Journal of Biomedical Science and Engineering
Vol.6 No.10(2013), Article ID:38272,6 pages DOI:10.4236/jbise.2013.610120
Analysis of malaria diagnosis discrepancies between RDTs and microscopy by nested PCR
![]()
Department of Parasitology-Mycology, Faculty of Medicine, Université des Sciences de la Santé, Libreville, Gabon
Email: *dpmawili@hotmail.com
Copyright © 2013 Denise Patricia Mawili-Mboumba et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received 24 July 2013; revised 25 August 2013; accepted 16 September 2013
Keywords: Plasmodium falciparum; RDTs; Microscopy; PCR; Gabon
ABSTRACT
A total of 415 samples from febrile children exhibiting either concordant (n = 108) or discordant (n = 307) results between microscopy, the gold standard diagnostic test, and two Rapid Diagnostic Tests (RDTs), OptiMAL-IT®(pLDH) and Acon®HRP2, were analysed using polymerase chain reaction (PCR) from May to December 2011 in Gabon. The aim of the study was to analyse these discrepancies using polymerase chain reaction (PCR). Nested PCR targeting the Plasmodium ssrRNA gene was used to distinguish P. falciparum, P. malariae and P. ovale. Plasmodium falciparum was the only malaria species identified. Discrepancies frequently involved samples that were negative by microscopy and positive by Acon®HRP2 (90%) or Optimal-it® (86%). The PCR assay detected submicroscopic infection in almost 23% of the microscopy-negative samples, whereas plasmodial DNA was not found in 77% of the Acon®HRP2 positivemicroscopy negative samples. Although results obtained with Optimal-it® were more frequently concordant with those of PCR genotyping, the low specificity of Optimal-iT® for non-falciparum malaria parasite detection resulted in a high proportion of false negative RDTs (90%) and a high frequency of tests with faint line intensity. The present study highlights the specific attributes of the different methods used to identify malaria parasite below the microscopy level of detection. RDT results that were discordant with either microscopy or PCR as the gold standard could represent a challenge for rapid, accurate fever case management in malaria endemic areas. It is necessary to pursue the development of more precise and more sensitive point-of-care diagnostic tools for malaria.
1. INTRODUCTION
Biological diagnosis of malaria is one of the major weapons for the management of infected patients [1,2]. The use of Rapid Diagnostic Tests (RDTs) for malaria diagnosis is currently recommended by the World Health Organization (WHO) as an alternative to microscopy in remote areas. These tests have been developed to improve care management of the febrile patients and to limit antimalarial drug prescription to parasitologically confirmed cases. Moreover, the decline of malaria prevalence observed in Sub-Saharan Africa following control efforts emphasises the need for an accurate biological diagnosis of true malarial infection [3-5]. Prompt diagnostic confirmation can be achieved through quality microscopy or quality rapid diagnostic tests that detect Plasmodium spp infections. Microscopy, the gold standard for malaria diagnosis, has a detection limit of 10 to 50 parasites/µL. Discrepancies with RDTs are frequently observed, leading to the misinterpretation of results, misdiagnosis and treatment errors [6-11]. Though no diagnostic tool is perfect, molecular techniques are highly sensitive assays compared to microscopy and RDTs that detect infections with different Plasmodium species [12, 13]. Detection of Plasmodium spp DNA based on nested polymerase chain reaction (nested PCR) is a diagnostic alternative that could facilitate the evaluation of RDTs’ performance [13-15]. In Gabon, a study assessing the performance of two RDTs, Acon® HRP2 and OptimalIt® (pLDH), revealed a sensitivity of 98% for parasite densities > 100 Parasites/µl compared to microscopy the gold standard method [16]. Nevertheless, almost 15% of 2125 samples analysed displayed discordant results between RDTs and microscopy. The aim of the present study was to analyse these discrepancies using Plasmodium species nested PCR.
2. MATERIALS AND METHODS
2.1. Study Site
The study was carried out between May and December 2011 in Gabon, where malaria transmission is perennial. Malaria is predominantly caused by Plasmodium falciparum, which has an estimated prevalence of 6% to as high as 40%, while the proportion of infections due to non-falciparum malaria species is below 5% [16,17].
