Advances in Microbiology
Vol. 3 No. 3 (2013) , Article ID: 33879 , 9 pages DOI:10.4236/aim.2013.33040
Optimization of Cultural Condition and Synergistic Effect of Lactose with Carboxymethyl Cellulose on Cellulase Production by Bacillus sp. Isolated from Fecal Matter of Elephant (Elephas maximus)
1Microbiology Laboratory, Department of Botany, the University of Burdwan, Burdwan, India
2Department of Marine Science, Calcutta University, Kolkata, India
Email: *tkmbu@yahoo.co.in
Copyright © 2013 Sangrila Sadhu et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received May 8, 2013; revised June 10, 2013; accepted June 25, 2013
Keywords: Avicelase; β-Glucosidase; Carboxymethyl Cellulase (CMCase); Lactose
ABSTRACT
A cellulase producing bacterium (E3 strain) was isolated from fecal matter of elephant and identified as Bacillus sp. using 16S rDNA sequenced based molecular phylogenetic approach. While studying the effect of substrates like Carboxymethyl cellulose (CMC), avicel, starch, maltose, sucrose, glucose, fructose, galactose and lactose on cellulase production, it was found that CMC was best carbon source induced cellulase production followed by lactose in this bacterial strain. A positive synergistic effect of lactose with CMC was also observed with enhancement of 5 - 6 times in cellulase production. The optimum cellulase production was recorded with 1% CMC and 1% lactose when added individually in the Omeliansky’s medium. The results showed that addition lactose with CMC greatly enhances the production and activity of various cellulase enzymes. The optimal fermentation conditions for the biosynthesis of cellulase by this strain were found to be temperature: 37˚C, pH 7.0. The nitrogen source NH4Cl at 0.15% was optimum for cellulase production by this bacterium.
1. Introduction
Cellulose is the most abundant and renewable natural biopolymer on earth available for bioconversion to valueadded bio-products. The bioconversion of cellulosic biomass is potentially sustainable approach to develop novel bioprocesses and products. Microbial cellulases have become the important biocatalyst due to their complex nature and wide spread applications. The potential applications of cellulose are in food, animal feed, textile, fuel and chemical industries [1,2]. The major industrial applications of cellulases are in textile industry for biopolishing of fabrics and producing stonewashed look of denims, as well as household laundry detergents for improving fabric softness and brightness [3]. Besides, they are used in animal feeds for improving the nutritional quality and digestibility, in processing of fruit juices and in baking, while de-inking of paper is an another emerging application. Application of enzymes in detergent, leather and paper industries demands identification of highly stable enzymes activity at extreme pH and temperature.
Cellulases contribute to 8% of the worldwide Industrial enzyme demands [4]. The cellulase market is expected to expand dramatically when cellulases are used to hydrolyzed pretreated cellulosic material to sugars, which can be fermented to bioethanol and biobased products on large scales. The cellulase market has been estimated in the United States to be as high as US $400 million per year [5]. In the period 2004-2014 an increase of approximately 100% in the use of cellulase as a specialty enzyme is expected [6]. The biotechnology companies Genencor International and Novozymes Biotech have reported the development of technology that has reduced the cellulase cost for the cellulose-to-ethanol process from US $5.40 per gallon of ethanol to approximately 20% per gallon of ethanol [7]. Three main strategies were: 1) an economical improvement in production of cellulase to reduce US $ per gram of enzyme by process and strain enhancement, e.g., cheaper medium from lactose to glucose and alternative inducer system; 2) isolation of new high-yielding strain; and 3) an improvement in the cellulase enzyme performance to reduce grams of enzyme for achieving equivalent hydrolysis by cocktails and component improvement [8].
Microbial conversion of cellulosic biomass into useful products is a complex process involving the combined action of at least three different enzymes namely endoglucanase (EC 3.2.1.4), exoglucanase (EC 3.2.1.91) or filter paperase (FPase), and β-glucosidase (EC 3.2.1.21) [9]. Amongst the cellulases, exoglucanase are found to have potential applications in the bioconversion of agricultural waste materials to useful products. Exoglucanases are of two types; 1,4-β-D-glucan cellobiohydrolase (EC 3.2.1.91) which removes cellobiose units and 1,4- β-D-glucan glucohydolase (EC 3.2.1.74) which removes glucose units, both acting from the non-reducing ends of oligosaccharides produced by the action of endoglucanase [10,11]. As, cellulases are found to be synthesized and excreted in the media extracellularly, it is also important to estimate the relative ability of a particular cellulase-producing strain to release all the three enzymes and the parameters affecting the synthesis.
Both fungi and bacteria have been exploited for their abilities to produce a wide variety of cellulases. Majority of studies on cellulase production have been studied on fungi. However, bacteria may serve as highly potent sources of industrially important enzymes for the conversion of cellulosic biomass due to their higher growth rate than fungi allowing for higher recombinant production of enzymes; more complex glycoside hydrolases with providing synergy and their extremely high natural diversity have the capability to produce thermostable, alkali stable enzymes. Thus, cellulolytic strains isolated from harsh environment are stable under extreme conditions, therefore, may be used for the bioconversion process which may increase rates of enzymatic hydrolysis, fermentation, and product recovery. Bacteria, due to their high natural diversity, faster growth have the capability to produce highly thermostable, alkali stable enzyme complement and may serve as highly potent sources of industrially important enzymes.
Source of cellulase system extraction is best suitable from microbial system found in the gut of microorganisms thriving on cellulosic biomass as their major feed. The rumen is a complex ecosystem in which feeds consumed by the ruminant animal are digested by an active and diverse micro flora. Cellulolytic bacteria are thought to be the primary agents in the rumen and are considered to be more important [12] for lingo-cellulose degradation. Considering the above fact, the present study was undertaken by isolation and characterization of a cellulolytic strain designated as E3 belonging to the genus Bacillus isolated from fecal matter of elephant. Attempts were also made to investigate how the different cultural conditions and various substrates influenced on enhancing cellulase production with special emphasis on the role of lactose as inducer.
