 Vol.3, No.8, 524-528 (2011) doi:10.4236/health.2011.38087 Copyright © 2011 SciRes. Openly accessible at http://www.scirp.org/journal/HEALTH/ Health Improved detection of the MUC1 cancer antigen CA 15-3 by ALYGNSA fluorimmunoassay Sinang Chourb, Brian Christopher Mackness, Leslie Ruth Farris, Melisenda Jean McDonald* Chemistry Department, University of Massachusetts Lowell, Lowell, USA; *Corresponding Author: Melisenda_Mcdonald@uml.edu Received 18 May 2011; revised 21 July 2011; accepted 28 July 2011. ABSTRACT Breast cancer is the second leading cause of cancer-related deaths in women worldwide; a prime cancer biomarker to aid in the diagnosis, directed treatment, clinical management, and reoccurrence of this cancer is a MUC1 peptide fragment: cancer antigen 15-3 (CA 15-3). Herein, an immuno-fluorescence assay for CA 15-3 was developed; this ALYGNSA system consists of a protein biolinker (Protein G’) adsorbed onto Poly (methyl methacrylate) (PMMA). The unique interaction of Protein G’ with PMMA, a thermo- plastic polymer has been demonstrated to im- prove human IgG capture antibody alignment/ orientation and result in greater assay sensi- tivity. Indeed a previous report (HEALTH 1 325 - 329, 2009) on the shed extracellular domain of HER-2/neu revealed a 10-fold increase in sensi- tivity of the ALYGSNA assay over a control ELISA assay. Results from this ALYGNSA assay study revealed that a 16-fold increase in dete- ction (≤0.94 U/mL) of CA 15-3 was found in comparison to a commercial control ELISA kit (≤15 U/mL). In conclusion, this enhanced sen- sitivity of the ALYGNSA assay for CA 15-3, may provide insights into the role/function of this biomarker in normal, as well as, breast cancer and other epithelial cancers. Keywords: CA 15-3; Cancer Antigen 15-3; Epithelial Tumor Antigen; MUC1; Breast Cancer Marker; ELISA; Fluorescent Immunoassay; ALYGNSA 1. INTRODUCTION Breast cancer (BC) is the second leading cause of can- cer-related deaths in women worldwide after lung cancer and is the most frequently diagnosed form of cancer among American women [1]. The most frequently used and best known biomarker is Her2 [2]; this oncoprotein is cell-membrane bound but its extracellular domain is shed into circulation making it a potent biomarker used to monitor the response to treatment, and to detect re- currences in patients with diagnosed breast carcinoma [3,4]. However, this biomarker is only expressed in 20% of all breast tumors [5]; therefore additional BC markers are rigorously being explored [6]. Mucins are promising candidates [7] in particular serum CA 15-3, a MUC1 Tumor Marker; this cancer antigen is overexpressed in >90% of breast carcinomas and metastases [8,9]. The mucin protein product encoded by the MUC1 gene contains approximately 50% carbohydrate by weight with a relative molecular mass of 400 kDa [10]. This cell surface mucin transmembrane glycoprotein, is expressed at the apical surface of most epithelia (e.g. mammary gland, female reproductive tract, stomach, etc.) in nor- mal tissue [11]. It is comprised of three structural do- mains: a large and heavily O-glycosylated extracellular segment (exo-domain), a hydrophobic type 1 transmem- brane region, and a short cytoplasmic tail domain in- volved in several signaling processes [12]. However, in cancerous tissue, this MUC-1 biomarker expression can be detected on the