Advances in Microbiology
Vol.4 No.10(2014), Article
ID:48851,12
pages
DOI:10.4236/aim.2014.410070
Effect of Heavy Metals on the Growth of Bacteria Isolated from Sewage Sludge Compost Tea
María Vela-Cano1,2*, Antonio Castellano-Hinojosa1, Antonia Fernández Vivas2, María Victoria Martínez Toledo1,2
1Institute of Water Research, University of Granada, Granada, Spain
2Department of Microbiology, University of Granada, Campus de Fuentenueva s/n, Granada, Spain
Email: *mvc@ugr.es, anthonych15@correo.ugr.es, mvmt@ugr.es, fvivas@ugr.es
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 30 June 2014; revised 26 July 2014; accepted 12 August 2014
ABSTRACT
The amount of urban sewage sludge is increasing daily. For this reason, it is necessary to have an environmentally friendly use for this residue. Land applications in olive grove soil, poor quality soil, as a compost or a compost tea could be considered as one of the best options, because it has a high content of organic matter. However, the presence of heavy metals in some sewage sludge could be an environmental problem. In this sense, the aim of this work is to check the tolerance of the culturable microbiota present in sewage sludge compost tea to four heavy metals, Cu, Cd, Pb and Zn. Among microbiota isolated, eight strains showed high resistance to Pb, Zn, Cu and Cd, and one microbe (Rhodococcus sp.) showed a special tolerance to every heavy metal. The strains were divided into six genrera: Rhodococcus, Virgibacillus, Leifsoni, Achromobacter, Cupriavidus and Oceanobacillus sp. Finally, Rhodococcus sp. strain 3 was able to remove different amounts of heavy metals from the culture media with intracellular and surface accumulation of Cu, Zn, Pb and Cd. The application of this bacterial strain in bioremediation processes is discussed.
Keywords:Wastewater Sludge, Compost Tea, Minimum Inhibitory Concentration, Heavy-Metal Removal, Rhodococcus sp.

1. Introduction
The olive oil grove surface in Andalusia is approximately 1,500,000 hectares, that is, 60% of the Spanish olive oil surface and 30% of Europe’s land area [1] . Previously, the olive cultivation system was environmentally sustainable, because there were few olive trees, and the productivity of those trees was also low [2] . However, nowadays, olive cultivation has become intensive, which has caused a loss of biodiversity [3] [4] . This fact, combined with the poor quality that olive grove soil traditionally has, results in a serious problem of soil degradation [5] . In this situation, it is essential to improve the bad quality of olive grove soil by adding organic amendments [6] .
On the other hand, new organic matter resources are being found [7] . The use of compost and compost tea from sewage sludge as a fertiliser is becoming popular practice [8] , because sewage sludge is a cheap source of a large number of nutrients such as N or P [9] [10] . Furthermore, various studies have shown how this organic matter improves the biological and physiochemical properties of soils [11] .
The amount of sewage sludge produced is higher each year; in order to obey the urban wastewater treatment directive [12] , a wastewater treatment plant must be installed in localities where the population is over 2000. For this reason, the production of this residue in Spain exceeded 1,000,000 tons in 2010. In addition, by having an agricultural use for sewage sludge, other environmentally unfriendly treatments such us incineration or dumping can be avoided [13] .
However, land applications of these products present some problems, owing to the presence of some toxic substances such as heavy metals [14] . Heavy metals are not biodegradable and can be harmful to health, so their presence in soils and, consequently, in plants and animals involve serious risks for the environment and human health [15] .
The toxicity and mobility of heavy metals depend on different physicochemical and environmental factors; it is necessary to consider the concentration, chemical form and metal properties as well as the pH value, type and organic-matter content of the soil [16] [17] .
To date, numerous physicochemical methods have been used to remove heavy metals from the environment. Chemical precipitation and ion exchange are the most used ones, but oxidation/reduction, filtration, electrochemical treatment, reverse osmosis, membrane technology and evaporation recovery are also common [18] . However, if the concentration is too low, these techniques have an excessive cost and are not efficient [19] . To solve this problem, biological processes have been carried out. Biosorption is the process in which some organisms, such as bacteria, fungi or algae, are used to remove environmental contaminants. Some microorganisms are able to remove heavy metals and others pollutants from the environment. In this context, it is widely known that microbes can tolerate and survive in extreme conditions. They have been found in areas where the temperature, salinity or pH values do not allow the existence of life [20] -[22] . When microorganisms are under stress conditions, such us living in a polluted environment with heavy metals, they develop survival mechanisms. This means that microorganisms may be used as a cheap and efficient bioremediation tool [22] . The processes implicated in heavy-metal detoxification include absorption, adsorption, ion exchange, surface complexation and precipitation [23] .
