Food and Nutrition Sciences
Vol. 4 No. 9A1 (2013) , Article ID: 36107 , 6 pages DOI:10.4236/fns.2013.49A1006
Exploring the Role of Fatty Acid on Transcription Factors Regulating Fatty Acid Metabolism with Emphasis on Trans Fatty Acid
![]()
1Department of Cardiac Biochemistry, All India Institute of Medical Sciences, Delhi, India; 2Department of Surgery, All India Institute of Medical Sciences, Delhi, India; 3Centre of Chronic Disease Control, Delhi, India; 4Department of Cardiology, All India Institute of Medical Sciences, Delhi, India.
Email: ransijob@gmail.com, *lakshmy_ram@yahoo.com, rparshad@yahoo.com, varnaseenu@hotmail.com, ccdc@ccdcindia.org, vkbahl2002@yahoo.com
Copyright © 2013 Ransi Ann Abraham et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received May 15th, 2013; revised June 15th, 2013; accepted June 22nd, 2013
Keywords: Transcription Factors; Trans Fatty Acid; Adipose Tissue; qPCR
ABSTRACT
Fatty acids are unique macromolecules as they act as biological modulators of transcription factors and regulate their own metabolism by controlling the activity or abundance of transcription factors of fatty acid metabolism either by RNA processing and RNA stability. Peroxisome Proliferator Activated Receptor (PPAR-γ) and Sterol Regulatory Element Binding Protein (SREBP-1c) are transcription factors expressed primarily in adipose tissue. We have studied the relation of fatty acid including trans fatty acid assessed in adipose tissue with the transcription factors. Adipose tissue was collected from 50 healthy subjects undergoing elective abdominal surgery. Fatty acid was assessed in the tissue by gas chromatography. The expressions of PPARγ and SREBP-1c were studied by real time RT-PCR. The expressions of PPARγ and SREBP1c were significantly correlated (r = 0.4 p < 0.005). The trans fatty acid did not show any significant correlation with expression but significant correlation was observed between DHA (Docosahexaenoic acid) and PPARγ expression (r = 0.33 p < 0.03) which remained significant (r = 0.87, p < 0.0001) after being adjusted for BMI and insulin. An upregulation of PPARγ led to decreased levels of SREBP1c. In conclusion, trans fatty acid did not affect the expressions of PPAR-γ and SREB1c in this study.
1. Introduction
Fatty acids are unique macromolecules, which act as biological modulators of transcription factors and thereby regulate gene expression and influence lipid, carbohydrate and protein metabolism [1]. This effect of fatty acid is cell specific and is determined by the number and position of double bonds which influence the function and metabolism of fatty acids, therefore changes in the fatty acid composition of diets may have profound effects on several physiological processes [2]. Trans fatty acids (TFA) are unsaturated fatty acids with at least one double bond in the trans configuration, and occur in small amounts in some foods from animals, however, majority of the trans fatty acids are artificially created during partial hydrogenation. The lipid and non lipid effects of TFA appear to be linked to its biological effects on hepatocytes, monocytes/macrophages, endothelial cells and adipocytes [3]. Although the molecular pathways whereby TFA influence these tissues are unclear, studies on other dietary fatty acids suggest that both cell membranes and gene transcription are likely to be effected.
The regulation of gene transcription by fatty acids is primarily by the changes in the activity of at least 4 transcription factors families namely PPAR (peroxisome proliferator activated receptor), LXR (liver X receptor), HNF-4α (hepatic nuclear factor 4) and SREBP (sterol regulatory element binding protein) [4,5]. Peroxisome proliferator-activated receptors (PPARs) belong to a ligand-activated nuclear hormone receptor superfamily and are known to regulate the expression of numerous genes involved in fatty acid metabolism and adipocyte differentiation. They exist in three isoforms: PPARα, PPARγ, and PPARδ [6]. PPARγ is mainly expressed in white adipose tissue [7]. SREBP 1a and SREBP-1c are encoded by a single gene through the use of alternative transcription start sites and differ by their first exon [8]. SREBP-1c are membrane-bound transcription factors that belong to the basic helix-loop helix leucine zipper family expressed in most of the tissues of mice and human with high levels in the liver, white adipose tissue, adrenal gland and brain [9].