entire cell membrane due to transfor- mation and loss of polarity [13,14]. After transport to the cell membrane, it undergoes proteolytic cleavage in which the soluble form of the large ectodomain is re- leased into circulation [15]. The tumor marker antigen CA15-3, which corresponds to an immuno-dominant epitope in the extracellular portion of the membrane bound mucin MUC1, is shed into the bloodstream. An increase in the serum CA 15-3 shed ectodomain is asso- ciated with progression of carcinoma in patients diag- nosed with breast cancer [16]. Levels of CA 15-3 meas- ured greater than 35 U/mL are indicative of the potential progression and recurrence of breast cancer [17-19]. In this present study, an assay was developed for the detection of the breast cancer biomarker CA 15-3 utiliz- ing the ALYGNSA system consisting of a protein bio- linker (Protein G’) adsorbed onto Poly (methyl metha- crylate) (PMMA). The unique interaction of Protein G’ with PMMA, a thermoplastic polymer, has been demon- strated to improve human IgG capture antibody align-  S. Chourb et al. / Health 3 (2011) 524-528 Copyright © 2011 SciRes. Openly accessible at http://www.scirp.org/journal/HEALTH/ 525 ment/orientation [20] and deliver greater sensitivity in the detection of several cancer biomarkers [21-24], in- cluding an additional breast cancer biomarker HER-2/ neu [21] and an additional mucin MUC16 (CA 125) [22]. Herein, this same ALYGNSA assay system will be em- ployed to measure CA 15-3 and compare the findings to a commercial control ELISA kit. 2. MATERIALS AND METHODS 2.1. CA 15-3 ELISA Assay This CA 15-3 ELISA assay utilizes the “sandwich” principle, where a capture antibody is directly adsorbed onto a substrate. The detector antibody is labeled with an enzyme, which upon addition of the substrate, produces a colored product quantifiable by absorbance analysis. The ELISA kit for detection of CA 15-3 purchased from BioQuant contained: microwells pre-coated with murine monoclonal anti-CA15-3 antibody, sample diluent, en- zyme conjugate diluent, enzyme conjugate concentrate, tetramethylbenzidine (TMB) solution and stop solution. A range of CA 15-3 reference standards (15 - 240 U/mL) provided were used directly (undiluted). This assay was performed, as closely as possible, to manufacturer’s in- structions. First, 200 μL of CA15-3 standards was dis- pensed into the appropriate microwells, gently mixed for 10 seconds, and incubated at 37˚C for 1 hour. The wells were rinsed 5 times with dH2O, and 200 μL of enzyme conjugate reagent was dispensed into each well followed by mixing (10 seconds) and incubation (37˚C for 1 hour). The wells were re-washed and 100 μL of TMB reagent was dispensed into each well and gently mixed for 10 seconds. The wells were incubated at room temperature in the dark for 20 minutes, and then finally, 100 μL of stop solution was added to each well to terminate the reaction. The plate was gently mixed for 30 seconds then read at 450 nm with a Bio-Rad microtiter plate reader. 