Food and Nutrition Sciences
Vol.4 No.5(2013), Article ID:31233,7 pages DOI:10.4236/fns.2013.45067
Herbicide-Resistant Mutations in Acetolactate Synthase Can Reduce Feedback Inhibition and Lead to Accumulation of Branched-Chain Amino Acids
![]()
1Plant Genome Engineering Research Unit, Agrogenomics Research Center, National Institute of Agrobiological Sciences, Tsukuba, Japan; 2Life Science Research Institute, Kumiai Chemical Industry Co. Ltd., Shizuoka, Japan; 3Department of Applied Biological Chemistry, Graduate School of Agricultural and Life Sciences, The University of Tokyo, Tokyo, Japan; 4Biotechnology Research Center, The University of Tokyo, Tokyo, Japan; 5Graduate School of Nanobioscience, Yokohama City University, Yokohama, Japan; 6Kihara Institute for Biological Research, Yokohama City University, Yokohama, Japan.
Email: stoki@affrc.go.jp
Copyright © 2013 Masaki Endo et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received February 28th, 2013; revised March 28th, 2013; accepted April 5th, 2013
Keywords: Rice; Acetolactate Synthase; Herbicide-Resistance; Branched-Chain Amino Acids
ABSTRACT
The branched-chain amino acids (BCAAs) valine, leucine and isoleucine are essential amino acids that are critical for animal growth and development. Animals need to obtain BCAAs from their diet because they cannot synthesize them. Plants are the ultimate source of these amino acids. Acetolactate synthase (ALS) is the first common enzyme in the biosynthesis of BCAAs. The metabolic control of BCAA biosynthesis involves allosteric regulation of ALS by the end-products of the pathway, i.e., valine, leucine and isoleucine. ALS holoenzyme seems to consist of two large catalytic subunits and two small regulatory subunits. In a previous study, using homologous recombination dependent gene targeting we created rice plants in which W548L and S627I mutations were induced into the endogenous gene encoding the ALS catalytic subunit. These two amino acid substitutions conferred hypertolerance to the ALS-inhibiting herbicide bispyripac-sodium. In this study, we revealed that feedback regulation by valine and leucine was reduced by these two amino acid substitutions. Furthermore, in leaves and seeds of ALS mutants with W548L and/or S627I substitution, a 2- to 3-fold increase in BCAAs was detected. Our results suggest that the ALS catalytic subunit is also involved in feedback regulation of ALS, and that judicious modification of the regulatory and catalytic subunits of ALS-coding genes by gene targeting can lead to the efficient accumulation of BCAA in plants.
1. Introduction
Branched-chain amino acids (BCAAs) are essential amino acids that play important roles in protein anabolism and neurotransmitter biosynthesis in animals [1,2]. Because animals lack the ability to synthesize BCAAs, they must acquire them from their diet. Plants are the ultimate source of these essential amino acids for animals. ALS is the first enzyme in the pathway for the biosynthesis of BCAAs [3]; ALS catalyzes the condensation of two pyruvate molecules to form acetolactate (a precursor of valine and leucine), and the condensation of pyruvate and α-ketobutyrate to yield acetohydroxybutyrate (a precursor of isoleucine) (Figure 1). Plant ALS is inhibited by all three BCAAs, with valine and leucine being particularly potent [4].
As the key enzyme in BCAA biosynthesis and a primary target site of action for at least four structurally distinct classes of herbicides (sulfonylureas, imidazolinones, triazolopyrimidine sulfonamides and pyrimidinyl carboxy herbicides; for review, see [5]), ALS has been well characterized in certain organisms. To date, most ALS enzymes that have been characterized have both a catalytic subunit (65 kDa) and a smaller regulatory subunit, which varies in size between 9 and 54 kDa, depending on the species of origin. The plant ALS regulatory subunit has been suggested to be able to stabilize and enhance catalytic subunit [6-8]. The ALS regulatory subunit also mediates end-product inhibition by BCAAs through a complex domain interaction, which in turn

Figure 1. The branched-chain amino acid (BCAA) biosynthetic pathway. ALS catalyzes first step of biosynthesis of BCAAs. ALS is a primary target site of action for at least four structurally distinct classes of herbicides. ALS is under feedback regulation by valine, leucine and isoleucine.
influences the catalytic subunits by an allosteric conformational change [9-13].