The first purpose of this work was to study the interaction between the microbes present in a sewage sludge compost tea with Pb, Zn, Cd and Cu. To reach this goal, the following analyses were carried out: 1) evaluation of the microorganism’s tolerance to Pb, Zn, Cd and Cu; 2) determination of the heavy metal minimum inhibitory concentration (MIC) for each microorganism and 3) selection of the most resistant strains, which could be used in bioremediation of heavy metals and their identification using molecular methods. In addition, the most resistant strain was selected in order to study its behaviour in the presence of these heavy metals by locating metals in the cell using high-resolution transmission electron microscopy (HRTEM) and determining the percentage sorption of each metal by inductively coupled mass spectrometry (ICP-MS) analysis.
2. Material and Methods
2.1. Culture Media
Two different culture media were used in this study: Tripticase soy agar (TSA) medium (Oxoid, Basingstoke, Hampshire, England) [24] and LPM medium. LPM medium was used as both a solid and a liquid medium; it was prepared as follows: 100 mL of salt stock solution (Table 1), 100 mL of peptone solution (10 g peptone per 1 L distilled water), 100 mL of CaCl2 (10 mM) and 5 mL of glycerol were added per litre of distilled water. The LPM media were then sterilised in an autoclave at 121˚C for 20 min. Solid media contained 16 g per litre of agar (DIFCO). After sterilisation, the LPM media were added (at 40˚C) to 0.4 mL thiamine solution (50 mg/mL) and an appropriate amount of metal solution. Thiamine and metal solutions were previously sterilised by filtration (0.45 μm) before addition.
2.2. Production of Compost Tea
The compost tea was made with compost from sewage sludge obtained from the composting plant Biomasas del Guadalquivir S.A., located in Santa Fe (Granada, Spain). It was prepared by mixing 100 L of distilled water with 1.5 kg of homogenised compost in a 120 L bioreactor (Figure 1). The mixture was continuously aerated using an aeration pump, maintaining aerobic conditions in the system throughout the process. The temperature was maintained at 23˚C using a thermostat. The time used to produce the compost tea was 5 days, because that is the time required for the physicochemical and biological properties to become stable.
2.3. Selection and Isolation of Cultivable Bacteria
Samples of compost tea were collected in sterilized glass bottles at 24 h intervals during the production process. The isolation of microorganisms was carried out using a serial dilution technique. Aliquots of 100 μL of different dilutions were spread onto plates of TSA medium. The plates were incubated at 28˚C for 5 days under aerobic conditions. Developed colonies were picked and isolated following morphological criteria. Purified isolates were obtained by repeatedly streaking colonies on a TSA medium and observing them using light microscopy. The identification and classification of the colony morphotypes were achieved using five parameters: colony size, form, colour, texture and margin.
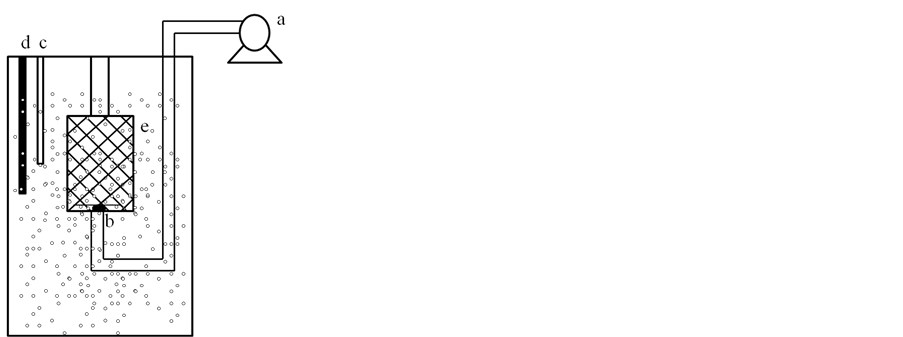
Figure 1. Diagram of the bench scale plant used for the production of compost tea. (a) Aeration pump; (b) Air diffuser; (c) Oxygen electrode (EcoScan DO6); (d) Termostat (Aqua Medic); (e) Membrane (pore size 1.5 mm) containing compost from sewage sludge.
Table 1. Chemical compositions of stock salt solutions used to prepare the LPM media.
The pH value of the final solutions was adjusted 7.0 by using 0.1 M HCl.
2.4. Evaluation of the Tolerance of the Microorganisms and Determination of Their Minimum Inhibitory Concentration (MIC)
To check the microorganism’s tolerance to Pb, Cd, Cu and Zn and to determinate their MIC, LPM medium was prepared and amended with increasing concentrations of heavy metal (0.25 - 16 mM for Cu, Zn and Pb and 0.015 - 4 mM for Cd). The metals were added as Pb(NO3)2·3H2O, Cu(NO3)2·3H2O, Cd(NO3)2·4H2O and N2O6Zn·6H2O. Each heavy-metal-doped plate was subdivided into eight equal sectors and 5 μL of a 24 h inoculum of selected strains was produced. The same method was carried out with control plates (plates without metal). Each sample was made in triplicate. Finally, the plates were incubated at 28˚C for 14 days to observe the growth of bacteria on the spotted area. The MIC was defined as the lowest concentration of each of the metals that inhibited visible growth of the test isolates.