2.2. CA 15-3 ALYGNSA Assay A “sandwich” fluoroimmunoassay which exploits uni- que polymer–protein noncovalent interactions between a recombinant Protein G’ biolinker and a poly (methyl methacrylate) (PMMA) surface has been termed the ALYGNSA antibody-orientation system. Protein and an- tibody reagents used in ALYGNSA assay were: recom- binant Protein G’ (Pierce), Cancer Antigen 15-3 (CA 15-3) monoclonal antibodies (capture and detector) (Fit- zgerald, clone# M002204, M002208) and CA 15-3 anti- gen also from Fitzgerald. The Pierce 660nm Protein As- say (Product #s: 22660, 22662) microplate format was utilized to verify protein concentrations of the CA 15-3 antibodies. The Thermo Scientific Micro BCA Protein Assay Kit (Product #: 23235) was used to verify recom- binant Protein G’ and CA 15-3 protein concentrations. Poly (methyl methacrylate) (PMMA) (Sigma) was used to coat polypropylene plates (Corning). The ALYGNSA assay protocol followed previously reported methods [25]. Briefly, the PMMA plates were coated with recom- binant Protein G’ (1 g/mL at 50 L/well) and incubated overnight at 4˚C. After washing once with TBST, the plates were coated with the capture CA 15-3 antibody at 5 g/mL at 50 L/well and incubated overnight at 4˚C. After washing, the plates were blocked with 5% non-fat dry milk (NFDM) for 1 hour at room temperature and washed. The CA 15-3 antigen was diluted to 60 U/mL in PBS and 100 L was applied to the top rows of each plate. A 1:1 serial dilution in PBS was performed and carried out to 0.94 U/mL. The plates were then incu- bated for 2 hours at room temperature, and then washed. The fluorescently labeled CA 15-3 detector antibody prepared by the DyLight 488 Microscale Antibody La- beling Kit as recently described [25] was diluted to 5 g/mL, and applied at 50 L/well. Following incubation for 1 hour at room temperature and a repeat washing, the plates were read for fluorescence at 485/523 nm on a BioTek Microplate Reader. 3. RESULTS and DISCUSSION 3.1. ELISA and ALYGNSA Immunoassays Enzyme-Linked Immunosorbant Assay (ELISA) is a common immunochemical colorimetric method used to detect cancer biomarkers in biological fluids, such as serum [16,26]. The CA 15-3 ELISA assay utilizes the “sandwich” principle, where a capture antibody is di- rectly adsorbed onto a substrate. The detector antibody is labeled with an enzyme, which upon addition of the sub- strate, produces a colored product quantifiable by absor- bance analysis. A comparison of 15 commercial immu- noassays for detection of CA15-3 (MUC1) in serum has been reported with manufacturer cut-off ranging from 23 – 39 (U/mL) [27]. A BioQuant ELISA kit was employed in the present study. After multiple runs of this com- mercial CA 15-3 ELISA kit assay, the Limit Of Detec- tion (LOD) for CA 15-3 protein, in our hands, was de- termined to be ≤15 U/mL (Figure 1; Ta bl e 1 ) ; it is in- teresting to note that this is below the cutoff of 17 U/mL (reported range 5 - 47 U/mL) for the non-cancerous state [28]. For more efficacy of the use of proteins and antibod- ies, an alternative form of this assay, a fluorescent ALY- GNSA assay was employed [20]. The ALYGNSA assay format is essentially the same as the colorimetric ELISA; to further enhance sensitivity of the assay, a properly 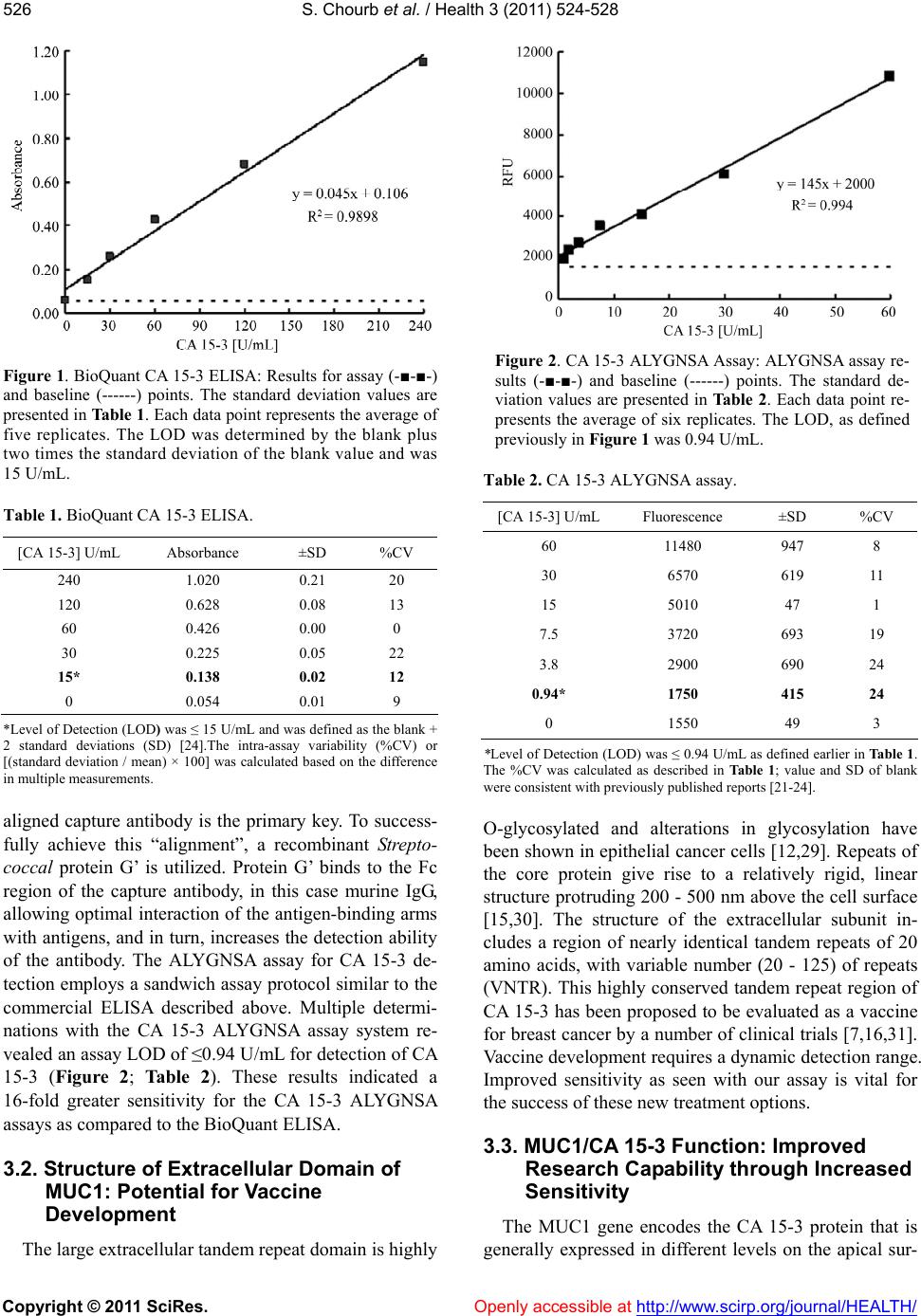 S. Chourb et al. / Health 3 (2011) 524-528 Copyright © 2011 SciRes. Openly accessible at http://www.scirp.org/journal/HEALTH/ 526 Figure 1. BioQuant CA 15-3 ELISA: Results for assay (-■-■-) and baseline (------) points. The standard deviation values are presented in Table 1. Each data point represents the average of five replicates. The LOD was determined by the blank plus two times the