2. Materials and Methods
2.1. Bacterial Strains, Culture Medium, and Screening of Cellulase Producers
The sample was collected from the fecal matter of elephant from Alipore Zoo, Kolkata in sterile container. It was placed on ice and was brought to the laboratory. One gram of fecal matter was diluted with 100 ml of distilled water and was homogenized by constant shaking using orbital shaker for 2 h. The strain was isolated on Omeliansky’s agar medium [g/l (W/Vol) (NH4)2SO4 1.0; K2HPO4 1.0; MgSO4·7H2O 0.5; NaCl 0.1; carboxymethyl cellulose (CMC) 1%, pH 7.0] by dilution plating of the homogenized sample followed by incubation at 37˚C for 4 days [13]. Screening for cellulase producers was carried out on CMC agar plate as per [14].
Strain presenting largest clearing zones in congo red test designated as E3 strain was used for enzyme production on basal salt medium containing 1% CMC as a sole carbon source.
2.2. Genomic DNA Extraction, PCR Amplification of 16S rDNA and Nucleotide Sequencing
Genomic DNA was extracted using Kit Xcelgen as well as quality and purity of DNA was checked by agarose gel electrophoresis [15]. PCR amplification was carried out by using both universal primer sequences (8F: 5’ AGA GTT TGA TCC TGG CTC AG 3’ and 1492R: 5’ ACG GCT ACC TTG TTA CGA CTT 3’. PCR was carried out in a final reaction volume of 25 µl in 200 µl capacity thin wall PCR tube in Eppendorf Thermal Cycler. Composition of reaction mixture of 25 µl contained DNase-RNase free water 7.5 µl, 2 X PCR master mixtures 12.50 µl, each primer concentration 1 µl and diluted DNA (30 ng/µl) of 3.0 µl. PCR tubes containing the mixture were tapped gently and spin briefly at 10,000 rpm. The PCR reaction was run for 30 cycles in a DNA thermal cycler. The following thermal profile was used for the PCR: denaturation at 95˚C for 1 min, primer annealing at 52˚C for 1 min and extension at 72˚C for 2 min. The final cycle included extension for 10 min at 72˚C to ensure full extension of the products. The amplified PCR products were then analyzed in a 1.0% (w/v) agarose gel, excised from the gel and purified. Purification of the amplicons was done by using Qiagen Mini elute Gel extraction Kit and sequencing of 16S rDNA was carried out automated DNA sequencing on ABI 3730 xl Genetic Analyzer (Applied Biosystems, USA). Sequencing was carried out using Big Dye® Terminator v3.1 Cycle sequencing kit.
2.3. Phylogenetic Study by 16S rDNA Sequence
The isolated E3 strain was identified as Bacillus sp. by 16S rDNA gene sequence analysis. A continuous stretch of 1376 nucleotides of 16S rRNA gene was used for phylogenetic analysis. Initially the sequence was edited by sequence scanner (Q value > 20) and the near related sequence were determined using BLAST program of NCBI (www.ncbi.nlm.nih.gov/Bast/nblast). The online programs from RDP database (http://rdp.cme.msu.edu/) like “Classifier” and “Sequence Match” were used to assess phylogenetic locus of culturable bacterial isolates. The “Taxon Passport” was used to retrieve 16S rRNA gene sequence from type strains of various Bacillus spp. During sequence analysis, the sequence similarity of all the sequence were calculated manually after aligning it in “align by muscle” programmed in MEGA 5.1 version and then looking for mismatches versus total number of nucleotides used in such comparison for phylogenetic analysis 16S rDNA sequences from type strains of different species of Bacillus showing sequence identity greater than 99% with the strain E3 were considered in this study. Phylogenetic tree was constructed according to the Kimura 2 parameter model [16] using the MEGA 5.1. The statistical significance of branch points was calculated by 1000 bootstrap re-samplings of the data [17].
2.4. Optimization of Cultural Condition for Enzymes Production
In order to develop a more suitable carbon sources, nitrogen sources for cellulose secretion, different carbon sources and nitrogen sources are tested in Omeliansky’s medium to improve the medium. Similarly, to detect more suitable cultural condition like incubation period, pH and temperature, different incubation period, pH and temperature are used. Cultures (50 ml in 250 ml flask) were incubated at 37˚C on a rotary shaker set at 150 rpm. This incubation condition was used throughout the studies reported here the except the temperature optimization experiment where different temperature were used. After cell growth, supernatant, also referred to as crude enzymes after centrifugation at 10,000 × g for 15 min from these cultures were examined for enzymes assay.
2.5. Enzyme Assay
The filter paper (FPase) activity of cellulase was measured as described by Ghosh [18]. The CMCase activity was assayed based on that of Miller [19]. 0.5 ml of culture supernatant fluid was incubated with 0.5 ml of 1% CMC in 0.05 M sodium acetate buffer, pH 4.8 at 40˚C for 1 h. Avicelase and β-glucosidase activity were determined under similar conditions, except that 1% avicel and 1% salicin were used as substrate. The reduced sugar released was analyzed using the dinitrosalicylic (DNS) method, using glucose as the standard sugar. A unit of enzyme activity was defined as the amount required producing 1 μ mole of reducing sugars per min. The specific activity is the number of units of enzyme activity per milligram of enzyme protein. The soluble protein concentration was determined by method of Lowry et al. [20] using BSA as standard.
2.6. Statistical Analysis
Values are the mean ± SEM of 3 replicates. All data were subjected to students t-test analysis with significance level of P < 0.05 using SPSS soft were package.
3. Result and Discussion
3.1. Isolation and Identification of Cellulolytic Bacteria
A cellulase-producing strain designated as E3 was isolated from fecal matter of Elephant (Elephas maximus ) and was identified to be a species of the genus Bacillus sp. by 16S rRNA gene sequence analysis. As evident from neighbor-joining phylogenetic (NJ) tree (Figure 1), the strain E3 together with Bacillus subtilis subsp. strain DSM 10 formed a cluster that is also well within the cluster represented by Bacillus subtilis subsp. spizizenii strain NRRL B-23049. The strain E3 also show closest sequence similarity 99% with Bacillus subtilis subsp. strain DSM 10 followed by Bacillus subtilis subsp. spizizenii strain NRRL B-23049. Such high percentage of sequence similarity and phylogenetic positioning clearly indicates that the strain E3 is definitely a species of Bacillus. However, in absence of detailed phenotypic characters, chemotaxonomy and overall genome sequence relatedness data, the E3 strain was concluded as Bacillus sp. pending further investigation of species status.