2.2. Samples Collection
Samples were selected from a previous RDT evaluation performed in Gabon comparing Acon®HRP2 and Optimal-It® [16]. They were distributed into two groups: concordant and discordant samples. Concordant samples had identical results with all methods and were classified as positive or negative. Samples with discordant results were distributed into two groups: those with discordant results between thick blood smears and Acon® HRP2 and those with discordant results between thick blood smears and Optimal-It® for the diagnosis of malaria in febrile children with suggestive symptoms. Dried blood spot (DBS) from all the samples with discordant results and 5% of randomly selected samples with concordant results were analysed. DBS were dried overnight and stored in sealed bags with desiccant at room temperature until molecular analysis.
2.3. DNA Extraction
DNA was extracted using methanol method (http://medschool.umaryland.edu/CVD/appendix1.asp, 12/06/2009). Briefly, each DBS was soaked in 200 μL of methanol. After incubation at room temperature for 20 minutes, the methanol was removed and the samples were dried before adding distilled water. The punches were crushed and heated at 95˚C for 20 minutes; then DNA was eluted with sterile water and stored at −20˚C.
2.4. Small Subunit Ribosomal (ssrRNA) Genes Amplification
Five microliters of DNA were included in the Polymerase Chain Reaction (PCR) assays. Nested PCR assays were performed using oligonucleotide primers designed to amplify the Plasmodium small subunit ribosomal RNA (ssrRNA) genes as previously described [18]. Specific primers rFAL 1 and 2, rMAL 1 and 2, rOVA 1 and 2 were used to detect P. falciparum, P. malariae, and P. ovale species, respectively. PCR products were analysed on a 1.5% agarose gel and visualised under UV light after ethidium bromide staining. A molecular size marker (100-base pair [bp] ladder) was employed for the validation of the fragments size. All samples from the two study sites were analysed in the same lab to exclude any difference that could be due to technical issues.
2.5. Line Intensity Analysis
According to the manufacturer’s instructions, RDTs results were considered positive (if both the control and test lines appeared) regardless of the intensity of the test line. Afterward, a visual assessment according to the intensity of the lines generated by each RDT was performed as described by Kyabayinze et al. (2008) [9]. RDTs positive samples were classified as following: those with a line intensity reported as “strong” if the test line was as intense as the control line and as “faint” if the line could be observed in good light.
2.6. Ethical Considerations
This study was a part of a project funded by the MNCP through Global Fund, which aimed at evaluating RDTs performance and parasite molecular analysis. Gabonese Ministry of Health approved the entire study. Children’s parents or guardians were informed about the study protocol, either for the malaria diagnostic method comparison or for the consecutive molecular analysis. The oral consent of the children’s parents or guardians was required prior to enrolment and sample collection for malaria diagnosis and parasite genotyping.
2.7. Statistical Analysis
All data were recorded on a CRF, entered and then cleaned using Epi-info 6 version 3.3.2 (2005 CDC Atlanta) to resolve discordances and delete double entries. Analysis was performed with Statview 5.0. (SAS Institute, Cary, NC, USA). Differences between groups were assessed using Chi-squared or Fisher’s exact tests for proportions. A p-value of less than 0.05 was considered significant.
3. RESULTS
A total of 415 blood samples were analysed by PCR: 307 with discordant results and 108 (60 positives and 48 negatives) with concordant results. P. falciparum was the only species identified either by microscopy or by PCR. Plasmodial DNA was not detected in any concordant negative sample but was detected in 98.3% (n = 59/60) of the positive samples.
3.1. Analysis of Samples with Discordant Results
Microscopy positive-RDT negative samples
PCR confirmed the microscopic P. falciparum infections in 52.6% (n = 10) of Acon®HRP2 and 33.3% (n = 8) of Optimal-It® negative samples (Table 1). Only three RDT and PCR negative samples had parasite densities greater than 100 asexual forms/µL.
Microscopy-negative RDT-positive samples
The majority of samples with discordant results were microscopy negative and RDT positive: 90% (n = 171/ 190) for Acon®HRP2 test and 86% (n = 147/171) for Optimal-It®. PCR detected the presence of P. falciparum in less than one quarter (n = 38/171) of Acon®HRP2 positive samples and more than half of the Optimal-It® P. falciparum ones (n = 24/44) (p < 0.01) (Table 1). None of the Optimal-It® positive samples was infected by non-P. falciparum malaria parasites according to PCR results. Indeed, only eight of the 103 non-P. falciparum Optimal-It® positive samples were successfully genotyped and identified as P. falciparum infection.