standard deviation of the blank value and was 15 U/mL. Table 1. BioQuant CA 15-3 ELISA. [CA 15-3] U/mL Absorbance ±SD %CV 240 1.020 0.21 20 120 0.628 0.08 13 60 0.426 0.00 0 30 0.225 0.05 22 15* 0.138 0.02 12 0 0.054 0.01 9 *Level of Detection (LOD) was ≤ 15 U/mL and was defined as the blank + 2 standard deviations (SD) [24].The intra-assay variability (%CV) or [(standard deviation / mean) × 100] was calculated based on the difference in multiple measurements. aligned capture antibody is the primary key. To success- fully achieve this “alignment”, a recombinant Strepto- coccal protein G’ is utilized. Protein G’ binds to the Fc region of the capture antibody, in this case murine IgG, allowing optimal interaction of the antigen-binding arms with antigens, and in turn, increases the detection ability of the antibody. The ALYGNSA assay for CA 15-3 de- tection employs a sandwich assay protocol similar to the commercial ELISA described above. Multiple determi- nations with the CA 15-3 ALYGNSA assay system re- vealed an assay LOD of ≤0.94 U/mL for detection of CA 15-3 (Figure 2; Table 2). These results indicated a 16-fold greater sensitivity for the CA 15-3 ALYGNSA assays as compared to the BioQuant ELISA. 3.2. Structure of Extracellular Domain of MUC1: Potential for Vaccine Development The large extracellular tandem repeat domain is highly Figure 2. CA 15-3 ALYGNSA Assay: ALYGNSA assay re- sults (-■-■-) and baseline (------) points. The standard de- viation values are presented in Ta ble 2 . Each data point re- presents the average of six replicates. The LOD, as defined previously in Figure 1 was 0.94 U/mL. Table 2. CA 15-3 ALYGNSA assay. [CA 15-3] U/mL Fluorescence ±SD %CV 60 11480 947 8 30 6570 619 11 15 5010 47 1 7.5 3720 693 19 3.8 2900 690 24 0.94* 1750 415 24 0 1550 49 3 *Level of Detection (LOD) was ≤ 0.94 U/mL as defined earlier in Table 1. The %CV was calculated as described in Ta bl e 1 ; value and SD of blank were consistent with previously published reports [21-24]. O-glycosylated and alterations in glycosylation have been shown in epithelial cancer cells [12,29]. Repeats of the core protein give rise to a relatively rigid, linear structure protruding 200 - 500 nm above the cell surface [15,30]. The structure of the extracellular subunit in- cludes a region of nearly identical tandem repeats of 20 amino acids, with variable number (20 - 125) of repeats (VNTR). This highly conserved tandem repeat region of CA 15-3 has been proposed to be evaluated as a vaccine for breast cancer by a number of clinical trials [7,16,31]. Vaccine development requires a dynamic detection range. Improved sensitivity as seen with our assay is vital for the success of these new treatment options. 3.3. MUC1/CA 15-3 Function: Improved Research Capability through Increased Sensitivity The MUC1 gene encodes the CA 15-3 protein that is generally expressed in different levels on the apical sur-  S. Chourb et al. / Health 3 (2011) 524-528 Copyright © 2011 SciRes. Openly accessible at http://www.scirp.org/journal/HEALTH/ 527 face of many normal and malignant epithelial cells (ex- pression over entire cell surface), including in the mam- mary gland, female reproductive tract, lung, kidney, sto- mach, gall bladder and pancreas, as well as, some non- epithelial cell types [7-9,31,32]. The precise function of CA 15-3 is unclear, although in general, the