Advances in Biological Chemistry
Vol. 2 No. 4 (2012) , Article ID: 24648 , 7 pages DOI:10.4236/abc.2012.24046
Elimination of a disulfide bridge in Aspergillus niger NRRL 3135 Phytase (PhyA) enhances heat tolerance and optimizes its temperature versus activity profile
![]()
1Southern Regional Research Center, ARS, USDA, New Orleans, USA
2Department of Animal Science, Cornell University, New York, USA
Email: *abul.ullah@ars.usda.gov
Received 1 August 2012; revised 3 September 2012; accepted 12 September 2012
Keywords: Phytase; Disulfide Bridge; Aspergillus niger; Site-Directed Mutagenesis
ABSTRACT
In this study, the optimum temperature was lowered while the residual phytase activity after heating to 70˚C was raised in a widely utilized phytase, Aspergillus niger NRRL 3135 PhyA. This was accomplished by site-directed mutagenesis of the cysteines that are involved in the formation of a single disulfide bridge (DB). When compared to wild type (WT), three of the four mutant phytases displayed a lower optimum temperature, 42˚C, and up to a four-fold increase in activity after heating. These findings have a potentially broad application to be incorporated along with other desirable features to engineer a phytase with superior physical and chemical attributes for animal feed applications.
1. INTRODUCTION
Several published studies have focused on engineering enhanced heat tolerance in histidine acid phytases (HAPhy) for use by the animal feed industry [1]. Much of the feed formulations now contain high levels of phytic acid originating from various plant components, and the addition of a phytase allows monogastric animals (swine, poultry, etc.) which lack the enzyme in their digestive tract to hydrolyze the phytic acid thus enabling nutritional benefit from a phytate rich diet [2]. Heat tolerance is desirable when the feed is pelletized because the extrusion process requires brief exposure to elevated temperature. Therefore, for any phytase to be commercially viable, it cannot be irreversibly denatured from heating even when exposed briefly.
To identify sources of and means to achieve thermostability, some of the earlier studies have included identification of new sources of microbial heat tolerant phytase [3-7] comprising amino acid (AA) sequences of known HAPhys and engineering a “consensus phytase” [8], adding a thermally stable protectant [9], immobilizing the enzyme [10], increasing the extent of glycosylation [11], modifying selected hydrogen bond and increasing ionic interaction [12], combining a number of thermostability mutations [13] and engineering of a structure-based chimeric HAPhy [14]. The primary objective of this research was not to increase the optimum temperature of the enzyme, but to develop strategies that would increase tolerance to a brief exposure to 70˚C - 90˚C and still maintain maximum phytase activity in 36˚C - 40˚C temperature range.
Aspergillus niger NRRL 3135 PhyA is a widely marketed phytase targeted for the animal feed industry. Like the other commercially available phytases, it is a histidine acid phosphatase and collectively termed HAPhy. As filamentous ascomycetes HAPhys, A. niger PhyA has five well conserved disulfide bridges (DB) that have roles in maintaining the proper three-dimensional structure of the catalytic center so that substrates can bind to the substrate-binding site while its phosphomonoester bond can be cleaved efficiently [15].
The vital function of DBs in fungal HAPhys has been previously established [15,16]. Some DB studies on phytases that do not belong in the histidine acid phosphatase class of enzyme have been conducted [17,18]; however, because of the structural differences, these studies probably only have minor or no relevance to the HAPhys. Site-directed mutants of all five of the A. niger NRRL 3135 PhyA DBs were investigated [16,19]. Elimination of the second bridge, which is very crucial for bringing the C-terminal end of the protein closer to the active center, resulted in a total loss of activity. Single cysteine mutants of the other four DB resulted in lower optimum temperature and modification of the pH profile in all the mutant phytases.
In that study, only a single cysteine in each pair was changed to eliminate the DB [16]. All the individual DB mutants displayed altered phytase activity except for the second DB mutant where activity was totally lost. The catalytic demise in the second DB mutant was not unexpected because the non-consecutive cysteines bring two remote areas of the enzyme in close contact with each other to form the active site. Among the altered phenotypes where activity was discernible, a decrease in optimum temperature was noticed. The optimum temperature for the native A. niger PhyA is 58˚C which is much higher than the body temperature of both poultry and swine. Therefore, lowering the optimum temperature of PhyA closer to the physiological body temperature of animals has a potential benefit if the DB mutant maintains the other desirable attributes of the WT phytase.