Overall, Acon® HRP2 positive tests showed less frequently a faint line intensity compared to Optimal-it® (p < 0.01) (Table 2). The proportion of P. falciparum infections detected by PCR was less than 10% for Optimal-It® (9.5%; n = 8/84) and Acon® HRP2 (7.7% n = 4/52). P. falciparum infections confirmed by PCR were found in only 28.6% of the Acon® HRP2 positive test with a strong line. This proportion was the lowest (15.4%) for non-P. falciparum positive samples with strong line.
4. DISCUSSION
Good RDTs performance is required for their use as diagnostic tool in remote malaria endemic areas where microscopy is lacking or poorly used.
In Gabon, studies assessing the performance of malaria RDTs are needed to guide the health authority decisions in the selection of RDTs with international quality
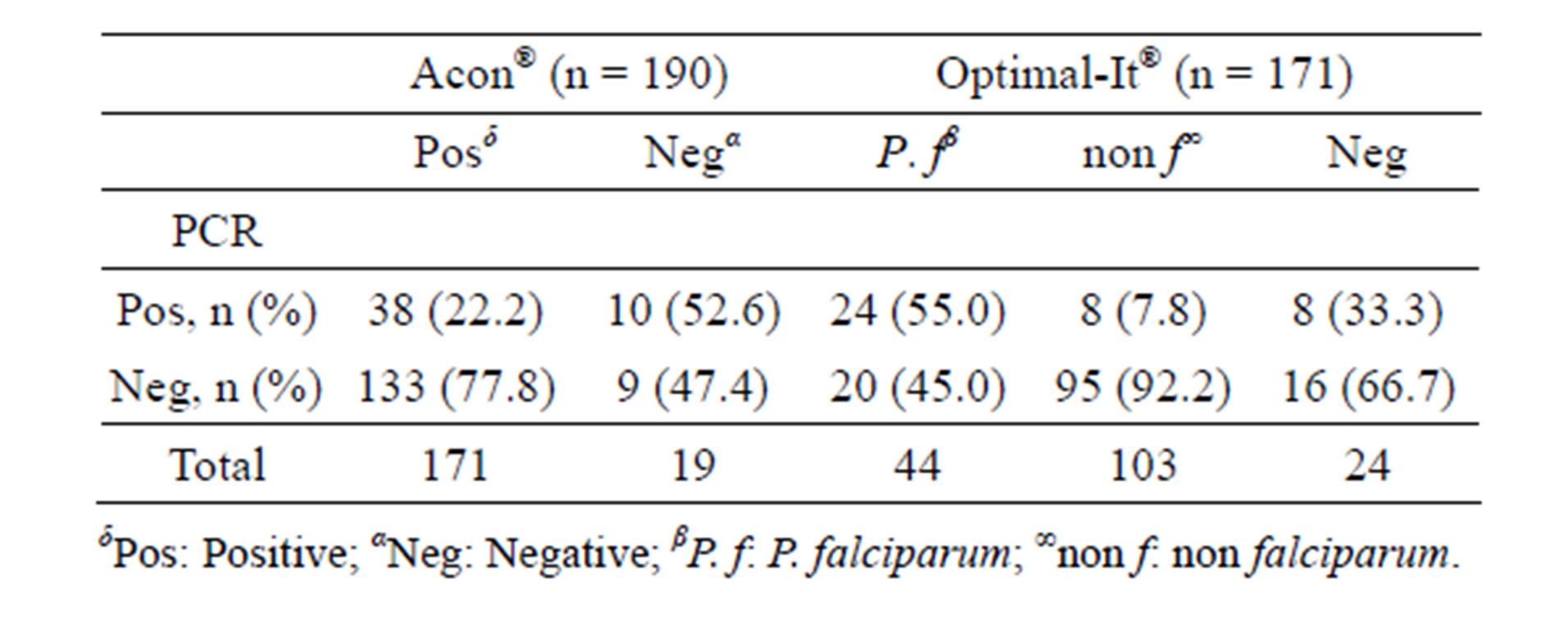
Table 1. Distribution of samples with discordant results according to the amplification rate.