physiologi- cal function of mucins is in lubrication and hydration of cell surfaces, protection of proteins and cells from prote- olysis and in the protection of tissues from microbial attack. However, other evidence suggests CA 15-3 ap- pears to play a role in cell-adhesion, where it modulates cell-cell and cell-extracellular matrix (ECM) interactions. In addition to its function as a protective barrier with adhesion-modulating properties, the CA 15-3 cytoplas- mic tail has the potential role in cell signal transduction and has been reported to contribute to metastases [32,33]. Increased assay sensitivity offers improved expression analysis for researchers working to identify the function of mucin protein MUC1. Further understanding of this protein may lead to the development of personalized treatment and preventative medicines. 4. CONCLUSIONS In summary, our work utilized a fluorescent ELISA incorporating the newly developed ALYGNSA antibody- orientation system has revealed a 16-fold increase in sensitivity (≤0.94 U/mL) of CA 15-3 when compared to a commercial ELISA kit (≤15 U/mL). The ALYGNSA assay could aid in evaluation and detection of CA 15-3 under normal conditions, be useful for surveillance of patients diagnosed with breast cancer and to monitor the course of therapy in advanced disease. Furthermore, an increase in sensitivity of the CA 15-3 assay for detection of the shed extracellular domain containing the VNTR region, may provide additional insights into the role/ function and clinical assessment of CA 15-3 in breast cancer and other epithelial expressing this protein. Fi- nally, this system has the potential to be incorporated into a cost-effective biosensor device [34]. This noninva- sive serum-based test platform would assist in earlystage cancer diagnosis allowing the clinician to respond in a proactive rather than a reactive manner. 5. ACKNOWLEDGEMENTS This work was supported by NSF award # 0425826. Special thanks to Adrianna Morris and Peter S. Chiev for their support and critical reading of this manuscript. REFERENCES [1] Jemal, A., Siegel, R., Xu, J. and Ward, E. (2010) Cancer Statics. CA Cancer Journal for Clinicians, 60, 277-300. doi:10.3322/caac.20073 [2] Moelans, C.B., de Weger, R.A., Van der Wall, E. and van Diest, P.J. (2011) Current technologies for HER2 testing in breast cancer. Critical Reviews in Oncology/Hemato- logy, Accessed May 5, 2011. doi:10.1074/mcp.R400001-MCP200 [3] Ross, J.S., Fletcher, J.A., Linette, G.P., Stec, J., Clark, E., Ayers, M., et al. (2004) Targeted therapy in breast cancer: The HER-2/neu gene and protein. Molecular & Cellular Proteomics, 3, 379-398. doi:10.1074/mcp.R400001-MCP200 [4] Bramwell, V.H.C., Doig, G.S., Tuck, A.B., Wilson, S.M., Tonkin, K.S., Tomiak, A., et al. (2009) Changes over time of extracellular domain of HER2 (ECD/HER2) se- rum levels have prognostic value in metastatic breast cancer. Breast Cancer Research and Treatment, 114, 503-511. doi:10.1007/s10549-008-0033-2 [5] Carney, W.P., Leitzel, K., Ali, S., Neumann, R. and Lip- ton, A. (2007) HER-2/neu diagnostics in breast cancer. Breast Cancer Research, 9, 207-217. doi:10.1186/bcr1664 [6] Jones, S.E. (2008) Metastatic breast cancer: The treat- ment challenge. Clinical Breast Cancer, 8, 224-233. doi:10.1038/nrc2761 [7] Mukhopadhyay, P., Chakraborty, S., Ponnusamy, M.P., Lakshmanan, I., Jain, M. and