. coli cell. (A) Viable colony numbers. An early stationary phase culture of wild type (ZK126) and mutant (ZK1056) E. coli cells in LB medium at 37˚C was diluted 1000-fold with fresh LB medium and the shaking culture at 160 rpm was continued at 37˚C until the 7th days. Cell growth, sample collection at various time points and the preparation for viable plate counting were described in the Materials and Methods section. The total number of viable colonies per ml was counted with a viable plate counting method in LB Agar plate. (B) Indirect immunofluorescent microscopic observation of E. coli cells. Exponentially growing (3.0 h), early stationary phase (6.0 h) and late stationary phase (72.0 h) wt and mutant cells were collected, fixed, stained and subjected to the indirect immunofluorescent microscopy as shown in top, middle and bottom panels, respectively. The distribution of Dps was visualized using the respective mono-specific rabbit antibody (prepared after absorption of nonspecific antibody using the respective null mutant cell extracts) as the primary antibody, and Cy3-labelled anti-rabbit IgG antibody as the secondary antibody. Photographs represent, from the left to right, PC (phase-contrast, the first column), DAPI-staining (the second column), the indirect immunofluorescent microscopy (Cy3, the third column), the combined image of phase-contrast and DAPI-staining (PC + DAPI, the forth column) and the combined image of phase-contrast, the DAPI-staining and the indirect immunofluorescent microscopy (PDC, the fifth column). The strain ZK126, wild-type Dps and ZK1056 is a Dps disrupted mutant. The scale bar at the bottom left represents 7 μm.
We have checked the location of E. coli Dps protein into the intact cell by indirect immunofluorescent microscope (Figure 2(B)). Collected cells were fixed, stained with various dyes. The location of nucleoid was identified by staining with DAPI, while cell shape and size was visualized by phase-contrast microscopy [16] . Intact cell nucleoid shapes, sizes and compactness change gradually when E. coli culture shifted gradually from the exponential growing phase to the stationary phase (compare 3.0 h culture with 6.0 to 72 h in Figure 2(B)). An average fluorescent area occupied by nucleoid reduces from exponential growing phase to late stationary phase is shown in Figure 2(B), which clearly indicate that the compaction of genomic DNA into nucleoid gradually form from the exponential phase to the late stationary phase in Dps wild type cell (ZK126). The accumulation level of Dps is also incleased gradually from the exponential growing to the late-stationary phase. However, this growth phase dependent compactness of nucleoid and accumulation level was not observed in Dps null mutant strain (ZK1056, lower panel of 3, 6 and 72 h cells). This result revealed that the Dps protein is involve in stationary phase dependent nucleoid compaction in E. coli.
3.3. Association in Vivo of Dps Protein into the E. coli Nucleoid
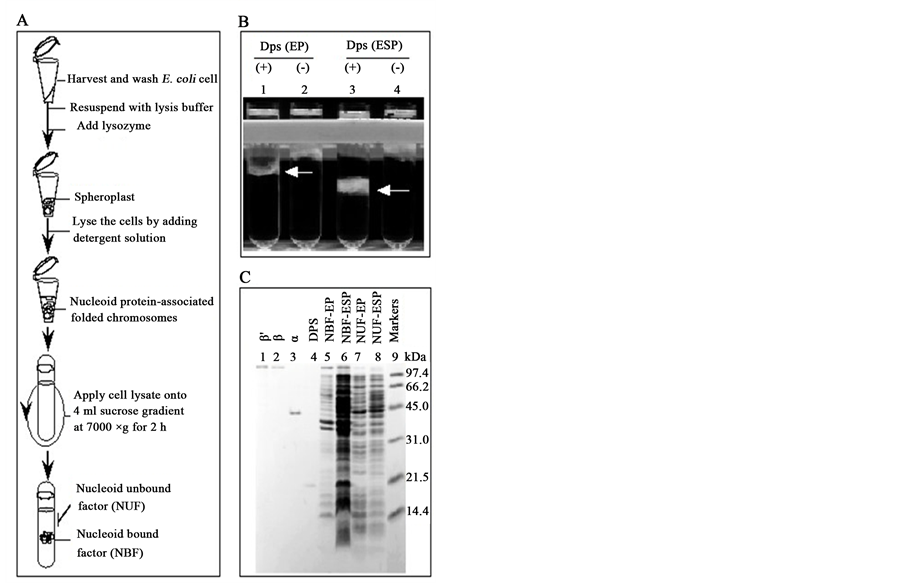
In order to check whether Dps protein is associated with the E. coli nucleoid, membrane-associated nuceloid was isolated from Dps wt (ZK126) and mutant (ZK1056) strain background by sucrose density gradient centrifugation as shown in Figure 3. The protocol for nucleoid separation from the whole cell extracts is shown in Figure 3(A). After run the centrifugation at 10,000 rpm for 2 hours, nucleoid bound fraction (NBF) and nucleoid unbound fractions (NUF) were separated easily by using 12% - 60% sucrose density gradient in a tube. The
