Open Journal of Genetics
Vol.4 No.1(2014), Article ID:44039,7 pages DOI:10.4236/ojgen.2014.41008
Prevalence of Human Leukocyte Antigen HLA-B*5701 in HIV-1 Infected Individuals in Brazil
Claudinéia de Araújo1*, Cristina Valetta de Carvalho1, Miriam Estela de Souza Freire2, Amanda Yamaguti3, Ivens Cuiabano Scaff4, Fernando José de Souza5, Flávia Galindo Silvestre Silva5, Ricardo Sobhie Diaz1, Ismael Dale Cotrim Guerreiro da Silva1
1Department of Gynecology, Molecular Gynecology Laboratory, Federal University of São Paulo, São Paulo, Brazil
2State Department of Health of Mato Grosso, Cuiabá, Brazil
3Unirondon University, Cuiabá, Brazil
4Infectious Diseases Division, Federal University of Mato Grosso, Cuiabá, Brazil
5Imunogenetics Laboratory, General University Hospital, Cuiabá, Brazil
Email: *claudiadna@gmail.com.br
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/


Received 10 February 2014; revised 2 March 2014; accepted 23 March 2014
ABSTRACT
This study was designed to establish the prevalence of HLA-B*5701 at HIV-1 infected individuals in Brazil. A total of 517 consecutive individuals were followed in this study from February 2009 through July 2011. The presence of HLA-B*5701 was determined by Nested-PCR with HLA-B*57 and HLA-B*5701 sequence-specific primers (PCR-SSP). The expression of HLA-B*57 was negative in the 385 (74.5%) and positive in the 103 (19.9%s) of infected individuals. Among these, the expression of HLA-B5701 was positive in the 29 (5.6%) of individuals. No demographic or ethnic differences were found between HLA-B*57/HLA-B*5701 HIV-1 negative patients, with a prevalence of Caucasians (57.1%) individuals. During the period of study, 68 patients were submited to an abacavir containing regimen. The HLA-B*5701 allele was observed in 7 (10.3%) patients, with a significant incidence of Hypersensitivity reactions at 4 of them (p < 0.001). Conclusions: Although Brazilian population consists of a mixture of individuals of Caucasian, African and Native American genetic background, prevalence of HLA-B*5701 in this population is similar to the one found in pure Caucasians.
Keywords:Immunogenetics; Human Immunodeficiency Virus; HLA-B*5701 Antigen Human; Abacavir

1. Introduction
Two and a half decades after the description of the first cases of Acquired Immunodeficiency Syndrome (AIDS), the human immunodeficiency virus remains as a major public health problems in Brazil and worldwide. In the present days, a significant progress to determine the immunogenetic factors that influence the progression of the disease provide new knowledge about virus-host interaction. The development of components of anti-HIV/AIDS multitherapy by drugs dramatically reduced mortality and morbidity related to the AIDS epidemic in many countries. Therefore, HIV/AIDS began to be treated as a manageable chronic condition and no longer a fatal disease. The rapid progress in the development of antiretroviral treatment (ART) by drugs that act by blocking the action of enzymes are important to the functioning of HIV replication and led to the introduction in 1996 of highly active antiretroviral treatment (HAART), which is a combination of at least three drugs, usually able to almost completly suppress the HIV replication [1] .
The effectiveness of HAART has been demonstrated in several clinical trials and observational studies of HIVinfected populations and provides opportunities to supplement and complement treatment efficacy in two ways. Firstly at the individual level and secondly at the population level, that evaluate rates mortality before this therapy. Despite of their high efficacy, the use of these drugs can promote a number of adverse events. It is estimated that 5% to 15% of patients treated with some of this drugs develops adverse reactions [2] . The low commitment to the treatment leads to the emergence of viral strains resistant to drugs that are very hard to treat, requiring careful monitoring of patients to have access to a response of treatment [3] .
Adverse reactions to drugs are significant complications in drug therapy. Previous studies showed that patients undergoing HAART usually have many side effects [4] [5] . Allergies are most common during therapy for HIV and occur with all non-nucleoside reverse transcriptase inhibitors (NNRTIs), as well as one of the nucleoside analogue, Abacavir, and the protease inhibitors (PIs) Aprenavir and Atazanavir.
