 American Journal of Anal yt ical Chemistry, 2011, 2, 639-649 doi:10.4236/ajac.2011.26073 Published Online October 2011 (http://www.SciRP.org/journal/ajac) Copyright © 2011 SciRes. AJAC Targeting Divalent Metal Ions at the Active Site of the HIV-1 RNase H Domain: NMR Studies on the Interactions of Divalent Metal Ions with RNase H and Its Inhibitors Jiangli Yan1*, Haihong Wu1, Tiffany Tom1, Oleg Brodsky1, Karen Maegley2 1Oncology Chemistry Pfizer Global Research and Development, La Jolla Laboratories, Science Center Drive, San Diego, USA 2Oncology Research Unit, Pfizer Global Research and Development, La Jolla Laboratories, Science Center Drive, San Diego, USA E-mail: jiangli.yan@pfizer.com Received June 27, 2011; revised July 3, 2011; accepted August 22, 2011 Abstract HIV-1 reverse transcriptase (RT) RNase H (HIV-RH) is a key target of anti-AIDS drugs. Metal-chelating compounds are an important class of chemicals in pharmacological drug discovery, especially in relation to HIV-RT and the highly-related HIV-integrase. The correlation between the metal-chelating properties and enzyme activities of the metal chelators is always of high scientific interest, as an understanding of this may accelerate the rational optimization of this class of inhibitors. Our NMR data show that Mg2+ and Ca2+ bind specifically to the active site of the RNase H domain and two Mg2+ ions sequentially bind one molecule of RNase H. We also demonstrate here, using saturated and unsaturated tricyclic N-hydroxypyridones designed to block the active site, that the primary binding sites and affinities of divalent metal ions are correlated with the structures of the chelating motifs. Chemical shift perturbation studies of protein/metal-ion/compound ternary complexes also indicate that divalent metal ions play important roles for the specific interaction of the compounds with the RNase H active site. Keywords: Metal Chelation, HIV-1, Reverse Transcriptase, RNase H, NMR Spectroscopy 1. Introduction HIV-1 reverse transcriptase [1] converts single-stranded retroviral RNA into double-stranded DNA, which is in- tegrated into the cellular genome [2-4]. HIV-1 RT is a multifunctional enzyme that has RNA-directed DNA polymerase, DNA-directed DNA polymerase and ribo- nuclease H (RNase H) activities [1]. HIV-1 RT is a het- erodimer composed of two peptide subunits, p66 and p51. The polymerase active site is located at the N-terminus of the p66 subunit, whereas its C-terminal end contains the RNase H active domain [5]. The RNase H domain of HIV-1 RT (HIV-RH) plays a role in many steps of re- verse transcription, such as the generation of an RNA primer for synthesis of the (+)-strand DNA, the degrada- tion of the viral genomic RNA in the intermediate RNA·DNA hybrid, and the removal of host tRNA and plus-strand primers [2,6,7]. HIV-RH forms a central five-stranded β-sheet surrounded by four α-helices. The core domain of the RNase H active site contains a highly-conserved DEDD motif that consists of four acidic residues, D443, E478, D498, and D549 [5,7]. The hydrolysis of the scissile phosphodiester bonds catalyzed by RNase H requires divalent metal ions, preferably Mg2+ [6,8]. However, there has been some controversy regarding the number of metal ions (one or two) involved in the catalysis. Crystallographic studies of both E. coli RNase H and HIV-RH have shown that the active site can bind two Mn2+ ions separated by approximately 4 Å and the authors proposed a two-metal-ion catalytic mechanism [9,10]. The ability of HIV-RH to bind to two metal ions, either two Mn2+ or two Mg2+, has also been confirmed by calorimetry and NMR, respectively [9,11]. However, this result was not supported by the crystal structure of the E. coli enzyme, in which only a single bound Mg2+ ion was identified (even though the crystal was obtained in high concentration of MgSO4) [12]. Re- cently, two Mg2+ ions were also observed ~4 Å apart in  J. L. YAN ET AL. 640 Bh-RNase HC-substrate complexes by Nowotny et al. [6,8]. Later analysis by the same group also strongly supported the two-metal-ion catalytic mechanism [6,13]. The RNase H activity of HIV-RH plays a crucial role in the retroviral life cycle [14]. Defective mutations of two key residues (E478Q and H539F) in the RNase H domain induce a marked reduction in viral proliferation [15], thus making HIV-RH an attractive chemothera- peutical target for anti-HIV drugs [16,17]. Designing metal-chelating compounds that are able to bind two divalent metal ions at the active site is one strategy that enables the direct blocking of the active site. Small N-hydroxyimide analogs that are optimized to bind two divalent metal ions at a 4 Ǻ distance between the ions inhibit HIV-RT activity in vitro with an IC50 < 1.0 uM [10]. Metal-chelating compounds are an important class of chemicals in pharmacological drug design, especially when targeted against proteins with metal ions at their active sites [4]. Metal-binding property of the metal- chelators affects the potency of enzyme inhibitors, and the correlation between metal-chelation affinities and enzyme inhibition is always of interest in the design of metal-chelating inhibitors. In this report, we applied NMR analytical methodol- ogy to study the interactions of the divalent metal ions Mg2+ and Ca2+ with HIV-RH and chemically-engineered metal-chelating compounds. When the pH and ionic strength are carefully maintained, we observed clean chemical shift perturbations at the active site of RNase H with both Mg2+ and Ca2+. We also used 1D 1H-NMR to characterize the interactions between the divalent metal ions and HIV-RH inhibitors. Two chemically similar series, saturated and unsaturated tricyclic N-hydroxypy- ridones have different chelating affinities depending on the structure of the primary metal-binding sites. To un- derstand the correlation between the metal-chelating properties and the inhibition of the enzyme by metal- chelating inhibitors, we also studied the HIV-RH/Mg2+/ inhibitor ternary complexes using a saturated tricyclic N-hydroxypyridone and -thujaplicinol as the tool com- pounds. 