Figure 3. Isolation and characterization of E. coli nucleoids. (A) The schematic procedure for preparation and isolation of E. coli nucleoids. (B) Sedimentation patterns of E. coli nucleoids. (C) Gel electrophoresis patterns of NBF and NUF from exponential and early-stationary phase cultures were compared with those of other known proteins on a SDS-15% polyacrylamide gel. Lanes 1 - 4 represent the standard known protein and β' (155 kDa), β (150 kDa), α (37.5 kDa) subunits of RNA polymerase and Dps (19.0 kDa). Lanes 5 - 6 and 7 - 8 represent the equal volume (2.0 μl) of exponential and early-stationary phase nucleoid bound fractions (NBF) and nucleoid unbound fractions (NUF), respectively. Lane 9 represent the standard molecular mass markers (molecular mass values in kDa shown from top to bottom: phosphorylase b, 97.4 kDa; bovine serum albumin, 66.2 kDa; ovalbumin, 45.0 kDa; carbonic anhydrase, 31.0 kDa; soybean trypsin inhibitor, 21.5 kDa; lysozyme, 14.4 kDa). Gels were stained with Coomassie brilliant blue (CBB), washed, dry and photographed.
tubes containing a single visible band are referred to as “nucleoid bound fraction” which contains more than 75% genomic DNA (data not shown). On the other hand, top 3 - 5 soluble fractions of each 4-tubes refer to as “nucleoid unbound fraction”, because more than 80% proteins are associated with these fractions (data not shown). The sedimentation profile of wt early stationary phase (ESP) NBF is faster than that of exponential phase (EP) NBF as indicated by arrows in Figure 3(B). However, this kind of variation was not observed in Dps mutant background (comparing lanes 1 and 3 with lanes 2 and 4 in Figure 3(B)). This result clearly indicated again that E. coli nucleoid compaction gradually takes place from exponential growth phase to the stationary growth phase and that the Dps protein is involved in the growth phase dependent compaction of nucleoid in E. coli.
We attempted to identify the proteins which are associated with E. coli nucleoids. Protein expression patterns from NBF were compared with those of NUF by SDS-polyacrylamide gel electrophoresis as shown in Figure 3(C). In lanes 5 - 6 and 7 - 8 we had applied 1 μl NBF and NUF, respectively, of two growth phases (EP and ESP). It is clear that the RNA polymerase components (β', β and α) are associated with the NBF (comparing lanes 1 - 3 with lane 5 or 6). Upon reaching ESP of cell growth at around 6.0 hours, several stationary-phase specific proteins including Dps were identified (comparing lane 4 with lane 5 or 6). However, under the conditions employed here by CBB staining, we could not detected Fis in NBF. The nucleoid components were analyzed further by quantitative Western immunoblot analysis as desorbed below.
Here, we have quantified 2 major nucleoid proteins from the NBF and NUF of EP and ESP cultures, respectively, as shown in Figure 4. One is Fis protein and another one is Dps protein. The former one is a major component of exponential phase nucleoid and the later one is responsible for final round of compaction of genomic DNA in to the nucleoid as suggested previously [11] [17] - [19] , because both protein accumulation level and the functions during nucleoid formations were varied and dependent on two specific growth phases [8] .