As a result, the treatment of HIV infection has become somewhat complicated between the benefits of a durable suppression of virus and the risks of toxicity of drugs. Nearby 25% of patients discontinue therapy within the first year of HAART due to side effects 5. There is about the same number of patients not taking the recommended doses of their medication because of concerns about their secondary effect. Interestingly, patients who report significant adverse effects are most often non-adherent to therapy [6] -[8] .
Abacavir (GW1592®) is a guanidine nucleoside analogue, which blocks the formation of genetic materials and inhibits virus replication. This reverse transcriptase inhibitor with activity against human immunodeficiency virus is available as a single drug, as part of combination pills of Abacavir and Lamivudine or combination pill of Zidovudine, Lamivudine and Abacavir. In all cases, this medicine has proved highly effective, but long term treatment may develop a toxicity profile. Hipersensitivity is the most important adverse effect of Abacavir that limits their use in therapy for HIV-positive patients. It is estimated that approximately 5% - 8% of patients are affects during the first six weeks of treatment. This numbers raises the level of medical supervision 9. Reactions to Abacavir (ABC HSR) also include combinations of fever, rash, headache, lethargy, myalgia and respiratory symdrome. Gastrointestinal disorders, as vomiting, diarrhea and abdominal pain, are further adverse effects of Abacavir. Respiratory symptoms such as breathlessness, cough and sputum are rare. Laboratorial characteristics as changes in blood counts, increased liver transaminases, alkaline phosphatase, creatinine and LDH have been shown to be associated with HSR. These symptoms become more severe and may even be fatal, especially in reexposure to drug [5] [9] . Advances to understand the molecular basis of diseases could be used to diagnosis, prevention and treatment of illness. The rate of likely hypersensitivity reaction is the main reason for early discontinuation of Abacavir use. The development of this hypersensitivity in patients infected with the HIV virus is not yet fully understood. Genetic tests to detect HLA-B*5701 status allow the exclusion of use of Abacavir in patients with this allele and contributes to decreased incidence of hypersensitivity among individuals with this virus [10] -[15] .
Brazil is a developing country that has successfully implemented antiretroviral treatments and monitoring tests, through its public health system, for all HIV infected individuals according to local guideline. In this study, we seek to determine prevalence of HLA-B*5701 in HIV positive patients under treatment in Mato Grosso State, located at the Central region of Brazil, to then establish the efficacy of screening of this allele to prevent the hypersensitivity reaction to Abacavir.
2. Materials and Methods
A total of 517 were consecutively recruited from outpatient’s clinics of the State Reference Center in High and Medium Complexity-CERMAC and forwarded to other medical care unit of Mato Grosso State. All the patients were being treated with antiretroviral drugs and gave written informed consent.
The prevalence of clinically suspected Abacavir hypersensitivity was investigated in all HIV-infected patients that received Abacavir therapy from February 2008 through July 2010. Because the HLAB*5701 screening test was not routinely performed at CERMAC until this study, HLA-B*5701 was tested in all patients already submitted to the Abacavir therapy in this period. Clinically suspected Abacavir hypersensitivity was defined with the occurrences of two or more of the following symptoms within six weeks after Abacavir treatment starting. Fever, rash, gastrointestinal disorders or respiratory symptoms that disappeared within three days after discontinuation of Abacavir treatment and that could not be explained by other causes.
Samples from each patient (200 mL) were collected with EDTA and genomic DNA extracted using GFXTM Genomic DNA Purification kit (Amersham Biosciences/GE®). The presence of HLA-B*5701 allele was determined by Nested-PCR with specific primers to HLA-B*57 and HLA-B*5701 as previously described [13] . Multiplex PCR was performed in a reaction mix (50 μl) containing 5 μl of 10× PCR buffer (Invitrogen®), 200 μM dNTP (GE Healthcare®), 2.5 U of Hot Start Taq polymerase (Quiagen®), 2.0 mM MgCl2, 5μl of DNA, and 0.2 μM of each primer HLAB*57 (1F:5’GTCTCACATCATCCAGG+T), (4R:5’CGCCTCCCACTTGCGCTGG+G) and primers: HGH-I (5’-CAGTGCCTTCCCAACCATTCCCTTA-3’) and HGH-II (5’ATCCACTCACGGATTTCTGTTGTGTTTC-3’) amplifying a region of 439 bp. This region was incorporated to ascertain DNA quality and uniform assay conditions: DNA samples from donors who had HLA-B*5701 were used as positive controls. PCR System 9600 (Applied Biosystems®), using an initial denaturation step at 96˚C for one minute followed by four cycles of 96˚C for 25 seconds, 70˚C for 45 seconds and 72˚C for 45 seconds; 24 cycles of 96˚C for 25 seconds, 65˚C for 50 seconds 72˚C for 45 seconds; eight cycles of 96˚C for 25 seconds, 55˚C for one minute, 72˚C for one minute. Cycling was completed by a final elongation step at 72˚C for seven minutes. The reaction products were determined by ethidium bromide staining (10 μg/mL) and examined by UV light. The presence of HLA-B*5701 allele was indicated by the size of the resultant product that also contained the HGH sequence (439 bp), the HLA-B57 subtype (193 bp) and the HLA-B*5701-specific amplification products (112 bp).