2. Material and Methods 2.1. Protein Preparation A cDNA fragment encoding the C-terminal domain of HIV-1 p66 protein (W426 to L560) was cloned into pET-28a in fusion with a C-terminal His6-tag. BL21 (DE3)-AI cells transformed with the recombinant plas- mid were grown at 37˚C in 2 L Celtone-N or Celtone-CN media (Cambridge Isotope Laboratories, Andover, MA) to produce 15N- or 13C/15N-enriched proteins, respec- tively. After three hours, the cultures were allowed to equilibrate to 15˚C - 18˚C. Expression of HIV-RH was then induced using 100 uM IPTG and 0.01% arabinose at 0.7 - 1.0 OD600 cell density. The cells were grown at 15˚C - 18˚C for 16 hours and harvested by centrifugation. The frozen cells were resuspended and lysed in 50 mM Tris-HCl buffer (pH 8.0) containing 250 mM NaCl, 100 ul/L benzonase (Novagen), 0.2 mg/ml lysozyme (Sigma, L-6876), and 0.25 mM TCEP (Pierce). The His6-tagged protein was purified from the cell lysate using immobi- lized metal affinity chromatography (ProBond resin, In- vitrogen). The eluate fraction (step elution, OmniPrep gravity column, BioRad) was further purified using size-exclusion chromatography (100 mL Phenomenex S3000 column) with a mobile phase containing 25 mM 1, 3-bis(tris(hydroxymethyl)methylamino)propane (bis-Tris propane, pH 6.5), 150 mM NaCl, 2% glycerol, and 0.25 mM TCEP. The desired fractions were pooled and con- centrated to 15 - 20 mg/ml using centrifugal concentra- tors (Millipore). 2.2. Backbone Resonance Assignment of HIV-RH All of the experiments for the resonances assignment were recorded at 30˚C on a Bruker Avance 700 MHz spectrometer equipped with a TCI cryo-probe. The NMR sample contained 0.9 mM 13C,15N-labeled HIV-RH in 25 mM bis-Tris-d14 (pH 6.5), 150 mM NaCl, 1 mM dithio- threitol-d6 (DTT) and 5% 2H2O. The backbone reso- nances were assigned using standard triple resonance experiments, including CBCA(CO)NH, HNCACB, HNCA, [18] and HN(CO)CA[19]. Sequential assignments were obtained using TopSpin 2.0 (Bruker Biospin) for spectral processing and SPARKY for sequential analysis [20]. 2.3. Metal Titration with HIV-RH and Its Inhibitors In the HIV-RH titration experiments, 0.5 M MgCl2 or Ca(NO3)2 was titrated into the protein solution. HIV-RH protein (70 uM) was prepared in buffer containing 25 mM bis-Tris-d14 (pH 6.5), 150 mM salt (NaCl for Mg2+ titration and NaNO3 for Ca2+ titration, respectively), 1 mM DTT, and 5% 2H2O. The pH of all reagents used in the titrations was carefully monitored and readjusted to 6.5 (if required) at each step of the titration. 1H-15N HSQC spectra were acquired on an Inova 600 MHz spectrometer (Varian Inc.) at 25˚C to monitor the chemi- cal shift changes as the metal ion concentrations increased. In the titrations with the inhibitors, each sample was prepared independently, using identical buffer conditions (except for the concentration of the metal ions). The final Copyright © 2011 SciRes. AJAC  J. L. YAN ET AL. Copyright © 2011 SciRes. AJAC 641 observed, most likely as a result of regional flexibility and/or signal overlapping. Most of those amino acids, i.e., N474-K476, E514-L517, A538-K540, and V548-L560, were located in loop regions and at the C-terminus. We observed and assigned three amino acids (D443, E478, and D498) of the DDED motif involved in metal binding, except D549 which is located in the flexible C-terminus of the protein. This is consistent with previous reports [22] that the C-terminus of isolated HIV-RH is highly dynamic and can adopt either an α-helical or random coil conformations in crystal structures, depending on the crystallization conditions and the space group [23]. In solution, the C-terminus is usually disordered, but can be stabilized at high concentration of Mg2+ (80 mM) [11,24]. concentration of the compound was 0.20 to 0.50 mM. 1D 1H-NMR experiments were performed using an Inova 600 MHz spectrometer (Varian Inc.) at 25˚C. All spectra were recorded with 128 transients, a 16-ppm sweepwidth using presaturation for water suppression. The NMR data were analyzed using ACD NMR Proc- essor (ACD-Labs, Inc) and the total chemical shift change Δ obs of the 1H-15N cross peak was calculated according to the formula, 22 0.17 obs NH (1) where and were the chemical shift changes in 15N and 1H dimensions, respectively. The co-crystal structure of HIV-RH domain protein [21] was used for chemical shift mapping. We observed that the conformation, stability and ac- tivity of HIV-RH are very sensitive to pH, ionic strength and divalent metal-ion concentration. A drop in the pH of the buffer from 7.0 to 5.0 induced dramatic and global shifts in the 1H-15N HSQC spectrum, and more HSQC signals were detectable