Journal of Agricultural Chemistry and Environment
Vol.1 No.1(2012), Article ID:24944,7 pages DOI:10.4236/jacen.2012.11007
In vitro antagonistic potential of Streptomyces sp. AM-S1 against plant and human pathogens@NolistTemp#
Email: sckang@daegu.ac.kr
1Department of Biotechnology, College of Engineering, Daegu University, Gyeongsan, South Korea
Received 5 October 2012; revised 8 November 2012; accepted 17 November 2012
ABSTRACT
In the present investigation, a total number of 132 different actinomycetes strains were isolated from the humus soil samples. Out of 132 isolates, 52 showed inhibitory activity against the fungal pathogen Rhizoctonia solani. Among the antagonists, the isolate designated as AM-S1 exhibited maximum inhibitory activity against the test pathogen R. solani (41 mm). Further, the light microscopic observations of the co-cultures showed severe structural alterations in the mycelia of R. solani near the zone of inhibition. The isolate AM-S1 was identified as Streptomyces sp. by morphological and 16S rDNA sequence analysis. The color of the aerial and substrate mycelia produced by the Streptomyces sp. AM-S1 varied with different media. The isolate Streptomyces sp. AM-S1 also effectively inhibited the growth of various plant and human pathogens. Further works are needed on optimization of this strain’s antagonistic activity, isolation and characterization of the antimicrobial metabolite.
Keywords: Actinomycetes; Streptomyces sp. AM-S1; Antagonistic activity; Rhizoctonia solani
1. INTRODUCTION
Losses in crop production due to plant disease average 13% worldwide and severely limit production, quality, and safety of food. Fungi are the major disease causing agents for the economically important plants in the field and during storage, especially in the subtropical and tropical regions. Many of synthetic pesticides can keep fungal infections at an acceptable level. However, they are associated with several drawbacks such as environmental pollution, lack of specificity, accumulation if biodegradation is slow or even missing and development of pathogen resistance [1]. In this scenario, the search continues for more environmentally friendly methods to control plant disease that will contribute to the goal of sustainability in agriculture. Microbe-based biocontrol methods are one alternative way to control diseases in place of agrochemicals.
Various kinds of microbial antagonists have been investigated as potential antifungal biocontrol agents for plant disease management. Actinomycetes are one of the important groups of soil microorganisms. They are important producers of bioactive compounds and constitute a potential group of biocontrol agents. Their population in soil depends on many factors, including soil type, aeration, organic matter, salinity, relative moisture content and temperature [2]. Actinomycetes have the capability to synthesize wide varieties of biologically active secondary metabolites such as antibiotics, pesticides, antiparasitic compounds and enzymes like cellulase, xylanase, proteinase and chitinase. Several descriptive researches have shown that actinomycetes are a promising group of fungus-antagonistic and root-colonizing microbes. They protect, to various degrees, several different plants from soil-borne fungal pathogens [3]. Consequently, biologists are becoming interested in using actinomycetes as agents to influence plant growth and for biological control of soil-borne root diseases of crop plants.
It is well known that actinomycetes produce 70% to 80% of bioactive secondary metabolites, where approximately 60% of antibiotics developed for agricultural use are isolated from Streptomyces spp. [4]. It has an enormous biosynthetic potential that remains unchallenged, without a potential competitor among other microbial groups [5]. Many reports have pointed out that since streptomycetes are frequently screened for antimicrobial activity, the existence of secondary metabolites with other activities may have been missed [6]. This microbial wealth from actinomycetes has yet to be investigated thoroughly.
Therefore, the present study was carried out to investigate the isolation of actinomycetes from forest humus soil samples and to determine the antagonistic activity against plant pathogen Rhizoctonia solani. Further, identification of most effective isolate by cultural, morphological and 16S rDNA sequencing method and to evaluate the broad spectrum antagonistic activity against certain plant and human pathogenic species.
2. MATERIALS AND METHODS
2.1. Isolation of Soil Actinomycetes
A total of 30 different humus soil samples were collected from the forest areas in Gyeongsan, Gyeongbuk, South Korea. Ten gram of soil samples were suspended in 95 mL of distilled water and these suspensions were considered as 10−1 dilution. Starch casein agar medium (SCA) was used for isolation of actinomycetes. From 10−1 suspension, 1 mL was transferred to 9 mL of sterile distilled water and subsequently serially diluted to 10−3, 10−4, etc. From the required dilution, 0.1 mL suspension was drawn and plated over the surface of SCA medium. All the plates were incubated at 28˚C ± 2˚C for 7 days. The actinomycetes were isolated as pure culture by routine microbiological methods and maintained on SCA slants.