Statistical analysis was performed using the (SPSS) Statistical Package for Social Sciences (v14.0). For frequencies and genotypic calculation and allelic proportions we used the chi-square test and Fisher’s exact test. The value of statistical significance was set at 5% (p < 0.05).
3. Results
A total of 517 patients were enrolled and randomly assigned to a study group. Of these, 385 (74.5%) were negative for HLA-B*57; 103 (19.9%) were positive for HLAB*57 and 29 (5.6%) were positive for HLA-B*5701 (Table 1).
Data of distribution by age, sex and racial are presented in Table 2. Of the HLA-B*57 and HLA-B*5701 negative patients, the average age was 40 years; 205 (53.2%) were male and 242 (62.9%) were Caucasians. Among the patients positive for allele HLAB*57, the average age were 38 years; 47 (45.6%) were male and 64 (62.1%) were Caucasians. Of the patients positive for allele HLA-B*5701, the average age was 40 years; 17 (58.6%) were men and 15 (57.7%) were Caucasian. During the study period, an Abacavir containing regimen was administered to 68 patients. The HLA-B*5701 allele was found in seven (10.3%) individuals and the hypersensitivity reaction clinically diagnosed in four of these patients, with a statistically significant incidence (p < 0.001) (Figure 1).
Table 1. Frequency of the allele HLA in positive HIV patients.
N = Number of patients.
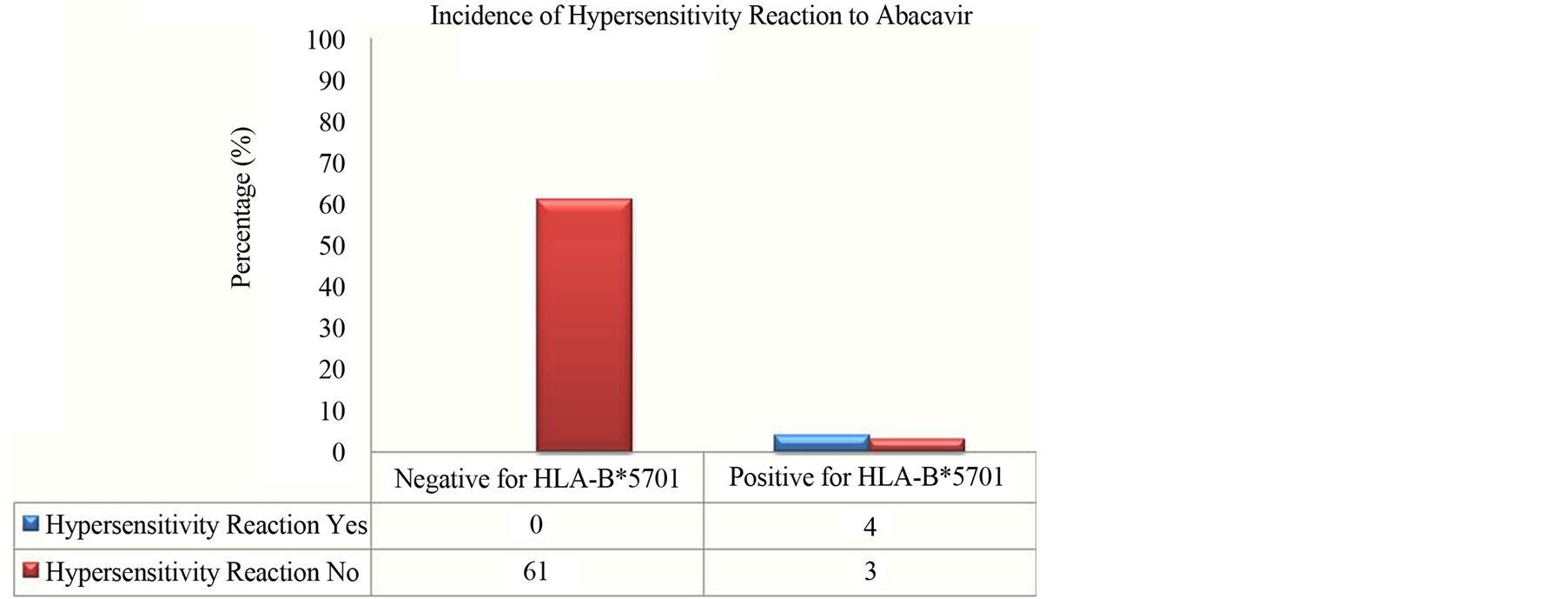
Figure 1. Incidence of hypersensitivity reaction to Abacavir in patients positive for HLA-B*5701.