3.2. Effect of Different Carbon Sources on Cellulase Production
It is apparent from literature that cellulose production depends on nature of carbon sources. Different fermentable sugars have been shown to either induce or inhibit cellulose production depending on individual species. To decipher the suitable sugar for the E3 strain, carboxymethyl cellulose (CMC), avicel, starch, maltose, sucrose, glucose, fructose, galactose and lactose were tested. Strain E3 utilized all carbon sources (Table 1) for growth and cellulase production. But CMC and lactose exhibited extensive role to enzyme yield and production. Among different carbon sources CMC was the best carbon
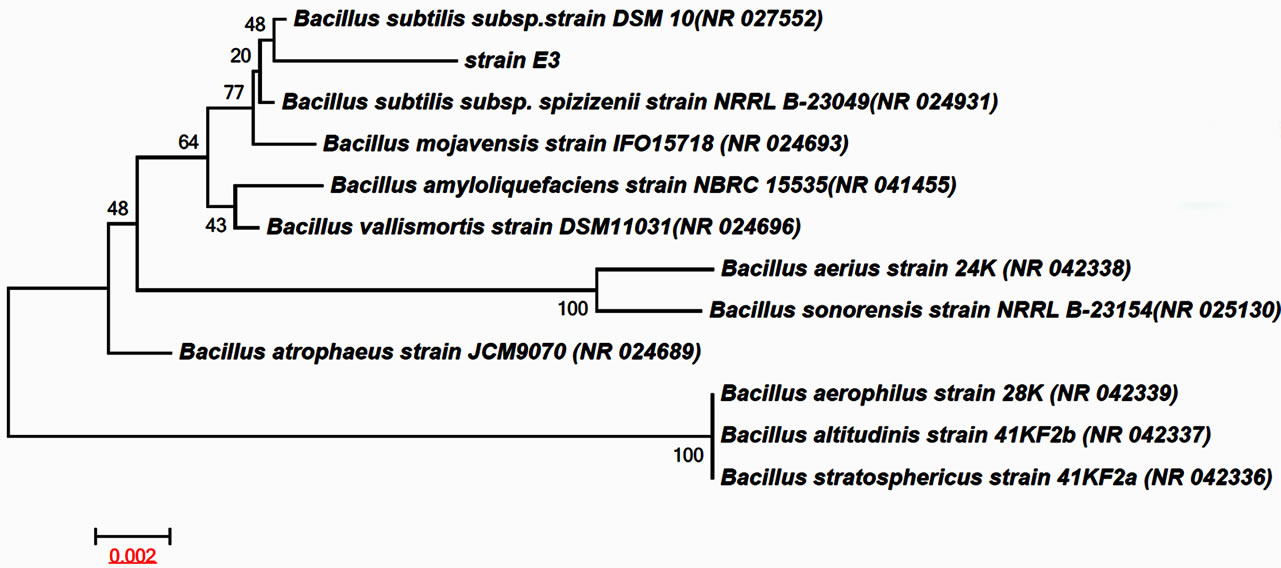
Figure 1. Neighbour-joining phylogenetic tree showing relationship of strain E3 with in different type strain of the genus Bacillus sp. The tree was generated using MEGA 5.1 software Kimura (1980) model. Bootstrap values of 1000 replications are shown at the nodes. Bar 0.002 substitutions per site.
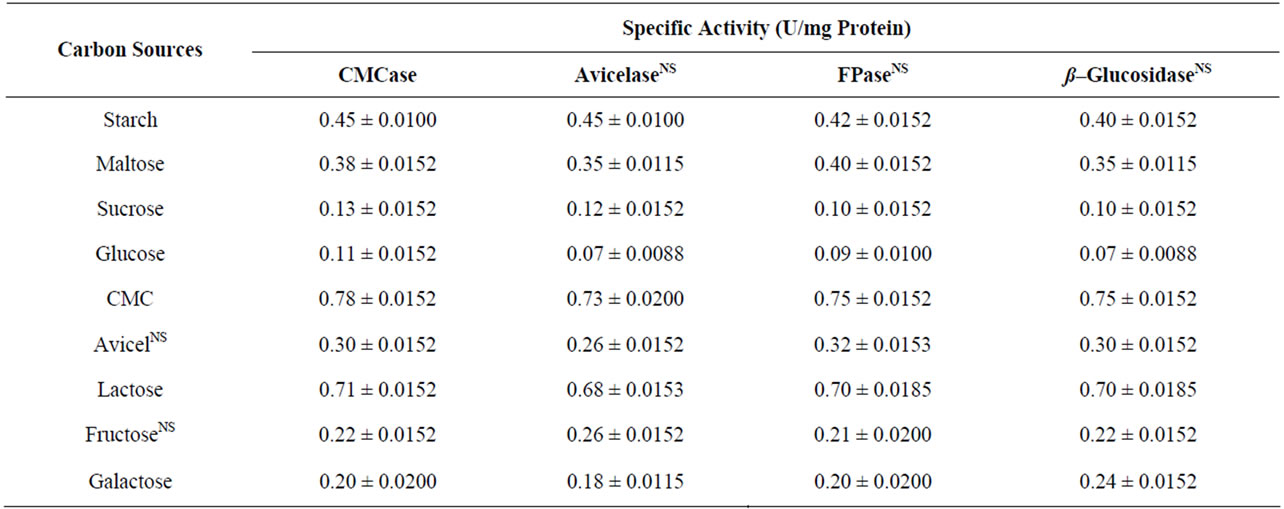
Table 1. Effect of different carbon sources (1% W/V) on cellulase production by E3 after 8 d of incubation at 37 ºC. The results of post hoc Tukey test showed significant differences between the paired variables under carbon sources and enzyme types except those presented with NS in superscript.
source followed by lactose for cellulose production. The data of Table 1 is in accordance with the result of Microbacterium sp. and and Bosea sp. [21,22].