2.2. Screening against Fungal Phytopathogens
All the isolated actinomycetes strains were subjected to primary screening against fungal plant pathogen Rhizoctonia solani. About 20 mL of potato dextrose agar (PDA) was poured into each Petri plate and allowed to solidify. Four actinomycetes isolates were streaked separately on the four corner of each Petri plate and incubated at 28˚C for 5 days. A mycelial disc of 8 mm diameter was taken from the mother culture of R. solani from actively growing region. Then the mycelial disc of the pathogen was placed in the center of actinomycetes isolates streaked plate; while the mycelial disc alone in PDA plate served as control. All the plates were incubated at 25˚C up to 4 days and the zone of inhibition was measured. Among the 132 actinomycetes isolates, a strain designated as AM-S1 isolate was selected for further studies as it effectively inhibited the mycelial growth of R. solani than other isolates.
2.3. Light Microscopy Study of AM-S1 and Fungal Pathogen Interaction
The interaction of AM-S1 and fungal pathogen was studied using light microscopy. In the dual plate assay, the samples were taken from the pathogen and antagonist interaction zone and observed under the light microscope.
2.4. Colony Morphology
The strain AM-S1 was grown on SCA in Petri plate for 10 days and the colony morphology, pigmentation, sporulation, etc. were recorded. The cover slip culture technique was used for studying the morphological characteristics of the AM-S1. Sterile glass cover slips were inserted at an angle of 45˚C into solidified SCA medium in Petri plate. A loopful of inoculum of the isolate was streaked along the line where the cover slip meets the agar and then the plates were incubated at room temperature for 7 days. The organism grows both on the medium and in a line across the upper surface of the cover slip. The cover slip was removed and examined under a light microscope and photographed. The morphological characters were observed and motility test was carried out using Gram’s staining and hanging drop method, respectively.
2.5. 16S rDNA Sequencing of the Isolate AM-S1
Genomic DNA was extracted from cultured cells following Pospiech and Neumann [7]. The 16S rDNA was amplified using the PCR method with Taq DNA polymerase and universal primers FD1 (5’-AGAGTTTGATCCTGGCTCAG-3’) and RP2 (5-ACGGCTACCTTGTTACGACTT-3’) following Kim et al. [8.] The PCR products were purified and sequenced by Geno Tech Company Limited (South Korea). The obtained sequence was compared for similarity with sequences present in the genomic database banks, using the “NCBI Blast” program available at the ncbi.nlm.nih.gov web site and highest matching sequences downloaded. The 16S rDNA sequence of AM-S1 was aligned with these sequences using the program CLUSTALX [9]. MEGA 4.0 software [10] was used to carry out phylogenetic analyses of the alignment using the Neighbor-Joining method.
2.6. Growth Characteristics of Streptomyces sp. AM-S1 in Different Solid Media
Starch Caesin Agar (SCA), Tryptone-yeast extract agar (ISP-1), Starch inorganic salt Agar (ISP-4), Glycerol Asparagine Agar (ISP-5), Glycerol Tyrosine Agar (ISP-7), Maltose Yeast Extract Agar (MYEA), Potato Dextrose Agar (PDA), Nutrient Agar (NA), Production Medium— III (PM-III) and Production Medium—IV (PM-IV) were prepared and poured into Petri plates. A loopful of Streptomyces sp. AM-S1 inoculum was streaked on the above media and incubated at room temperature. After 7 days, the growth, pigmentation of aerial and substrate mycelia were observed and the results were documented.
2.7. Antagonistic Activity of Streptomyces sp. AM-S1 against Different Fungal Phytopathogens
The isolate Streptomyces sp. AM-S1 was tested for antagonistic activity against a wide range of fungal phytopathogens viz., Rhizoctonia solani, Botryodiplodia theobromae, Sclerotium rolfsii, Fusarium oxysporum, Curvularia lunata, Helminthosporium oryzae, Penicillium expansum and Aspergillus niger on PDA medium employing dual culture technique. All the said phytopahtogenic isolates were obtained from Department of Biotechnology, College of Engineering, Daegu University, Gyeongsan, South Korea.