Table 2. Baseline characteristics of the study poulation.
4. Discussion
Hypersensitivity reactions to drugs are commonly reported with a variety of frequency, severity, and clinical manifestations. Some studies have suggested the presence of familial clustering of drug hypersensitivity as an important genetic predisposition for adverse reactions.
As related to different forms of drugs response, predisposition to drug hypersensitivity reactions is a multifactorial and multigenic process [13] . The present study demonstrated that HLAB*5701 as well the hypersensitivity were significantly frequent (p < 0.001) at HIV-infected Brazilian individuals.
Several evidence strongly suggest the association between occurrence of hypersensitivity reaction to Abacavir and the HLA-B*5701 allele [14] -[16] . The involvement of genetic factors in determining susceptibility to Abacavir hypersensitivity has been previously reported in a multisystem inflammatory syndrome that affected a specific proportion of susceptibility of Abacavir-treated patients during earliest phases of treatment [13] -[20] . Therefore, the identification of individuals harboring the HLA-B*5701 may help in identifying the individuals that will present a slow pace of disease progression [15] .
There are also functional considerations that influence the widespread implementation of a pharmacogenetic approach to Abacavir prescription. The main of these, is the diagnostic methods to HLA-B*5701 that involve the use of a molecular typing technique for determining HLA alleles within the B17 serological family (e.g. HLA-B*5701, HLA-B*5702, HLA-B*5703 and HLA-B*5801).
All high resolution typing assays designed for this purpose must therefore achieve appropriate specificity and must be rigorously and continually exposed to quality assurance processes to ensure that HLA-B*5701 is accurately diagnosed owing to the potential harm that may caused by prescribing Abacavir to a patient who carries the HLA-B*5701 allele but who has been given a negative test result. In addition, systems must be put into place where the information provided by such screening is available to be evaluated before a drug prescription is filled by hospital pharmacies [7] [9] [19] . In Western Australia, the pharmacy system databases provide the information about results of pharmacogenetic testing to the allergy field to ensure that Abacavir will not distribute without prior explicit knowledge of risks and consent of the treating clinician.
As described in a recent review of this topic, screening should promote a more intelligent pharmacovigilance that incorporates knowledge of the genetic screening result along with ongoing monitoring for evidence of drug hypersensitivity in Abacavir-treated patients [21] .
Initially, it was found that the HLA-B*5701, HLA-DR7 and HLA-DQ3 haplotype was associated with Abacavir hypersensitivity [10] [22] . The 3.4% rate of clinically diagnosed hypersensitivity reaction that was not immunologically confirmed in the prospective-screening group is similar to the false positive rates of hypersensitivity reaction (2% to 7%) among patients not receiving Abacavir in double-blind comparative treatment studies [3] [14] [23] . Supporting this notion is the 2% - 7% rates of hypersensitivity reaction to Abacavir that have been reported consistently in the non-Abacavir therapy arms in randomized, double-blinded controlled studies [11] [18] [20] .