at lower pH. Interestingly, most early structural studies of HIV-RH by NMR [25-28] and x-ray crystallography [29-31] were carried out in acidic conditions, suggesting that the isolated HIV- RH domain is more stable at lower pH. 2.4 Calculation of the Dissociation Constants (Kd) In the single binding mode, one metal ion binds with one molecule of protein or metal-chelating inhibitor. When the reaction occurs under the fast exchange conditions, the observed chemical shift ( obs) is the weighted average of the chemical shifts of the free and bound species. Therefore, the observed chemical shift change (Δ obs) is a function of the dissociation constant Kd [2] (Equation (2)), 3.2. Determination of the Binding Constants (Kd) of Divalent Metal Ions with HIV-RH where f and b are the fractions and f and b are the chemical shifts of the free and bound protein or compound, respectively. M0 and L0 are the initial concentrations of the metal ion and the chelator (protein or ligand). The data were fit using GraphPad Prism (GraphPad Software) to determine the Kd from the chemical shift changes. Divalent metal ions are crucial for RNase H activity and contribute to the conformation and stability of the protein, especially at the C-terminus [11,28]. In our NMR bind- ing study of the divalent metal ions to HIV-RH, we chose 25 mM bis-Tris (pH 6.5) containing 150 mM NaCl as our buffer to maintain pH and ionic strength, therefore to mitigate the effects on conformation, stability and ac- tivity of HIV-RH. The buffer was also compatible with those used in the biochemical assays. Every reagent used in the experiment was prepared in the buffer described, and the pH was checked in each step and adjusted when necessary. 3. Results and Discussion 3.1. NMR Assignments of the 1H-15N HSQC Spectra and Flexibility at the C-Terminus of HIV-RH An isolated, C-terminal His6-tagged protein that contains the C-terminal domain of HIV-RT p66 protein (W426- L560) was expressed and purified in its 15N-labeled or 13C/15N-labeled forms for our NMR studies. The protein was mostly well-folded as the 1H-15N HSQC peaks were well resolved and dispersed. Backbone resonance as- signments of most amino acids were obtained from the standard triple resonance experiments (data available if required). The 1H- 15N HSQC peaks of 26 amino acids, not including those of the His6-tag and prolines, were not In the metal titration experiments, we dialyzed 15N- labeled HIV-RH in the described buffer. MgCl2 (1.0 M, prepared in the same buffer) was titrated into the protein solution to increase the Mg2+ concentration from 0 to 80 mM. Approximately 24 peaks showed obvious shifts on the 1H-15N HSQC spectra (Figure 1(a)). The Mg2+ dose- dependent chemical shift changes (∆ ), calculated as weighted sums of the changes in both the 15N 2 0000 00 4 obs obs fb fdd0 2 KLMKLML L (2)  J. L. YAN ET AL. Copyright © 2011 SciRes. AJAC 642 (a) 010 20 30 40 50 60 70 80 9 0 20 40 60 80 100 120 140 160 180 G444 A4 4 5 T477 E4 7 8 Q500 M ++ Conce ntr ation mM Chemical Shift Change (Hz) (b) D549 D443 E478 D498 (c) Figure 1. (a) Overlay of the HSQC spectra shows the chemical shift perturbations of HIV-RH by Mg2+ at 0 mM (purple), 2.5 mM (pink), 5 mM (cyan), and 10 mM (blue). (b) The Mg2+ titration curve of the chemical shift changes vs. the Mg2+ concentration (0, 2.5, 5, 10, 20, 40, and 80 mM) for five selected amino acids. (c) Mapping of the shift per- turbations of HIV-RH by Mg2+ at a concentration of 5.0 mM on the HIV-RH structure. Red: ∆ 50 Hz, pink: ∆ = 35 - 50 Hz, light pink: ∆ ≤ 35 Hz, Yellow: not observed or assigned. and 1H dimensions (Equation (1)), were fitted by Equa- tion (2) using GraphPad Prism (GraphPad Software) as shown in Figure 1(b). It was clear that the titration curves reached a maximum plateau at about 80 mM of Mg2+. The binding constant (Kd) of Mg2+/HIV-RH were determined by the titration data of 17 amino acids that showed ∆ > 10 Hz with at least 6 data points. The cal- culated Kd ranged from 8.4 to 16.6 mM with a goodness of fit (R2) from 0.984 to 0.999. The averaged Kd was 13 4 mM, very similar to the value reported previously [11]. We also carried out titration experiments with step- wise increases in Ca2+ concentrations from 0 to 15 mM in a similar fashion to those using Mg2+. We obtained a very similar pattern of chemical shift perturbations. In the plots of ∆ vs. the Ca2+ concentration, the titration curves reached plateaus at 15 mM Ca2+. The Kd values of 21 amino acids were calculated to be in the range from 1.5 to 3.3 mM, yielding reasonable values of R2 from 0.992 to 1.000. The average Kd for Ca2+ to HIV-RH is 2.9 0.5 mM, indicating that the binding of Ca2+ to HIV-RH is four times tighter than that of Mg2+. Figure 1(c) showed the chemical shift perturbation map of Mg2+ on the HIV-RH domain protein. Amino acids at the catalytic center showed the greatest chemical shift perturbations. All three observable residues in the metal chelating DDED motif, i.e., D443, E478, and D498 stood out in the chemical shift perturbations in- duced by Mg2+. Ca2+ binds to the same site as Mg2+ and amino acids located at the active site also showed the largest chemical shift changes in response to Ca2+. Our NMR titration data demonstrated small and localized chemical shift perturbation patterns in HIV-RH in re- sponse to the divalent metal ions, Mg2+ and Ca2+, when the buffer pH and ionic strength were carefully main- tained. It is clear that Mg2+ and Ca2+ bind specifically to the active site of the RNase H domain, but do not sig- nificantly change the global conformation and dynamics of the protein. However, the C-terminus of our construct is still mostly disordered even at high concentrations of divalent metal ions at the testing buffer conditions. Most of the C-terminal residues of HIV-RH were not stable enough to produce measurable signals in the HSQC spectra in the presence of 80 mM Mg2+ or 15 mM Ca2+. 