A major advantage of ALS-inhibiting herbicide compounds is that they are non-toxic to animals, highly selective, and very potent, thereby requiring only low application rates. Thus, ALS-inhibiting herbicides are an essential part of the multibillion-dollar weed-control market. Because ALS-inhibiting herbicides control a broad spectrum of grass and broadleaf weeds, including weeds that are closely related to the crop itself and some key parasitic weeds, several herbicide-resistant plant mutants have been screened (e.g. [14,15]) and herbicide-resistant mutations are well studied. Because herbicide-resistance and BACC accumulation are favorable traits for cultivation and nutrition, respectively, modification of ALS is currently a hot topic in crop molecular breeding, and detailed analysis of ALS-modified crops is important.
In a previous study, we succeeded in creating herbicide-resistant rice plants in which point mutations—a tryptophan (TGG) to leucine (TTG) change at amino acid 548 (W548L), and a serine (AGT) to isoleucine (ATT) change at amino acid 627 (S627I)—were introduced into the endogenous ALS catalytic subunit using a homologous recombination-dependent gene targeting system [16]. These double mutations in the rice ALS gene represented a novel combination of spontaneous mutations. Although each individual amino acid change in ALS resulted in a phenotype tolerant to the sulfonylurea herbicide bispyripac-sodium, conversion of both amino acids conferred increased tolerance to bispyripac-sodium when expressed in Escherichia coli [17]. To demonstrate the effects of W548L and S627I mutations on BCAA synthesis and feedback regulation by BCAAs in rice, we investigated the activity of the ALS holoenzyme in the presence of BCAAs and measured amino acid content in ALS modified rice plants.
2. Results
2.1. Feedback Regulation Is Impaired by W548L and S627I Double Mutations
To understand the effect of the W548L/S627I double mutation on feedback regulation of ALS by BCAAs, we analyzed ALS enzymatic activity in the presence of valine and leucine. Protein extracts from wild-type and ALS mutants [W548L/S627I (m)], in which W548L and S627I double mutations were introduced into the ALS catalytic subunit [16], as well as extracts of plants overexpressing ALS harboring the double mutation [W548L/ S627I (ox)] were examined for their sensitivities to valine and leucine. The ALS activity of wild-type was inhibited by 65% in the presence of 1 mM valine and leucine, compared with the activity apparent in the absence of these amino acids (Figure 2). In contrast, the inhibition rates of ALS activity in W548L/S627I (m) and W548L/S627I (ox) were 22% and 53%, respectively.
In W548L/S627I (ox) plants, the mutated ALS gene is driven by a strong 35S promoter, thus the transcriptional level of this mutated ALS gene is much higher than that in wild-type or W548L/S627I (m) (approximately 20- fold higher than wild-type, data not shown). Regardless of the fact that the proportion of wild-type ALS transcript may be less than 5% that in W548L/S627I (ox) plants,
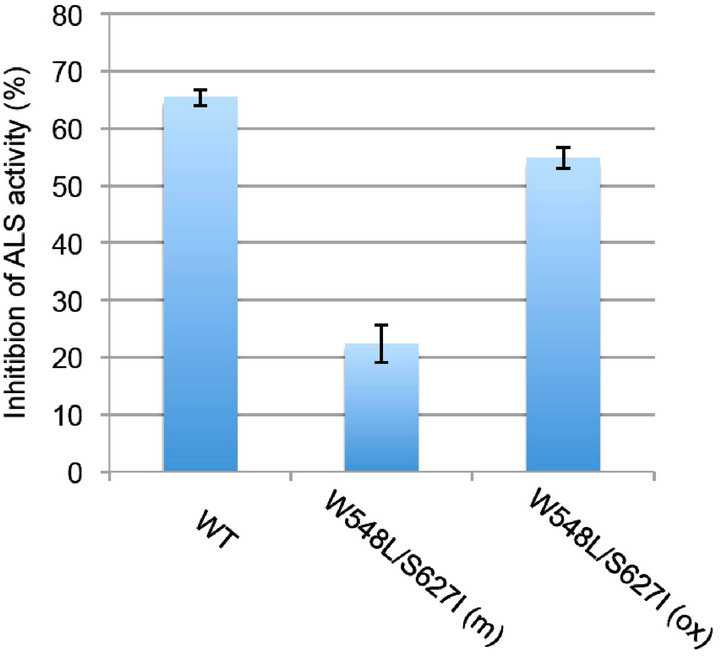
Figure 2. Inhibition of ALS activity by 1 mM valine and leucine. Decreased ALS sensitivity to amino acid feedback is revealed. WT, wild-type; W548L/S627I (m), mutant rice plants with W548L and S627I mutations in endogenous ALS gene; W548L/S627I (ox), rice plants overexpressing ALS gene with W548L and S627I mutations. Data are means ± standard error (N = 2).