Study of the influence of trans fatty acids on transcription factors expression in adipocytes would likely help us to understand the molecular mechanism by which trans fatty acid causes dyslipidemia. It is against this background that an attempt was made to see the affect of trans fatty acid, which are structurally similar to polyunsaturated fatty acid (PUFA) but are known to behave as saturated fatty acid (SFA), on the expression of transcription factors, PPARγ and SREBP1c, in adipose tissue.
2. Methods
50 healthy subjects of 20 - 60 years of age undergoing elective abdominal surgery were recruited for the study. Subjects who had a previous history of coronary artery disease, cancer, and endocrine disorder and those who had gained or lost 5 Kg weight in previous two years and on changed diet due to clinical condition in the same duration were excluded. Written informed consent was obtained from all subjects. Ethics committee of the All India Institute of Medical Sciences, New Delhi, approved the study. Height, weight, waist circumference and hip circumference of all subjects were measured by standard procedures.
10 ml blood was collected from subjects who were overnight fasting, centrifuged and serum was separated and stored at −70˚C till analysis. Adipose was collected from subjects during the surgery, washed in saline to remove blood and one portion stored in a amber coloured vial containing a solution of hexane: isoproponol (3:2 v/v) and kept at −70˚C. BHT (Butylated Hydroxytoluene) as antioxidant was added to all the samples before storage. Another portion of the adipose tissue was weighed and stored in RNA later (RNA stabilization solution from Ambion) at −70˚C for transcription studies.
2.1. Fatty Acid Estimation
The lipids from adipose tissue were isolated by Folch method [10]. The extracted lipids were esterified using 500 μl methylating reagent containing methanol-acetyl chloride 20:1 (v/v) by heating in water-bath at 50˚C for one hour. The esterified fatty acids evaporated under stream of nitrogen, dissolved in isooctane and stored in amber coloured vials till separation of fatty acids. The esters were re-dissolved in iso-octane, C17:0 was added as a internal standard, mixed and 1 μl sample was taken in Hamilton syringe and injected into the fused silica capillary cis/trans column SP 2560, 100 m × 250 μm internal diameters × 0.20 μm film (Supelco, Belefonte, Pennsylvania) attached to the gas chromatography (GC) instrument. Nitrogen gas was used as a carrier. A four temperature programme was found ideal for separation of all the fatty acids including the trans fatty acid isomers. The temperature conditions were as follows: The port temperatures of both the injector and the detector were set at 250˚C. The oven temperature was initially set at 90˚C for 4 min and was then increased 15˚C/min until a temperature of 150˚C was reached and held for 10 min, the temperature was further increased at 1˚C/min till 170˚C after which rate of temperature change was 5˚C/min until a temperature of 230˚C was reached and maintained for 30 minutes. A split ratio of 1:10 and an injection volume of 1 μl were used. The separated fatty acid methyl esters were detected by Flame-ionization detector attached to the GC. Each fatty acid peak in the unknown samples was identified with reference to C17:0 as well as by comparing with known standards both individual as well as mixed (FattyAcid Methyl Esters from SUPELCO). For confirmation of fatty acid some unknown samples were spiked with known standards. Peaks were quantified by calculating the area under the peak using software from AIMIL (Nucon Technologies).
The fatty acids were also recalculated and grouped together in different categories as per their degree of unsaturation. The fatty acids included among total saturated fatty acids (TSFA) are C12:0, C14:0, C15:0, C16:0 and C18.0, and those in total monounsaturated fatty acids (TMUFA) are cis C14:1 cis, C16:1 cis and C18:1 cis. Total polyunsaturated fatty acids (TPUFA) (cis) include C18:2 cis, cis, C18:3n-6, C18:3n-3. Total trans fatty acid (TTFA) comprises of C16:1 9-trans, C18:1 9-trans and C18:1 11-trans, C18:2 trans, trans, C18:2 trans, cis and C18:2 cis, C18:3 trans trans trans, C18:3 trans trans cis, C18:3 trans cis cis, C18:3 cis cis trans, C18:3 trans cis trans, C18:3 cis trans cis, C18:3 cis trans trans. Total long chain fatty acid (TLCFA) comprises of cis C20:4, cis C20:5 and cis C22:6. To calculate the interassay and intraassay coefficient of variation fatty acid methyl esters were prepared from pooled serum in multiple tubes and run on GC. The interassay coefficient of variation was calculated from the variation observed in three GC runs of the sample on the same day from the same vial which was carefully diluted for the runs and utmost care was taken to prevent evaporation of diluted sample. The methyl esters from pooled serum were run on three consecutive days to calculate the intra-asay coefficient of variation.