Analysis of Polish HIV-infected patients, showed the HLA-B*5701 variant at 11 of 234 (4.7%). In another study at United Kingdom, 54 patients (7.3%) from a total 739 presented HLA-B*5701 status [21] . Three recent cohort studies [5] [11] [24] reported that screening for HLA-B*5701 allele significantly reduces the incidence of hypersensitivity reaction to Abacavir. A Randomized Evaluation of DNA Screening in a Clinical Trial (PREDICT)-1 showed that the absence of HLA-B*5701 allele had a negative predictive value of 100% for skin patch-test confirmed hypersensitivity reaction to Abacavir, and negative predictive value of 96% for clinically suspected hypersensitivity reaction to Abacavir only adverse reactions were reported to be less severe [11] [23] [25] . A study with patients from Taiwan, showed that the occurrence of Abacavir hypersensitivity was only 0.9% in HIV-infected patients [6] . Furthermore, a studied have indicated that prospective screening to exclude patients with HLA-B*5701 allele to receive Abacavir therapy results in a decrease in the incidence of hypersensitivity reaction to Abacavir [17] .
The frequency of allele HLA-B*5701 varies by ethnic group, therefore observations of an increased risk of hypersensitivity reaction to Abacavir in patients of Caucasian versus African descent is likely related to higher expression of HLA-B*5701 in Caucasians [11] [15] [16] . Indeed, our analysis showed that 62.9% of the patients HLA-B*5701 positive were Caucasian. Interestingly, although the Brazilian population consist of a mixture of Caucasians, African, and Native American genetic background [20] , the frequency of HLA-B*5701 detected in our study was similar at the same proportion as seen in the pure Caucasian individuals. Even more, in a recent study 41 (5.7%) of 725 individuals, were HLA-B*5701 positive. Among these, 7.2% were caucasian, 2.8% blacks and 5.6% of other ethnic group. The retrospective study of hypersensitivity to Abacavir and Pharmacogenetic Evaluation study, conducted in 130 caucasian and 69 black individuals from the USA with a previous diagnosis of hypersensitivity reaction to Abacavir, demonstrated that 42 caucasian and five Black individuals with positive skin patch (47/199), 100% carried the HLA-B*5701 allele [8] . Given the potential severity of Abacavir hypersensitivity’s, it has been suggested that only a test that is 100% of predictive success in all populations would be clinically useful. However, it has to be conceded that even the current test characteristics are striking and exceed those of many other tests that are widely used today, and perhaps we may have to be more modest in our requirements to introduce pharmacogenetics into clinical practice [8] .
Several observation from a cost-effectiveness study [15] -[17] on the basis of retrospective data from three study cohorts support that HLA-B*5701 allele may be useful in defining Abacavir Hypersensitivity reaction. The model utilized in this analysis incorporates estimates of the HLA-B*5701 test’s sensitivity and specificity and included the incremental cost of avoiding an HSR as the economic outcome, taking into account the probability of HSR, the probability of testing positive for HLA-B*5701, the costs of treatment of Abacavir HSR, the costs of the Abacavir containing regimen and the costs of alternative regimens required if a positive test occurred. They concluded that pre-treatment screening would be a cost-effective use of health-care resources [7] .
The ability to identify patients at risk of a severe adverse event as a result of HAART or those that may gain a specific benefit from a treatment modality is the current goal of pharmacogenetics within the field of HIV medicine. The pharmacogenetic screening for the HLA-B*5701 allele in targeted populations has the potential to significantly improve HIV-1 patient care by allowing for a more informed use of Abacavir treatment [24] .
After colonization, miscegenation between Amerindians, Africans and Europeans, originated the Brazilian population, one of the most heterogeneous in the world [26] . According to IBGE (Brazilian Institute of Geography and Statistics), the percentage distribution of the second Brazilian population ethnicity is distributed in 47.7% self-reported white, 7.6% black, 43.1% brown, yellow and 1.1% only 0.4% called themselves indigenous. The Mato Grosso State shows a considerable contingent of brown and white, 52.76% and 37.22% respectively. These data are based on self-designation of ethnic factors. Therefore, the grouping of individuals according to phenotypic or social characteristics is extremely complicated due to the high level of miscegenation presented for most of people, the example of what happens to the people of our state and has been demonstrated by several authors that are most common intra differences than inter-population in the various groups have studied molecular markers indicative of ancestry in Brazil [26] [27] . Accordingly that, we are conducting a study with emphasis on population genetics, which focuses on the biological variability of the same subjects assessed, seeking to fill this important gap present in this scientific study. This is the first report describing the HLA-B*5701 profile among Brazilians. Furthermore, in the lights of the actual prevalence of HLA-B*5701 found here, we understand that it is of great importance to make this test available through the Brazilian Public system to HIV-1 infected individuals candidates to use Abacavir.