the inhibition rate of ALS activity of W548L/S627I (ox) (50%) was close to that of wild-type (65%), while the inhibition rate of W548L/S627I (m) was 22%.
2.2. Increased BCAA Accumulation in ALS Mutated Rice Plants
As the W548L and S627I mutations in the ALS catalytic subunit seem to reduce feedback regulation by valine and leucine, it is possible that ALS mutants containing these mutations may accumulate BCAAs. To test this hypothesis, the free amino acids contents were measured. In addition to wild-type and W548L/S627I (m), we prepared ALS mutants named W548L (m) and S627I (m), in which single mutations (either W548L or S627I) were introduced in the ALS catalytic subunit by tissue-cultureinduced somaclonal mutation. Amino acids were extracted from hydroponic cultured seedlings and threshed mature seeds of wild-type and ALS mutants [W548L/ S627I (m), W548L (m), S627I (m)) and analyzed using CE-MS. Amounts of free amino acids varied greatly according to the type of amino acid in both leaves (Figure 3) and seeds (Figure 4). Although the proportion of BCAAs in total free amino acids was not as high, the overall amounts of BCAAs were increased significantly in all three ALS mutants in both tissues (Figure 5). In the case of herbicide-resistance, each individual amino acid change resulted in a BS-tolerant phenotype, and conversion of both amino acids conferred increased tolerance to BS [17]. However, the amount of BCAAs was almost the same in all 3 ALS mutants, and W548L and S627I double mutations did not show a synergistic effect for the accumulation of BCAAs. Apart from BCAAs, an increase of more than 1.5-fold in phenylalanine, methionine and citrulline was detected in ALS mutant leaves (Figure 3) and levels of asparagine, glutamate, glutamic acid, phyenylalanine, arginine, aspartic acid were increased in ALS mutant seeds (Figure 4).
3. Discussion
Because plants are the main source of BCAAs for animals, it is of great interest to generate plants that produce high levels of BCAAs. Meanwhile, from the viewpoint

Figure 3. Contents of free amino acids in leaves. Free amino acid contents in leaves were measured by CE-MS. WT, wild-type; W548L/S627I (m), mutant rice plants with W548L and S627I mutations in endogenous ALS gene; W548L (m), mutant rice plants with W548L mutation in endogenous ALS gene; S627I (m), mutant rice plants with S627I mutation in endogenous ALS gene. Data are the means and standard errors from three independent biological repeats.

Figure 4. Contents of free amino acids in seeds. Free amino acid contents in seeds were measured by CE-MS. Abbreviations as in Figure 3. Data are the means and standard errors from three independent biological repeats.