3.3. Effect of Incubation Days on Cellulase Production
Incubation time is necessary for optimal production of enzymes. In the present study the enzyme production was increasing steadily from the beginning and reached the maximum at 10 d of incubation, then the level start decreasing (Figure 2). Similar result was found in Microbacterium sp. [21]. It was observed that Streptomyces sp. BRC1 and BRC2 gradually raised endoglucanase synthesis and reached maximum activity at 3 d, after that enzyme activity slowly decreased [23]. In Bosea sp., the maximum activity was reached at 8 d of incubation [22].
3.4. Optimum Concentration of CMC
The different concentrations of CMC were tested for cellulase production, among which 1% CMC was optimum for this strain (Figure 3). Above this concentration cellulase production was inhibited. Similarly cellulase production was inhibited by 1% cellulose in Thermomonospora curvata isolated from municipal solid waste
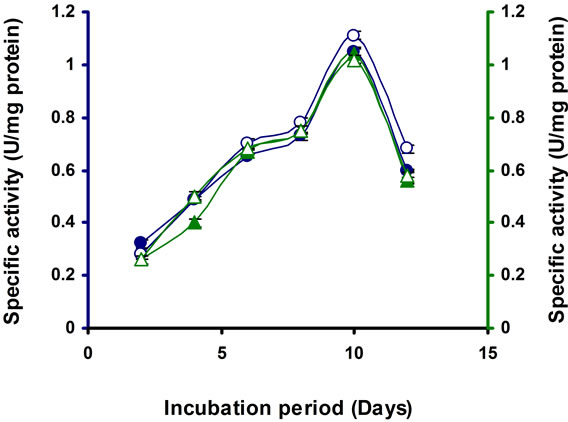
Figure 2. Time course of CMCase, Avicelase, FPase and β-glucosidase of crude enzyme produced by culture medium containing 1% CMC at 37˚C by Bacillus sp. Symbols: Open circles, CMCase; closed circles, Avicelase; open triangles, FPase; closed triangles, β-glucosidase.
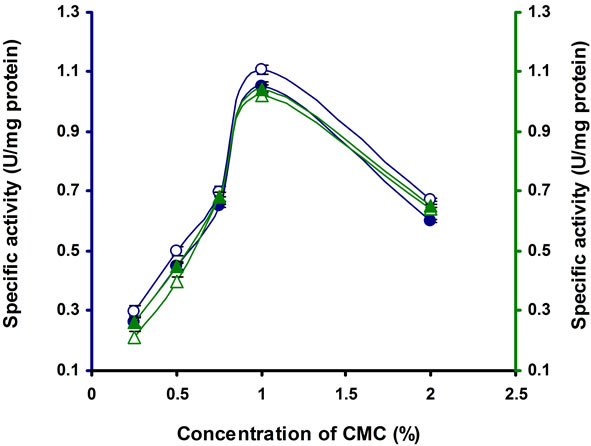
Figure 3. CMCase, Avicelase, FPase and β-glucosidase of crude enzyme produced by culture medium containing different concentrations of CMC at 37˚C after 10 d by Bacillus sp. Symbols: Open circles, CMCase; closed circles, Avicelase; open triangles, FPase; closed triangles, β-glucosidase.
compost [24]. The optimum concentrations of CMC were found to be 3% and 0.75% in Microbacterium sp. and Bosea sp. respectively [21,22].
3.5. Effect of Temperature on Cellulase Production
Temperature is the important factors for growth of microorganism and production of enzyme. It was found from Figure 4, the strain E3 gradually raised cellulase synthesis and reached maximum activity at 37˚C, after that cellulase production slowly decreased. Similar result was found in Bosea sp. [22]. The optimum temperature is 26˚C in Streptomyces sp. BRC1 and BRC2 [23] but 52˚C in Micromonospora chalcae [25]. So the different bacterial strains able to produced this enzyme at different op-
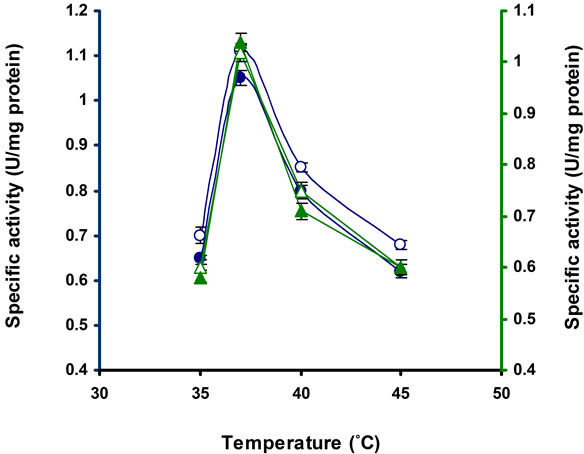
Figure 4. Production of cellulases by Bacillus sp. in broth culture containing 1% CMC after 10 d of incubation at different temperatures. Symbols: Open circles, CMCase; closed circles, Avicelase; open triangles, FPase; closed triangles, β-glucosidase.
timal temperature.
3.6. Effect of pH on Cellulase Production
As shown in Figure 5 the pH optima for the cellulase productivity was 7.0 of the E3 strain. Beyond this limit of pH, the enzyme productivity was decreased. Similar results have been found in Bosea sp. [22], Streptomyces sp. F2621 [26], Streptomyces BRC1 and BRC2 [23], Clostridium sp. [27], Streptomyces sp. AT7 [28] and Anoxybacillus flavithermus EHP1 [29].