2.2. Real Time PCR to Study Gene Expression
RNA was isolated by RNeasy Lipid Tissue Mini Kit (QIAGEN) as per the manufacturer’s instructions. Adipose tissue stored in RNA later was removed from storage, weighed carefully and homogenized by a mechanized tissue ruptor under ice before isolation. RNA was quantified using picodrop and gene expression was studied by real-time two-step RT-PCR. cDNA was synthesized using Superscript III first-strand synthesis super mix (Invitrogen) commercially available kit. Real time PCR was carried out using 2 × Quatitect SYBR green PCR master mix. The reaction was carried out in a 25 µl reaction mix containing 12.5 µl of PCR master mix, 1 µl of forward and reverse primer 10nm concentration and nuclease free water. The PCR was followed by melt curve analysis carried out between 65˚C and 95˚C. The amplification curve gives data regarding the kinetics of amplification of the target sequence which is the cycle time (Ct) at which fluorescence reach the threshold levels.
Biochemical parameters such as glucose, cholesterol, triglycerides were estimated by standard kits from RANDOX, UK. High Density Lipoprotein cholesterol (HDL-c) was estimated after precipitating low density lipoproteins and chylomicrons with the precipitating reagent. Very low density lipoprotein (VLDL) and low density lipoprotein (LDL) values were derived from frieldwald equation. Insulin was measured using Enzyme Linked Immunosorbent Assay (ELISA) method using commercially available kit (Mercodia, Sweden) as per manufacturer’s protocol.
2.3. Data Analysis
To control for experimental error introduced during the multistage process required to extract and process the RNA two internal controls 18sRNA (18S ribosomal RNA) and Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) were used, which were made to undergo real time PCR (qPCR). The expression was quantified by relative quantitation method which measures changes in the steady-state levels of a gene of interest relative to an invariate control gene known which consisted of a 4-fold dilution series of cDNA and a NTC (no template control). The standard curve, covering 5 orders of magnitude of template concentration showed good linearity with a correlation coefficient of >0.99 for each reference and gene studied. Quantities interpolated from the resulting standard curve were used to calculate relative mRNA levels in each unknown samples. Standard curves were prepared for both the target and the endogenous reference. The target amount was divided by the endogenous reference amount to obtain a normalized target value.
Significance of the correlations between the fatty acid measured in adipose tissue and the expression of transcription factors was examined by the nonparametric Spearman’s rank correlation test. Correlations were also computed between the biochemical parameters and transcription factors. The threshold for significance was set at p ≤ 0.05. The analysis was done on SPSS version 16.0 software (SPSS, Chicago, IL, USA).
3. Results
The baseline characteristics of subjects are given in Table 1. The mean age of the subjects (N = 50), (29 females and 21 males) was 39 (8.8) yrs with the mean BMI of 23.7 (9.5) kg/m2 which was not significantly different in males and females. The yield of RNA from adipose tissue stored in RNA later was 40.23 ng/µl (24.54 - 58.59) with a 260/280 ratio ranging from 1.7 to 2.0. 0.5 µg of RNA was linearly converted to cDNA. The expression for PPARγ ranged from 0.66 to 2.41 while the expression of SREBP1c ranged from 0.03 to 1.09. The distribution of fatty acid in adipose tissue expressed as median and intraquartile range is given in Table 2. Saturated fatty acid was the most abundant fatty acid with palmitic acid forming the bulk, contributing 36 percent in adipose followed by stearic acid. The distribution of monounsaturated fatty acid and trans fatty acid was 21.96% 3.56% respectively while cis polyunsaturated fatty acid was 20%. The coefficient of variation of fatty acid measurement on the same day was 1.05% for saturated fatty acid, 0.77% for mono unsaturated fatty acid, 1.03% for poly unsaturated fatty acid and 6.55% for total trans fatty acid. The Inter day coefficient of variation for saturated fatty acid was 1.52%, for monounsaturated fatty acid it was 1.55%, 0.66% for PUFA and 13.25% for total trans fatty acids. Duplicate samples of adipose tissue were run to assess the repeatability of the method. The Intra class
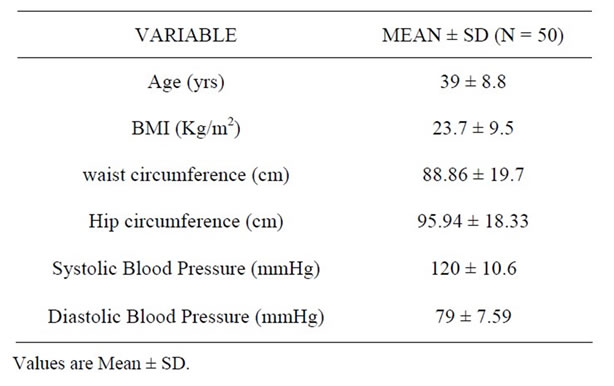
Table 1. The baseline characteristics of subjects.