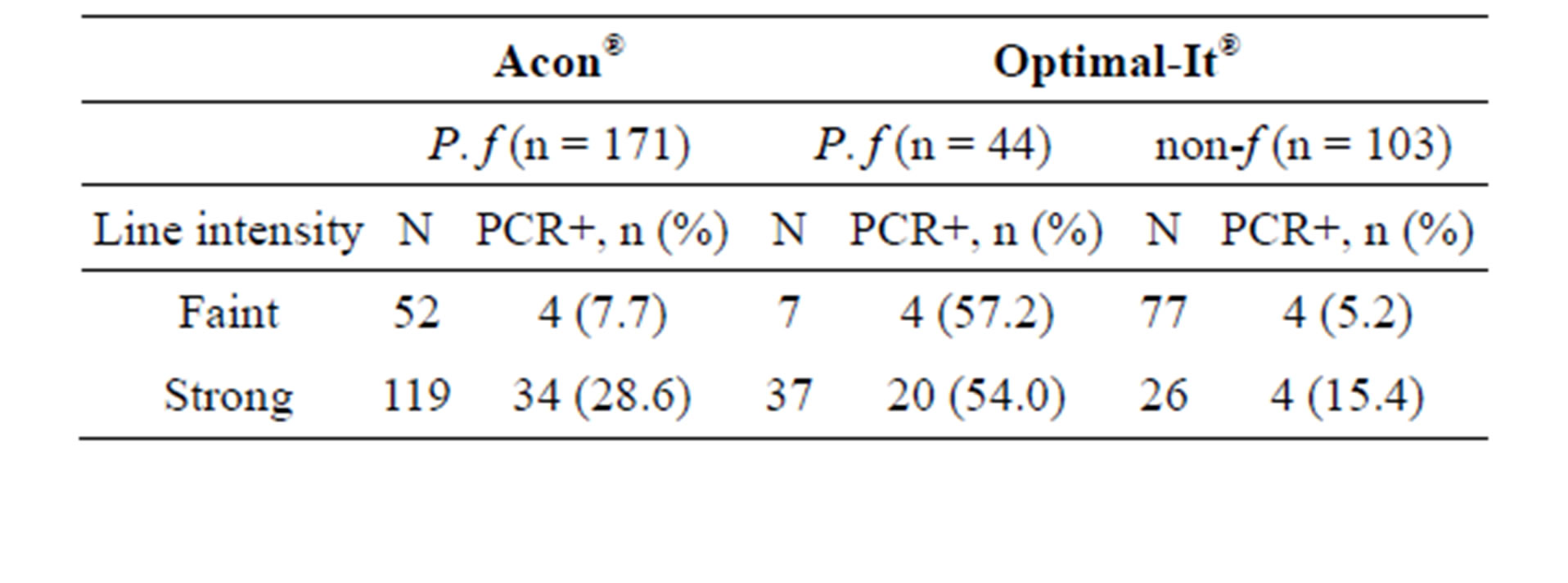
Table 2. Relationship between RDT positive line intensity and PCR results.
standard. Currently, only two studies have evaluated RDTs performance in Gabon [16,19]. Although good sensitivity and good negative predictive values were observed with these tests, frequent discrepancies were found when microscopy was used as the gold standard. These discrepancies were mainly due to a high proportion of false positive results (>20%). Indeed, as found elsewhere, when RDTs were compared to microscopy, false positive was more frequently found with those detecting HRP2 proteins than with those detecting pLDH [9,20,21].
PCR analysis of microscopy-negative TDR-positive samples highlighted the ability of RDTs to detect submicroscopic P. falciparum infections. Hopkins et al found almost 12% of subpatent parasitemia among samples with discordant results [22]. Parasite antigens that are produced by peripheral circulated or sequestered parasites can be detected in case of low peripheral parasitemia [23,24]. Persistent release of P. falciparum HRP2 protein as previously reported by others authors is confirmed in 77% of the Acon® HRP2 positive discordant samples that were negative by PCR and microscopy in the present study [9,18]. This protein is known to have a long half-life and is produced in large quantities by all stages of P. falciparum [25].