3.7. Effect of Different Nitrogen Sources on Cellulase Production
The enzyme production is affected significantly under different concentration of the organic and inorganic nitrogen sources. The production media incorporated with different inorganic nitrogen source, supported to bring more cellulase activity by Bacillus sp. Among the different nitrogen sources tested, the enzyme activity was higher with NH4Cl (Table 2). To find out the suitable concentration of NH4Cl, different concentrations of NH4Cl tested, among which 0.15% NH4Cl was optimum for this strain (Figure 6). Similar results have been found in Bosea sp. [22]. Meat extract and tryptone (1%) served as intensive sources to Streptomyces sp. BRC1 and yeast extract (1%) suited for Streptomyces sp. BRC2 [23]. Organic N source were found to be more suitable than inorganic N sources for optimizing cellulose production Bacillus sp. [30,31]. Rajoka [30] reported KNO3 and NH4NO3 as the best N sources for cellulose production in Cellulomonas flavigena. NH4 compounds were considered as most favourable N sources for cellulase synthesis as noted in Thermomonospora fusca [32].
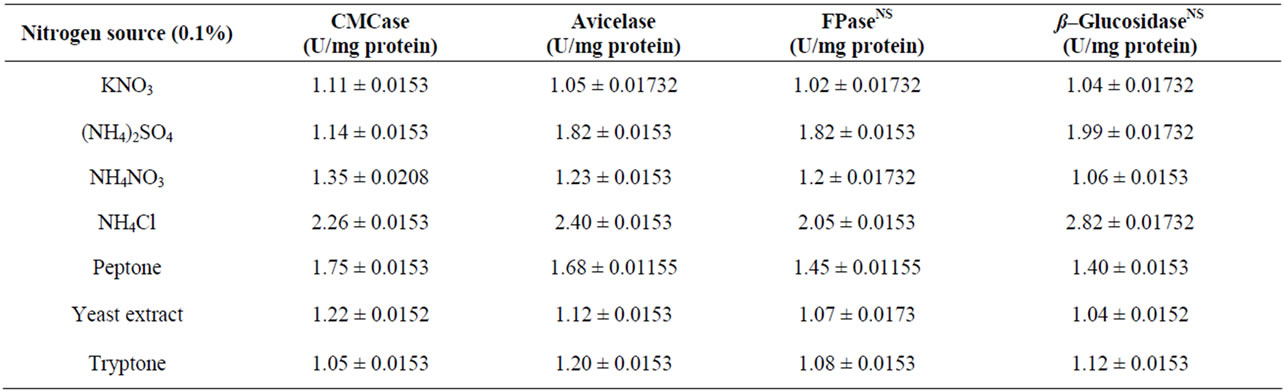
Table 2. Effect of different Nitrogen sources (0.1% W/V) on cellulase production by Bacillus sp after 10 d of incubation at 37˚C and pH 7.0. The results of post hoc Tukey test showed significant differences between the paired variables under nitrogen sources and enzyme types except those presented with NS in superscript.
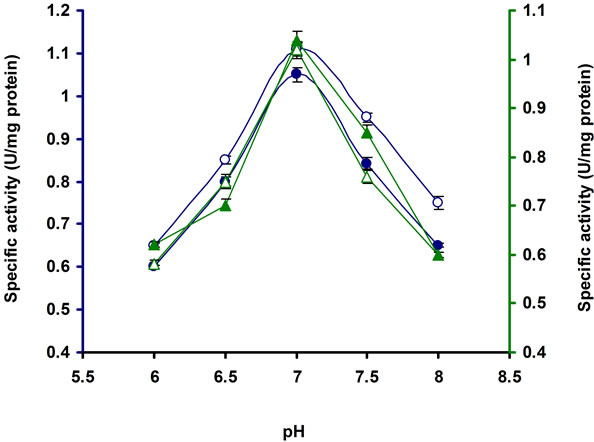
Figure 5. Production of cellulases by E3 in broth culture containing 1% CMC after 10 days of incubation at 37˚C at different pH. Symbols: open circles, CMCase; closed circles, Avicelase; open triangles, FPase; closed triangles, β-glucosidase.
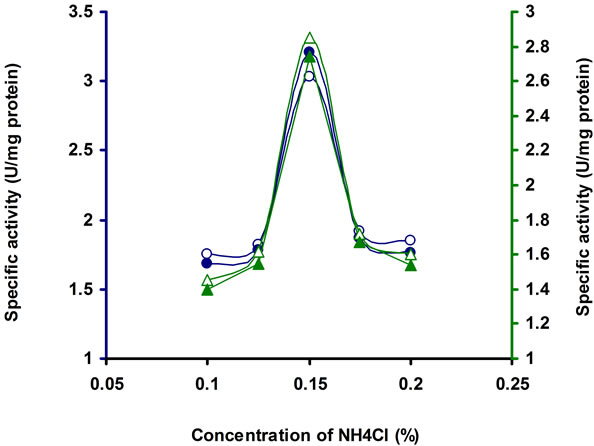
Figure 6. CMCase, Avicelase, FPase and β-glucosidase of crude enzyme produced by culture medium containing different concentrations of NH4Cl at 37˚C after 10 d by Bacillus sp. Symbols: open circles, CMCase; closed circles, Avicelase; open triangles, FPase; closed triangles, β-glucosidase.
3.8. Optimum Concentration of Lactose
It was observed from the Table 1 the lactose was the penultimate carbon source for cellulase production after CMC for this strain. It was observed that 1% lactose was optimum for cellulase production of this strain (Figures 7(a) and (b)). A repeated measure ANOVA justified the differences on the supplement added to the medium, the time (in days) of incubation and the activity of the different enzyme by this Bacillus sp. strain E3. It is apparent from the multivariate test that significant differences were obvious between the enzyme types and their peak activity in respect to the incubation periods and the media.