3.3. Sequential Binding of Two Mg2+ Ions to the Active Site In the Mg2+ titration experiments, a few peaks, including G444, Q500, W535ω, and V536, became broader and broader as the Mg2+ concentration increased. More inter- estingly, they split into two sets of peaks at 80 mM Mg2+ (Figure 2(a)). The splitting of these signals indicated the  J. L. YAN ET AL.643 existence of two slowly-exchanging protein conforma- tions, and the occupancy of the second metal binding site at high Mg2+ concentrations (80 mM or higher). At 80 mM Mg2+, the second metal-binding site was approxi- mately half-occupied by Mg2+, whereas the first metal-binding site was almost fully occupied. Interest- ingly, the affinity of the second Mg2+ ion for HIV-RH was reported to be ~35 mM [11]. Figure 2(b) shows four residues with split HSQC signals at 80.0 mM Mg2+ in the HIV-RH structure. Three of these, Q500, W535ω, and V536, are close to D498, while D444 is on the opposite side of the active site, adjacent to D443. When the con- centration of Mg2+ was 5.0 mM, large chemical shift changes were observed for D443 (Figure 2(c)). Our data suggest that two Mg2+ ions bind sequentially to the active site of HIV-RH. The first Mg2+ binds to site 1 and inter- acts with D443 and E478, whereas the second Mg2+ binds to site 2 and interacts with D498 and D549 with a slightly lower (~3 times) affinity (Figure 2(c)). The weaker metal binding affinity of site 2 is probably due to the flexibility of the C-terminus, where D549 is located. However, the peak splitting was not observed in the Ca2+ titration experiments. The distance between the two Mg2+ ions in the active site is 4 Å, as shown in the crystal structure [8,10,11]. Ca2+ has a much larger atomic radius than Mg2+; therefore, it is unlikely that two Ca2+ ions can 8.40 8.35 8.30 8.25 8.20 108.5 109.0 109.5 110.0 110.5 111.0 7.8 7.7 7.6 7.5 123.5 124.0 124.5 125.0 125.5 126.0 126.5 127.0 E478 1 H, ppm 15 N, ppm 1 H, ppm 15 N, ppm (a) D549 D443 E478 D498 D549 D443 E478 D498 (b) (c) Figure 2. (a) Regions of the HSQC spectra of HIV-RH showing peak shifting with increasing Mg2+ concentrations, at 0, 2.5, 5.0, 10, 20, 40, and 80 mM, in the direction indicated by the arrows. The dashed circles show peak splitting for V536 and G444 and broadening for E478 and G453 at 80 mM Mg2+. (b) Mapping of splitting HSQC signals on the HIV-RH structure. Red: AA with splitting HSQC signals, Yellow: not observed or assigned. (c) Mapping of the shift perturbations of HIV-RH by .0 mM Mg2+ on the HIV-RH structure as shown in Figure 2(c) in a different orientation. 5 Copyright © 2011 SciRes. AJAC  J. L. YAN ET AL. Copyright © 2011 SciRes. AJAC 644 fit simultaneously into the active site. Although we saw evidence of the sequential binding of two Mg2+ ions in our experiments, we were not able to obtain enough data points to determine the binding constant of the second Mg2+ ion through curve fitting. 3.4. Interaction of Metal-Ions to Metal-Chelating Compounds Divalent metal ions, especially Mg2+, are directly in- volved in the catalytic activities of HIV-RH, and two Mg2+ ions bind to one molecule of HIV-RH at the active site. To target the active site, it is a common strategy to design metal-chelating compounds binding to two diva- lent metal ions at the active site [2,16]. The effect of compound/metal binding property on enzyme inhibition is always of interest in the lead generation and optimiza- tion of metal-chelating compounds. In this study, we utilized 1D 1H-NMR experiments to identify the metal- binding sites of our lead compounds through chemical shift perturbations induced by metal ions. Again, similar to the protein/metal interaction, protons that are near the metal-binding site generally experience larger perturba- tions. Based on Equation (2), we determined the binding affinities of the lead compounds to metal ions through chemical shift changes. The sample preparation, data acquisition and data processing were all automated for a rapid measurement of Kd. Derivatives of unsaturated and saturated tricyclic N-hydroxypyridones, shown in Figures 3(a) and (b) re- spectively, were designed and synthesized in-house for targeting HIV-RH. The unsaturated tricyclic N-hydroxy- pyridone derivatives contain a flat and rigid three-ring core and two hypothetical metal-binding motifs (Figure 3(a)). In the Mg2+ titration experiments, all protons on the tricyclic N-hydroxypyridone core showed significant and incremental chemical shift changes as the Mg2+ con- centration increased; the example of compound U-1 is shown in Figure 3(c). Meanwhile, only slight chemical shift changes were observed with the other protons, which were located in the R1 and R2 substitute groups. H-13 gave the largest chemical shift changes (up to 130 Hz) when the Mg2+ concentration changed. Therefore, H-13 is the proton nearest