In an investigation of the role of protein-disulfide isomerase, DsbC, in Escherichia coli phytase (AppA), a series of DB mutants were tested [20]. The authors reported that all the double DB mutants displayed higher specific enzyme activity than the corresponding single mutant for each bridge. In fact, one mutant (C200S/ C210S) had over 120% of wild type (WT) AppA phosphatase activity while the single mutant C200S displayed only approximately 70% of the WT phytase activity [20].
To determine if a similar activity enhancement could be achieved in A. niger PhyA, a series of additional single and double mutations for the first DB were constructed and the resultant mutants were evaluated for activity. The first such DB was selected for this study, which is located towards the N-terminal end of the enzyme and pairs two consecutive cysteines, C31 and C40 (NCBI # AAA32705) (Figure 1). The distance between this DB and the beginning of the active site (R81) was computed to be 2.2 nm. The selection of the first DB was based on it having the best kinetic perfection among all the previous DB mutants [16] and it also being the least conserved DB among all the fungal HAPhys [21]. The earlier study also revealed that this DB is not critical for
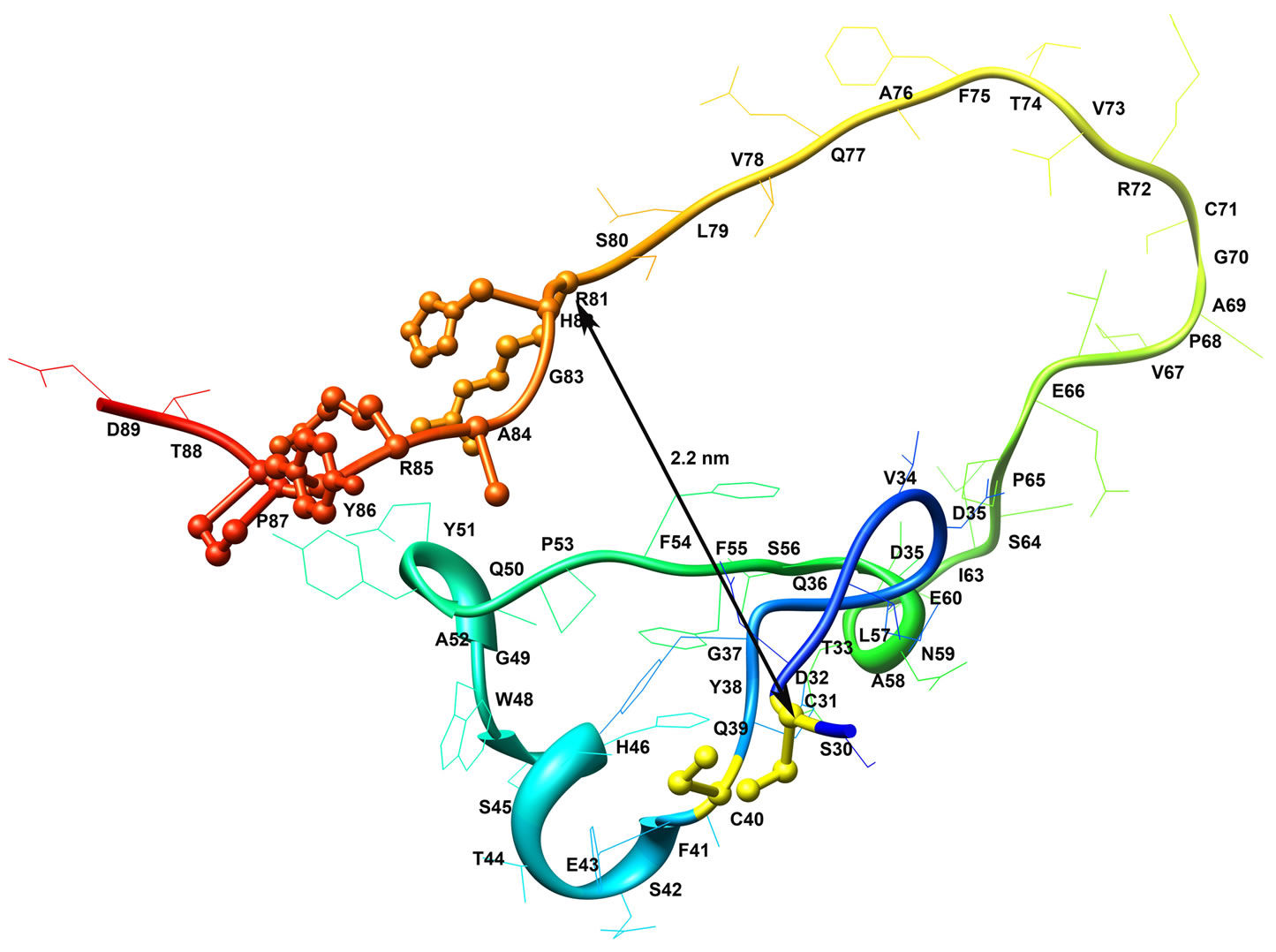
Figure 1. Molecular model of the first 89 residues of A. niger PhyA phytase starting from residue number 30, which is based on three-dimensional structure (1IHP). The first disulfide bridge is located in the N-terminal region, which is formed between C31 and C40 cysteines. The distance between the first disulfide bridge and the beginning of the active site (R81) was computed to be 2.2 nm.