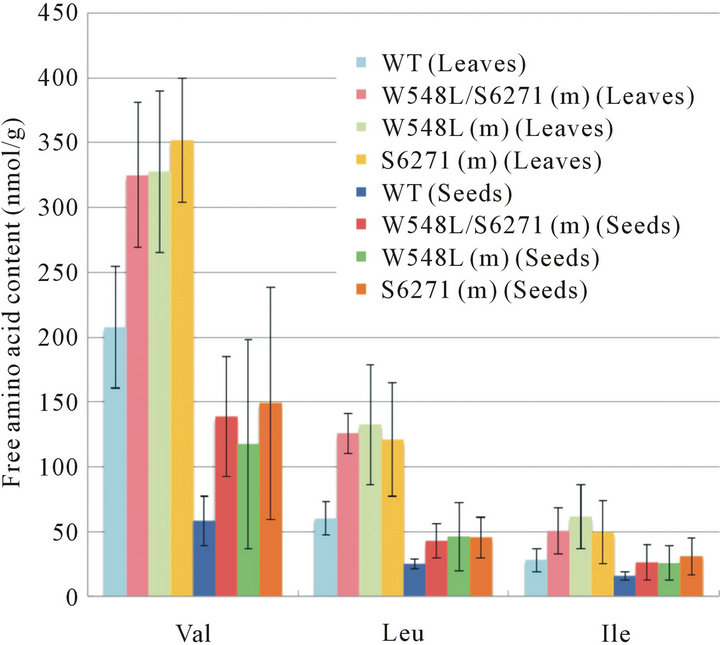
Figure 5. BCAA content in wild-type and ALS mutant rice. Data of BCAA accumulation extracted from Figures 3 and 4.
of developing herbicide-resistant crops and weed control, the effect of mutations on ALS enzymatic activity and fitness must also be investigated thoroughly. In this context, Yu et al. [18] investigated the effect on ALS functionality and plant development of various ALS-inhibiting herbicide-resistance mutations in Lolium rigidum (Ryegrass). They remarked that the P197A, P197R, P197G, P197S and W574L mutations in the ALS catalytic subunit that were detected frequently in weeds had no major impact on ALS functionality. Furthermore, in most cases, herbicide-resistance mutations do not drastically change ALS substrate affinity, but rather alter ALS sensitivity to BCAA feedback inhibition, resulting in accumulation of these amino acids (e.g., A205V in eastern black nightshade (Solanum ptychanthum) [19]; P197H in Prickly Lettuce (Lactuca serriola) [20,21]; P197T in Prickly Lettuce (Lactuca serriola) [22].
In a previous study, we succeeded in creating bispyripac-sodium hyper-resistant rice plants by introducing W548L and S627I mutations into the catalytic subunit of ALS by gene targeting [16]. Because mutant plants in which several herbicide-resistant mutations are induced spontaneously in the endogenous ALS gene appear only rarely, there were no previous reports on the effect of multiple mutations on BCAA feedback regulation in plants. In this study, we revealed that rice plants with W548L and S627I double mutations in the ALS catalytic subunit were not only hyper-resistant to herbicide but also resistant to feedback inhibition. Interestingly, the inhibition rate of ALS activity of W548L/S627I (ox) (50%) by valine and leucine was closer to that of wildtype (65%) than to that of W548L/S627I (m) (22%) (Figure 2). Considering that the main target of feedback inhibition is the ALS regulatory subunit, one hypothesis for explaining this result is that W548L/S627I mutations in the ALS catalytic subunit might change the conformation of the ALS holoenzyme and inhibit BCAA binding to the ALS regulatory subunit. Because wild-type ALS remains in W548L/S627I (ox) rice, holoenzymes including wild-type ALS catalytic subunit may be inhibited by BCAAs. This is in contrast to W548L/S627I (m), in which all ALS holoenzymes are BCAA insensitive. Further studies, including ALS holoenzyme reconstitution experiments and analysis of feedback inhibition in W548L (m) and S627I (m) are needed to prove this hypothesis and determine the contribution of each mutation on feedback regulation.
For BCAA accumulation, direct modification of the ALS regulatory subunit may be more effective. In fact, several Arabidopsis mutants exhibiting enhanced resistance to BCAAs were screened, and the mutations in the ALS regulatory subunit responsible for the phenotype were identified [23]. Because amino-acid resistance is a selectable phenotype, a gene targeting strategy similar to that used to induce herbicide-resistant mutations [16] can be applied to induce amino-acid resistant mutations in the ALS regulatory subunit. Indeed, generation of hightryptophan accumulating rice by introducing precise mutations in OASA2—an α-subunit of anthranilate synthase, a key enzyme of tryptophan biosynthesis—via gene targeting was successful [24]. Structural biology may help predict effective mutations for BCAA accumulation and herbicide-resistance, and advancement of molecular breeding technology including gene targeting may help create novel and valuable crops.