2.8. Antagonistic Activity of Streptomyces sp. AM-S1 against Human Pathogens
The Streptomyces sp. AM-S1 was streaked on NA at the left corner of Petri plates and incubated at 28˚C. After 5 days, the human pathogens viz., Candida albicans (KCTC 27241), Staphylococcus aureus (KCTC 1916), Bacillus subtilis (KCTC 1027), Salmonella typhimurium (KCTC 2515) and Escherichia coli (KCTC 1045) were streaked at right angle to the Streptomyces sp. AM-S1 colonies. All the said human pathogens were procured from Koreran Collection for Type Cultures (KCTC), South Korea. The pathogens alone streaked in NA plates served as control. The plates were incubated at 28˚C ± 2˚C up to 48 h and observed for the inhibition of pathogen’s growth.
2.9. Statistical Analysis
The data were subjected to a one-way analysis of variance (ANOVA) and the significance of the difference between means was determined by Duncan’s multiple range test (P < 0.05) using statistica (Statsoft Inc., Tulsa, OH, USA). Values expressed are means of 3 replicate determinations ± standard deviations (SD).
3. RESULTS
3.1. Isolation and Screening of Actinomycetes against Rhizoctonia solani
A total of 30 different humus soil samples were collected from the forest areas in Gyeongsan, South Korea. A total of 132 morphologically different actinomycetes strains were isolated from humus soil samples by soil dilution technique using SCA medium.