Batra, S. K. (2011) Mucins in the pathogenesis of breast cancer: Implications in di- agnosis, prognosis and therapy. Biochimica et Biophysica Acta, 1815, 224-240. [8] Kufe, D.W. (2009) Mucins in cancer: Function, prognosis and therapy. Nature Reviews Cancer, 9, 874-885. doi:10.1016/j.tibs.2009.10.003 [9] Senapati, S., Das, S. and Batra, S.K. (2010) Mu- cin-interacting proteins: From function to therapeutics. Trends in Biochemical Sciences, 35, 236-245. doi:10.1016/j.tibs.2009.10.003 [10] Duffy, M.J. (1999) CA 15-3 and related mucins as cir- culating markers in breast cancer. Annals of Clinical Bi- ochemistry, 36, 579-586. [11] Bafna, S., Kaur, S. and Batra, S.K. (2010) Mem- brane-bound mucins: The mechanistic basis for altera- tions in the growth and survival of cancer cells. Onco- gene, 29, 2893-2904. doi:10.1038/onc.2010.87 [12] Lagow, E., DeSouza, M.M. and Carson, D.D. (1999) Mammalian reproductive tract mucins. Human Repro- duction, 15, 280-292. [13] Agrawal, B., Gendler, S.J. and Longenecker, B.M. (1998) The biological role of mucins in cellular interactions and immune regulation: Prospects for cancer immunotherapy. Molecular Medicine Today, 4, 397-403. doi:10.1016/S1357-4310(98)01322-7 [14] Brayman, M., Thathiah, A. and Carson, D.D. (2004) MUC1: A multifunctional cell surface component of re- productive tissue epithelia. Reproductive Biology and Endocrinology, 2, 1-9. doi:10.1186/1477-7827-2-4 [15] Parry, S., Silverman, H.S., McDermott, K., Willis, A., Hollingsworth, M.A. and Harris, A. (2001) Identification of MUC1 proteolytic cleavage sites in vivo. Biochemical and Biophysical Research Communications, 283, 715- 720. doi:10.1006/bbrc.2001.4775 [16] Welsh, J.B., Sapinoso, L.M., Kern, S.G., Brown, D.A., Liu, T., Bauskin, A.R., et al. (2003) Large-scale delinea- tion of secreted protein biomarkers overexpressed in  S. Chourb et al. / Health 3 (2011) 524-528 Copyright © 2011 SciRes. http://www.scirp.org/journal/HEALTH/Openly accessible at 528 cancer tissue and serum. Proceedings of the National Academy of Sciences USA, 100, 3410-3415. doi:10.1073/pnas.0530278100 [17] Duffy, M.J., Duggan, C., Keane, R., Hill, A.D.K., McDermott, E., Crown, J. and O’Higgins, N. (2004) High preoperative CA 15-3 concentrations predict ad- verse outcome in node-negative and node-positive breast cancer: Study of 600 patients with histologically con- firmed breast cancer. Clinical Chemistry, 50, 559-563. doi:10.1373/clinchem.2003.025288 [18] Tampellini, M., Berruti, A., Gerbino, A., Buniva, T., Tor- ta, M., Gorzegno, G., et al. (1997) Relationship between CA 15-3 serum levels and disease extent in predicting overall survival of breast cancer patients with newly di- agnosed metastatic disease. British Journal of Cancer, 75, 698-702. doi:10.1038/bjc.1997.124 [19] Kim, H.S., Park, Y.H., Park, M.J., Chang, M.H., Jun, H.J., Kim, K.H., et al. (2009) Clinical significance of a se- rum CA15-3 surge and the usefulness of CA15-3 kinetics in monitoring chemotherapy response in patients with metastatic breast cancer. Breast Cancer Research and Treatment, 118, 89-90. doi:10.1007/s10549-009-0377-2 [20] Clarizia, L.-J.A., Sok, D., Wei, M., Mead, J., Barry, C. and McDonald, M.J. (2009) Antibody orientation en- hanced by selective polymer–protein noncovalent inter- actions. Analytical and Bioanalytical Chemistry, 393, 1531-1538. doi:10.1007/s00216-008-2567-x [21] Chourb, S., Mackness, B.C., Farris, L.R. and McDonald, M.J. (2009) Enhanced immuno-detection of shed ex- tracellular domain of HER-2/neu. Health, 1, 325-329. doi:10.4236/health.2009.14053 [22] Sok, D., Clarizia, L.