to the Mg2+ ion (Figure 3(a)). Three more unsaturated tricyclic N-hydroxypyridone compounds were tested and H-13 showed the largest chemical shift change in all cases. The chemical shift changes of the core protons at the compound/Mg2+ ratio of 1:20 when the titration curves reached the plateaus were listed in Table 1. The chemical perturbations of the other three core protons H-5, H-8 and H-12 showed only half of the change of H-13. This indicates that Mg2+ preferably binds unsaturated tricyclic N-hydroxy-pyri- done derivatives at site I by interacting with the two oxygen atoms of the O14∸N1∸C2∸O15 motif (Fig- ure 3(a)). The Kd values of the two unsaturated com- pounds (U-1 and U-4) were determined through chemical shift changes, and equivalent with each other within the measurement deviation range. The chelator-Mg2+ com- plexes of the other two compounds (U-2 and U-3) had poor solubility. As the Mg2+ concentration was approxi- mately equivalent to the compound concentration, we saw precipitates in the NMR tubes and the proton signals became too weak to measure. Consequently, we were not able to calculate the Kd due to the lack of points. Inter- estingly, when [Mg2+] was higher than 20 times of the compound concentration, the compound proton signals became stronger and therefore measurable. The similar phenomena were also observed with compound U-1 and U-4. As shown in Figure 3(c), proton signals of U-1 became broader and broader and then sharper and sharper as the Mg2+ concentration increased gradually from 0 to 80 mM. In addition, when [Mg2+] was over 40 times of the compound concentration, the compound proton sig- nals shifted to an opposite direction. We believe this was related to the second metal-ion binding and the ternary complexes (compound/(Mg2+)2) had better solubility than the binary complexes (compound/Mg2+). The chemical shift change “turn-over” occured all at a Mg2+ concentra- tion of 16 - 20 mM, suggesting that with ~20 mM Mg2+, the second metal-binding site is approximately half-oc- cupied by Mg2+, whereas the first metal-binding site was almost fully occupied and therefore the Kd of the second Mg2+ to this series compound was estimated as ~10mM. In our NMR study, we could not obtain sufficient data to determine Kd of the second Mg2+ binding using the dou- ble binding curve fit due to the maximum concentration of MgCl2 in stock solution of 0.5 M. The observations were different when the tricyclic N-hydroxypyridone core was saturated at the car- bon-carbon bond between C-12 and C-13 (Figure 3(b) ). Saturation of the C-C bond makes the six-member-ring more flexible and no longer flat. Figure 3(d) shows the shift of the proton signals of 0.20 mM compound S-5 in the Mg2+ titration measurements. Table 1 lists the chemical shift perturbations of the core protons of four saturated tricyclic N-hydroxypyridones at a concentration ratio of compound: Mg2+ of 1:100 when the ∆ vs. [Mg2+] titration curves reached the plateaus. All four saturated derivatives showed the same interaction mapping, which obviously was different from the mapping of the unsatu- rated analogs. Instead of H-13 in the unsaturated cases, proton H-5 of saturated compounds presented the largest chemical shift change, indicating Mg2+ preferably boun- dat site II and interacts with the nitrogen and oxygen at-  J. L. YAN ET AL.645 oms of the N4∸C3∸C2∸O15 motif when the C12-C13 bond was saturated. In the NMR spectra, only one set of triplet peaks was observed for either of the two H-12 or two H-13 (Figure 3(d)), indicating the equivalence of the 12 2 N 1 13 3 11 5N 4 N 7 6 10 9 8O 15 O 14 R1 R2 Mg2+ I II 12 2 N 1 13 3 11 5N 4 N 7 6 10 9 8O 15 O 14 R1 R2 Mg2+ I II (a) (b) 7.30 7.25 7.20 7.15 7.10 Chemical Shift (ppm) 7.75 7.70 Chemical Shift (ppm) 8.825 8.775 Chemical Shift (ppm) 1 H, ppm 1 H , p pm 1 H , p pm H* H-13 H-12 H* H-5 (c) 8.775 8.725 Chemical Shift (ppm)3.0503.000 2.950 Chemical Shift (ppm)3.450 3.400 Chemical Shift (p... 1 H , p p m 1 H , pp m 1 H , p p m H* H-5 H-13 H-12 (d) Figure 3: (a) The core structure of unsaturated tricyclic N-hydroxypyridones. (b) The core structure of saturated tricyclic N-hydroxypyridones. (c) 1D 1H-NMR spectra of 0.50 mM U-1 with Mg2+ at concentrations of 0.0, 0.05, 0.10, 0.25, 0.50, 1.00, 2.5, 5.0, 10.0, 35.0, and 50.0 mM. (d) 1D 1H-NMR spectra of 0.20 mM S-5 with Mg2+at concentrations of 0.0, 0.2, 0.4, 0.8, 1.6, 3.2, 5.0, 10.0, 20.0, 40.0, and 80.0 mM. R1 and R2 are substituent groups. Two potential metal binding motifs are labeled as I O14∸N1∸C2∸O15 and II N4∸C3∸C2∸O15. Mg2+ prefers binding to saturated tricyclic N-hydroxypyridones at motif II, but to the saturated tricyclic N-hydroxypyridones at I. The arrows indicate the shift of the proton signals as the Mg2+ concentration increases. H* are protons on the R1 or R2 substituents. Copyright © 2011 SciRes. AJAC  J. L. YAN ET AL. 646 -1 to U-4) and saturated (S-5 to S-8) tricyclic N-hydroxypyridones by Chem shift changes ( ), Hz Table 1. Chemical shift perturbations of unsaturated (U Mg2+ chelating. Cpd Cpd: Mg2+ ratio H5 H8 H12 H13 Mg2+ binding Kd, mM RNase H IC50, μM U-1 1:20 38 36 34 130 0.2 0.46 U-2 1:20 46 64 42 152 N.A. N.A. U-3 1:20 56 -- 43 137 N.A. 2.2 U-4 1:20 41 45 38 126 0.2 0.49 S-5 1:100 31 3 15 12 4.4 2.2 S-6 1:100 26 19 21 10 4.5 0.22 S-7 1:100 26 19 20 9 4.4 0.038 13 * * S-8 1:100 33 6 10 2.2 0.29 o protons at C-12 or