Among 132 isolates screened using dual plate assay, 52 showed inhibitory activities against R. solani. The zone of inhibition towards R. solani ranged from 3 mm to 41 mm. The present in vitro tests indicated that, 39.3% of strains were antagonistic against R. solani. Among the antagonists, the isolate designated as AM-S1 exhibited the maximum inhibitory activity (Figure 1). Owing to its superior performance against the plant pathogens, the isolate AM-S1 was selected for further studies.

Figure 1. Mycelial inhibition of R. solani by Streptomyces sp. AM-S1 in dual culture method. (A) Control R. solani, (B) R. solani and Streptomyces sp. AM-S1.
3.2. Light Microscopic Study of AM-S1 and Fungal Phytopathogens
Light microscopic observations on mycelial growth of R. solani near the zone of inhibition from dual culture plates showed considerable changes in hyphal morphology compared to normal growth in control plate. There was a significant interaction between the AM-S1 and R. solani. Microscopic observations showed marked morphological changes and severe structural alterations of the fungal mycelium. The hyphae became thinner, vacuolated and showed stunted growth with proliferating branches near the growing tip (Figures 2(A) and (B)).
3.3. Morphology
Colonies of the strain AM-S1 are grey or pale brown and wrinkled on SCA. They revealed the presence of the vegetative hyphae (Figure 3(A)). In the observation with light microscopy on the SCA, spore-bearing hyphae of the strain appeared as spiral chains (Figure 3(B)). The number of the spores per chain was higher than 20, which indicated that they are long chains of spores. The taxonomic status of the AM-S1, according