catalytic activity. Along with the lowering of the optimum temperature, a significant enhancement in heat tolerance was achieved in the double mutants. This optimization of temperature to match the physiological temperature of the animals and elevated thermostability may have commercial potential either alone or when incorporated with other desirable features for next generation of engineered HAPhys.
2. MATERIALS AND METHODS
2.1. Media and Reagents
Escherichia coli JM 109 cells (Stratagene, Cedar Creek, TX) or E. coli Dα competent cells (Invitrogen, San Diego, CA) were cultured at 37˚C in Luria Broth (LB) medium. Pichia pastoris strain X33 from Invitrogen (Carlsbad, CA) was grown at 30˚C as previously described [12]. The antibiotic Zeocin (Invitrogen, San Diego, CA) was added at 100 μg·mL–1 to yeast extractpeptone-dextrose (YPD) or Buffered Minimal Glycerol media (BMGY) or Buffered Minimal Methanol (BMMY) media for yeast and at 25 μg·mL–1 to LB for E. coli.
2.2. Oligonucleotides and DNA Sequencing
Oligonucleotides were synthesized by Sigma Genosys (Woodlands, TX) or Integrated DNA Technologies (San Diego, CA). The oligonucleotide primers employed in this study, C31G, C40G, C40S, C40T, and C40Y are listed in Table 1. DNA sequencing was performed with the dye terminator cycle sequencing (DTCS) Quick Start Kit (Beckman Coulter, Fullerton, CA) using a CEQ 8000 Genetic Analysis System (Beckman Coulter) to confirm all the mutations.
2.3. Site-Directed Mutagenesis
A pGAP vector containing the phyA gene [22] was utilized to produce the mutant phytases. Three oligonucleotide primers were employed to generate the necessary mutants, C31G, C40G, C40S (Table 1) Depending on the oligonucleotide used mutations were created using thermocycler cycling parameters of: 1 cycle 95˚C for 1 min and 30 cycles of 95˚C (1 min), 55˚C (1 min), 65˚C (10 min); or 1 cycle 95˚C for 30 s and 20 cycles of 95˚C (40 s), 60˚C (40 s) and 68˚C (5 min) and the QuickChange Multi Site-Directed Mutagenesis kit (Strategene, Cedar, TX).
2.4. Protein Expression
P. pastoris X33 was used to express the proteins. The pGAP vector containing the phyA gene was linearized by the restriction enzymes AvrII or BspHI (New England Biolabs, Beverly, MA) and transformed in Pichia cells by electroporation using an ECM 630 Electro Cell Manipulator (Gentronics, Inc., BTX Instrument Division, San Diego, CA). The transformed cells were immediately incubated for 2 hours without shaking in 1 ml of YPD-Plus media (Zymo Research, Orange, CA) and then plated on YPD plates plus zeocin and incubated at 30˚C for 3 days. Single colony transformants were then transferred into liquid YPD media at 30˚C for 2 days and then assayed for phytase activity.