3.9. Effect of Lactose on Cellulase Production
Most microbial cellulases are induced in presence of cellulose but cellulose itself cannot directly trigger the induction as it is insoluble. A basal level of cellulases production occurs in the absence of glucose. The soluble saccharides such as cellobiose, sorphorose, lactose, trehalose, sorbose, and galactose might serve as inducers for cellulase synthesis as reported in Clostridium papyrosolvens [33], Acidothermus cellulolyticus [34]. Cellobiose serves as inducer in Bacillus sp. [35] and Anoxybacillus sp. 527 [36]. We therefore, investigated the synergistic effect of lactose with CMC on induction of cellulase production by E3 strain as lactose was the penultimate suitable carbon source for this E3 strain (Table 1). The specific activity of CMCase, Avicelase, FPase and β-glucosidase were recorded highest when 1% CMC supplemented with 1.0% lactose (Table 3, Figure 8). Activities of same enzymes under similar condition were also lower when the substrates (CMC or Lactose) were added individually. This result indicated that Lactose synergistically enhanced the cellulase synthesis. Similar results reported earlier with a fungus Hypocrea jecorina [37] and in bacteria Microbacterium [21]. However, the
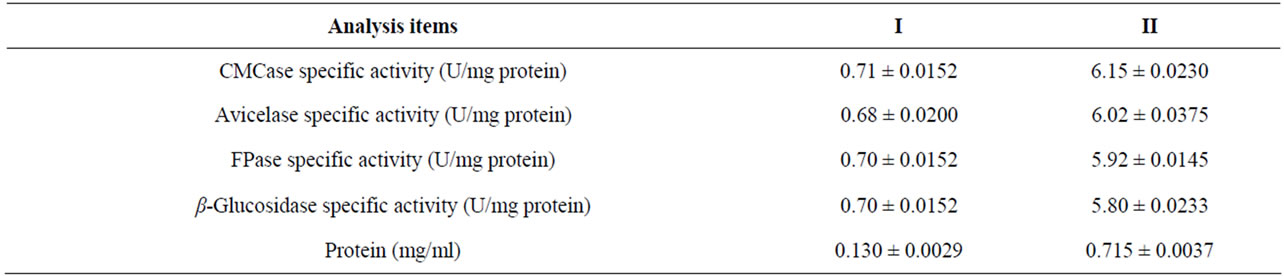
Table 3. Effect of Lactose: Culture medium containing (I) 1% (W/L) CMC as carbon source over 8d or (II) 1% CMC + 1% Lactose over 10 d at 37˚C by E3 strain. All t-values are significantly different at P < 0.001 level.
 (a)
(a)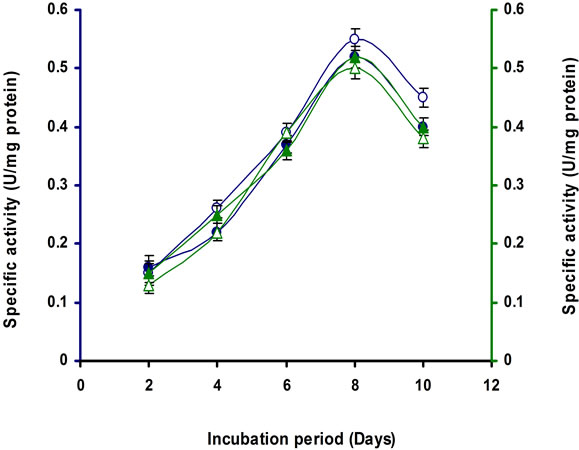 (b)
(b)
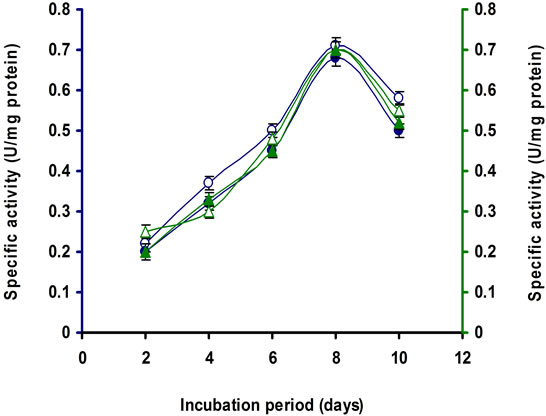
Figure 7. Time course of CMCase, Avicelase, FPase and β- glucosidase of crude enzyme produced by culture medium containing (a) 1% Lactose; (b) 2% Lactose at 37˚C by Bacillus sp. Symbols: open circles, CMCase; closed circles, Avicelase; open triangles, FPase; closed triangles, β-glucosidase.
mechanism by which lactose triggers the formation of cellulase is unknown; Seiboth et al. stated that lactose might act as an inducer of cellulase formation rather than promoting cellulase biosynthesis by relieving the carbon catabolite repression.
Lactose consists of D-galactose and D-glucose. Cellu-
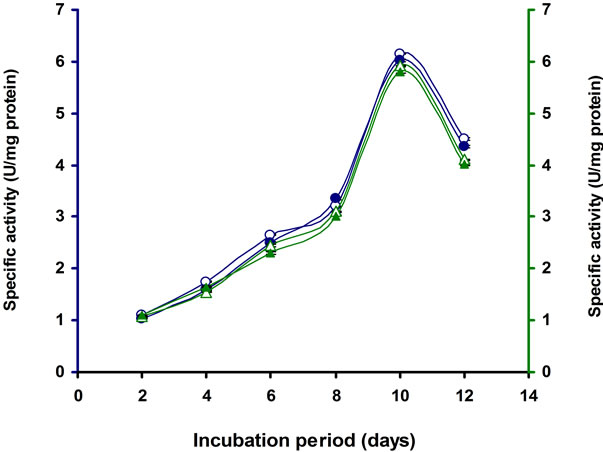
Figure 8. Time course of CMCase, Avicelase, FPase and β-glucosidase of crude enzyme produced by culture medium containing 1% CMC + 1% Lactose at 37˚C by Bacillus sp. Symbols: open circles, CMCase; closed circles, Avicelase; open triangles, FPase; closed triangles, β-glucosidase.
lase synthesis cannot be induced by galactose or glucose individually (Table 1). This is reflected through the two tailed paired t-test carried out on the specific activity of the enzymes under the two different media –1% CMC and 1% CMC + 1% Lactose, with significantly higher activity in lactose containing medium. Protein yield was also higher in the medium containing lactose. Reports from literature suggested that glucose inhibited cellulase synthesis [38] and lactose induces significantly higher cellulase levels than D-galactose in T. reesei [39]. The induction mechanism of cellulase formation is studied in fungus by many workers [40] and in bacteria [21]. We therefore consider that lactose is a water soluble disaccharide in the mixed medium with CMC and was fast taken up by Bacillus sp. and produced cellulase initially which would enable an initial attack on CMC followed by more cellulase biosynthesis. Thus, the cellulase production was remarkably increased when lactose was added with CMC as a carbon source. Lactose is a powerful inducer that generally enhances the cellulase yield in this organism by stimulating secretion of various proteins with cellulase activity.