In contrast, the pLDH enzyme is short-lived in the blood but is a good indicator of current infection, enabling more frequent detection of submicroscopic P. falciparum infections with Optimal-It®.
However, this test displayed a low specificity for nonP. falciparum malaria parasite detection; at least 90% of non-P. falciparum RDT positive samples were negative by PCR. This lack of detection could be observed because pLDH is produced in small quantities. Therefore, pLDH-based RDT sensitivity is limited, especially in the case of non-P. falciparum species detection. These species usually have low parasitaemias and can be associated with low P. falciparum parasitaemias, below the limit of detection [26]. This latter point could partly explain the fact that only P. falciparum DNA was detected in these positive samples. Moreover, several studies highlighted the low sensitivity of pLDH and aldolase RDTs for the detection of non P. falciparum malaria parasites [27-29].
Both RDTs failed to detect two samples with high parasitaemia. This finding might be related to several factors, such as the variability of the target antigen, which is not uncommon in Africa. Indeed, Koita et al. (2012) showed that a deletion of the repeat region of the HRP2 gene is linked to false negative results [30]. Another factor can be the prozone effect observed in case of high P. falciparum or non P. falciparum malaria parasitaemia [28,31-33]. Thus, additional analysis involving sequencing or multiple real-time PCR is required.
According to RDTs line intensity, few samples with faint bands had a true malaria infection according to PCR genotyping. False positive RDTs results with faint bands were frequently observed when Optimal-it® results identified non-P. falciparum malaria infection. Allen and colleagues, who also found high proportions of false positive results with samples with a faint line on RDTs, suggested that such results should be considered negative [34]. Nevertheless, a recent study evaluating a pan HRP2/aldolase RDT revealed a lower proportion of false positive results linked to faint line intensity, underlining the necessity of understanding the utility of band intensity interpretation [19]. Line intensity interpretation seems to be dependent on several factors such as transmission level [9,34,35]. Despite its high sensitivity, PCR may fail to reveal the presence of Plasmodium in microscopically infected samples, especially in those with low parasitaemia, as observed in the present study. The slide positivity was indisputable taking into account the quality of the method of reading. Such lack of amplification of nucleic acid by nested PCR, real-time PCR or multiplex PCR is reported for samples with PD < 500 p/µl, even higher (PD > 12,000 p/µl) [14,36-38]. Indeed, DNA amplification is more sensitive than antigen detection and microscopy, but the final performance depends on the original sample and how much sample goes into the PCR reaction and the quality of the sample.
These findings confirm the complexity of choosing malaria diagnostic tools in the era of malaria control and elimination when several factors, such as the threshold parasite detection level, parasite target of the different techniques and malaria endemicity, must be considered. It is obvious that the interpretation of non-P. falciparum positive RDT should take into account the antigen detected and the prevalence of Plasmodium species in endemic areas. Nevertheless, all methods used are necessary because of their specificities either for malaria parasite detection and quantification, species identification and disease burden, even in areas where P. falciparum is predominant, as it is frequently observed in Gabon. In any case, accurate, specific and sensitive diagnostic tools that are able to improve both malaria detection and febrile patient care management must be continuously developed.
REFERENCES
- World Health Organization (2006) Guidelines for treatment of malaria. 2006.
- Perkins, M.D. and Bell, D.R. (2008) Working without a blindfold: The critical role of diagnostics in malaria control. Malaria Journal, 7, S5. http://dx.doi.org/10.1186/1475-2875-7-S1-S5
- D’Acremont, V., Lengeler, C., Mshinda, H., Mtasiwa, D., Tanner, M. and Genton, B. (2009) Time to move from presumptive malaria treatment to laboratory-confirmed diagnosis and treatment in African children with fever. PLOS Medicine, 6, e252. http://dx.doi.org/10.1371/journal.pmed.0050252
- Drakeley, C. and Reyburn, H. (2009) Out with the old, in with the new: The utility of rapid diagnostic tests for malaria diagnosis in Africa. Transactions of the Royal Society of Tropical Medicine and Hygiene, 103, 333-337. http://dx.doi.org/10.1016/j.trstmh.2008.10.003
- Murray, C.K., Gasser, R.A., Magill, A.J. and Miller, R.S. (2008) Update on rapid diagnostic testing for malaria. Clinical Microbiology Reviews, 21, 97-110. http://dx.doi.org/10.1128/CMR.00035-07
- Wongsrichanalai, C., Barcus, M.J., Muth, S., Sutamihardja, A. and Wernsdorfer, W.H. (2007) A review of malaria diagnostic tools: Microscopy and rapid diagnostic test (RDT). The American Journal of Tropical Medicine and Hygiene, 77, 119-127.