2.5. Purification and Characterization of Expressed Phytase
A 500 mL crude culture filtrate of each mutant strain was used to purify the genetically altered phytase. The steps used to purify the protein include ammonium sulfate precipitation followed by three chromatographic steps. By adding 2.0 M HCl while mixing gently, the pH of the culture filtrate was lowered from 6.0 to 4.6. The ammonium sulfate precipitation of phytase was achieved first by saturating the crude culture filtrate to 45% and then removing the precipitate by centrifugation, which does not contain phytase activity. Additional ammonium sulfate was added to the supernatant to raise the saturation level to 90%. The solution was stirred for about 30 min at 4˚C followed by centrifugation to isolate the precipitated phytase, which was subsequently dissolved in 15 mL of glycine buffer (25 mM, pH 3.5). The traces of ammonium sulfate were removed from the re-suspended phytase solution by rapid dialysis over a period of 4 hours with three changes of glycine buffer.

Table 1. Disulfide bridge mutation oligonucleotide primers. Codons underlined and [ ] denote nucleotide change to produce the mutant.
A 10 ml UNOsphereTM S column (2.5 × 2 cm) was equilibrated in the glycine buffer and the dialyzed protein sample was applied to it at a flow rate of 2.0 mL per min. A linear sodium chloride gradient (0.2 to 1.0 M) was developed in 15 min using the same buffer. The enzyme was eluted as a single component; the active fractions were pooled and dialyzed against 25 mM imidazole buffer (pH 7.0). For the second chromatographic step, a 4 mL UNOsphereTM DEAE column (1.5 × 2.3 cm) was equilibrated in the imidazole buffer. The dialyzed protein from the previous step was passed through the column at a flow rate of 2.5 mL per min. A salt gradient (0 to 0.3 M sodium chloride) was developed in 60 mL. The bound phytase activity was eluted as a single component. The active fractions were pooled and dialyzed against the glycine buffer.
For the final step, a 2 mL UNOsphereTM S column (1.5 × 1.2 cm) was equilibrated in glycine buffer. The dialyzed protein from the previous step was applied to this column at a flow rate of 1.5 mL per min. A linear sodium chloride gradient 0.2 to 0.75 M was developed in 15 min in the same buffer. The purified enzyme was eluted as a single peak.
2.6. Phytase Activity and pH Profile Determination
Phytase assays were carried out in 1.0 mL volume at optimum temperatures in 25 mM glycine or 25 mM sodium acetate buffers [23]. The liberated inorganic orthophosphate was quantitated spectro photometrically [24] using a freshly prepared acetone-molybdate-acid (AMA) reagent consisting of anhydrous acetone (2 parts), 10 mM ammonium molybdate (1 part), and 2.5 M sulfuric acid (1 part). Adding 2.0 mL of AMA per assay tube terminated the enzyme assay. After 30 s 100 μL of citric acid (1.0 M) was added to each tube to fix the color. Absorbance was read at 355 nm after blanking the spectrophotometer with the appropriate control. Phytase activity was expressed as ηkat/mL (nmoles ortho-phosphate released per sec). The pH profiles were established based on the previously employed method [25].
2.7. Determination of Thermostability
The purified native PhyA and DB mutants with a protein content of 0.4 mg in 100 µL were individually sealed in a narrow glass tube with ParafilmTM and heated at 70˚C for 5 min in a water bath. Samples were then equilibrated at room temperature for 10 min before placing on ice. The control experiments were done in an identical way except they were maintained at room temperature for 5 min. Phytase activity was measured at optimum pH of each phytases at 37˚C, 42˚C, 52˚C and 58˚C. Establishing the activity of the control as 100%, the activity of the samples after heating at 70˚C was expressed as a percentage.
3. RESULTS
The phytases of the DB mutants were isolated, purified and characterized. The pH profile of each single or double cysteine mutations for the first disulfide bridge are shown in Figure 2. Except for C40G, all mutants lost significant activity in the acidic pH range. The two mutants with the lowest activity in the acidic range were C31G/C40S and C31G. An intermediate profile is seen in the double mutant C31G/C40G. Purification necessary for characterization of the single mutants C40T and C40Y was not possible due to diminished enzyme activity.
All the DB mutants that were characterized exhibited a lower optimum temperature than the native PhyA (Figure 3). In the 36˚C - 40˚C temperature range, all DB mutants except for C31G featured higher activity than the WT phytase. The optimum temperature recorded for the DB mutants were; C31G 53˚C, C40G 52˚C, C31G/ C40G 42˚C, and C31G/C40S 42˚C.