C-13. The saturated N-hydro- 3.5. Ternary Complex of HIV-RH, Mg2+, and The use of metal chelation as an anchor to the active site -thujaplicinol to the first Mg2+ ion, as determined from und s-7), which has a weak bind- in tw xypyridone ring is relatively flexible and exchanges be- tween chair and boat conformations. Four saturated N- hydroxypyridones had very similar Kd values, in a tight range of 2 - 5 mM (Table 1) which was 10 to 20 times larger than those of the unsaturated analogs. The overall chemical shift changes and the differences between the largest and the other perturbations of the core protons of the saturated compounds were both smaller than those observed with the unsaturated ones. We also observed the sharp-broad-sharp line-shape changes as the Mg2+ concentration increased, but didn’t see the second bind- ing of Mg2+ as well, probably because the Mg2+ concen- tration was not high enough. Metal-Chelating Compounds is a common strategy in designing inhibitors to target HIV-RH and the highly- related HIV integrase enzymes [2,16,32]. The final product of the design is a ternary complex (compound/ metal-ion/protein). Therefore, the correlation between compound/metal or protein/metal binding affinity and enzyme inhibition is always of high interest. Sometimes, metal ions are critical for inhibition by the designed compound. -thujaplicinol, a selective inhibitor of HIV RT, is a typical metal chelator [33]. Our study demonstrated that it binds to two Mg2+ions sequen- tially. Its proton signals incrementally shifted downfield and became broader and broader as the concentration of Mg2+ increased. When the concentration of Mg2+ reached 8 mM, the signals started to shift in the opposite direc- tion. As the Mg2+ concentration increased further, the signals shifted upfield and became narrower. The Kd of the proton chemical shift changes, is 0.64 0.10 mM. Interestingly, -thujaplicinol introduced no chemical shift perturbations in the HSQC signals of the HIV-RH domain when metal ions were absent. On the contrary, in the presence of 8 mM Mg2+, significant chemical shift perturbations were observed in the HSQC spectrum of the HIV-RH protein attributable to 500 M -thujap- licinol. The experiments indicated that the presence of a metal ion is necessary for -thujaplicinol to interact with the HIV-RH domain, in agreement with the crystallo- graphic results [21]. On the other hand, 500 M saturated tricyclic N-hy- droxypyridone (compo g affinity (Kd 4.4 mM) to Mg2+, induced less than 25 Hz of chemical shift changes in the signals of 14 residues of HIV-RH in the presence of 8 mM Mg2+. Three de- tectable chelating residuals of the DEDD motif, i.e., D443, E478 and D498, were among them. The chemical shift changes were 14, 10 and 13 Hz, respectively. We also observed line-shape-broadening of the E478 signal. In comparison with Figure 1(c), Figure 4 indicates that the compound interacted with the active site of HIV-RH through Mg2+ chelation in the buffer containing 8 mM Mg2+. Interestingly, in the absence of divalent metal ions, this compound was found to bind HIV-RH at an alterna- tive binding site through our NMR and crystallographic studies (unpublished data). Although compound s-7 in- duced small chemical shift perturbations in the HIV-RH domain, it showed strong inhibition against HIV-RH with an IC50 of 0.038 M. One reason could be that the experimental conditions used for the IC50 measurement were very different from those used in the NMR experi- ments. In our biochemical assay, we used substrate Copyright © 2011 SciRes. AJAC 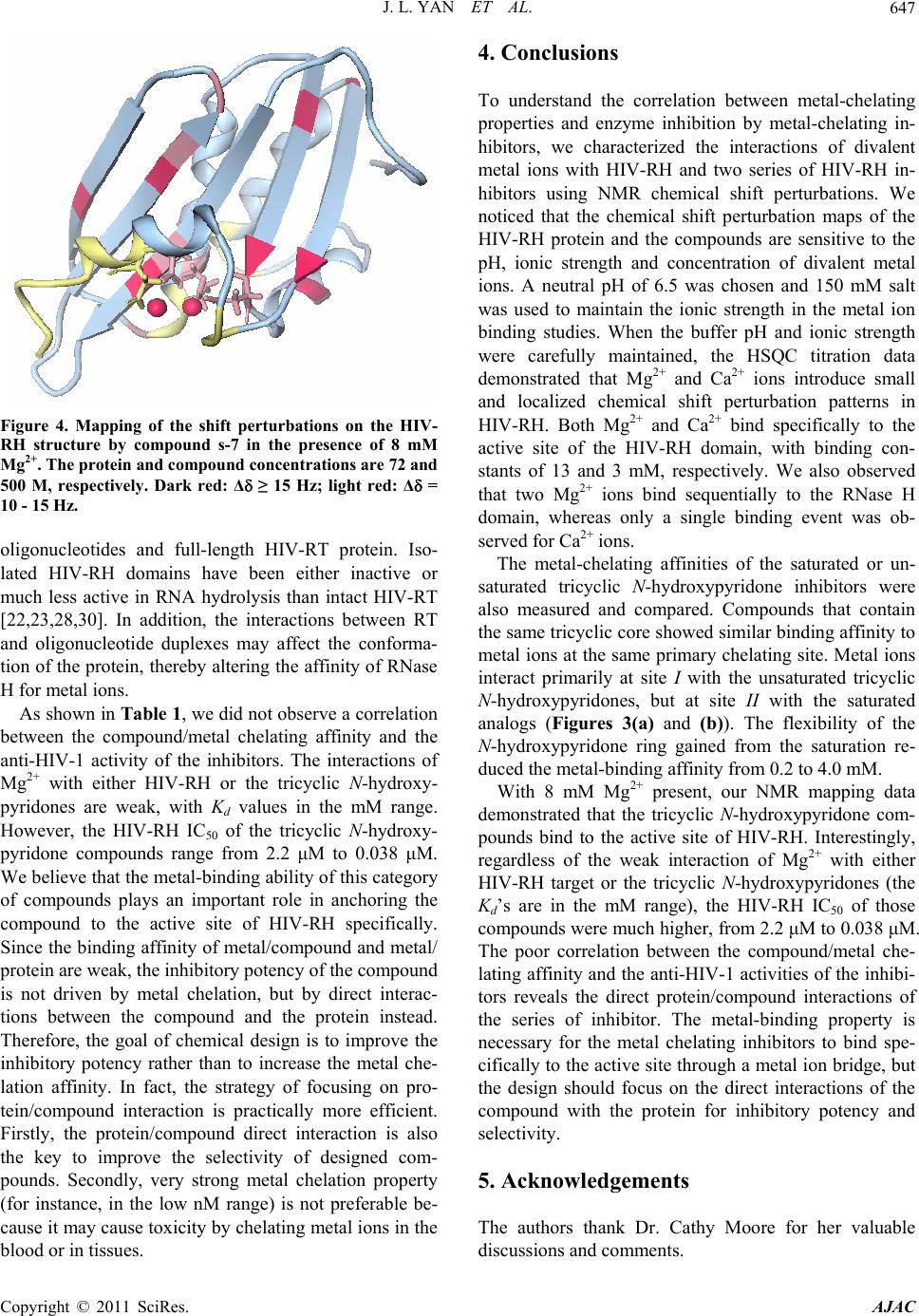 J. L. YAN ET AL.647 Figure 4. Mapping of the shift perturbations on the HIV- RH structure by compound s-7 in the presence of 8 mM tides and full-length HIV-RT protein. Iso- ted HIV-RH domains have been either inactive or nd/metal chelating affinity and the an To understand the correlation between metal-chelating me inhibition by metal-chelating in- hibitors, we characterized the interactions of divalent ompared. Compounds that contain th ly, re The authors thank Dr. Cathy Moore for her valuable Mg2+. The protein and compound concentrations are 72 and 500 M, respectively. Dark red: Δ ≥ 15 Hz; light red: Δ = 10 - 15 Hz. oligonucleo la much less active in RNA hydrolysis than intact HIV-RT [22,23,28,30]. In addition, the interactions between RT and oligonucleotide duplexes may affect the conforma- tion of the protein, thereby altering the affinity of RNase H for metal ions. As shown in Table 1, we did not observe a correlation between the compou ti-HIV-1 activity of the inhibitors. The interactions of Mg2+ with either HIV-RH or the tricyclic N-hydroxy- pyridones are weak, with Kd values in the mM range. However, the HIV-RH IC50 of the tricyclic N-hydroxy- pyridone compounds range from 2.2 μM to 0.038 μM. We believe that the metal-binding ability of this category of compounds plays an important role in anchoring the compound to the active site of HIV-RH specifically. Since the binding affinity of metal/compound and metal/ protein are weak, the inhibitory potency of the compound is not driven by metal chelation, but by direct interac- tions between the compound and the protein instead. Therefore, the goal of chemical design is to improve the inhibitory potency rather than to increase the metal che- lation affinity. In fact, the strategy of focusing on pro- tein/compound interaction is practically more efficient. Firstly, the protein/compound direct interaction is also the key to improve the selectivity of designed com- pounds. Secondly, very strong metal chelation property (for instance, in the low nM range) is not preferable be- cause it may cause toxicity by chelating metal ions in the blood or in tissues. 