Fis is a small basic DNA-binding protein in E. coli, which was identified as a factor for site-specific DNA recombination [17] . Several lines of evidence indicate that Fis also participates in other processes such as transcription of the growth-related genes and DNA replication [19] [20] . The intracellular level of Fis protein in exponential phase was estimated to be about 60,000 molecules per cell, but the protein became undetectable in the stationary phase [8] . Therefore, Fis is a most abundant sequence specific nucleoid-associated protein in the growing cells and covers every 200 - 300 bp stretch along the genome DNA. When highly expressed, Fis represses its own synthesis by binding to the Fis promoter region. Upon entry into the stationary phase, the synthesis of Fis is switched off, resulting in the decrease in intracellular level by 500 to 1000 fold [21] [22] . The growth-dependent change in the Fis level is in good agreement with the fact that Fis is needed for transcription of the growth-related genes such as those for rRNA and tRNA or for DNA replication [21] .
Dps was identified as a starvation inducible DNA-binding protein in E. coli [7] . It is a small peptide of 19 kDa and forms a dodecameric complex [13] . High-level expression of Dps also occurs under various stress conditions. The purified Dps binds to DNA with no apparent sequence specificity [10] , and thus Dps is classified as a member of the bacterial nucleoid associated or histone-like protein family, which includes HU, H-NS, IHF and Fis [7] - [17] [21] . Dps exists about 6,000 molecules per cell in exponential phase and thereafter gradually increases up to about 180,000 molecules per cell at late stationary phase [8] . Dps is thus the main abundant stationary phase nucleoid proteins in E. coli.
Figure 4(A) and Figure 4(B) represent the protein purification steps of Fis and Dps, respectively, as described previously [12] . These purified proteins were used to prepare antibody for Western immunoblot analysis as shown in Figure 4(C). Nucleoid bound fraction (NBF) and nucleoid unbound fractions (NUF) were separated by sucrose density gradient centrifugation. One μg of each of above 4 fractions of EP and ESP was loaded onto the polyacrylamide gel in lanes 1 - 4 of Figure 3(C) and their Fis and Dps levels were visualized by Western immunoblot analysis. For the estimation of 2 nucleoid proteins from 2 different fractions, we always have used standard known proteins (lanes 5 - 13 in Figure 4(C)). Our results clearly indicate that the accumulation level of Fis and Dps in the nucleoid bound fractions varies and is growth phase dependent (comparing lane 1 with lane 2 in Figure 4(C)). NBF and NUF bands were further used to estimate the total amount of Fis or Dps proteins (ng/μg) present in 2 diffferent growth phases as shown in Figure 4(D). Finally, percentage distribution ratios of Fis or Dps in 2 distinct growth phases are calculated and shown in Figure 4(E), in addition to other nucleoid proteins estimated previously [8] . Altogether, our results demonstrate that the FIs is a major component of exponential phase nucleoid, whereas Dps occupies more than half of early stationary phase nucleoid in E. coli.
Figure 4. Growth phase dependent variation in the protein composition of E. coli nucleoids. (A) and (B) represent the protein purification steps of Dps and Fis, respectively in (A) (lanes 1 - 5 represent cell extract, Sup (supernatant), DEAE column, Heparin column fractions and known protein markers) and in (B) (lanes 1 - 5 represent cell extract, Pcs (precipitation), Heparin column, Mono-S column and known protein markers). The detail protocol for purification of Dps and Fis protein were reported previously [12] . (C) Western immunoblot analysis of E. coli Dps and Fis protein. One μg nucleoid bound fraction (NBF) and nucleoid unbound fraction (NUF) of two growth phases (EP and ESP) were applied to lanes 1 - 4 of the SDS-containing, 15% polyacrylamide gels for the measurement of Dps and Fis protein level. Lanes 1 and 2 represent the nucleoid bound fractions in two different growth phases (EP and ESP), while lanes 3 and 4 represent the nucleoid unbound fractions of EP and ESP which were separated by sucrose density gradient centrifugation. Remaining lanes 5-13 represent various standard concentrations of know proteins either Dps or Fis for the quantification of each 2 proteins in various fractions (lanes 1 - 4). Proteins were Electro-blotted onto PVSF membranes. The blots were probed with either anti-Dps or anti-Fis antibodies. After immuno-staining, the intensity of stained bands (chemiluminescence) was quantified with a lumino-image analyzer LAS-1000 (Fuji, Japan). (D) Cellular concentrations of Dps and Fis, which were measured from NBF and NUF of EP and ESP cultures, respectively, as shown in (C) and (E) (possible models for the growth-dependent changes in the structure and protein composition of E. coli nucleoid). The percentage (%) distribution ratios of nucleoid proteins in E. coli were estimated from Western immunoblot analysis as shown in (C). Intracellular concentrations of CbpB, Hfq, H-NS, HU, IHF and StpA were previously estimated [8] . The major components of exponential phase nucleoid are Fis, HU, H-NS, StpA and Hfq, while Dps occupies more than half of the stationary-phase nucleoid. All the nucleoid proteins are assumed to be associated with the DNA.