Acknowledgements
This study was supported by grants from Research Support Foundation of the State of Mato Grosso (FAPEMAT) and National Counsel of Technological and Scientific Development (CNPq). We thank State Department of Health of the State of Mato Grosso (SES/MT) for assistance.
References
- Demoly, P. and Gomes, E.R. (2005) Drug Hypersensitivities: Definition Epidemiology and Risk Factors. Allergy and Immunology, 37, 202-206.
- Carr, A. and Cooper, D.A. (2000) Adverse Effects of Antiretroviral Therapy. Lancet, 356, 1423-1430. http://dx.doi.org/10.1016/S0140-6736(00)02854-3
- Pirmohamed, M. and Back, D.J. (2001) The Pharmacogenomics of HIV Therapy. The Pharmacogenomics Journal, 1, 243-253. http://dx.doi.org/10.1038/sj.tpj.6500069
- Vidal, F., Gutiérrez, F., Gutiérrez, M., Olona, M., Sánchez, V., Mateo, G., et al. (2010) Pharmacogenetics of Adverse Effects Due to Antiretroviral Drugs. AIDS, 12, 15-30.
- Mallal, S., Phillips, E., Carosi, G., Molina, J.M., Workman, C., Tomazic, J., et al. (2008) HLA B*5701 Screening for Hypersensitivity to Abacavir. The New England Journal of Medicine, 358, 568-579. http://dx.doi.org/10.1056/NEJMoa0706135
- Young, B., Squires, K., Patel, P., Dejesus, E., Bellos, N., Berger, D., et al. (2008) First Large, Multicenter, Open-Label Study Utilizing HLA-B*5701 Screening for Abacavir Hypersensitivity in North America. AIDS, 20, 1673-1675. http://dx.doi.org/10.1097/QAD.0b013e32830719aa
- Park, W.B., Choe, P.G., Song, K.H., Lee, S., Jang, H.C., Jeon, J.H., et al. (2009) Should HLA-B*5701 Screening Be Performed in Every Ethnic Group before Starting Abacavir? HIV/AIDS, 48, 365-367.
- Martin, A., Nolan, D., Gaudieri, S., Almeida, C.A., Nolan, R., James, I., et al. (2004) Predisposition to Abacavir Hypersensitivity Conferred by HLA-B*5701 and a Haplotypic Hsp70-Hom Variant. Proceedings of the National Academy of Sciences of the United States of America, 101, 4180-4185. http://dx.doi.org/10.1073/pnas.0307067101
- Martin, A.M., Nolan, D. and Mallal, S. (2005) HLA-B*5701 Typing by Sequence-Specific Amplification: Validation and Comparison with Sequence-Based Typing. Tissue Antigens, 65, 571-574. http://dx.doi.org/10.1111/j.1399-0039.2005.00401.x
- Pirmohamed, M. (2006) Genetic Factors in the Predisposition to Drug-Induced Hypersensitivy Reactions. AAPS Journals, 8, E20-E26. http://dx.doi.org/10.1208/aapsj080103
- Hughes, C.A., Foisy, M.M., Dewhurst, N., Higgins, N., Robinson, L., Kelly, D.V., et al. (2008) Abacavir Hypersensitivity Reaction: An Update. Annals of Pharmacotherapy, 42, 387-396. http://dx.doi.org/10.1345/aph.1K522
- Lucas, A., Nolan, D. and Mallal, S. (2007) HLA-B*5701 Screening for Susceptibility to Abacavir Hypersensitivity. Journal of Antimicrobial Chemotherapy, 59, 591-593. http://dx.doi.org/10.1093/jac/dkl557
- Phillips, E.J. (2006) Genetic Screening to Prevent Abacavir Hypersensitivity Reaction: Are We There Yet. Clinical Infectious Diseases, 43, 103-105. http://dx.doi.org/10.1086/504878
- Mallal, S., Nolan, D., Witt, C., Masel, G., Martin, A.M., Moore, C., et al. (2002) Association between Presence of HLA-B*5701, HLA-DR7 and HLA-DQ3 and Hypersensitivity to HIV-1 Reverse-Transcriptase Inhibitor Abacavir. Lancet, 359, 727-732. http://dx.doi.org/10.1016/S0140-6736(02)07873-X
- Phillips, E., Rauch, A., Nolan, D., Martin, A., Almeida, C., Knowles, S., et al. (2007) Genetic Characterization of Patients with MHC Class I Mediated Abacavir Hypersensitivity Reaction. 4th IAS Conference on HIV Pathogenesis, Treatment and Prevention, Sydney, 22-25 July 2007, Abstract No. MOPEB001.