4. Materials and Methods
4.1. Enzymatic Assay of ALS
Extraction of protein and assay of ALS activity were performed as described previously [25] with the following modifications. Proteins including ALS were extracted from the upper part of rice seedlings; shoots were homogenized in 5 volumes of 0.1 M potassium phosphate buffer (pH 7.5) containing 0.5 mM MgCl2, 10% (v/v) glycerol. The homogenates were filtered through one layer of nylon filter gauze and centrifuged at 15,000 g for 20 min. ALS was precipitated from the supernatant with ammonium sulfate at 50 % saturation. After centrifugation, the pellets were dissolved in 0.1 M potassium phosphate buffer (pH 7.5) containing 10% glycerol and 0.5 mM MgCl2. The solution was centrifuged at 15,000 g for 20 min to remove insoluble substances. The supernatant fluids were then desalted on a Sephadex G-25 column equilibrated with the same buffer and stored at −80˚C. All operations were carried out at 0 to 4˚C. Enzyme activity of ALS was assayed at 37˚C in 0.5 ml of assay mixture containing 20 mM potassium phosphate buffer (pH 7.5), 20 mM sodium pyruvate, 0.5 mM thiamine pyrophosphate (TPP), 0.5 mM MgCl2, 10 μM FAD and 1 mM valine and leucine. Assays were initiated by adding 100 μl of crude enzyme solution and terminated after 30 min by the addition of 50 μl 6 N H2SO4. The amount of acetolactate produced by the enzyme reaction was determined as described previously [26] with the following modifications. The acidified reaction mixtures were heated for 10 min at 60˚C, after which 0.5 ml of 0.5% (w/v) creatine and 0.5 ml of 2-naphthol solution (5% in 2.5 N NaOH), both freshly prepared, were added to the solution and then heated for 10 min at 60˚C. The absorbance at 525 nm was then determined with a double beam spectrophotometer.
4.2. Quantification of Amino Acids
Samples were ground into a fine powder in liquid N2 and further homogenized with ice-cold 50% (v/v) methanol containing PIPES and Met sulfone as internal standards. Homogenates were then centrifuged at 13,000 g for 5 min, the supernatant was filtered through a Millipore 5-kDa cutoff filter (Amicon), and the filtrates were used for analysis. Organic acids, nucleotides, and amino acids were separated and quantified by CE-MS (Agilent Technologies), as described previously [27,28]. Amino acids were separated in an uncoated fused-silica capillary using 1 M formic acid as a run buffer. Accuracy was verified with known concentrations of reference standard compounds.
5. Acknowledgements
We thank A. Nagashii, F. Suzuki, M. Amagai for their technical help. This work was supported by a grant from the Ministry of Agriculture, Forestry and Fisheries of Japan (Genomics for Agricultural Innovation, GMC- 0001). This work was also supported by a PROBRAIN (Program for Promotion of Basic Research Activities for Innovative Biosciences) grant to S. T. from the BioOriented Technology Research Advancement Institution (BRAIN) of Japan. A part of this study was also supported financially by the Budget for Nuclear Research of the Ministry of Education, Culture, Sports, Science and Technology, based on screening and counseling by the Atomic Energy Commission.
REFERENCES
- J. D. Fernstrom, “Branched-Chain Amino Acids and Brain Function,” The Journal of Nutrition, Vol. 135, No. 6, 2005, pp. 1539-1546.
- K. S. Nair and K. R. Short, “Hormonal and Signaling Role of Branched-Chain Amino Acids,” Journal of Nutrition, No. 135, Vol. 6, 2005, pp. 1547S-1552S.