The results of this study and the several lines of recent evidence suggest that Dps plays a key role in the final round of compaction of genomic DNA into the form of nucleoid at stationary phase. This tight compaction state of DNA protects genome from various internal and external stresses arising at stationary phase. Indeed, the Dps mutant cells are very sensitive to environmental stresses [14] . Our results and several mutational analyses demonstrated that the stationary phase specific tight compaction of the nucleoid required a protein, Dps [22] . Moreover, overexpression of Dps induced an intracellular crystalline structure in vivo and purified Dps proteins were co-crystallized with DNA [23] . The highly compacted nucleoids observed in stationary phase appear to have similar characteristics to a bio-crystal [23] -[25] . Therefore, Dps protein appears to be crucial for achieving the formation of nucleoid and for protecting various environmental stresses at stationary phase.
Acknowledgements
The authors thank S. Hiraga and N. Fujita for valuable suggestions, demonstrating immuniflorescent microscopy and comments during the preparation of the manuscript, respectively. This work was supported by Grants-in- Aid from the Ministry of Education, Science and Culture of Japan and CREST (Core Research for Evolutional Science and Technology) of Japan Science and Technology Corporation (JST).
References
- Rouviere-Yaniv, J., Yaniv, M. and Germond, J.E. (1979) E. coli DNA Binding Protein HU Forms Nucleosome Like Structure with Circular Double-Stranded DNA. Cell, 17, 265-274. http://dx.doi.org/10.1016/0092-8674(79)90152-1
- Hecht, R.M., Taggart, R.T. and Pettijohn, D.E. (1975) Size and DNA Content of Purified E. coli Nucleoids Observed by Fluorescence Microscopy. Nature, 253, 60-62. http://dx.doi.org/10.1038/253060a0
- Ishihama, A. (1999) Modulation of the Nucleoid, the Transcription Apparatus and the Translation Machinery for Stationary Phase Survival. Genes to Cells, 4, 135-143.
- Stonington, O.G. and Pettijohn, D.E. (1971) The Folded Genome of Escherichia coli Isolated in a Protein-DNA-RNA Complex. Proceedings of Natural Academy of Sciences of the United States of America, 68, 6-9. http://dx.doi.org/10.1073/pnas.68.1.6
- Makinoshima, H., Nishimura, A. and Ishihama, A. (2002) Fractionation of Escherichia coli Cell Populations at Different Stages during Growth Transition to Stationary Phase. Molecular Microbiology, 43, 269-279. http://dx.doi.org/10.1046/j.1365-2958.2002.02746.x
- Jishage, M. and Ishihama, A. (1997) Variation in RNA Polymerase Sigma Subunit Composition within Different Stocks of Escherichia coli W3110. Journal of Bacteriology, 179, 959-963.
- Almiron, M., Link, A.J., Furlong, D. and Kolter, R. (1992) A Novel DNA-Binding Protein with Regulatory and Protective Roles in Starved Escherichia coli. Genes & Development, 6, 2646-2654. http://dx.doi.org/10.1101/gad.6.12b.2646
- Ali Azam, T., Iwata, A., Nishimura, A., Ueda, S. and Ishihama, A. (1999). Growth Phase-Dependent Variation in Protein Composition of the Escherichia coli Nucleoid. Journal of Bacteriology, 181, 6361-6370.
- Martinez, A. and Kolter, R. (1997) Protection of DNA during Oxidative Stress by the Nonspecific DNA-Binding Protein Dps. Journal of Bacteriology, 179, 5188-5194.
- Talukder, A.A. (2005) Survival and Death in Bacteria. In: Yamada, M., Ed., Structure and Composition of Escherichia coli Nucleoid, Research Signpost, Kerala, Vol. 2, 77-101.