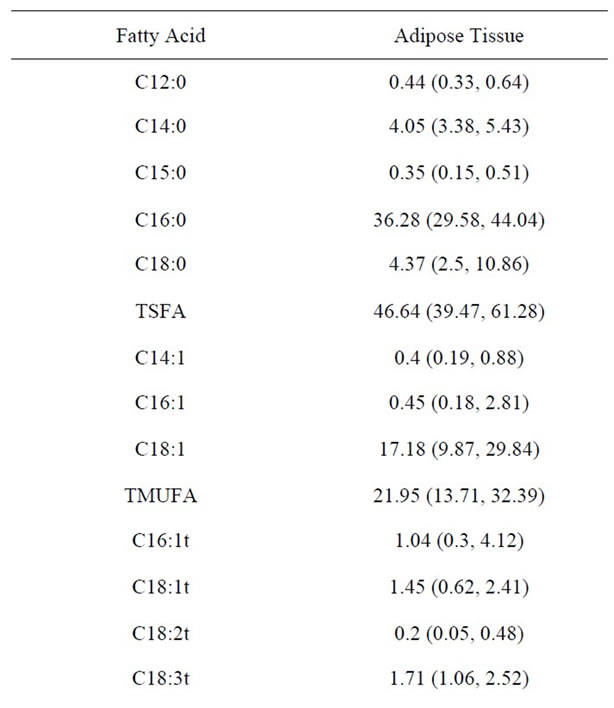
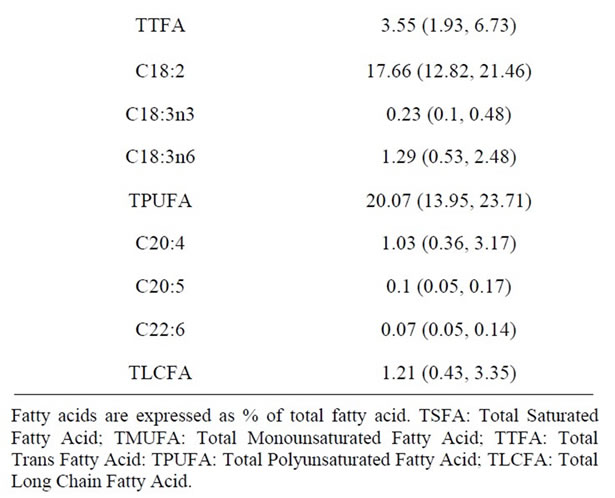
Table 2. Distribution of fatty acid in adipose tissue.
correlation (ICC) was 0.93, 0.74, 0.86 and 0.94 for saturated fatty acid, mono unsaturated fatty acid, poly unsaturated fatty acid and total trans fatty acid respectively.
The expression of PPARγ and SREBP1c was significantly correlated (r = 0.4 p < 0.005). The spearman correlation of adipose fatty acids with the transcription factors PPARγ and SREBP1c is given in Table 3. The correlations of individual trans fatty acids with transcription factors are given in Table 4. Both the transcription factors were negatively correlated with total saturated fatty acid (TSFA) but the correlation was not significant. A significant correlation was observed for adipose tissue level of DHA and PPARγ expression (r = 0.33 p < 0.03).
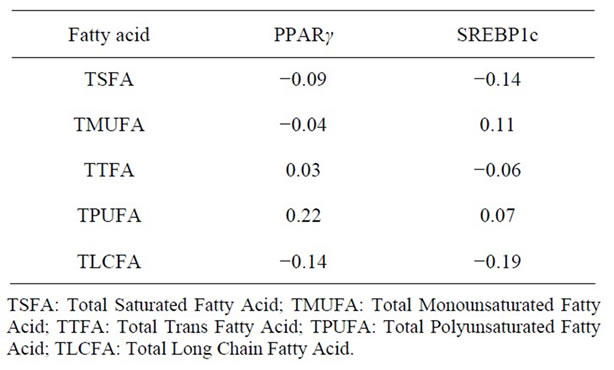
Table 3. Spearman correlation of adipose tissue fatty acid and transcription factors.