2.5. Identification of Microorganisms
DNA was extracted from the solid cultures (one colony per strain) using Qiagen columns (QIAamp DNA mini kit; Qiagen, Hilden, Germany), according to the manufacturer’s protocol. After extraction, the polymerase chain reaction (PCR) was performed for the amplification of the 16S-rDNA sequence (primer sequence are in Table 2). 1 µL (2 - 5 ng) of extracted DNA was used as a template for PCR, using universal primers fD1 and rD1 to amplify the 16S-rRNA gene in almost its full length [25] . Conditions for each of the PCR reactions were kept as previously described [26] . DNA sequences were analysed using bio-computing tools provided online by the European Bioinformatics Institute (http://www.ebi.ac.uk). The BLASTn program [27] was used to analyse the sequence similarity. The ClustalX v. 1.8 software [28] was used to align the DNA sequences. Phylogenetic and molecular evolutionary analyses were conducted using MEGA version 3.1 [29] . Gaps were excluded from the analysis.
2.6. Removal Experiments
Batch studies were conducted using 250 mL Erlenmeyer flasks that contained 50 mL LPM medium with heavymetal concentrations of 4 mM for Cd and 8 mM for Zn, Cu and Pb. All flasks were inoculated with 1 mL of a cell suspension of the selected strain (strain 3), which was previously grown in TSA medium at 28˚C for 24 h under aerobic conditions.
Batch experiments were performed at 28˚C and 100 rpm in an Incubator Shaker (HWY-211). In order to know the amount of heavy metal retained by the cells, the concentration of each heavy metal was determined by ICP-MS (NEXION 300D) after 24 h of incubation. Control media without heavy metals were also included for comparison.
2.7. Transmission Electron Microscopy (TEM)
Samples of 24 h from the batch culture were taken and fixed with 2.0% glutaraldehyde and 1.0% formaldehyde at pH 7.4 (sodium cacodylate buffer, 0.05 M) and 4˚C for 4 h. Then, the samples were washed and post-fixed in OsO4 before being dehydrated in an ethanol gradient. Samples were embedded in Epon resin and polymerised. Ultra-thin sections (70 nm) were prepared with an Ultracut E ultramicrotome and stained with uranyl acetate and Pb. Samples originally doped with Pb were only stained with uranyl.
3. Results and Discussion
3.1. Physicochemical and Biological Characterisation of Compost Tea
The physicochemical properties (including heavy-metal concentrations) and microbiological characteristics of
the compost tea are shown in Table3 In our case, the concentration of heavy metals in the compost tea was relatively low. This was probably a consequence of the low level of industrial activity in the geographic area where the sewage sludge was generated. Moreover, it was evident that, although the compost tea showed a significant microbial activity (5.3 × 106 CFU/mL), the concentration of nutrients such as N, P and K was also low.
3.2. Molecular Identification of Heavy-Metal-Tolerant Bacteria
30 different morphological colony types were isolated from the sewage sludge compost tea samples. Among those isolates, eight colony types showed resistance to Pb, Zn, Cu and Cd, and one of them showed a special tolerance to every heavy metal (Table 4). The taxonomic affiliation of the selected strains, based on partial sequencing of the 16S-rRNA gene (V1 to V3 hyper-variable regions, ca. 650 nt), are shown in Table4 The
aEC = electrical conductivity, OM = Organic matter. bConcentration of heavy metals in compost tea (μg/L).
Table 4. Identity and PCR product length for target sequences of microbial isolates investigated in this study.
strains were divided in six genera of Gram-positive and Gram-negative bacteria. Sequence comparison with databases demonstrated the affiliation of strain 3 to Rhodococcus sp. (100% identity), strain 4 to Virgibacillus sp. (97% identity), strain 14 to Leifsonia shinshuensis (99% identity), strain 18 to Achromobacter xylosoxidans (99% identity), strain 24 to Cupriavidus sp. (99% identity), strain 25 to Achromobacter sp. (99% identity), strain 28 to Achromobacter sp. (99% identity) and strain 29 Oceanobacillus sp. (100% identity).
The phylogenetic tree (Figure 2) shows that most of the identified populations were evolutionarily related to beta-Proteobacteria (four strains). Firmicutes accounted for two strains and were related to Bacilli. Phylogenetic alpha-Proteobacteria and Actinobacteria were represented by only one strain each. According to our results, Proteobacteria were identified as the dominant group of bacteria with tolerance to heavy metals in the compost tea. Proteobacteria are widespread components of the bacterial community in compost, where members of betaand alpha-Proteobacteria are responsible for the processes of biodegradation of organic matter during the composting process. Our data also identified Firmicute members as prevalent populations in the compost tea, with the capacity to resist high concentrations of heavy metals. Two identified Bacilli sequences were phylogenetically close to Bacillales (Figure 2). They are commonly present in compost [30] and these finding are consistent with the well-known ability to Bacillales to hydrolyse macromolecules.
