Applied Mathematics
Vol.09 No.04(2018), Article ID:84308,30 pages
10.4236/am.2018.94031
Models of Cancer Growth Revisited
Jens Christian Larsen
Vanlø se Alle 50 2. mf. tv, 2720 Vanløse, Copenhagen, Denmark
Copyright © 2018 by author and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY 4.0).
http://creativecommons.org/licenses/by/4.0/



Received: April 2, 2018; Accepted: April 27, 2018; Published: April 30, 2018
ABSTRACT
In the present paper we study models of cancer growth, initiated in Jens Chr. Larsen: Models of cancer growth [1] . We consider a cancer model in variables C cancer cells, growth factors , (oncogene, tumor suppressor gene or carcinogen) and growth inhibitor (cells of the immune system or chemo or immune therapy). For this says, that cancer grows if (1) below holds and is eliminated if the reverse inequality holds. We shall prove formulas analogous to (1) below for arbitrary . In the present paper, we propose to apply personalized treatment using the simple model presented in the introduction.
Keywords:
Cancer, Mass Action Kinetic System, Immunity

1. Introduction
Cancer grows if and
(1)
and is eliminated if the reverse inequality holds. Here and are initial conditions in , see section three for definitions and also [1] . So if you have many (few) growth inhibitors compared to growth factors, cancer is eliminated (cancer grows).
In [1] we proved, that Formula (1) when implied that cancer grows and is eliminated if the reverse inequality holds. In the present paper we prove, that cancer grows if and
(2)
and is eliminated if the reverse inequality holds. In [1] we also considered a mass action kinetic system with vector field f like the one in Section 4 with and proved, that there is a relationship between such a model and the model T of Section 3. Namely if you linearize f at a singular point and then discretize the flow then you get a mapping T of Section 3. See section 4 for details.
Consider now the cancer model from [1]
(3)
Here , where T denotes a transpose. If you fit my model to measurements, you will get some information about the particular cancer. is the cancer agressiveness parameter. If this parameter is high cancer initially proliferates rapidly. is the carcinogen severity. is the fitness of the immune system, its response to cancer. are decay rates. g is a vector of birth rates. gives the growth factor response to cancer and gives the growth inhibitor response to cancer. So fitting my model may have prognostic and diagnostic value. If we have a toxicology constraint for chemo therapy or immune therapy with a suitable safety margin
(4)
then we can keep the system at the toxicology limit by requiring
(5)
which is equivalent to
(6)
If , then we can give chemo therapy at this rate. Then we get the induced system
(7)
(8)
We shall prove that this treatment benefits the patient in section 2. To get the system to the toxicology limit P assume that we have
(9)
Then looking at the third coordinate of T we see that we shall require
(10)
which implies that
(11)
We can also fit the ODE model of section 4 with , by defining the Euler map
(12)
. Iterating this map will give an approximation to the flow. Then is the rate at which you give chemo therapy. If we have the constraint
(13)
then looking at the third coordinate of H we see that to keep the system at the toxicology limit with a suitable safety margin, we must have
(14)
Solving for we get
(15)
Since the are positive we can give the chemo therapy at this rate. To get this system to the toxicology limit we shall require
(16)
which means, that
(17)
I felt I had to suggest this. If you want to try this you may want to do it stepwise.
In Figure 1, I have plotted a fit of T to three Gompertz functions
(18)
Figure 1. A fit to Gompertz functions. The upper curve is , the middle and the lower curve is . The solid curves are the Gompertz functions and the dots the model T.
(19)
(20)
From a paper from 1964 [2] we know that solid tumors grow like Gompertz functions. That is the cancer burden is approximately a Gompertz function. Define the error functions
(21)
(22)
(23)
where are measurements of at equidistant time points . We set , , Then solve the equations
(24)
(25)
(26)
(27)
in unknowns and
(28)
(29)
(30)
in unknowns and
(31)
(32)
(33)
in unknowns . For instance
(34)
gives
(35)
The result is
(36)
(37)
(38)
(39)
(40)
(41)
(42)
(43)
(44)
(45)
I have also fitted S to two Gompertz functions
(46)
(47)
See Figure 2 and Figure 3. Define error functions
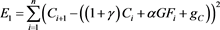 (48)
(48)
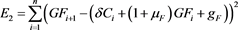 (49)
(49)
Figure 2. A fit of S to a Gompertz functions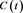 . The solid curve is the Gompertz function and the dots are the model S.
. The solid curve is the Gompertz function and the dots are the model S.
Figure 3. A fit of S to a Gompertz functions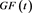 . The solid curve is the Gompertz function and the dots are the model S.
. The solid curve is the Gompertz function and the dots are the model S.
and measurements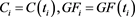 . Solve
. Solve
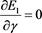 (50)
(50)
 (51)
(51)
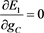 (52)
(52)
in unknowns 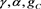 and
and
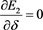 (53)
(53)
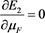 (54)
(54)
 (55)
(55)
in unknowns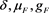 . The result is
. The result is
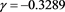 (56)
(56)
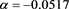 (57)
(57)