Figure 2. The pH profile of purified PhyA WT phytase and disulfide bridge mutants expressed in Pichia pastoris. Assays were performed at the optimum temperature for each phytase.
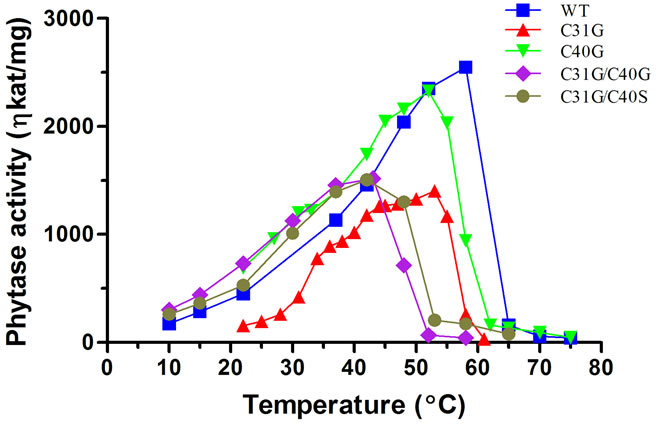
Figure 3. The temperature profile of purified PhyA WT phytase and disulfide bridge mutants expressed in Pichia pastoris. The pH was adjusted for each enzyme to maximize activity.
To mimic heating conditions during feed processing, all DB mutants and the native PhyA were heated to 70˚C for five minutes and then tested for phytase activity. When the results were compared, all the DB mutants retained a higher percentage of activity at 37˚C than the WT. More specifically the WT PhyA retained 21.3%, while C31G, C40G, 31G/C40G and C31G/C40S retained 24.8%, 39.3%, 49.6% and 84.2% respectively (Figure 4). Interestingly, both the double DB mutants retained more activity than the two single DB mutants at the physiologically important temperature of 37˚C.
4. DISCUSSION
Previously, DB mutants were produced and biochemically characterized in a bacterial HAPhy [20,25]. In the former study one cysteine, C200, involved in a DB in E. coli phytase along with specific neighboring amino acids (AA) were replaced to produce an additional N-glycosylation site. Mutant C200N/D207N/S211N, which eliminated the second E. coli DB displayed an altered pH profile with 54% higher specific activity for phytase, and retained higher (P < 0.05) residual phytase activity after heat treatments at 80˚C and 90˚C [23]. In a separate E. coli phytase study where DBs were probed, neither the optimum temperature nor pH profile of the generated DB mutants was determined [20]. The emphasis of the study however was to elucidate the role of the protein-disulfide isomerase, DsbC, in the formation of nonconsecutive DB and for that reason both the single and double cysteine DB mutants were tested. The substrate for the activity assay was also p-nitrophenyl phosphate at 37˚C and not sodium phytate. It is still interesting to note that all the AppA single cysteine DB mutants in the study exhibited reduced enzymatic activity; most displayed less than 10% activity of the WT AppA. However, all the double DB mutants displayed higher activity than the corresponding single mutant. A possible explanation for this

Figure 4. The percentage of activity after heat treatment of PhyA WT phytase and disulfide bridge mutants at 70˚C. The percentage activity remaining at 37˚C, 42˚C, 52˚C, and 58˚C before and after heating. The pH was adjusted to maximize activity of each phytase.
observation was that unpaired cysteines in the single mutant could form incorrect DBs resulting in a misfolded protein. This would render them more susceptible to protease digestion. However, this would not be possible when both of the paired cysteines are mutated. The one DB single mutation in the DsbC study with the highest activity, C200S, corresponds to the C200N DB containing mutant in the previous study [24] also displayed higher than WT activity. The double consecutive DB mutant C200S/C210S even had greater activity (124%) than WT E. coli phytase [20]. In the current study it is important to note that the double DB mutants had higher thermostability than the single DB mutants or WT.