3.3. MIC Values
The MICs of the four heavy metals assayed for the 30 strains of isolates from compost tea are shown in Table5 The experiments were conducted to determine the precise concentrations of heavy metals that inhibit visible growth of the tested strains. The tolerance of the tested strains was observed to be in order of Cu > Pb > Zn > Cd. Similar results were reported by Çolak et al. [31] using several strains of Bacillus.
Our results showed that Rhodococcus strain 3 was the microorganism with the highest tolerance to the different heavy metals assayed. In this context, strains of Rhodococcus have been reported [32] with the ability to catabolise a wide range of harmful environmental pollutants, such as toluene, naphthalene, herbicides and polychlorinated biphenyls. Moreover, this microorganism has been reported to have an increased resistance to different heavy metals such as Cd and Cr, suggesting that Rhodococcus microorganisms may be useful for the bioremediation of sites contaminated with high concentrations of them [33] [34] .
However, we cannot forget that all our results have been obtained in artificial culture media and their translatation to real condition must be considered with care. Thus, future work must be done to confirm these preliminary conclusions.
In our study, Cd was the heavy metal that showed the greatest toxicity. In this sense, most of the strains were unable to grow at Cd concentrations higher than 4 mM. These results were similar to other studies carried out
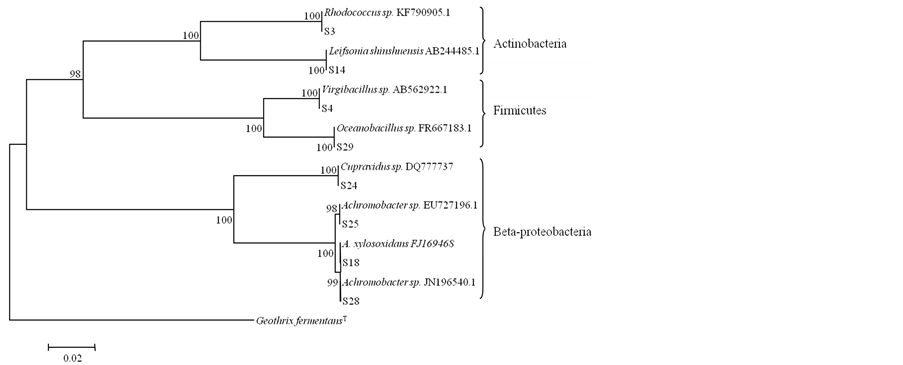

Figure 2. Phylogenetic tree showing the taxonomic location of each strain based on 16S rRNA sequences. The tree was obtained using the neighborjoining method. Geothrix fermentans was used as rooting out group. The significance of each branch is indicated by a bootstrap values calculated for 1000 subsets. Bar, 2 substitutions per 100 n.
Table 5. Tolerance of the bacterial strains isolated from compost tea to heavy metals expressed as Minimal Inhibitory Concentration (MIC).
previously [35] [36] . Thus, Karakagh et al. [36] isolated three Cd-resistant strains from an agricultural soil in Iran, Bacillus sp., Actinomyces sp. and Streptomyces sp., whose MICs were fixed at 3.5, 4.0 and 4.0 mM, respectively. Similarly, Malik and Jaiswal [37] reported that different Pseudomonas strains were unable to tolerate Cd concentrations higher than 1.6 mM. However, Rhodococcus strain 3 showed an unusual ability to resist high concentrations of Cd (Table 4), suggesting that this microorganism can grow in environments polluted with high concentrations of this heavy metal. Obviously, this biological property could be considered of importance for future biotechnological applications, particularly in bioremediation processes.
The tolerance of the bacterial strains assayed in the present study to the Zn was similar to that detected in the case of Cd. Thus, the MIC values were between 2 - 4 and 4 - 8 mM for the majority (85%) of isolations. Similar tolerances have been reported in other bacterial strains (Klebsiella ornithinolytica, Klebsiella oxytoca, Pseudomonas veronii and Ralstonia taiwanensis) isolated from urban wastewaters [19] , suggesting that the resistance to the Zn could be a biological capacity that is relatively widespread in the microbial world.
Rhodococcus strain 3 was the microorganism that showed increased resistance to high Zn concentrations (Table 4). In this context, the biosorption abilities of Rhodococcus strains for Cd and Zn removal have previously been tested [38] , suggesting that these microorganisms are a potential engineering biosorbent for environmentally and extractively sustainable applications. Obviously, more experimental work is needed to confirm the application of strain 3 in bioremediation processes.
Almost 50% of the selected strains showed an important tolerance to Pb, with MICs in the range between 8 and 16 mM (Table 4), suggesting that most of the microbiota present in the compost tea can tolerate high concentrations of Pb. Other authors [22] [31] have also reported different microbial strains, identified as members of the genus Bacillus, Enterobacter and Klebsiella, with tolerance to Pb. However, the reported strains always showed lower MIC values than those observed in our study. This statement was especially evident in the case of Rhodococcus strain 3, because this microorganism showed a MIC value greater than 16 mM, confirming the high resistance of this bacterial strain to different heavy metals. Moreover, as indicated above for Cd and Zn, biosorption of Pb ions using Rhodococcus strains has been investigated in detail [39] , suggesting that this biological activity is a spontaneous process in nature and that there is a potential application for these microorganisms in bioremediation technologies.