In Maple, there is a command QPSolve that minimizes a quadratic error function with constraints on the signs of the parameters estimated. There are several important monographs relevant to the present paper, see [3] - [8] . There are several publications by the author impacting on the present paper, see [9] - [15] .
2. The Routh Hurwitz Criterion for Maps
We shall derive a well known criterion for stability of a fixed point of a map. To this end define the Möbius transformation

which maps the left hand plane 


implies



implies

Define

Then

when 



This shows, that g is a bijective map with inverse

denote the characteristic polynomial of the two by two matrix A in (7). Note, that if 






Define

So if 

If this polynomial is a Routh Hurwitz polynomial, i. e. the roots lie in

lie in the interior 

and

Also

If

and 


But this implies, by adding these two inequalities, that

However, then

A contradiction to (80). So if 




by the Routh Hurwitz criterion. We shall find the fixed points of S, with

Then the first coordinate gives

But the denominator is positive if 


and we are lowering 

Now suppose

The assumption 
Since

We claim that 




Clearly both

and

have unique fixed points 


We need the following definition.
Definition. A fixed point 






for all
Now observe, that


Notice that

in the max norm

because

But now stability follows from the estimate

and this implies that 


If

as



3. Models of Cancer Growth
Consider the mapping

where

and

The matrix here is denoted A. T maps 









C is cancer 


Proposition 1 The characteristic polynomial 


Proof. With


Decompose 








Suppose henceforth, that

Then the characteristic polynomial of A is

So the eigenvalues are 

is a factor of

are eigenvalues of A, where


For the moment assume

We shall find formulas for the complements

of D and the determinant of D,
Proposition 2 For 

For

Proof. Suppose












which is what we wanted to prove.
Now suppose that


Now consider the case




Decompose after rows

which gives

The proposition follows, because

Proposition 3

Proof. We have

Initially let

when 

when

If

(

Now we shall use induction over q to prove the formula in the statement of the proposition. Decompose after the last row




In B we have decomposed after 



The proposition follows.
The aim of our computations is to show that there exists an affine vector field X on 

Let 


First notice that

if

and

which we assume. We have used that

So the eigenvectors in D are linearly independent, hence

Now define when

where


where we denote the last vector




Now we get



It follows that 

that is

We shall require

because then the time one map of Y is

Now define

Then the flows of X and Y are related by

But then the time one map of X is

which is what we wanted.
Theorem 4 Assume, that

and

We have the formula




Proof. We use the formula



We have

and then

for 

We shall write

and then we have

When 

while for 

Notice that

Continuing from (184)