In the removal of a DB, one or both of the cysteines necessary for the formation of a DB are mutated by sitedirected mutagenesis to another amino acid. Consequently, the removal of the bridge allows a new molecular configuration for the enzyme and also the novel interactions between the side chain of the newly introduced amino acid and the side group of the neighboring amino acid(s). Properties of the DB mutants are therefore the results of the combination of both the loss of the structural stability afforded by the DB and the interaction of the AA(s) replacing the cysteine(s) and the neighboring AAs. The results of this study suggest that the replacement of both cysteines pairs in the first DB of A. niger PhyA yielded mutants with more desirable attributes for animal feed applications than mutants from replacement of a single cysteine. Just as all the single cysteine replacements had lower enzyme activity than the corresponding double mutant in E. coli phytase [20], a single free cysteine in PhyA may contribute to the enzyme’s misfolding or other structural instability, which is manifested in a lower catalytic rate. In this study, the higher heat tolerance of C31G/C40S compared with C31G/ C40G also indicates the importance of selecting an amino acid that may form unique hydrogen bonds with the side groups of neighboring amino acids. The importance of which amino acid is selected to replace the cysteine is underscored by the differences in the activity levels of C40S (Figure 4) and the lack of enzyme activity in C40T and C40Y. The specific cause for the paucity of enzyme activity for C40T and C40Y is unknown and must await further study. It is interesting to note the propinquity of the first DB to the active site (Figure 1) which did not yield all mutants lacking phytase activity, but in some instances a phytase with higher activity at 37˚C and superior heat tolerance than WT PhyA. This is quite fortuitous for A. niger PhyA phytase where the first DB is located not too far from the active site (Figure 1). The E. coli AppA phytase lacks a corresponding DB [20] and for this reason a different strategy is needed to lower the optimum temperature of activity to match the body temperature of most simple stomached animals.
5. CONCLUSION
The results presented in this study clearly demonstrated that by eliminating a crucial disulfide bridge in PhyA phytase, the mutant biocatalysts have acquired new temperature optima after subjected to 70˚C for a brief period. This enhanced thermostabilty in disulfide bridge mutants could be commercially exploited to design a superior phytase, which is already being used in the poultry and swine feed industry to lower phosphate in the waste product.
![]()
![]()
REFERENCES
- Mullaney, E., Daly, C.B. and Ullah, A.H.J. (2000) Advances in phytase research. Advances in Applied Microbiology, 47, 157-199. doi:10.1016/S0065-2164(00)47004-8
- Wodzinski, R.J. and Ullah, A.H.J. (1996) Phytase. Advances in Applied Microbiology, 42, 263-302. doi:10.1016/S0065-2164(08)70375-7
- Berka, R.M., Rey, M.W., Brown, K.M., Byun, T. and Klotz, A.V. (1998) Molecular characterization and expression of a phytase gene from the thermophilic fungus Thermomyces lanuginosus. Applied and Environmental Microbiology, 64, 4423-4427.
- Mitchell, D.B., Vogel, K., Weimann, B., Pasamontes, L. and van Loon, A.P.G.M. (1997) Phytase subfamily of histidine acid phosphatases: Isolation of genes for two novel phytases from fungi Aspergillus terreus and Myceliophthora thermophila. Microbiology, 143, 245-252. doi:10.1099/00221287-143-1-245
- Pasamontes, L., Haiker, M., Wyss, M., Tessier, M. and van Loon, A.P.G.M. (1997) Gene cloning, purification and characterization of a heat-stable phytase from the fungus Aspergillus fumigates. Applied and Environmental microbiology, 63, 1996-1700.
- Segueilha, L., Lambrechts, C., Boze, H., Moulin, G. and Galzy, P. (1992) Purification and properties of the phytase from Schwanniomyces castelli. Journal of fermentation and Bioengineering, 74, 7-11. doi:10.1016/0922-338X(92)90259-W
- Yamada, K., Minoda, Y. and Yamamoto, S. (1968) Phytase from Aspergillus terreus Part I: Production, purification and some general properties of the enzyme. Agricultural and Biological Chemistry, 32, 1275-1282. doi:10.1271/bbb1961.32.1275
- Lehman, M. (1988) Consensus phytase. International patent Application, No EP0897985A2.
- Zhang, L., Wang, Y., Zhang, C., Wang, Y., Zhu, D., Wang, C. and Nagata, S. (2006) Supplementation effect of Ecotoine on thermostability of phytase. Journal of Biosciences and Bioengineering, 102, 560-563. doi:10.1263/jbb.102.560
- Greiner, R. and Konietzny, U. (1996) Conctruction of a bioreactor to produce special breakdown products of phytate. Journal of Biotechnology, 48, 153-159. doi:10.1016/0168-1656(96)01505-2
- Han, Y. and Lei, X.G. (1999) Role of glycosylation in the functional expression of an Aspergillus niger phytase gene (phyA) in Saccharomyces cerevisiae. Applied Microbiology and Biotechnology, 65, 1915-1918.