- World Health Organization (2010) Malaria rapid diagnostic test performance; results of WHO product testing of malaria RDTs: Round 2. http://www.wpro.who.int/internet/files/rdt/RDTMalariaRd2_FINAL.pdf
- Abeku, T.A., Kristan, M., Jones, C., Beard, J., Mueller, D.H., Okia, M., Rapuoda, B., Greenwood, B. and Cox, J. (2008) Determinants of the accuracy of rapid diagnostic tests in malaria case management: Evidence from low and moderate transmission settings in the East African highlands. Malaria Journal, 7, 202-211. http://dx.doi.org/10.1186/1475-2875-7-202
- Kyabayinze, D.J., Tibenderana, J.K., Odong, G.W., Rwakimari, J.B. and Counihan, H. (2008) Operational accuracy and comparative persistent antigenicity of HRP2 rapid diagnostic tests for Plamodium falciparum malaria in a hyperendemic region of Uganda. Malaria Journal, 7, 221-231. http://dx.doi.org/10.1186/1475-2875-7-221
- Chandler, C.I., Jones, C., Boniface, G., Juma, K., Reyburn, H. and Whitty, C.J. (2008) Guidelines and mindlines: Why do clinical staff over-diagnose malaria in Tanzania? A qualitative study. Malaria Journal, 7, 53-65. http://dx.doi.org/10.1186/1475-2875-7-53
- Reyburn, H., Mbatia, R., Drakeley, C., Carneiro, I., Mwakasungula, E., Mwerinde, O., Saganda, K., Shao, J., Kitua, A., Olomi, R., Greenwood, B.M. and Whitty, C.J. (2004). Overdiagnosis of malaria in patients with severe febrile illness in Tanzania: A prospective study. British Medical Journal, 329, 1212-1217. http://dx.doi.org/10.1136/bmj.38251.658229.55
- Cnops, L., Jacobs, J. and Van Esbroeck, M. (2011) Validation of a four-primer real-time PCR as a diagnostic tool for single and mixed Plasmodium infections. Clinical Microbiology and Infection, 17, 1101-1107. http://dx.doi.org/10.1111/j.1469-0691.2010.03344.x
- Mangold, K.A., Manson, R.U., Koay, E.S., Stephens, L., Regner, M., Thomson, R.B. Jr., Peterson, L.R. and Kaul, K.L. (2011). Real-time PCR for detection and identification of Plasmodium spp. Journal of Clinical Microbiology, 43, 2435-2440. http://dx.doi.org/10.1128/JCM.43.5.2435-2440.2005
- Bashir, I.M., Otsyula, N., Awinda, G., Spring, M., Schneider, P. and Waitumbi, J.N. (2013) Comparison of PfHRP- 2/pLDH ELISA, qPCR and microscopy for the detection of plasmodium events and prediction of sick visits during a malaria vaccine study. PLoS One, 8, e56828. http://dx.doi.org/10.1371/journal.pone.0056828
- Rakotonirina, H., Barnadas, C., Raherijafy, R., Andrianantenaina, H., Ratsimbasoa, A., Randrianasolo, L., Jahevitra, M., Andriantsoanirina, V. and Ménard, D. (2008) Accuracy and reliability of malaria diagnostic techniques for guiding febrile outpatient treatment in malaria-endemic countries. The American Journal of Tropical Medicine and Hygiene, 78, 217-221.