So this gives the first term in

Note that

Now we have



Hence the 


and the 

So




The theorem follows.
Now suppose that




The first column is denoted







Now as in [1] we get




and similarly



So

because we assume that
Proposition 5 For

and for

Also

Finally

since

Proof. The proposition follows immediately from proposition 2 and 3.
The flow of

is, for

We want to have that this equals for

Thus


Remember the formula

Define when

where


where the last vector is denoted


Now we also get


So 

because then the time one map of Y is

Then define

The flows are related by

But then the time one map of X is

which is what we intended to find.
Theorem 6 When 



Proof. We have the following computation


And we want to have


that is

But we have arranged that

so we get


Denote the two by two matrix in the last line

We can also compute the first term in

omitting the factor




hence the first term in

Now


The 




The 




where

The theorem follows.
Now assume that

Proposition 7 For q odd and

and for 

For q even and

and for 

Also

when q is odd and

when q is even.
Proof. (277) q odd. We are deleting the row r with



The first sign here is the sign when decomposing after row








We have the sign

on

on column 




For 

We have, decomposing after the last column




Here

For q even we get

Now decompose after the last column

Now we get decomposing after the last column




where

In the determinant B, we have decomposed after row 2 to

Define for 

From Proposition 7, we get
Proposition 8 For q odd and

and for q odd and

For q even and

and for q even and

Also for q odd and

and for q odd and

For q even and

and for q even and

Finally for q odd

and

for q even.
4. An ODE Model
In [1] we also considered a three dimensional ODE model of cancer growth in the variables 






Here the complexes are






the forward reaction rate is denoted 




We shall find a polynomial giving candidates of singular points of this vector field.
From

where


for


We can then multiply with

and define the constants

to obtain the polynomial of degree


if we assume that

and the discrete dynamical system of section three, see also [1] . Linearize the vector field at a singular point 

Also define the Euler map

for






and


then you obtain a discrete model T of section three.
Example Let 



5. Summary
In this paper, we considered a discrete mathematical model and an ODE model of cancer growth in the variables 


then cancer grows, and if the reverse inequality holds, cancer is eliminated. We also proposed personalized treatment using the simple model of cancer growth in the introduction and the ODE model of section four.
Cite this paper
Larsen, J.C. (2018) Models of Cancer Growth Revisited. Applied Mathematics, 9, 418-447. https://doi.org/10.4236/am.2018.94031
References
- 1. Larsen, J.C. (2017) Models of Cancer Growth. Journal of Applied Mathematics and Computing, 53, 615-643.
- 2. Laird, A.K. (1964) Dynamics of Cancer Growth. British Journal of Cancer, 18, 490-502. https://doi.org/10.1038/bjc.1964.55
- 3. Adam, J.A. and Bellomo, N. (1997) A Survey of Models for Tumor-Induced Immune System Dynamics. Birkh?user, Boston.
- 4. Geha, R. and Notarangelo, L. (2012) Case Studies in Immunology. Garland Science, New York, London.
- 5. Marks, F., Klingmüller, U. and Müller-Decker, K. (2009) Cellular Signal Processing. Garland Science, Heidelberg.
- 6. Molina-Paris, C. and Lythe, G. (2011) Mathematical Models and Immune Cell Biology. Springer Verlag, Beilin. https://doi.org/10.1007/978-1-4419-7725-0
- 7. Murphy, K. (2012) Immuno Biology. 8th Edition, Garland Science, New York, London.
- 8. Rees, R.C. (2014) Tumor Immunology and Immunotherapy. Oxford University Press, Oxford.
- 9. Larsen, J.C. (2016) Hopf Bifurcations in Cancer Models. JP Journal of Applied Mathematics, 14, 1-31.
- 10. Larsen, J.C. (2017) A Mathematical Model of Adoptive T Cell Therapy. JP Journal of Applied Mathematics, 15, 1-33.
- 11. Larsen, J.C. (2016) Fundamental Concepts in Dynamics. Survey Article.
- 12. Larsen, J.C. (2017) The Bistability Theorem in a Cancer Model. International Journal of Biomathematics. (To appear)
- 13. Larsen, J.C. (2016) The Bistability Theorem in a Model of Metastatic Cancer. Applied Mathematics, 7, 1183-1206. https://doi.org/10.4236/am.2016.710105
- 14. Larsen, J.C. (2017) A Study on Multipeutics. Applied Mathematics, 8, 746-773. https://doi.org/10.4236/am.2017.85059
- 15. Larsen, J.C. (2017) A Mathematical Model of Immunity. JP Journal of Applied Mathematics. (To appear)




