Table 4. Spearman correlation of individual trans fatty acid adipose tissue and transcription factors.
Correlation between DHA and PPAR-γ expression remained significant (r = 0.87, p < 0.0001) even after adjusting for BMI and insulin. No significant correlation was observed between trans fatty acids and PPARγ or SREB1c. Trans isomer of α linolenic acid (18:3t) assessed in adipose tissue showed correlation of r = 0.27 with SREBP1c levels while the correlation with trans isomer of oleic acid [18:1t] was r = 0.22. Table 5 shows the correlation of the transcription factor with the biochemical variables analyzed. Triglyceride showed a significant correlation with PPAR-γ (0.52), which was also reflected in VLDL.
4. Discussion
Many of the effects of fatty acids in both cell biology and human health and disease relate to their abilities to regulate gene expression and subsequent downstream events. Studies and reviews on the nature of the transcription factors and their expression in the presence of polyunsaturated fatty acid have been carried out but their expressions in relation to presence of trans fatty acid are sparse. Trans fatty acids leads to lower levels of n-3 fatty acid and changes in PUFA content [11].
Trans isomer of α linoleic acid (18:3t) and trans isomer of oleic acid (18:1t) assessed in adipose tissue showed
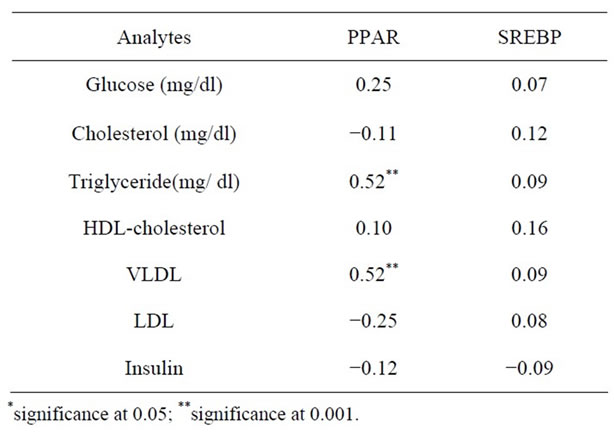
Table 5. Correlation of biochemical analytes and transcription factors.
the correlation of r = 0.27 and r = 0.22 with SREBP1c though not significant. In an experimental study, SREBP- 1c expression was 130% higher in trans-fed mice than in PUFA or SFA fed mice. PPARγ mRNA was over 4-fold higher in trans-fed mice than in PUFAand SFA-fed mice. However, the groups did not differ in expression of genes related to fatty acid oxidation [12]. Effect of TFA and palmitic acid on increased cholesterol synthesis and plasma LDL-C concentration is likely to be mediated by SREBP-1c which is known to activate enzymes of triglyceride and cholesterol synthesis [13]. In our study also we found a good correlation (r = 0.58) between trans isomer of oleic acid (18:1t) and LDL which is probably mediated through SREBP-1c.
The correlation of 0.24 and 0.22 for linoleic acid and total PUFA was observed with PPARγ. A significant positive correlation was found between DHA (0.33: p ≤ 0.03) and PPARγ. The correlation remained significant after adjusting for BMI and insulin. In an invitro study where adipocytes were incubated with fatty acid a significant increase in PPARγ1 mRNA was observed in the presence of eicosapentaenoic acid (EPA) (C20:5n-3). These effects were independent of gender, age, BMI, fasting glycemia, and type of sugar. Neither linolenic acid (C18:3n-3), or PUFAs from the n-6 series affected PPARγ expression in human adipocytes [14,15]. When the PPARγ mRNA levels in adipose tissue biopsy was studied in the same subjects a strong positive correlation (r = 0.76; p < 0.016) was found for EPA as well as DHA (r =0.86; p < 0.006). In contrast to these n-3 PUFAs, there was no correlation between PPARγ mRNA and concentrations of other fatty acids such as linoleic acid (C18:2n-6; r = 0.21, p < 0.95), arachidonic acid (C20: 4n-6; r = 0.60, p < 0.12), oleic acid (C18:1n-9; r = 0.61, p < 0.08), or palmitic acid (C16:0 r = 0.17, p < 0.65) [16]. This is similar to correlations observed in our study. EPA and DHA activate PPAR which leads of upregulation of gene of fatty acid metabolism and down regulates genes of inflammation [17]. A significant correlation of r = 0.5 was observed between PPARγ and triglyceride concentration. It has been shown that feeding SFA or TFA increases TG levels in plasma which may be due to increased lipolysis as a result of decreased insulin sensitivity. It has been also shown that infusion of a triglyceride mixture for 5 hours is able to promote a marked increase in PPARγ mRNA expression in human subcutaneous adipose tissue [18]. PPARγ activation by agonists favours the storage of lipids in adipose tissue, thereby protecting the rest of the body from lipid overload and insulin resistance [19] suggesting a mechanism for beneficial effect of dietary PUFA. Linoleic acid (n-6) was the main PUFA consumed by the subjects in our study.