4. Conclusions properties and enzy metal ions with HIV-RH and two series of HIV-RH in- hibitors using NMR chemical shift perturbations. We noticed that the chemical shift perturbation maps of the HIV-RH protein and the compounds are sensitive to the pH, ionic strength and concentration of divalent metal ions. A neutral pH of 6.5 was chosen and 150 mM salt was used to maintain the ionic strength in the metal ion binding studies. When the buffer pH and ionic strength were carefully maintained, the HSQC titration data demonstrated that Mg2+ and Ca2+ ions introduce small and localized chemical shift perturbation patterns in HIV-RH. Both Mg2+ and Ca2+ bind specifically to the active site of the HIV-RH domain, with binding con- stants of 13 and 3 mM, respectively. We also observed that two Mg2+ ions bind sequentially to the RNase H domain, whereas only a single binding event was ob- served for Ca2+ ions. The metal-chelating affinities of the saturated or un- saturated tricyclic N-hydroxypyridone inhibitors were also measured and c e same tricyclic core showed similar binding affinity to metal ions at the same primary chelating site. Metal ions interact primarily at site I with the unsaturated tricyclic N-hydroxypyridones, but at site II with the saturated analogs (Figures 3(a) and (b)). The flexibility of the N-hydroxypyridone ring gained from the saturation re- duced the metal-binding affinity from 0.2 to 4.0 mM. With 8 mM Mg2+ present, our NMR mapping data demonstrated that the tricyclic N-hydroxypyridone com- pounds bind to the active site of HIV-RH. Interesting gardless of the weak interaction of Mg2+ with either HIV-RH target or the tricyclic N-hydroxypyridones (the Kd’s are in the mM range), the HIV-RH IC50 of those compounds were much higher, from 2.2 μM to 0.038 μM. The poor correlation between the compound/metal che- lating affinity and the anti-HIV-1 activities of the inhibi- tors reveals the direct protein/compound interactions of the series of inhibitor. The metal-binding property is necessary for the metal chelating inhibitors to bind spe- cifically to the active site through a metal ion bridge, but the design should focus on the direct interactions of the compound with the protein for inhibitory potency and selectivity. 5. Acknowledgements discussions and comments. Copyright © 2011 SciRes. AJAC  J. L. YAN ET AL. 648 [1] E. J. Artsa and S. F. Le Grice, “Interaction of Retrovir criptase with Template-Primer Duplexes Progress in Nucleic Acid Research 6. References al Reverse Trans during Replication,” and Molecular Biology, Vol. 58, 1997, pp. 339-393. doi:10.1016/S0079-6603(08)60041-0 [2] V. Goldschmidt, J. Didierjean, B. Ehresmann, et al., “Mg2+ Dependency of HIV-1 Reverse Transcription hibition by Nucleoside Analogues an , In- d Resistance,” Nu- cleic Acids Research, Vol. 34, No. 1, 2006, pp. 42-52. doi:10.1093/nar/gkj411 [3] M. Götte, “Inhibition of HIV-1 Reverse Transcription: Basic Principles of Drug Action and Resistance,” Exp Review of Anti-Infectivert e Therapy, Vol. 2, 2004, pp. 707-716. doi:10.1586/14789072.2.5.707 [4] M. Götte, S. Fackler, T. Hermann, et al., “HIV-1 Reverse Transcriptase-Associated Rnase H Cleaves RNA/RNA in Arrested Complexes: Implications for the Mechanis us Research, Vol. 134, No. 1-2, 2008, pp m by Which Rnase H Discriminates between RNA/RNA and RNA/DNA,” EMBO Journal, Vol. 14, No. 4, 1995, pp. 833-841. [5] S. J. Schultz and J. J. Champoux, “RNase H Activity: Structure, Specificity, and Function in Reverse Transcrip- tion,” Vir . 86-103. doi:10.1016/j.virusres.2007.12.007 [6] M. Nowotny and W. Yang, “Stepwise Analyses of Metal Ions in Rnase H Catalysis from Substrate Destabilization to Product Release,” EMBO Journal, Vol. 25, No. 9, 2006, pp. 1924-1933. doi:10.1038/sj.emboj.7601076 [7] W. Yang, J. Y. Lee and M. Nowotny, “Making and Break- ing Nucleic Acids: Two-Mg2+-Ion Catalysis and Substrate Specificity,” Molecular Cell, Vol. 22, No. 1, 2006, pp. 5-13. doi:10.1016/j.molcel.2006.03.013 [8] M. Nowotny, S. A. Gaidamakov, R. J. Crouch, et al., “Crystal Structures of RNase H Bound to an RNA/DNA Hybrid: Substrate Specificity and Metal-Dependent Ca- talysis,” Cell, Vol. 121, No. 7, 2005, pp. 1005-1016. doi:10.1016/j.cell.2005.04.024 [9] J. A. Cowan, T. Ohyama, K. Howard, et al., “Metal-Ion Stoichiometry of the HIV-1 RT Ribonuclease H Dom Evidence for Two Mutually E ain: xclusive Sites Leads to New Mechanistic Insights on Metal-Mediated Hydrolysis in Nucleic Acid Biochemistry,” Journal of Biological In- organic Chemistry, Vol. 5, No. 1, 2000, pp. 67-74. doi:10.1007/s007750050009 [10] J. Q. Hang, S. Rajendran, Y. L. Yang, et al., “Activity of the Isolated HIV Rnase H Domain and Specific Inhi by N-Hydroxyimides,” Bioch bition emical and Biophysical Re- criptase in the Presence of Magnesium,” Biochem search Communications, Vol. 317, No. 2, 2004, pp. 321- 329. [11] K. Pari, G. A. Mueller, E. F. Derose, et al., “Solution Structure of the Rnase H Domain of the HIV-1 Reverse Trans , Vol. 42, No. 3, 2003, pp. 