4. Acknowledgements
Financial support for the first author provided by University Grant Commission through The University of Burdwan is gratefully acknowledged.
REFERENCES
- P. Beguin and J. P. Aubert, “The Biological Degradation of Cellulose,” FEMS Microbiology Reviews, Vol. 13, No. 1, 1994, pp. 25-58. doi:10.1111/j.1574-6976.1994.tb00033.x
- M. P. Coughlan, “Cellulose Degradation by Fungi, in Microbial Enzymes and Biotechnology,” In: W. Fogarty and C. Kely, Eds., Microbial Enzymes and Biotechnology, Elsevier, London, 1990, pp. 1-36. doi:10.1007/978-94-009-0765-2_1
- A. Cavaco-Paulo, “Mechanism of Cellulose Action in Textile Processes,” Carbohydrate Polymers, Vol. 37, No. 3, 1998, pp. 273-277. doi:10.1016/S0144-8617(98)00070-8
- E. P. S. Bon and A. F. Maria, “Bioethanol Production via Enzymatic Hydrolysis of Cellulosic Biomass,” FAO Seminar on the Role of Agricultural Biotechnologies for Production of Bioenergy in Developing Countries, Rome, 12 October 2007, pp. 1-11. http://www.fao.org/biotech/seminaroct 2007.htm
- Y. H. P. Zhang, M. E. Himmel and J. R. Mielenz, “Outlook of cellulase Improvement: Screening and Selection Strategies,” Biotechnology Advances, Vol. 24, No. 5, 2006, pp. 452-481. doi:10.1016/j.biotechadv.2006.03.003
- R. B. Costa, M. V. A. Silva, F. C. Freitas, V. S. F. Leita˜o, P. S. B. Lacerda, M. A. Ferrara and E. P. S. Bon, “Mercado e Perspectivas de Uso de Enzimas Industriais e Especiais no Brasil,” In: E. P. S. Bon, M. A. Ferrara, M. L. Corvo, A. B. Vermelho, C. L. A. Paiva, R. B. Alencastro and R. R. R. Coelho, Eds., Enzimas em Biotecnologia, Produc¸a˜o, Aplicac ¸o˜es e Mercados, Intercieˆncia, Rio de Janeiro, 2008, pp. 463-488.
- N. Moreira, “Growing Expectations: New Technology Could Turn Fuel into a Bump Crop,” Science News Online, Vol. 168, No.14, 2005, pp. 209-224.
- M. Knauf and M. Moniruzzaman, “Lignocellulosic Biomass Processing: A Perspective,” International Sugar Journal, Vol. 106, No. 1263, 2004, pp. 147-150.
- K. E. Erikson and B. Patterson, “Extra Cellular Enzyme System Produced by the Fungus Sporotrichum pulverulentum,” Biotechnology and Bioengineering, Vol. 20, No. 3, 1975, pp. 317-332.
- R. Mullings, “Measurement of Sacharification by Cellulases,” Enzyme and Microbial Technology, Vol. 7, No. 12, 1985, pp. 586-591. doi:10.1016/0141-0229(85)90025-0
- Y. H. P. Zhang and L. R. Lynd, “Toward an Aggregated Understanding of Enzymatic Hydrolysis of Cellulose Systems,” Biotechnology and Bioengineering, Vol. 88, No. 7, 2004, pp. 797-824. doi:10.1002/bit.20282
- C. S. Stewart, H. J. Flint and M. P. Bryant, “The Rumen Bacteria,” In: P. N. Hob-son and C. S. Stewart, Eds., The Rumen Microbial Ecosystem, Blackie Academic and Professional Publishers, London, 1997, pp. 10-72.
- V. Omeliansky, “Ueber die Garung der Cellulose,” Proceedings of Indiana Academy of Science, Vol. 8, 1902, pp. 225-231.
- R. M. Teather and P. J. Wood, “Use of Congo Red-Polysaccharide Interactions in Enumeration and Characterization of Cellulolytic Bacteria from the Bovine Rumen,” Applied and Environmental Microbiology, Vol. 43, No. 4, 1982, pp. 777-780.
- J. Sambrook and D. W. Russell, “Molecular Cloning: A Laboratory Manual,” 3rd Edition, Cold Spring Harbor Laboratory Press, New York, 2001.
- M. Kimura, “A Simple Method for Estimating Evolutionary Rates of Base Substitutions through Comparative Studies of Nucleotide Sequences,” Journal of Molecular Evolution, Vol. 16, No. 2, 1980, pp. 111-120. doi:10.1007/BF01731581
- J. Felsenstein, “Confidence Limits on Phylogenies: An Approach Using the Bootstrap,” Evolution, Vol. 39, No. 4, 1985, pp. 783-791. doi:10.2307/2408678
- T. K. Ghosh, “Measurement of Cellulase Activities,” Pure and Applied Chemistry, Vol. 59, No. 2, 1987, pp. 257-268. doi:10.1351/pac198759020257
- G. L. Miller, “Use of Dinitrosalicylic Acid Reagent for Determination of Reducing Sugars,” Journal of Analytical Chemistry, Vol. 31, No. 3, 1959, pp. 426-428. doi:10.1021/ac60147a030
- O. H. Lowry, N. J. cRosebrough, A. L. Farr and R. J. Randall, “Protein Measurement with the Folin-Phenol Reagent,” The Journal of Biological Chemistry, Vol. 193, No. 1, 1951, pp. 265-275.
- S. Sadhu, P. Saha, S. Mayilraj and T. K. Maiti, “LactoseEnhanced Cellulase Production by Microbacterium sp. Isolated from Fecal Matter of Zebra (Equus zebra),” Current Microbiology, Vol. 62, No. 3, 2011, pp. 1050-1055. doi:10.1007/s00284-010-9816-x
- S. Sadhu, P. Saha, S. Mayilraj and T. K. Maiti, “Characterization of a Bosea sp. Strain SF5 (MTCC 10045) Isolated from Compost Soil Capable of Producing Cellulase,” The Journal of Microbiology, Biotechnology and Food Sciences, Vol. 2, No. 2, 2012, pp. 576-591.