- Mawili-Mboumba, D.P., Bouyou Akotet, M.K., Ngoungou, E.B. and Kombila, M. (2010) Evaluation of rapid diagnostic tests for malaria case management in Gabon. Diagnostic Microbiology and Infectious Disease, 66, 162- 168. http://dx.doi.org/10.1016/j.diagmicrobio.2009.09.011
- Mawili-Mboumba, D.P., Akotet, M.K., Kendjo, E., Nzamba, J., Medang, M.O., Mbina, J.R., Kombila, M. and MCORU team (2013) Increase in malaria prevalence and age of at risk population in different areas of Gabon. Malaria Journal, 12, 3. http://dx.doi.org/10.1186/1475-2875-12-3
- Singh, B., Bobogare, A., Cox-Singh, J., Snounou, G., Abdullah, M.S. and Rahman, H.A. (1999) A genusand species—Specific nested polymerase chain reaction malaria detection assay for epidemiologic studies. The American Journal of Tropical Medicine and Hygiene, 60, 687-692.
- Bouyou Akotet, M.K., Mawili-Mboumba, D.P., Madoungou, B. and Kombila, M. (2013) Performances of Malaria P.f/Pan rapid test device Acon® (Pf HRP2/ pan aldolase) and malaria Pf rapid test device Acon® (Pf HRP2) for the diagnosis of malaria in adults and children living in Gabon, Central Africa. Diagnostic Microbiology and Infectious Disease, 77, 58-63. http://dx.doi.org/10.1016/j.diagmicrobio.2013.05.007
- Hopkins, H., Kambale, W., Kamya, M.R., Staedke, S.G., Dorsey, G. and Rosenthal, P.J. (2007) Comparison of HRP2- and pLDH-based rapid diagnostic tests for malaria with longitudinal follow-up in Kampala, Uganda. The American Journal of Tropical Medicine and Hygiene, 76, 1992-1997.
- Rubio, J.M., Buhigas, I., Subirats, M., Baquero, M., Puente, S. and Benito, A. (2001) Limited level of accuracy provided by available rapid diagnosis tests for malaria enhances the need for PCR-based reference laboratories. Journal of Clinical Microbiology, 39, 2736-2737. http://dx.doi.org/10.1128/JCM.39.7.2736-2737.2001
- Hopkins, H., Bebell, L., Kambale, W., Dokomajilar, C., Rosenthal, P.J. and Dorsey, G. (2008) Rapid diagnostic tests for malaria at sites of varying transmission intensity in Uganda. Journal of Infectious Diseases, 197, 510-518. http://dx.doi.org/10.1086/526502
- Desakom, V., Dondorp, A.M., Silamut, K., Pongtavornpinyo, W., Sahassananda, D., Chotivanich, K., Pitisuttithum, P., Smithyman, A.M., Day, N.P. and White, N.J. (2005). Stage dependent production and release of histidin-rich protein 2 by Plasmodium falciparum. Transactions of the Royal Society of Tropical Medicine and Hygiene, 99, 517-524. http://dx.doi.org/10.1016/j.trstmh.2004.11.014
- Endeshaw, T., Graves, P.M., Ayele, B., Mosher, A.W., Gebre, T., Ayalew, F., Genet, A., Mesfin, A., Shargie, E.B., Tadesse, Z., Teferi, T., Melak, B., Richards, F.O. and Emerson, P.M. (2012) Performance of local light microscopy and the ParaScreen Pan/Pf rapid diagnostic test to detect malaria in health centers in Northwest Ethiopia. PLoS One, 7, e33014. http://dx.doi.org/10.1371/journal.pone.0033014
- Moody, A. (2002) Rapid diagnostic tests for malaria parasites. Clinical Microbiology, 15, 66-78. http://dx.doi.org/10.1128/CMR.15.1.66-78.2002
- World Health Organisation (2012) Characteristics of evaluation panel used for round 4 of WHO malaria RDT product testing at US CDC. http://www.who.int/tdr/publications/rapid_diagnostic/en/index.html
- Maltha, J., Gillet, P., Cnops, L., Bottieau, E., Van Esbroeck, M., Bruggeman, C. and Jacobs, J. (2011) Evaluation of the rapid diagnostic test SDFK40 (Pf-pLDH/pan-pLDH) for the diagnosis of malaria in a non-endemic setting. Malaria Journal, 10, 7. http://dx.doi.org/10.1186/1475-2875-10-7
- Wongsrichanalai, C., Arevalo, I., Laoboonchai, A., Yingyuen, K., Miller, R.S., Magill, A.J., Forney, J.R. and Gasser Jr., R.A. (2003) Rapid diagnostic devices for malaria: Field evaluation of a new prototype immunochromatographic assay for the detection of Plasmodium falciparum and non-falciparum Plasmodium. The American Journal of Tropical Medicine and Hygiene, 69, 26-30.