An upregulation of PPARγ often leads to decreased levels of SREBP1c [20] and this was observed in our study also. The lower expression of SREBP-1c could be mediated through LXR. PPARγ further lower SREBP-1c level by influencing proteolytic processing which is mediated by liver X receptors (LXRs). The adipose level of TFA in our study is low and PUFA was high. High PUFA are known to inhibit SREBP-1c which may be another explanation for the low SREBP-1c expression seen in our study.
5. Conclusion
This study showed a higher expression of PPARγ as compared to SREBP-1c. We found significant correlations of PPARγ with only DHA. Trans fatty acids showed some trends in correlation but were not significant statistically. We found significant correlation between PPARγ and SREBP-1c. Cross-talk between these nuclear factors could be important in nutritional regulation of fatty acid metabolism [21].
6. Acknowledgements
This study was funded by the institutional grant from All India Institute of Medical Sciences.
REFERENCES
- D. B. Jump, “Dietary Polyunsaturated Fatty Acids and Regulation of Gene Transcription,” Current Opinion in Lipidology, Vol. 13, No. 2, 2002, pp. 155-164. doi:10.1097/00041433-200204000-00007
- G. Mann, “Metabolic Consequences of Dietrary Trans Fatty Acids,” Lancet, Vol. 343, No. 8908, 1994, pp. 1268-1271. doi:10.1016/S0140-6736(94)92157-1
- R. Micha and D. Mozaffarian, “Trans Fatty Acids: Effects on Metabolic Syndrome, Heart Disease and Diabetes,” Nature Reviews Endocrinology, Vol. 5, 2009, pp. 335- 344. doi:10.1038/nrendo.2009.79
- T. T. Lu, J. J. Repa and D. J. Mangelsdorf, “Orphan Nuclear Receptors as eLiXiRs and FiXeRs of Sterol Metabolism,” The Journal of Biological Chemistry, Vol. 276, 2001, pp. 37735-37738.
- J. M. Olefsky, “Nuclear Receptor Minireview Series,” The Journal of Biological Chemistry, Vol. 276, 2001, pp. 36863-36864. doi:10.1074/jbc.R100047200
- B. P. Kota, T. H. Huang and B. D. Roufogalis, “An Overview on Biological Mechanisms of PPARs,” Pharmacological Research, Vol. 51, No. 2, 2005, pp. 85-94. doi:10.1016/j.phrs.2004.07.012
- P. Tontonoz, E. Hu and B. M. Spiegelman, “Stimulation of Adipogenesis in Fibroblasts by PPAR γ2, a Lipid-Activated Transcription Factor,” Cell, Vol. 79, No. 7, 1994, pp. 1147-1156. doi:10.1016/0092-8674(94)90006-X
- X. Hua, J. Wu, J. L. Goldstein, M. S. Brown and H. H. Hobbs, “Structure of the Human Gene Encoding Sterol Regulatory Element Binding Protein-1(SREBPF1) and Localization of SREBPF1 and SREBPF2 to Chromosome 17p11.2 and 22q13,” Genomics, Vol. 25, No. 3, 1995, pp. 667-673. doi:10.1016/0888-7543(95)80009-B
- I. Shimomura, H. Shimano, J. D. Horton, J. L. Goldstein and M. S. Brown, “Differential Expression of Exons 1a 1nd 1c in mRNA for Sterol Regulatory Element Binding Protein-1 in Human and Mouse Organs and Cultured Cells,” Journal of Clinical Investigation, Vol. 99, No. 5, 1997, pp. 838-845. doi:10.1172/JCI119247
- J. Folch, M. Lees and G. H. S. Stanley, “A Simple Method for the Isolation and Purification of Total Lipids from Animal Tissues,” The Journal of Biological Chemistry, Vol. 226, 1957, pp. 497-509.