to morphological observations, that bacterium belongs to the genus Streptomyces. Substrate mycelia of the isolate Streptomyces sp. (AM-S1) grew actively on all tested media. Further, the colour of the aerial and substrate mycelia produced by the Streptomyces sp. varied with different media (Table 1).
3.4. 16S rDNA Sequence Analysis
After PCR amplification from strain AM-S1, the 16S rDNA gene was sequenced. The obtained sequence was compared for similarity with sequences present in the genomic database banks, using the “NCBI Blast” program
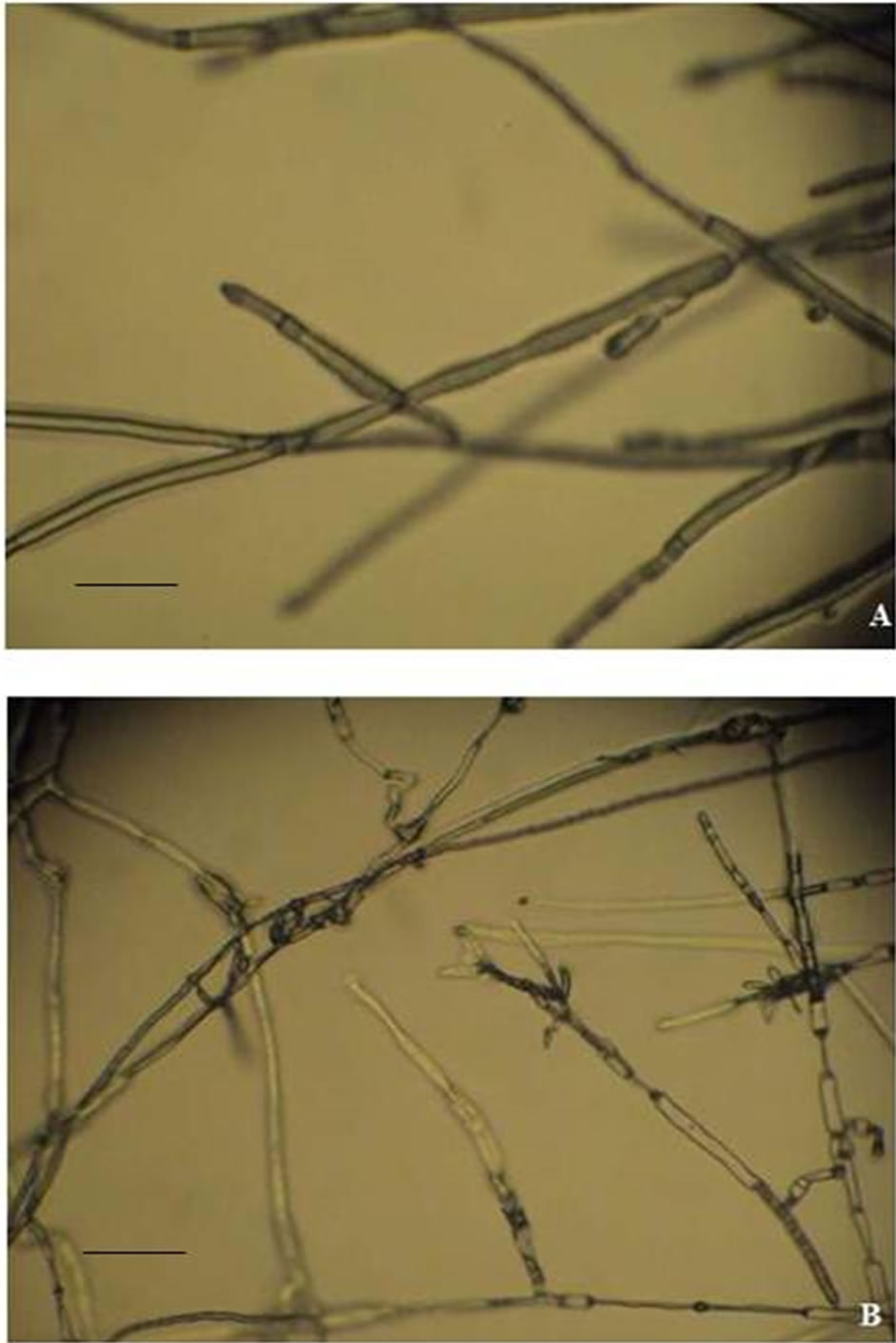
Figure 2. Mycelial changes in the pathogen R. solani from the zone of interaction. (A) Control mycelium; (B) Mycelium from inhibition zone. The bars represent 20 µm (400×).

Table 1. Culture characteristics of Streptomyces sp. AM-S1.
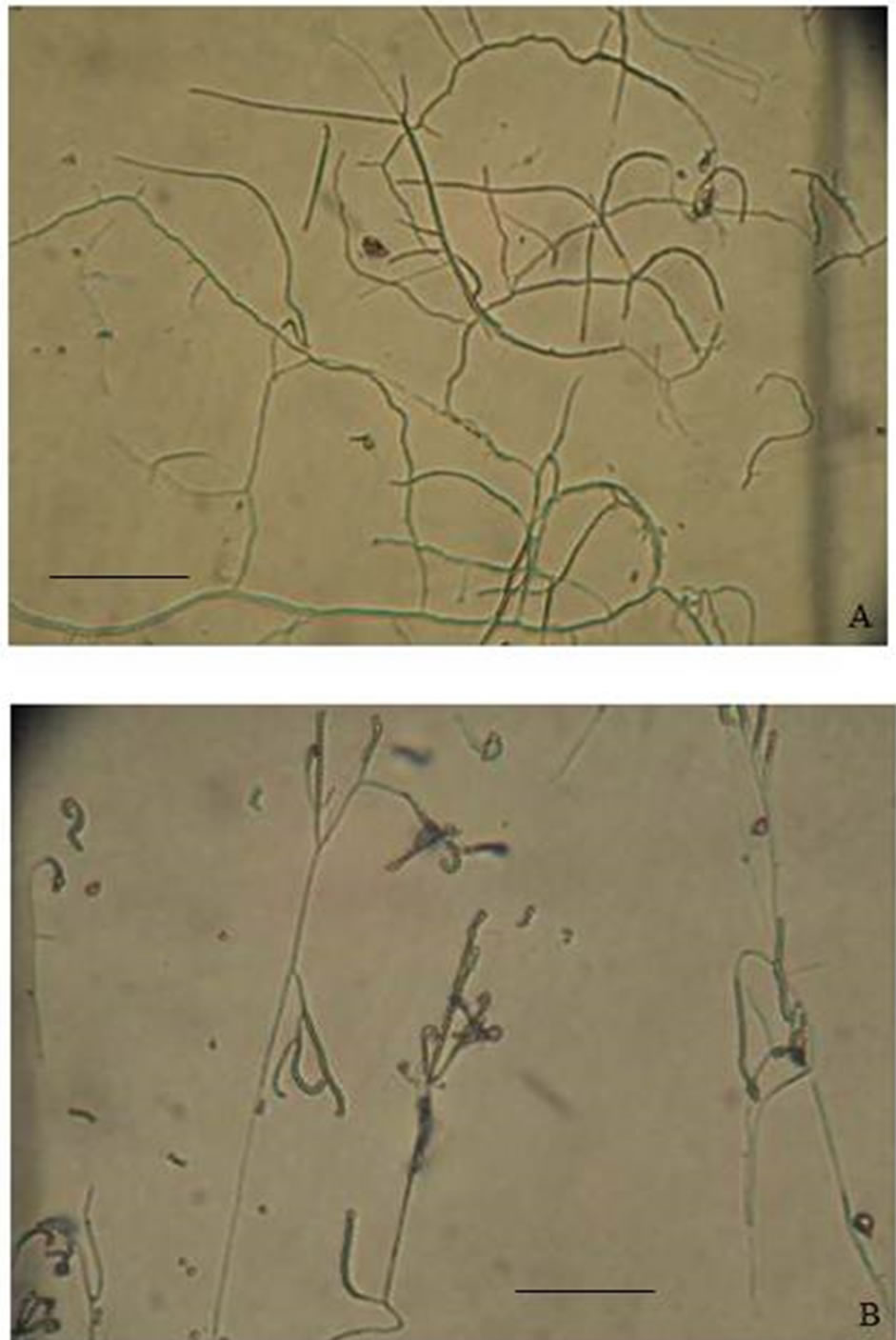
Figure 3. Morphology of Streptomyces sp. AM-S1. (A) Mycelium; (B) Spore chain. The bars represent 10 µm (400×).
available at the ncbi.nlm.nih.gov web site. The 16S rDNA gene sequence of AM-S1 had 99% sequence identity with the 16S rDNA gene sequences from several Streptomyces sp. (Figure 4). The sequence of Steptomyces sp. AM-S1 has been submitted to Genbank with the accession number JX444563. This result clearly suggests that the AM-S1 belongs to the genus Streptomyces.
3.5. Broad Spectrum Antagonistic Activity of Streptomyces sp. AM-S1 against Plant and Human Pathogens