Cu was the best-tolerated heavy metal (Table 4). Thus, with the exception of strains 1 and 23, all strains tested showed resistance to Cu at concentrations up to 4 mM, and many of them up to 8 mM. This suggests that the microbiota present in the compost tea can grow in the presence of high concentrations of Cu without evident modification in their biological activity. A similar suggestion was indicated by Vullo et al. [19] , although these authors reported lower MIC values.
Rhodococcus strain 3 was also resistant to the presence of Cu (MIC value up to 8 mM). It is evident that this strain presented the highest resistance ability with respect to the other strains studied under our experimental conditions. As previously indicate [38] [39] , species of the genus Rhodococcus have the ability to tolerate different heavy metals, but in our study, we found, for the first time, a Rhodococcus strain with the ability to grow in the presence of high concentrations of Cd, Zn, Pb and Cu.
3.4. Removal Studies
The results obtained from the MIC analyses revealed that Rhodococcus sp. (strain 3) was the most tolerant bacteria to Cd, Zn, Pb and Cu. For this reason, the biosorption capacities of Rhodococcus sp. in culture media amended with Zn, Pb and Cu at a concentration of 8 mM, or Cd at a concentration of 4 mM, were studied. It is known that the biosorption depends largely on the pH value; consequently, this parameter was carefully controlled throughout the experiments.
Figure 3 shows that Rhodococcus sp. strain 3 was able to remove different amounts of heavy metals. The maximum removal occurred with Pb, for which a reduction higher than 63% was observed. Cu and Zn were also removed by as much as 43.35% and 27.95%, respectively. While only 12% of Cd was removed. These percentages are comparable with results previously reported by others researchers [18] [38] [39] . However, for our study, it should be noted that the biosorption of the four heavy metals was produced by the same bacterial strain, whereas in the other cases, different microorganisms were utilised for each heavy metal. From our point of view, this increases the significance of Rhodococcus sp. (strain 3) that was used in our experiments.
In order to fully understand the interaction between Rhodococcus strain 3 and heavy metals, TEM analysis was performed. Figure 4 shows images of living cells of Rhodococcus sp. (strain 3) grown in the presence of Cu (A), Cd (B), Zn (C) and Pb (D). The results suggest that Cu was mainly deposited on the cell surface structures, although small amounts of Cu were also detected inside the cells. A similar result was observed in culture media containing Cd (Figure 4(B)), although, in this case, more metal could be detected inside the cells. Finally, TEM
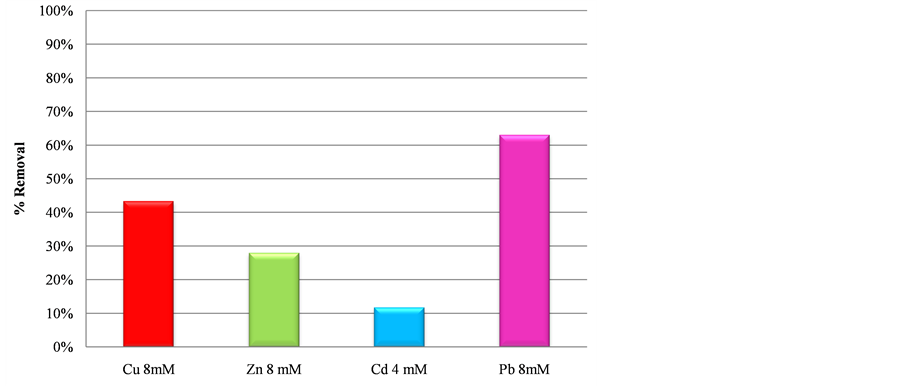
Figure 3. Removal capacity of heavy metals by Rhodococcus sp.
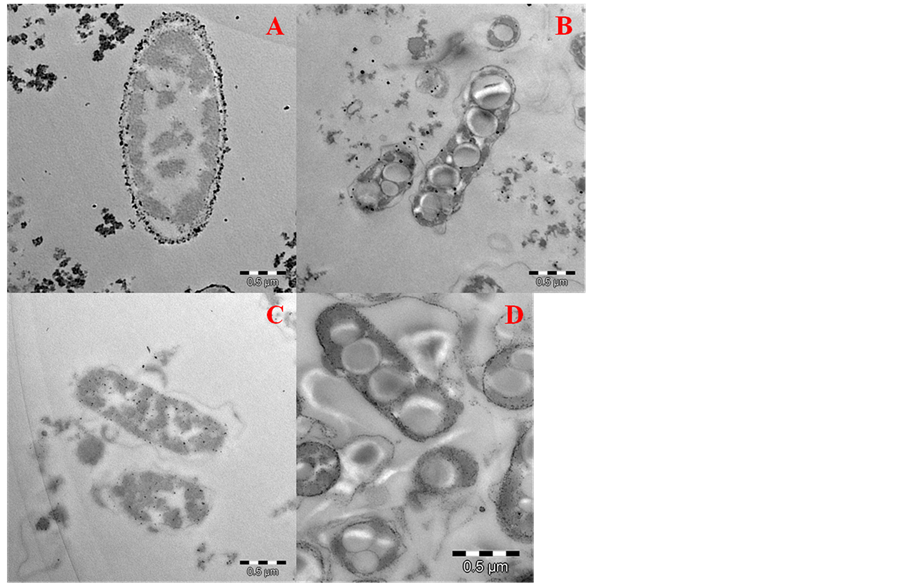
Figure 4. Transmission Electron Microscopy (TEM) of Rhodococcus sp. strain 3, growth in the presence of 8 mM of copper (A); 4 mM of cadmium (B); 8 mM of zinc (C); and 8 mM of lead (D).