- D. Chipman, Z. Barak and J. V. Schloss, “Biosynthesis of 2-Aceto-2-Hydroxy Acid; Acetolactate Synthases and Acetohydroxyacid Synthases,” Biochimica et Biophysica Acta (BBA)—Protein Structure and Molecular Enzymology, Vol. 1385, No. 2, 1998, pp. 401-419. doi:10.1016/S0167-4838(98)00083-1
- B. J. Miflin and P. R. Cave, “The Control of Leucine, Isoleucine and Valine Biosynthesis in a Range of Higher Plants,” Journal of Experimental Botany, Vol. 23, No. 75, 1972, pp. 511-516. doi:10.1093/jxb/23.2.511
- C. A. Corbett and F. J. Tardif, “Detection of Resistance to Acetolactate Synthase Inhibitors in Weeds with Emphasis on DNA-Based Techniques: A Review,” Pest Management Science, Vol. 7, No. 62, 2006, pp. 584-597. doi:10.1002/ps.1219
- M. Vyazmensky, C. Sella, Z. Barak and D. M. Chipman, “Isolation and Characterization of Subunits of Acetohydroxy Acid Synthase Isozyme III and Reconstitution of the Holoenzyme,” Biochemistry, Vol. 35, No. 32, 1996, pp. 10339-10346. doi:10.1021/bi9605604
- D. M. Chipman, R. G. Duggleby and K. Tittmann, “Mechanisms of Acetohydroxyacid Synthases,” Current Opinion in Chemical Biology, Vol. 9, No. 5, 2005, pp. 475- 481. doi:10.1016/j.cbpa.2005.07.002
- R. G. Duggleby, J. A. McCourt and L. W. Guddat, “Structure and Mechanism of Inhibition of Plant Acetohydroxyacid Synthase,” Plant Physiology and Biochemistry, Vol. 46, No. 3, 2008, pp. 309-324. doi:10.1016/j.plaphy.2007.12.004
- Y. T. Lee and R. G. Duggleby, “Identification of the Regulatory Subunit of Arabidopsis thaliana Acetohydroxyacid Synthase and Reconstitution with Its Catalytic Subunit,” Biochemistry, Vol. 40, No. 23, 2001, pp. 6836- 6844. doi:10.1021/bi002775q
- Y. T. Lee and R. G. Duggleby, “Regulatory Interactions in Arabidopsis thaliana Acetohydroxyacid Synthase,” FEBS Letter, Vol. 512, No. 1-3, 2002, pp. 80-184. doi:10.1016/S0014-5793(02)02253-6
- S. Mendel, T. Elkayam, C. Sella, V. Vinogradov, M. Vyazmensky, D. M. Chipman and Z. Barak, “Acetohydroxyacid Synthase: A Proposed Structure for Regulatory Subunits Supported by Evidence from Mutagenesis,” Journal of Molecular Biology, Vol. 307, No. 1, 2001, pp. 465-477. doi:10.1006/jmbi.2000.4413
- S. Mendel, M. Vinogradov, M. Vyazmensky, D. M. Chipman and Z. Barak, “The N-Terminal Domain of the Regulatory Subunit Is Sufficient for Complete Activation of Acetohydroxyacid Synthase III from Escherichia coli,” Journal of Molecular Biology, Vol. 325, No. 2, 2003, pp. 275-284. doi:10.1016/S0022-2836(02)01142-7
- A. Kaplun, M. Vyazmensky, Y. Zherdev, I. Belenky, A. Slutzker, S. Mendel, Z. Barak, D.M. Chipman and B. Shaanan, “Structure of the Regulatory Subunit of Acetohydroxyacid Synthase Isozyme III from Escherichia coli,” Journal of molecular Biology, Vol. 357, No. 3, 2006, pp. 951-963. doi:10.1016/j.jmb.2005.12.077
- G. W. Haughn and C. R. Somerville, “A Mutation Causing Imidazolinone Resistance Maps to the Csr1 Locus of Arabidopsis thaliana,” Plant Physiology, Vol. 92, No. 4, 1986, pp. 1081-1085. doi:10.1104/pp.92.4.1081
- K. Sathasivan, G. W. Haughn and N. Murai, “Molecular Basis of Midazolinone Herbicide-Resistance in Arabidopsis thaliana var Columbia,” Plant Physiology, Vol. 97, No. 3, 1991, pp. 1044-1050. doi:10.1104/pp.97.3.1044
- M. Endo, K. Osakabe, K. Ono, H. Handa, T. Shimizu and S. Toki, “Molecular Breeding of a Novel Herbicide-Tolerant Rice by Gene Targeting,” The Plant Journal, Vol. 52, No. 1, 2007, pp. 157-166. doi:10.1111/j.1365-313X.2007.03230.x
- T. Shimizu, K. Kaku, K. Kawai, T. Miyazawa and Y. Tanaka, “Molecular Characterization of Acetolactate Synthase in Resistant Weeds and Crops,” In: J. M. Clark and H. Ohkawa, Eds., ACS Symposium Series 899: Environmental Fate and Safety Management of Agrochemicals, American Chemical Society, Washington DC, 2005, pp. 255-271.