- Ali Azam, T., Hiraga, S. and Ishihama, A. (2000) Two Types of Localization of the DNA-Binding Proteins within the Escherichia coli Nucleoid. Genes to Cells, 5, 613-626. http://dx.doi.org/10.1046/j.1365-2443.2000.00350.x
- Ali Azam, T. and Ishihama, A. (1999) Twelve Species of the Nucleoid-Associated Protein from Escherichia coli. Sequence Recognition Specificity and DNA Binding Affinity. Journal of Biological Chemistry, 274, 33105-33113. http://dx.doi.org/10.1074/jbc.274.46.33105
- Grant, R.A., Filman, D.J., Finkel, S.E., Kolter, R. and Hogle, J.M. (1998) The Crystal Structure of Dps, a Ferritin Homolog That Binds and Protects DNA. Nature Structural Biology, 5, 294-303. http://dx.doi.org/10.1038/nsb0498-294
- Kim, J., Yoshimura, S.H., Hizume, K., Ohniwa, R.L, Ishihama, A. and Takeyasu, K. (2004) Fundamental Structural Units of the Escherichia coli Nucleoid Revealed by Atomic Force Microscopy. Nucleic Acid Research, 32, 1982-1992. http://dx.doi.org/10.1093/nar/gkh512
- Kornberg, T., Lockwood, A. and Worcel, A. (1974) Replication of the Escherichia coli Chromosome with a Soluble Enzyme System. Proceedings of the National Academy of Sciences of the United States of America, 71, 3189-3193. http://dx.doi.org/10.1073/pnas.71.8.3189
- Murphy, L.D. and Zimmerman, S.B. (1997) Isolation and Characterization of Spermidine Nucleoids from Escherichia coli. Journal of Structural Biology, 199, 336-346. http://dx.doi.org/10.1006/jsbi.1997.3884
- Ball, C.A., Osuna, R., Ferguson, K.C. and Johnson, R.C. (1992) Dramatic Changes in Fis Levels upon Nutrient Upshift in Escherichia coli. Journal of Bacteriology, 174, 8043-8056.
- Sato, Y.T., Watanabe, S., Kenmotsu, T., Ichikawa, M., Yoshikawa, Y., Teramoto, J., Imanaka, T., Ishihama, A. and Yoshikawa, K. (2013) Structural Change of DNA Induced by Nucleoid Proteins: Growth Phase-Specific Fis and Stationary Phase-Specific Dps. Biophysics Journal, 105, 1037-1044. http://dx.doi.org/10.1016/j.bpj.2013.07.025
- Pettijohn, D.E. (1996) The Nucleoid. In: Neidhardt, F.C., et al., Eds., Escherichia coli and Salmonella, American Society for Microbiology Press, Washington DC, 158-166.
- Ghosh, S., Mallick, B. and Nagaraja, V. (2014) Direct Regulation of Topoisomerase Activity by a Nucleoid-Associated Protein. Nucleic Acids Research, 42, 11156-11165. http://dx.doi.org/10.1093/nar/gku804
- Ishihama, A. (2009) The Nucleoid: An Overview. In: Neidhardt, F.C., et al., Eds., Escherichia coli and Salmonella, 2nd Version, American Society for Microbiology Press, Washington DC.
- Cournac, A. and Plumbridge, J. (2013) DNA Looping in Prokaryotes: Experimental and Theoretical Approaches. Journal of Bacteriology, 195, 1109-1119. http://dx.doi.org/10.1128/JB.02038-12
- Wolf, S.G., Frenkiel, D., Arad, T., Finkel, S.E., Kolter, R. and Minsky, A. (1999) DNA Protection by Stress-Induced Biocrystallization. Nature, 400, 83-85. http://dx.doi.org/10.1038/21918
- Meyer, A.S. and Grainger, D.C. (2013) The Escherichia coli Nucleoid in Stationary Phase. Advance in Applied Microbiology, 83, 69-86. http://dx.doi.org/10.1016/B978-0-12-407678-5.00002-7
- Ohniwa, R.L., Muchaku, H., Saito, S., Wada, C. and Morioka, K. (2013) Atomic Force Microscopy Analysis of the Role of Major DNA-Binding Proteins in Organization of the Nucleoid in Escherichia coli. PLoS ONE, 8, e72954. http://dx.doi.org/10.1371/journal.pone.0072954
NOTES
*Corresponding author.