analysis showed that Pb and Zn were mainly deposited inside cells, suggesting the bioaccumulation of these heavy metals.
4. Conclusion
Fundamentally, biosorption of heavy metals by bacterial cells is based on non-enzymatic processes such as adsorption [40] . The metal uptake by microbial cells may be an active or passive process or it could exhibit both active and passive processes, depending on the microbial strain [41] [42] . However, bioaccumulation is an active process dependent upon the metabolic energy of the microorganisms. In other words, bioaccumulation can be considered as an energy-dependent heavy-metal transport system. In our study, TEM analysis of Rhodocuccus sp. (strain 3) demonstrated intracellular and cell surface accumulation of Cu, Zn, Pb and Cd, suggesting the active bioaccumulation of these heavy metals. Therefore, the metal resistance of Rhodococcus sp. strain 3 could be exploited in bioremediation, biosorption and bioaccumulation processes. In this sense, complementary experiments are in progress.
References
- BOJA (Boletín Oficial de la Junta de Andalucía) (2011) Ley 5/2011, de 6 de octubre, del olivar de Andalucía, 205, 6-13.
- Duarte, F., Jones, N. and Fleskens, L. (2007) Traditional Olive Orchards on Sloping Land: Sustainability or Abandonment? Journal of Environmental Management, 89, 86-98.
http://dx.doi.org/10.1016/j.jenvman.2007.05.024 - de Graaff, J., Duran, V.H., Jones, N. and Fleskens, L. (2008) Olive Production Systems on Loping Land: Prospects and Scenarios. Journal of Environmental Management, 89, 129-139.
http://dx.doi.org/10.1016/j.jenvman.2007.04.024 - Areal, F.J. and Riesgo, L. (2013) Farmers’ Views on the Future of Olive Farming in Andalusia, Spain. Land Use Policy, 36, 543-553. http://dx.doi.org/10.1016/j.landusepol.2013.10.005
- Milgroom, J., Soriano, M.A., Garrido, J.M., Gómez, J.A. and Fereres, E. (2005) The Influence of a Shift from Conventional to Organic Olive Farming on Soil Management and Erosion Risk in Southern Spain. Renewable Agriculture and Food Systems, 22, 1-10. http://dx.doi.org/10.1017/S1742170507001500
- Serramiá, N., Sánchez-Monedero, M.A., Roig, A., Contin, M. and de Nobili, M. (2013) Changes in Soil Humic Pools after Soil Application of Two-Phase Olive Mill Waste Compost. Geoderma, 192, 21-30. http://dx.doi.org/10.1016/j.geoderma.2012.08.032
- Fernández, J.M., Senesi, N., Plaza, C., Brunetti, G. and Polo, A. (2009) Effects of Composted and Thermally Dried Sewage Sludges on Soil and Soil Humic Acid Properties. Pedosphere, 193, 281-291. http://dx.doi.org/10.1016/S1002-0160(09)60119-3
- Ingelmo, F., Molina, M.J., Soriano, M.D., Gallardo, A. and Lapena, L. (2012) Influence of Organic Matter Transformations on the Bioavailability of Heavy Metals in a Sludge Based Compost. Journal of Environmental Management, 95, 104-109. http://dx.doi.org/10.1016/j.jenvman.2011.04.015
- He, M., Tian, G. and Liang, X. (2009) Phytotoxicity and Speciation of Copper, Zinc and Lead during the Aerobic Composting of Sewage Sludge. Journal of Hazardous Materials, 163, 671-677.
http://dx.doi.org/10.1016/j.jhazmat.2008.07.013 - Theodoratos, P., Moirou, A., Xenidis, A. and Paspaliaris, I. (2000) The Use of Municipal Sewage Sludge for the Stabilization of Soil Contaminated by Mining Activities. Journal of Hazardous Materials, B77, 177-191. http://dx.doi.org/10.1016/S0304-3894(00)00243-0
- Epstein, E., Chaney, R.L., Henry, C. and Logan, T.J. (1992) Trace Elements in Municipal Solid Waste Compost. Biomass and Bioenergy, 3, 227-238. http://dx.doi.org/10.1016/0961-9534(92)90028-O
- Council Directive 91/271/EEC (1991) Concerning Urban Waste-Water Treatment.