Further screening against a wide range of fungal phytopathogens revealed that the, Streptomyces sp. effectively inhibited the mycelial growth of R. solani, B. theobromae, S. rolfsii, F. oxysporum, C. lunata, H. oryzae, P. expansum and A. niger with the inhibition zones ranging from 22 mm to 41 mm (Table 2). Table 3 shows broad spectrum antimicrobial activities of the Streptomyces sp. AM-S1 against C. albicans, S. aureus, B. subtilis, S. typhimurium and E. coli. The result suggests that a single streptomycetes strain will be able to provide broad-spectrum antibiosis to control diverse plant and
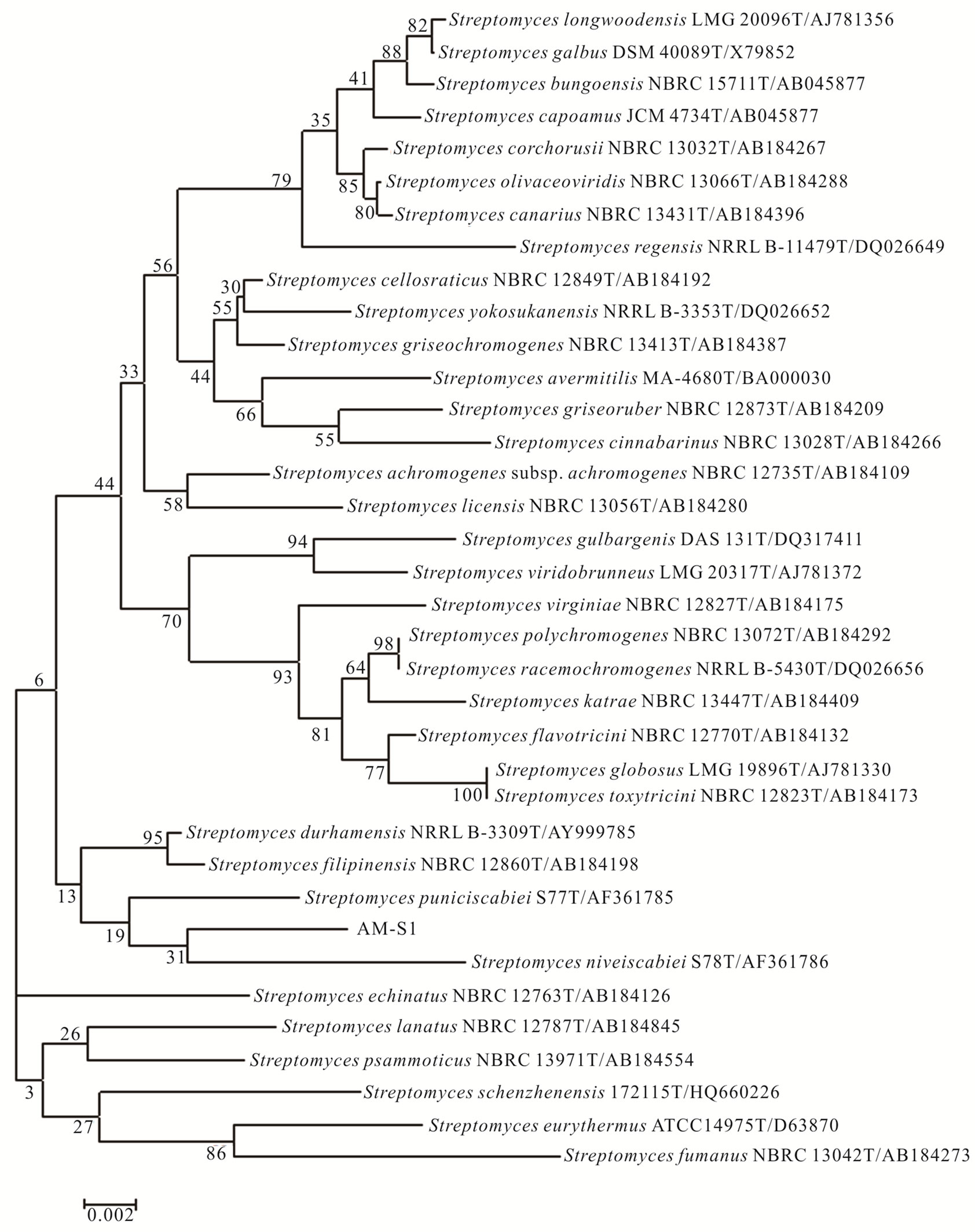

Figure 4. Phylogenetic tree showing the position of Streptomyces sp. AM-S1. Neighbor-Joining tree showing the position of strain AM-S1 compared to related organisms in a 16S rRNA gene tree. Numbers at nodes indicate levels of bootstrap support (%) based on 1000 resampled datasets. The scale bar indicates the number of nucleotide substitutions per site.
human pathogens. These pathogenic species demonstrated differences in their sensitivity to the antagonistic effect of the isolate Streptomyces sp. AM-S1.
4. DISCUSSION
Actinomycetes are Gram-positive filamentous bacteria. The majority of this group is saprophytic and found widely distributed in the soil. They are grow extensively in soils rich in organic matter and previous studies have shown that the number of actinomycetes in soil is positively correlated with the level of organic matter and moisture content [11]. It is generally accepted that in

Table 2. Antagonistic activity of Streptomyces sp. AM-S1 against certain fungal phytopathogens.