- Eibach, D., Traore, B., Bouchrik, M., Coulibaly, B., Coulibaly, N., Siby, F., Bonnot, G., Bienvenu, A.L. and Picot, S. (2013) Evaluation of the malaria rapid diagnostic test VIKIA malaria Ag Pf/Pan™ in endemic and non-endemic settings. Malaria Journal, 12, 188. http://dx.doi.org/10.1186/1475-2875-12-188
- Koita, O.A., Doumbo, O.K., Ouattara, A., Tall, L.K., Konaré, A., Diakité, M., Diallo, M., Sagara, I., Masinde, G.L., Doumbo, S.N., Dolo, A., Tounkara, A., Traoré, I. and Krogstad, D.J. (2012) False-negative rapid diagnostic tests for malaria and deletion of the histidine-rich repeat region of the hrp2 gene. The American Journal of Tropical Medicine and Hygiene, 86, 194-198. http://dx.doi.org/10.4269/ajtmh.2012.10-0665
- Farcas, G.A., Zhong, K.J., Lovegrove, F.E., Graham, C.M. and Kain, K.C. (2003) Evaluation of the Binax NOW ICT test versus polymerase chain reaction and microscopy for the detection of malaria in returned travelers. The American Journal of Tropical Medicine and Hygiene, 69, 589- 592.
- Fernando, S.D., Karunaweera, N.D. and Fernando, W.P. (2004) Evaluation of a rapid whole blood immunochromatographic assay for the diagnosis of Plasmodium falciparum and Plasmodium vivax malaria. Ceylon Medical Journal, 49, 7-11.
- Gillet P, Mori M, Van Esbroeck M, Van den Ende, J. and Jacobs J. (2009) Assessment of the prozone effect in malaria rapid diagnostic tests. Malaria Journal, 8, 271. http://dx.doi.org/10.1186/1475-2875-8-271
- Allen, L.K., Hatfield, J.M., DeVetten, G., Ho, J.C. and Manyama, M. (2011) Reducing malaria misdiagnosis: The importance of correctly interpreting ParaChek Pf “faint test bands” in low transmission area of Tanzania. BMC Infectious Diseases, 11, 308-314. http://dx.doi.org/10.1186/1471-2334-11-308
- McMorrow, M.L., Masanja, M.I., Abdulla, S.M., Kahigwa, E. and Kachur, S.P. (2008). Challenges in routine implementation and quality control of rapid diagnostic tests for malaria—Rufiji District, Tanzania. The American Journal of Tropical Medicine and Hygiene, 79, 385- 390.
- Coleman, R.E., Sattabongkot, J., Promstaporm, S., Maneechai, N., Tippayachai, B., Kengluecha, A., Rachapaew, N., Zollner, G., Miller, R.S., Vaughan, J.A., Thimasarn, K. and Khuntirat, B. (2006). Comparison of PCR and microscopy for the detection of asymptomatic malaria in a Plasmodium falciparum/vivax endemic area in Thailand. Malaria Journal, 5, 121-127. http://dx.doi.org/10.1186/1475-2875-5-121
- Nicastri, E., Bevilacqua, N., Sañé Schepisi, M., Paglia, M.G., Meschi, S., Ame, S.M., Mohamed, J.A., Mangi, S., Fumakule, R., Di Caro, A., Capobianchi, M.R., Kitua, A., Molteni, F., Racalbuto, V. and Ippolito, G. (2009) Accuracy of malaria diagnosis by microscopy, rapid diagnostic test, and PCR methods and evidence of antimalarial overprescription in non-severe febrile patients in two Tanzanian hospitals. The American Journal of Tropical Medicine and Hygiene, 80, 712-717.
- Proux, S., Suwanarusk, R., Barends, M., Zwang, J., Price, R.N., Leimanis, M., Kiricharoen, L., Laochan, N., Russell, B., Nosten, F. and Snounou, G. (2011) Considerations on the use of nucleic acid-based amplification for malaria parasite detection. Malaria Journal, 10, 323-330. http://dx.doi.org/10.1186/1475-2875-10-323
NOTES
*Corresponding author.