- Xu, C., Chen, W. and Hong, J. (2013) Life-Cycle Environmental and Economic Assessment of Sewage Sludge Treatment in China. Journal of Cleaner Production, 67, 79-87.
- Cai, Q., Mo, C., Wu, Q., Zeng, Q. and Katsoyiannis, A. (2007) Concentration and Speciation of Heavy Metals in Six Different Sewage Sludge-Composts. Journal of Hazardous Materials, 147, 1063-1072. http://dx.doi.org/10.1016/j.jhazmat.2007.01.142
- Paradelo, M., Devesa, R., Moldes, A.B. and Barral, M.T. (2007) Physiologically Based Extraction of Heavy Metals in Compost: Preliminary Results. Journal of Trace Elements in Medicine and Biology, 21, 83-85. http://dx.doi.org/10.1016/j.jtemb.2007.09.016
- Nyamangara, J. (1998) Use of Sequential Extraction to Evaluate Zinc and Copper in a Soil Amended with Sewage Sludge and Inorganic Metal Salts. Agriculture, Ecosystems and Environment, 69, 135-141. http://dx.doi.org/10.1016/S0167-8809(98)00101-7
- Rodríguez, L., Ruiz, E., Alonso-Azcárate, J. and Rincón, J. (2009) Heavy Metal Distribution and Chemical Speciation in Tailings and Soils around a Pb-Zn Mine in Spain. Journal of Environmental Management, 90, 1106-1116. http://dx.doi.org/10.1016/j.jenvman.2008.04.007
- Ahluwalia, S. and Goyal, D. (2007) Microbial and Plant Derived Biomass for Removal of Heavy Metals from Wastewater. Bioresource Technology, 98, 2243-2257.
http://dx.doi.org/10.1016/j.biortech.2005.12.006 - Vullo, D.L., Ceretti, H.M., Daniel, M.A., Ramírez, S.A.M. and Zalts, A. (2008) Cadmium, Zinc and Copper Biosorption Mediated by Pseudomonas veronii 2E. Bioresource Technology, 99, 5574-5581. http://dx.doi.org/10.1016/j.biortech.2007.10.060
- Munoz, A.J., Ruiz, E., Abriouel, H., Gálvez, A., Ezzouhri, L., Larini, K. and Espínola, F. (2012) Heavy Metal Tolerance of Microorganisms Isolated from Wastewaters: Identification and Evaluation of Its Potential for Biosorption. Chemical Engineering Journal, 210, 325-332.
http://dx.doi.org/10.1016/j.cej.2012.09.007 - Vijayaraghavan, K. and Yun, Y. (2008) Bacterial Biosorbents and Biosorption. Biotechnology Advances, 26, 266-291. http://dx.doi.org/10.1016/j.biotechadv.2008.02.002
- Wang, J. and Chen, C. (2009) Biosorbents for Heavy Metals Removal and Their Future. Biotechnology Advances, 27, 195-226. http://dx.doi.org/10.1016/j.biotechadv.2008.11.002
- Munoz, A.J., Ruiz, E., Abriouel, H., Gálvez, A., Ezzouhri, L., Larini, K. and Espínola, F. (2012) Heavy Metal Tolerance of Microorganisms Isolated from Wastewaters: Identification and Evaluation of Its Potential for Biosorption. Chemical Engineering Journal, 210, 325-332.
http://dx.doi.org/10.1016/j.cej.2012.09.007 - Fomina, M. and Gadd, G.M. (2014) Biosorption: Current Perspectives on Concept, Definition and Application. Bioresource Technology, 160, 3-14. http://dx.doi.org/10.1016/j.biortech.2013.12.102
- Avidano, L., Gamalero, E., Cossa, G.P. and Carraro, E. (2005) Characterization of Soil Health in an Italian Polluted Site by Using Microorganisms as Bioindicators. Applied Soil Ecology, 30, 21-33. http://dx.doi.org/10.1016/j.apsoil.2005.01.003
- Weisburg, W.G., Barns, S.M., Pelletier, D.A. and Lane, D.J. (1991) 16S Ribosomal DNA Amplification for Phylogenetic Study. Journal of Bacteriology, 173, 697-703.
- Vinuesa, P., Rademaker, J.L.W., de Bruijn, F.J. and Werner, D. (1998) Genotypic Characterization of Bradyrhizobium Strains Nodulating Endemic Woody Legumes of the Canary Islands by PCR-Restriction Fragment Length Polymorphism Analysis of Genes Encoding 16S rRNA (16S rDNA) and 16S-23S rDNA Intergenic Spacers, Repetitive Extragenic Palindromic PCR Genomic Fingerprinting, and Partial 16S rDNA Sequencing. Applied and Environmental Microbiology, 64, 2096-2104.