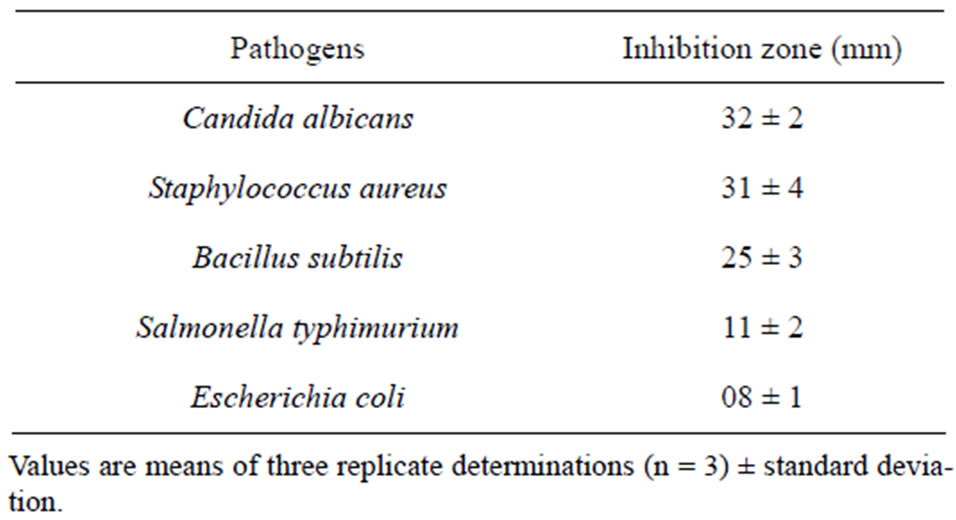

Table 3. Antagonistic activity of Streptomyces sp. AM-S1 against certain human pathogens.
relatively dry, humic, calcareous soils, actinomycetes form the dominant fraction of the microflora with viable counts reaching 106 g−1 dry weight soil [12]. Therefore, in the present study 132 different actinomycetes were isolated from 30 forest humus soils samples. Soil microbial diversity is considered to be critical to the maintenance of soil health and quality, and agriculture practices influence soil microbial diversity and the ability to suppress plant diseases [13]. Among the antagonists, the isolate designated as AM-S1 registered higher antagonistic activity against R. solani. R. solani is an important fungal phytopathogen that lives in the soil in the form of sclerotia and does not generates asexual spores. It has a wide host range and distribution and causes sheath blight in some field crops, such as corn, rice, lawn grass and cucumber [14]. Further, the microscopic studies revealed mycelial deformities near the zone of inhibition. Actinomycetes isolates were effective in restricting radial growth of phytopathogenic fungi. There are many reports related to antibiotic substances which induced malfunctions such as stunting, distortion, swelling, hyphal protuberances or highly branched appearance of fungal germ tubes [15].
Streptomycetes have been determined largely because of morphological and biochemical criteria, resulting in the arrangement of strains into cluster groups [16]. On the basis of the aerial and substrate mycelia, development of spiral spore chains and smooth spore surface of the strain, they are placed under the genus Streptomyces. The taxonomic identity of the strain AM-S1 was ascertained by various physicochemical parameters and 16S rDNA sequencing as Streptomyces sp. and hence the isolate was designated as Streptomyces sp. AM-S1. Further, the growth characteristic study demonstrated that the nutrient compositions of the medium greatly influence the growth and morphology of organisms.
Apart from its antagonistic activity against R. solani, the present isolate Streptomyces sp. AM-S1 exhibited broad antimicrobial spectrum against various plant and human pathogens. Many studies refer to the role of actinomycetes, mostly Streptomyces strains, in plant protection which are antagonists against most of the phytopathogenic fungi. Actinomycetes are especially significant in inhibiting phytopathogens for the following reasons: they can survive in soils of various types, have long survival due to their spore forming abilities, are prolific producers of various bioactive compounds such as antibiotics, siderophores, chitinases, and phytohormones and have phosphate solubilizing abilities [17,18]. The potential antagonistic activity of the streptomycetes found in this study against plant pathogenic fungi highlights their potential in bio-control programs. This study confirmed that the Streptomyces are able to produce a wide variety of antibiotics with antifungal activity.
From the present study, it could be demonstrated that humus soil samples provided a rich source of diversity of actinomycetes. The morphological and 16S rDNA sequencing studies clearly demonstrated that the present the isolate AM-S1 is belongs to the genus Streptomyces. The Streptomyces sp. AM-S1 had the strongest inhibitory effects on in vitro growth of various plant and human pathogens. Further work on optimization of this strain’s antagonistic activity, mechanism of action and evaluation for its disease control potential under field conditions is under progress.
5. ACKNOWLEDGEMENTS
This work was supported by a grant from the Next-Generation BioGreen 21 Program (No. PJ008157), Rural Development Administration, Republic of Korea.
REFERENCES
- Oskay, M. (2009) Antifungal and antibacterial compounds from Streptomyces strains. African Journal of Biotechnology, 8, 3007-3017.
- McCarthy, A.J. and Williams, S.T. (1990) Methods for studying the ecology of actinomycetes. Methods in Microbiology, 22, 533-563.