- Zhang, W., Mullaney, E.J. and Lei, X.G. (2007) Adopting selected hydrogen bonding and ionic interactions from Aspergillus fumigates phytase structure improves the thermostability of Aspergillus niger PhyA phytase. Applied and Environmental Microbiology, 73, 3069-3076. doi:10.1128/AEM.02970-06
- Kim, M.S., Weaver, J.D. and Lei, X.G. (2008) Assembly of mutations for improving thermostability of Escherichia coliAPPA2 phytase. Applied Microbiology and Biotechnology, 79, 751-758. doi:10.1007/s00253-008-1478-2
- Jermutus, L., Tessier, M., Pasamontes, L., van Loon, A.P.G.M. and Lehman, M. (2001) Structure-based chimeric enzymes as an alternative to directed enzyme evolution: Phytase a test case. Journal of Biotechnology, 85, 15-24. doi:10.1016/S0168-1656(00)00373-4
- Wang, X.Y., Meang, F.G. and Zhou, H.M. (2004) The role of disulfide bonds in the conformational stability and catalytic activity of phytase. Biochemistry and cell biology, 82, 329-334. doi:10.1139/o03-082
- Mullaney, E., Locovare, H., Sethumadhavan, K., Boone, S., Lei, X.G. and Ullah, A.H.J. (2010) Site-directed mutagenesis of disulfide bridges in Aspergillus niger NRRL phytase (PhyA), their expression in Pichia pastoris and catalytic characterization. Applied Microbiology and Biotechnology, 87, 1367-1372. doi:10.1007/s00253-010-2542-2
- Cheng, C., Wong, K.B. and Lim, L.B. (2007) The effect of disulfide bond on the conformational stability and catalytic activity of beta-propeller phytase. Protein and Peptide Letters, 14, 175-183. doi:10.2174/092986607779816069
- Olczak, M., Morawiecka, B. and Watorek, W. (2003) Plant purple acid phosphatase: Genes, structural and biological function. Acta Biochimica Poland, 50, 1245-1256.
- Lei, X., Mullaney, E.J. and Ullah, A.H.J. (2007) Using mutations to improve Aspergillus phytases. US Patent No. 7309505 B2.
- Berkman, M., Boyd, D. and Beckwith, J. (2005) The nonconsecutive disulfide bond of Escherichia coli phytase (AppA) renders it dependent on the protein-disulfide isomerase, DsbC. Journal of Biological Chemistry, 280, 11387-11394.doi:10.1074/jbc.M411774200
- Mullaney, E. and Ullah, A.H.J. (2005) Conservation of cysteine residues in fungal Histidine acid phytases. Biochemical and biophysical Research Communications, 328, 404-408.doi:10.1016/j.bbrc.2004.12.181
- Kim, T., Mullaney, E.J, Porres, J.M. Roneker, K.R., Crowe, S., Rice, S., Ko, T. Ullah, A.H.J., Daly, C.B., Welch, R. and Lei, X.G. (2006) Shifting the pH profile of Aspergillus niger PhyA phytase to match the stomach pH enhances its effectiveness as an animal feed additive. Applied and Environmental Microbiology, 72, 4397-4403. doi:10.1128/AEM.02612-05
- Ullah, A.H.J. and Gibson, D.M. (1987) Extracellular phytase (E.C. 3.1.3.8) from Aspergillus ficuum NRRL 3135: Purification and characterization. Preparative Biochemistr, 17, 63-91. doi:10.1080/00327488708062477
- Heinonen, J.K. and Lahti, R.J. (1981) A new convenient colorimetric determination of inorganic orthophosphate and its application to the assay of inorganic pyrophosphatase. Analytical Biochemistry, 113, 313-317. doi:10.1016/0003-2697(81)90082-8
- Rodriguez, E., Wood, Z.A., Karplus, A. and Lei, X.G. (2000) Site-directed mutagenesis improves catalytic efficiency and thermostability of Escherichia coli pH 2.5 acid phosphatase/phytase expressed in Pichia pastoris. Archive for Biochemistry and Biophysics, 382, 105-112. doi:10.1006/abbi.2000.2021.
NOTES
*Corresponding author.