639-650. doi:10.1021/bi0204894 [12] K. Katayanagi, M. Okumura and K. Morikawa, “Crystal Structure of Escherichia Coli Rnase Mg2+ at 2.8 a Reso HI in Complex with lution: Proof for a Single Mg(2+)-Binding Site,” Proteins, Vol. 17, No. 4, 1993, pp. 337-346. doi:10.1002/prot.340170402 [13] M. Nowotny, S. A. Gaidamakov, R. Ghirlando, et al., “Structure RNA/DNA Hybrid: Insight in of Human Rnase H1 Complexed with an to HIV Reverse Transcrip- tion,” Molecular Cell, Vol. 28, No. 2, 2007, pp. 264-276. doi:10.1016/j.molcel.2007.08.015 [14] O. Schatz, F. V. Cromme, F. Grüninger-Leitch, et al., “Point Mutations in Conserved Amino Acid Residues within the C-Terminal Domain of HIV-1 Reverse Tran- scriptase Specifically Repress RNase H Function,” FEBS Letters, Vol. 257, No. 2, 1989, pp. 311-314. doi:10.1016/0014-5793(89)81559-5 [15] S. F. Le Grice, T. Naas, B. Wohigensinger, et al., “Sub- unit-Selective Mutagenesis Indicates Min merase Activity in Heterodimer-Ass imal Poly- ociated P51 HIV-1 esign, Vol. 12, No. 15, 2006, Reverse Transcriptase,” The EMBO Journal, Vol. 10, No. 12, 1991, pp. 3905-3911. [16] K. Klumpp and T. Mirzadegan, “Recent Progress in the Design of Small Molecule Inhibitors of HIV RNase H,” Current Pharmaceutical D pp. 1909- 1922. doi:10.2174/138161206776873653 [17] E. Tramontano, “HIV-1 RNase H: Recent Progress in an Exciting, Yet Little Explored, Drug Target,” Mini-Reviews in Medicinal Chemistry, Vol. 6, No. 6, 2006, pp. 727-737. doi:10.2174/138955706777435733 [18] G. M. Clore and A. M. Gronenborn, “Multidimensional Heteronuclear Nuclear Magnetic Resonance of Proteins,” Methods in Enzymology, Vol. 239, 1994, pp. 349-363. doi:10.1016/S0076-6879(94)39013-4 [19] S. Grzesiek and A. Bax, “Amino Acid Type Determina- tion in the Sequential Assignment Procedure of U formly 13C/15N-Enriched Proteins,” ni- Journal of Bio- et al., t the RNase H Active molecular NMR, Vol. 3, No. 2, 1993, pp. 185-204. [20] D. Goddard and D. G. Kneller, “SPARKY 3,” University of California, San Francisco, 2000. [21] D. M. Himmel, K. A. Maegley, T. A. Pauly, “Structure of HIV-1 Reverse Transcriptase with the In- hibitor Beta-Thujap- licinol Bound a Site,” Structure, Vol. 17, No. 12, 2009, pp. 1625-1635. doi:10.1016/j.str.2009.09.016 [22] A. Jacobo-Molina, J. Ding, R. G. Nanni, et al., “Crystal Structure of Human Immunodeficiency Virus Type 1 Reverse Transcriptase Comple xed with Double-Stranded al of Biological Chem- DNA at 3.0 a Resolution Shows Bent DNA,” Proceed- ings of the National Academy of Sciences of the USA, Vol. 90, No. 13, 1993, pp. 6320-6324. [23] J. L. Keck and S. Marqusee, “The Putative Substrate Recognition Loop of Escherichia Coli Ribonuclease H Is Not Essential for Activity,” Journ istry, Vol. 271, No. 33, 1996, pp. 19883-19887. doi:10.1074/jbc.271.33.19883 [24] G. A. Mueller, K. Pari, E. F. Derose, et al., “Backbone Dynamics of the Rnase H Domain of HIV-1 Transcriptase,” Biochem, Vol Reverse . 43, No. 29, 2004, pp. Copyright © 2011 SciRes. AJAC  J. L. YAN ET AL. Copyright © 2011 SciRes. AJAC 649 r 2D NMR,” Journal of Biomolecular NMR, 9332-9342. [25] Y. Oda, H. Nakamura, S. Kanaya, et al., “Binding of Metal Ions to E. coli Rnase HI Observed by 1H-15N Heteronuclea Vol. 1, No. 3, 1991, pp. 247-255. doi:10.1007/BF01875518 [26] Y. Oda, H. Nakamura and S. Kanaya, “Role of Histidine 124 in the Catalytic Function of R Escherichia coli,” Journalibonuclease HI from of Biological Chemistry, Vol. se Transcriptase in Solution n Immunodeficiency Virus Reverse Tran- 268, No. 1, 1993, pp. 88-92. [27] R. Powers, G. M. Clore, A. Bax, et al., “Secondary Struc- ture of the Ribonuclease H Domain of the Human Im- munodeficiency Virus Rever Using Three-Dimensional Double and Triple Resonance Heteronuclear Magnetic Resonance Spectroscopy,” Journal of Molecular Biology, Vol. 221, No. 4, 1991, pp. 1081- 1090. [28] R. Powers, G. M. Clore, S. J. Stahl, et al., “Analysis of the Backbone Dynamics of the Ribonuclease H Domain of the Huma scriptase Using 15N Relaxation Measurements,” Biochem, Vol. 31, No. 38, 1992, pp. 9150-9157. doi:10.1021/bi00153a006 [29] D. Chattopadhyay, B. C. Finzel, S. H. Munson, et al., “Crystallographic Analyses of an Active HIV-1 Ribonu- clease H Domain Show Structural Features That Distin- guish It from the Inactive Form,” Acta Crystallographica Section D: Biological Crystallography, Vol. 49, 1993, pp. 423-427. doi:10.1107/S0907444993002409 [30] J. F. Davies, Z. Hostomska, J. Hostomsky, et al., “Crystal Structure of the Ribonuclease H Domain of HIV-1 Re- verse Transcriptase,” Science, Vol. 252, No. 5002, 1991, pp. 88-95. doi:10.1126/science.1707186 [31] J. Jäger, S. J. Smerdon, J. M. Wang, et al., “Comparison of Three Different Crystal Forms Shows HIV-1 Reverse Transcriptase Displays an Internal Swivel Motion,” Structure, Vol. 2, No. 9, 1994, pp. 869-876. doi:10.1016/S0969-2126(94)00087-5 [32] J. A. Grobler, K. Stillmock, B. Hu, et al., “ Inhibitor Mechanism and HIV-1 Inte Diketo Acid grase: Implications for Metal Binding in the Active Site of Phosphotrans- ferase Enzymes,” Proc. Natl. Acad. Sci. USA, Vol. 99, No. 10, 2002, pp. 6661-6666. doi:10.1073/pnas.092056199 [33] S. R. Budihas, I. Gorshkova, S. lective Inhibition of HIV-1 R Gaidamakov, et al., “Se- everse Transcriptase-Asso- ciated Ribonuclease H Activity by Hydroxylated Tro- polones,” Nucleic Acids Research, Vol. 33, No. 4, 2005, pp. 1249-1256. doi:10.1093/nar/gki268
|