- P. Chellapandi and M. J. Himanshu, “Production of endoglucanase by the native strains of Streptomyces isolates in submerged fermentation”, Brazilian Journal of Microbiology, Vol. 39, 2008, pp. 122-127. doi:10.1590/S1517-83822008000100026
- E. Stackebrandt and B Goebel, “Taxonomic Note: A Place for DNA-DNA Reassociation and 16S rRNA Sequence Analysis in the Present Species Definition in Bacteriology”, International Journal of Systematic Bacteriology, Vol. 44, No. 4, 1994, pp. 846-849. doi:10.1099/00207713-44-4-846
- J. Gallagher, A. Winters, N. Barron, L. McHale and A.P McHale, “Production of Cellulase and β-Glucosidase Activity during Growth of the Actinomycete Micromonospora chalcae on Cellulose-Containing Media,” Biotechnology Letters, Vol. 18, No. 5, 1996, pp. 537-540. doi:10.1007/BF00140199
- M. Tuncer, A. Kuru, M. Isikli, N. Sahin and F. G. Celenk, “Optimization of Extra-Cellular Endoxylanase, Endoglucanase and Peroxidase Production by Streptomyces sp. F2621 Isolated in 17 Turkey,” Journal of Applied Microbiology, Vol. 97, No. 4, 2004, pp. 783-791. doi:10.1111/j.1365-2672.2004.02361.x
- B. H. Lee and T. H. Blackburn, “Cellulase Production by a Themophilic Clostridium sp.,” Applied Microbiology, Vol. 30, No. 3, 1975, pp. 346-353.
- M. A. T. Amira, A. R. Shatha, S. A. A. Siham and A. N. Basima, “Cellulase Production from Actinomycetes Isolated from Iraqi Soils: II Cell Growth and Cellulase Activity of Streptomyces sp. Strain AT7 at Different Temperatures,” Journal of Islamic Academy of Sciences, Vol. 2, No. 3, 1989, pp. 185-188.
- A. S. S. Ibrahim and I. Ahmed, “Isolation and Identification of New Cellulases Producing Thermophilic Bacteria from an Egyptian Hot Spring and Some Properties of the Crude Enzyme,” Australian Journal of Basic and Applied Science, Vol. 1, No. 4, 2007, pp. 473-478.
- M. I. Rajoka, “Influence of Various Fermentation Variable on Exoglucanase Production in Cellulomonas flavigena,” Electronic Journal of Biotechnology, Vol. 7, No. 3, 2004, pp. 259-266. doi:10.2225/vol7-issue3-fulltext-2
- S. Acharya and A. Choudhury, “Effect of Nutritional and Environmental Factors on Cellulose Activity by Thermophillic Bacteria Isolated from Hot Spring,” Journal of Scientific and Industrial Research, Vol. 70, No. 2, 2011, pp. 142-148.
- N. A. Spiridonov and D. B. Wilson, “Regulation of Biosynthesis of Individual Cellulases in Thermomonospora fusca,” Journal of Bacteriology, Vol. 180, No. 4, 1998, pp. 3529-3532.
- S. Thirumale, R. D. Swaroopa and K. Nand, “Control of Cellulose Formation by Trehalose in Clostridium Papyrosolvens CFR-703,” Process Biochemistry, Vol. 37, No. 3, 2001, pp. 241-245. doi:10.1016/S0032-9592(01)00212-6
- M. Shiang, J. C. Liden, A. Mohagheghi, K. Grohmann and M. E. Himmel, “Regulation of Cellulose Synthesis in Acidothermus cellulolyticus,” Biotechnology Progress, Vol. 7, No. 4, 1991, pp. 315-322. doi:10.1021/bp00010a005
- L. M. Robson and G. H. Chambliss, “Characterization of the Cellulolytic Activity of a Bacillus Isolate,” Applied and Environmental Microbiology, Vol. 47, No. 5, 1984, pp. 1039-1046.
- L. Y. Liang, Z. Feng, J. Yesuf and J. W. Blackburn, “Optimization of Growth Medium and Enzyme Assay Conditions for Crude Cellulases Produced by a Novel Thermophilic and Cellulolytic Bacterium Anoxybacillus sp. 527.” Applied Biochemistry and Biotechnology, Vol. 160, No. 6, 2010, pp. 1841-1852. doi:10.1007/s12010-009-8677-x
- B. Seiboth, G. Hofmann and C. P. Kubicek, “Lactose Metabolism and Cellulase Production in Hypocrea jecorina: The Gal 7 Gene, Encoding Galactose-1-phosphate Uridylyltransferase, Is Essential for Growth on Galactose but Not for Cellulase Induction,” Molecular Genetics and Genomics, Vol. 267, No. 1, 2002, pp. 124-132. doi:10.1007/s00438-002-0654-9
- M. Ilman, A. Saloheimo, M. L. Onnela and M. E. Penttila, “Regulation of Cellulase Gene Expression in the Filamentous Fungus Trichoderma reesei,” Applied and Environmental Microbiology, Vol. 63, No. 4, 1997, pp. 1296-1306.
- L. Karaffa, E. Fekete, C. Gamauf, A. Szentirmai, C. P. Kubicek and B. Seiboth, “Dgalactose Induces Cellulase Gene Expression in Hypocrea jecorina at Low Growth Rates,” Microbiology, Vol. 152, No. 5, 2006, pp. 1507-1514. doi:10.1099/mic.0.28719-0
- M. Schmoll and C.P. Kubicek, “Regulation of Trichoderma Cellulose Formation: Lessons in Molecular Biology from an Industrial Fungus,” Acta Microbiologica et Immunologica Hungarica, Vol. 50, No. 2, 2003, pp. 125-145. doi:10.1556/AMicr.50.2003.2-3.3
NOTES
*Corresponding author.