- A. Ascherio and W. C. Willett, “Metabolic and Atherogenic Effects of Trans Fatty Acids,” Journal of Internal Medicine, Vol. 238 No. 2, 1995, pp. 93-96. doi:10.1111/j.1365-2796.1995.tb00906.x
- R. M. Machado, et al., “Intake of Trans Fatty Acids Causes Nonalcoholic Steatohepatitis and Reduces Adipose Tissue Fat Content,” The Journal of Nutrition, Vol. 140, No. 6, 2010, pp. 1127-1132. doi:10.3945/jn.109.117937
- K. Sundaram, M. A. French and T. A. Clandinin, “Exchanging Partially Hydrogenated Fat for Palmitic Acid in the Diet Increases LDL-Cholesterol and Endogenous Cholesterol Synthesis in Normocholesterolemic Women,” European Journal of Nutrition, Vol. 42, No. 4, 2003, pp. 188-194. doi:10.1007/s00394-003-0411-9
- D. Auboeuf, J. Rieusset, L. Fajas, et al., “Tissue Distribution and Quantification of the Expression of mRNAs of Peroxisome Proliferator-Activated Receptors and Liver X Receptor-Alpha in Humans: No Alteration in Adipose Tissue of Obese and NIDDM Patients,” Diabetes, Vol. 46, No. 8, 1997, pp. 1319-1327. doi:10.2337/diabetes.46.8.1319
- J. Rieusset, F. Andreelli, D. Auboeuf, et al., “Insulin Acutely Regulates the Expression of the Peroxisome Proliferator Activated Receptor in Human Adipocytes,” Diabetes, Vol. 48, No. 4, 1999, pp. 699-705. doi:10.2337/diabetes.48.4.699
- C. Chambrier, J.-P. Bastard, J. Rieusset, et al., “Eicosapentaenoic Acid Induces mRNA Expression of Peroxisome Proliferator Activated Receptor,” Obesity Research, Vol. 10, 1999, pp. 518-525.
- M. Bouwens, O. van de Rest, N. Dellschaft, et al., “Fish Oil Supplementation Induces Anti-Inflammatory Gene Expression Profiles in Human Blood Mononuclear Cells,” The American Journal of Clinical Nutrition, Vol. 90, No. 2, 2009, pp. 415-424. doi:10.3945/ajcn.2009.27680
- E. Nisoli, M. O. Carruba, C. Tonello, C. Macor, G. Federspil and R. Vettor, “Induction of Fatty Acid Translocase/CD36, Peroxisome Proliferator-Activated Receptor-2, Leptin, Uncoupling Proteins 2 and 3, and Tumor Necrosis Factor-Gene Expression in Human Subcutaneous Fat by Lipid Infusion,” Diabetes, Vol. 49, No. 3, 2000, pp. 319-324. doi:10.2337/diabetes.49.3.319
- P. Ferre, “The Biology of Peroxisome Proliferator-Activated Receptors: Relationship with Lipid Metabolism and Insulin Sensitivity,” Diabetes, Vol. 53, Suppl. 1, 2004, pp. S43-S50. doi:10.2337/diabetes.53.2007.S43
- R. J. Deckelbaum, T. S. Worgall and T. Seo, “n-3 Fatty Acids and Gene Expression,” The American Journal of Clinical Nutrition, Vol. 83, Suppl. 6, 2006, pp. 1520S- 1525S.
- T. Ide, H. Shimano, T. Yoshikawa, et al., “Cross-Talk between Peroxisome Proliferator-Activated Receptor (PPAR) Alpha and Liver X Receptor (LXR) in Nutritional Regulation of Fatty Acid Metabolism. I. PPARs Suppress Sterol Regulatory Element Binding Protein-1c Promoter through Inhibition of LXR Signaling,” Molecular Endocrinology, Vol. 17, No. 7, 2003, pp. 1240-1254.
NOTES
*Corresponding author.