-J.A., Farris, L.R. and McDonald, M.J. (2009) Novel fluoroimmunoassay for ovarian can- cer biomarker CA-125. Analytical and Bioanalytical Chemistry, 393, 1521-1523. doi:10.1007/s00216-008-2569-8 [23] Mackness, B.C., Chourb, S., Farris, L.R. and McDonald, M.J. (2010) Polymer-protein-enhanced fluoroimmunoas- say for prostate-specific antigen. Analytical and Bioana- lytical Chemistry, 396, 681-686. doi:10.1007/s00216-009-3234-6 [24] Mackness, B.C. and McDonald, M.J. (2010) Serum- -Based ALYGNSA immunoassay for the prostate cancer biomarker, total prostate-specific antigen (tPSA). Ana- lytical and Bioanalytical Chemistry, 397, 3151-3154. doi:10.1007/s00216-010-3827-0 [25] Chourb, S. (2010) Enhanced immuno-detection of breast cancer biomarkers: Shed extracellular domain of Her-2/ neu and CA 15-3. Master’s Thesis, University of Massa- chusetts Lowell, Lowell. [26] Sturgeon, C.M., Duffy, M.J., Stenman, U.H., Lilja, H., Brünner, N., Chan, D.W., et al. (2008) National Academy of Clinical Biochemistry Laboratory Medicine Practice Guidelines for use of tumor markers in prostate, colorec- tal, breast and ovarian cancers. Clinical Chemistry, 54, e11-e79. doi:10.1373/clinchem.2008.105601 [27] Pichon, M.F., Brun, G.L., Hacene, K., Basuyau, J.P., Rie- dinger, J.M., Eche, N., Fulla, Y. and Charlier-Bret, N., (2009) Comparison of fifteen immunoassays for the measurement of serum MUC-1/CA 15-3 in breast cancer patients. Clinical Chemistry and Laboratory Medicine, 47, 985-992. doi:10.1515/CCLM.2009.213 [28] Bon, G.G., Kenemans, P., Dekker, J.J., Hompes, P.G., Ver- straeten, R.A., van Kamp, G.J. and Schoemaker, J. (1999) Fluctuations in CA 125 and CA 15–3 serum concentra- tions during spontaneous ovulatory cycles. Human Re- production, 14, 566-570. doi:10.1093/humrep/14.2.566 [29] Jonckheere, N. and Van Seuningen, I. (2010) The mem- branebound mucins: From cell signaling to transcrip- tional regulation and expression in epithelial cancers. Biochimie, 92, 1-11. doi:10.1016/j.biochi.2009.09.018 [30] Hattrup, C.L. and Gendler, S.J. (2008) Structure and function of the cell surface (tethered) mucins. Annual Review of Physiology, 70, 431-457. doi:10.1146/annurev.physiol.70.113006.100659 [31] Yuan, S., Shi, C., Ling, R., Wang, T., Wang, H. and Han, W. (2010) Immunization with two recombinant bacillus calmette-guerin vaccines that combine the expression of multiple tandem repeats of mucin-1 and colony stimulat- ing-factor suppress breast tumor growth in mice. Journal of Cancer Research and Clinical Oncology, 136, 1359- 1367. doi:10.1007/s00432-010-0787-x [32] Thie, H., Toleikis, L., Li, J., von Wasielewski, R., Bastert, G., Schirrmann, T., et al. (2011) Rise and fall of an Anti- -MUC1 specific antibody. PLoS ONE, 6, Article Number: e15921. doi:10.1371/journal.pone.0015921 [33] Hanash, S.M., Baik, C.S. and Kallioniemi, O. (2011) Emerging molecular biomarkers—Blood-based strategies to detect and monitor cancer. Nature Reviews Clinical Oncology, 8, 142-150. doi:10.1038/nrclinonc.2010.220 [34] Bohunicky, B. and Mousa, S.A. (2011) Biosensors: The new wave in cancer diagnosis. Nanotechnology, Science and Applications , 4, 1-10.
|