- Q. Yu, H. Han, M. M. Vila-Aiub and S. B. Powles, “AHAS Herbicide-Resistance Endowing Mutations: Effect on AHAS Functionality and Plant Growth,” Journal of Experimental Botany, Vol. 61, No. 14, 2010, pp. 3925- 3934. doi:10.1093/jxb/erq205
- J. Ashigh and F. J. Tardif, “An Ala205Val Substitution in Acetohydroxyacid Synthase of Eastern Black Nightshade (Solanum ptychanthum) Reduces Sensitivity to Herbicides and Feedback Inhibition,” Weed Science, Vol. 55, No. 6, 2007, pp. 558-565. doi:10.1614/WS-07-054.1
- C. V. Eberlein, M. J. Guttieri, P. H. Berger, J. K. Fellman, C. A. Mallory-Smith, D. C. Thill, R. J. Baerg and W. R. Belknap, ”Physiological Consequence of Mutation for ALS-Inhibitor Resistance,” Weed Science, Vol. 47, No. 4, 1999, pp. 383-392.
- C. V. Eberlein, M. J. Guttieri, C. A. Mallory-Smith, D. C. Thill and R. J. Baerg, “Altered Acetolactate Synthase Activity in ALS-Inhibitor Resistant Prickly Lettuce (Lactuca serriola),” Weed Science, Vol. 45, No. 2, 1997, pp. 212- 217.
- C. Preston, L. M. Stone, M. A. Rieger and J. Baker, “Multiple Effects of a Naturally Occurring Proline to Threonine Substitution within Acetolactate Synthase in Two Herbicide-Resistant Populations of Lactuca serriola,” Pesticide Biochemistry and Physiology, Vol. 84, No. 3, 2006, pp. 227-235. doi:10.1016/j.pestbp.2005.07.007
- H. Chen, K. Saksa, F. Zhao, J. Qiu and L. Xiong “Genetic Analysis of Pathway Regulation for Enhancing BranchedChain Amino Acid Biosynthesis in Plants,” The Plant Journal, Vol. 63, No. 4, 2010, pp. 573-583. doi:10.1111/j.1365-313X.2010.04261.x
- H. Saika, A. Oikawa, F. Matsuda, H. Onodera, K. Saito and S. Toki, “Application of Gene Targeting to Designed Mutation Breeding of High-Tryptophan Rice,” Plant Physiology, Vol. 156, No. 3, pp. 1269-1277. doi:10.1104/pp.111.175778
- T. Shimizu, I. Nakayama, T. Nakao and H. Abe, “Partial Purification and Properties of Acetolactate Synthase and Etiolated Pea Seedlings,” Journal of Pesticide Science, Vol. 19, No. 3, 1994, pp. 187-196. doi:10.1584/jpestics.19.3_187
- T. B. Ray, “Site of Action of Chlorsulfuron; Inhibition of Valine and Isoleucine Biosynthesis in Plants,” Plant Physiology, Vol. 75, No. 3, 1984, pp. 827-831. doi:10.1104/pp.75.3.827
- H. Takahashi, M. Hayashi, F. Goto, S. Sato, S. Soga, T. Nishioka, M. Tomita, M. Kawai-Yamada and H. Uchimiya, “Evaluation of Metabolic Alteration in Transgenic Rice Overexpressing Dihydroflavonol-4-Reductase,” Annual of Botany, Vol. 98, No. 4, 2006, pp. 819-825. doi:10.1093/aob/mcl162
- H. Takahashi, A. Watanabe, A. Tanaka, S.N. Hashida, M. Kawai-Yamada, K. Sonoike and H. Uchimiya, “Chloroplast NAD Kinase Is Essential for Energy Transduction through the Xanthophyll Cycle in Photosynthesis,” Plant Cell Physiology, Vol. 47, No. 12, 2006, pp. 1678-1682. doi:10.1093/pcp/pcl029