- El-Tarabily, K. and Sivasithamparam, K. (2006) Nonstreptomycete actinomycetes as biocontrol agents of soilborne fungal plant pathogens and as plant growth promoters. Soil Biology and Biochemistry, 38, 1505-1520. doi:10.1016/j.soilbio.2005.12.017
- Ilic, S.B., Konstantinovic, S.S., Todorovic, Z.B., Lazic, M.L., Veljkovic, V.B., Jokovic, N. and Radovanovic, B.C. (2007) Characterization and antimicrobial activity of the bioactive metabolites in streptomycete isolates. Microbiology, 76, 421-428. doi:10.1134/S0026261707040066
- Solanki, R., Khanna, M. and Lal, R. (2008) Bioactive compounds from marine actinomycetes. Indian Journal of Microbiology, 48, 410-431. doi:10.1007/s12088-008-0052-z
- Garcia, I., Job, D. and Matringe, M. (2000) Inhibition of p-hydroxyphenylpyruvate dioxygenase by the diketonitrile of isoxaflutole: A case of half-site reactivity. Biochemistry, 3, 7501-7507. doi:10.1021/bi000135h
- Pospiech, A. and Neumann, B. (1995) A versatile quickprep of genomic DNA from grampositive bacteria. Trends in Genetics, 11, 217-218. doi:10.1016/S0168-9525(00)89052-6
- Kim, H.J., Lee, S.C. and Hwang, B.K. (2006) Streptomyces cheonanensis sp. Nov., a novel streptomycete with antifungal activity. International Journal of Systematic and Evolutionary Microbiology, 56, 471-475. doi:10.1099/ijs.0.63816-0
- Jeanmougin, F., Thompsona, J.D., Gouy, M., Higgins, D.G. and Gibson, T.J. (1998) Multiple sequence alignment with Clustal X. Trends in Biochemical Sciences, 23, 403-405. doi:10.1016/S0968-0004(98)01285-7
- Kumar, S., Nei, M., Dudley, J. and Tamura, K. (2008) MEGA: A biologist-centric software for evolutionary analysis of DNA and protein sequences. Briefings in Bioinformatics, 9, 299-306. doi:10.1093/bib/bbn017
- Hayakawa, M., Otoguro, M., Takeuchi, T., Yamazaki, T. and Iimura, Y. (2000) Application of a method incorporating differential centrifugation for selective isolation of motile actinomycetes in soil and plant litter. Antonie van Leeuwenhoek, 78, 171-185. doi:10.1023/A:1026579426265
- Goodfellow, M. and Williams, S.T. (1983) Ecology of actinomycetes. Annual Review of Microbiology, 37, 189- 216. doi:10.1146/annurev.mi.37.100183.001201
- Garbeva, P., Postma, J., Veen, J.A. and Elsas, J.D. (2006) Effect of above-ground plant species on soil microbial community structure and its impact on suppression of Rhizoctonia solani AG3. Environmental Microbiology, 8, 233-246. doi:10.1111/j.1462-2920.2005.00888.x
- Huang, X., Zhang, N., Yong, X., Yang, X. and Shen, Q. (2012) Biocontrol of Rhizoctonia solani damping-off disease in cucumber with Bacillus pumilus SQR-N43. Microbiological Research, 167, 135-143. doi:10.1016/j.micres.2011.06.002
- Getha, K. and Vikineswary, S. (2002) Antagonistic effects of Streptomyces violaceusniger strain G10 on Fusarium oxysporum f.sp. cubense race 4: Indirect evidence for the role of antibiosis in the antagonistic process. Journal of Industrial Microbiology and Biotechnology, 28, 303-310. doi:10.1038/sj.jim.7000247
- Williams, S.T., Goodfellow, M., Alderson, G., Wellington, E.M.H., Sneath, P.H.A. and Sackin, M.J. (1983) Numerical classification of Streptomyces and related genera. Journal of General Microbiology, 129, 1743-1813.
- Jain, P.K. and Jain, P.C. (2007) Isolation, characterization and antifungal activity of Streptomyces sampsonii GS 1322. Indian Journal of Experimental Biology, 45, 203- 206.
- Hamdali, H., Hafidi, M., Virolle, M.J. and Ouhdouch, Y. (2008) Rock phosphatesolubilizing Actinomycetes: Screening for plant growth-promoting activities. World Journal of Microbiology and Biotechnology, 24, 2565-2575. doi:10.1007/s11274-008-9817-0