- Altschul, S.F., Madden, T.L., Schaffer1, A.A., Zhang, J., Zhang, Z., Miller, W. and Lipman, D.J. (1997) Gapped BLAST and PSI-BLAST: A New Generation of Protein Database Search Programs. Nucleic Acids Research, 25, 3389-3402. http://dx.doi.org/10.1093/nar/25.17.3389
- Jeanmougin, F., Thompson, J.D., Gouy, M., Higgins, D.G. and Gibson, T.J. (1998) Multiple Sequence Alignment with Clustal X. Trends in Biochemical Sciences, 23, 403-405.
http://dx.doi.org/10.1016/S0968-0004(98)01285-7 - Kumar, S., Tamura, K., Jakobsen, I.B. and Nei, M. (2001) MEGA2: Molecular Evolutionary Genetics Analysis Software. Bioinformatics, 17, 1244-1245.
http://dx.doi.org/10.1093/bioinformatics/17.12.1244 - Lin, Y., Du, D., Si, C., Zhao, Q., Li, Z. and Li, P. (2014) Potential Biocontrol Bacillus sp. Strains Isolated by an Improved Method from Vinegar Waste Compost Exhibit Antibiosis against Fungal Pathogens and Promote Growth of Cucumbers. Biological Control, 71, 7-15.
http://dx.doi.org/10.1016/j.biocontrol.2013.12.010 - Colak, F., Atarb, N., Yazicioglu, D. and Olgunb, A. (2011) Biosorption of Lead from Aqueous Solutions by Bacillus Strains Possessing Heavy-Metal Resistance. Chemical Engineering Journal, 173, 422-428. http://dx.doi.org/10.1016/j.cej.2011.07.084
- Van der Geize, R. and Dijkhuizen, L. (2004) Harnessing the Catabolic Diversity of Rhodococci for Environmental and Biotechnological Applications. Current Opinion in Microbiology, 7, 255-261.
- Fleck, L.C., Correa, F. and Zachia, M.A. (2000) Physiological Aspects of Hydrocarbon Emulsification, Metal Resistance and DNA Profile of Biodegrading Bacteria Isolated from Oil Polluted Sites. Biotechnology Letters, 22, 285-289. http://dx.doi.org/10.1023/A:1005607112566
- Belimov, A.A., Hontzeas, N., Safronova, V.I., Demchinskaya, S.V., Piluzza, G., Bullitta, S. and Glick, B.R. (2005) Cadmium-Tolerant Plant Growth-Promoting Bacteria Associated with the Roots of Indian Mustard (Brassica juncea L. Czern.). Soil Biology & Biochemistry, 37, 241-250.
http://dx.doi.org/10.1016/j.soilbio.2004.07.033 - Esposito, A., Pagnanelli, F., Lodi, A., Solisio, C. and Vegliò, F. (2001) Biosorption of Heavy Metals by Sphaerotilus natans: An Equilibrium Study at Different pH and Biomass Concentrations. Hydrometallurgy, 60, 129-141. http://dx.doi.org/10.1016/S0304-386X(00)00195-X
- Karakagh, R.M., Chorom, S., Motamedi, H., Kalkhajeh, Y.K. and Oustan, S. (2012) Biosorption of Cd and Ni by Inactivated Bacteria Isolated from Agricultural Soil Treated with Sewage Sludge. Ecohydrology & Hydrobiology, 12, 191-198. http://dx.doi.org/10.1016/S1642-3593(12)70203-3
- Malik, A. and Jaiswal, R. (2000) Metal Resistance in Pseudomonas Strains Isolated from Soil Treated with Industrial Wastewater. World Journal of Microbiology & Biotechnology, 16, 177-182. http://dx.doi.org/10.1023/A:1008905902282
- Gisset, T., Botero, A.E., de Mesquita, L.M. and Torem, M.L. (2007) Biosorptive Removal of Cd and Zn from Liquid Streams with a Rhodococcus opacus Strain. Minerals Engineering, 20, 939-944. http://dx.doi.org/10.1016/j.mineng.2007.03.014
- Bueno, B.Y.M., Torem, M.L., de Carvalho, R.J., Pino, G.A.H. and de Mesquita, L.M.S. (2011) Fundamental Aspects of Biosorption of Lead (II) Ions onto a Rhodococcus opacus Strain for Environmental Applications. Minerals Engineering, 24, 1619-1624.
http://dx.doi.org/10.1016/j.mineng.2011.08.018 - Areco, M.M., Hanela, S., Duran, J. and Afonso, M.S. (2012) Biosorption of Cu(II), Zn(II), Cd(II) and Pb(II) by Dead Biomasses of Green Alga Ulva lactuca and the Development of a Sustainable Matrix for Adsorption Implementation. Journal of Hazardous Materials, 213-214, 123-132.
http://dx.doi.org/10.1016/j.jhazmat.2012.01.073 - Jing, H., Mezgebe, B., Hassan, A.A., Sahle-Demessie, E., Sorial, G.A. and Bennett-Stamper, C. (2014) Experimental and Modeling Studies of Sorption of Ceria Nanoparticle on Microbial Biofilms. Bioresource Technology, 161, 109-117. http://dx.doi.org/10.1016/j.biortech.2014.03.015
NOTES

*Corresponding author.