- Phillips, E.J., Wong, G.A., Kaul, R., Shahabi, K., Nolan, D.A., Knowles, S.R., et al. (2005) Clinical and Immunogenetic Correlates of Abacavir Hypersensitivity. AIDS, 19, 979-981. http://dx.doi.org/10.1097/01.aids.0000171414.99409.fb
- Parczewski, M., Leszczyszyn-Pynka, M., Wnuk, A., Urbañska, A., Fuksiñska, K., Bander, D., et al. (2010) Introduction of Pharmacogenetic Screening for the Human Leucocyte Antigen (HLA) B*5701 Variant in Polish HIV-Infected Patients. HIV Medicine, 11, 345-348. http://dx.doi.org/10.1111/j.1468-1293.2009.00787.x
- Waters, L.J., Mandalia, S., Gazzard, B. and Nelson, M. (2007) Prospective HLA-B*5701 Screening and Abacavir Hypersensitivity: A Single Centre Experience. AIDS, 21, 2533-2549. http://dx.doi.org/10.1097/QAD.0b013e328273bc07
- Rauch, A., Nolan, D., Martin, A., McKinnon, E., Almeida, C. and Mallal, S. (2006) Prospective Genetic Screening Decreases the Incidence of Abacavir Hypersensitivity Reactions in the Western Australian HIV Cohort Study. Clinical Infectious Diseases, 43, 99-102. http://dx.doi.org/10.1086/504874
- Zucman, D., Truchis, P., Majerholc, C., Stegman, S. and Caillat-Zucman, S. (2007) Prospective Screening for Human Leukocyte Antigen-B*5701 Avoids Abacavir Hypersensitivity Reaction in the Ethnically Mixed French HIV Population. Journal of Acquired Immune Deficiency Syndromes, 45, 1-3. http://dx.doi.org/10.1097/QAI.0b013e318046ea31
- Hammond, E., Mamotte, C., Nolan, D. and Mallal, S. (2007) HLA-B*5701 Typing: Evaluation of an Allele-Specific Polymerase Chain Reaction Melting Assay. Tissue Antigens, 70, 58-61. http://dx.doi.org/10.1111/j.1399-0039.2007.00840.x
- Saag, M., Balu, R., Phillips, E., Brachman, P., Martorell, C., Burman, W., et al. (2008) High Sensitivity of Human Leukocyte Antigen-b*5701 as a Marker for Immunologically Confirmed Abacavir Hypersensitivity in White and Black Patients. Clinical Infectious Diseases, 46, 1111-1118. http://dx.doi.org/10.1086/529382
- Sun, H.Y., Hung, C.C., Lin, P.H., Chang, S.F., Yang, C.Y., Chang, S.Y., et al. (2007) Incidence of Abacavir Hypersensitivity and Its Relationship with HLA-B*5701 in HIV-Infected Patients in Taiwan. Journal of Antimicrobial Chemotherapy, 60, 599-604.
- Hughes, S., Parry-Billingsc, K., Givens, N., Wong, D., Loeschel, E., Parada, P., et al. (2006) Genetic Screening Test for Abacavir Hypersensitivity—PREDICT-1: Description of a Novel Randomised Prospective Study Design to Determine the Clinical Utility of Prognostic Screening for HLA-B*5701 in 8th International Congress Drug Therapy in HIV Infection. Glaugow, Scotland UK.
- Orkin, C., Wang, J., Bergin, C., Molina, J.M., Lazzarin, A., Cavassini, M., et al. (2010) An Epidemiologic Study to Determine the Prevalence of the HLA-B*5701 Allele among HIV-Positive Patients in Europe. Pharmacogenetics and Genomics, 20, 307-314. http://dx.doi.org/10.1097/FPC.0b013e3283390666
- Bonilla, C., et al. (2004) Admixture in the Hispanics of the San Luis Valley, Colorado, and Its Implications for Complex Trait Gene Mapping. Annals of Human Genetics, 68, 139-153.
NOTES

*Corresponding author.


