Natural Resources
Vol.3 No.4(2012), Article ID:26267,5 pages DOI:10.4236/nr.2012.34027
Tissue-Specific Isoenzyme Variations in Tilapia Fish, Oreochromis niloticus
![]()
1Department of Biology, Faculty of Science, Taif University, Taif, Saudi Arabia; 2Department of Zoology, Faculty of Science, Cairo University, Giza, Egypt.
Email: *yasser92us@yahoo.com
Received September 15th, 2012; revised October 21st, 2012; accepted November 7th, 2012
Keywords: Oreochromis niloticus; Malate Dehydrogenase; Acid Phosphatase; Peroxidase; Isoenzymes
ABSTRACT
Native polyacrylamide gel electrophoresis have been used to analyze malate dehydrogenase (MDH), acid phosphatase (Acph) and peroxidase (Px) isoenzymes in different tissues (liver, kidney, muscle and heart) of the tilapia fish, Oreochromis niloticus in order to study the tissue specificity of these isoenzymes. Three, two and one fractions have been recorded respectively for the three isoenzymes in different studied tissues. The MDH-1 and MDH-2 have been expressed only in muscle and heart while MDH-3 has been expressed in all studied tissues. The percentage amount of MDH in general varied significantly between muscle and different studied tissues. With respect to acid phosphatase, the percentage amount of the total enzyme showed significant difference between liver and muscle and that this variation may be due to higher gene activity in liver. Peroxidase isoenzyme was recorded in liver and heart only with significant increase in liver. The kidney was the least among the studied tissues in showing gene expression for the studied isoenzymes and therefore, liver, heart and muscle tissues are better applicable in studying the isoenzymatic profiles for fish physiology and systematics.
1. Introduction
Tilapiini is a highly diverse tribe with more than 70 species belonging to the order Perciformes, family Cichlidae [1] to which the commonly called tilapia belongs. Tilapia is a generic term used to designate a group of comercially important Cichlid fish, which consists of three genera: Oreochromis, Sarotherodon and Tilapia [2]. The Nile tilapia O. niloticus is the most widely farmed species [3], recognized as one of the most important species in tropical and subtropical aquaculture because of their production potential [4].
Several investigations have been concerned with the characterization of tissue-and organ-specific isoenzyme patterns [5-12] among which little were concerned with fishes. Few studies have concerned with the isoenzymatic profiles in the Nile tilapia [4,13].
LDH and MDH isoenzymes are major system found in fishes. They are classified in different groups on the basis of their possession in different tissues or cell organelles [14,15]. Chaudhuri and Krishna [16] studied the tissue specificity and the degree of polymorphism of five enzyme systems in Labeo rohita from Yamuna namely in liver, muscle, heart and brain tissues.
Acid phosphatases often occur in multiple forms differing in molecular sizes [17]. In animals its biological role is not yet clear but it is involved in many biological systems which are linked to energy metabolism, metabolic regulation and cellular signal transduction pathways [18].
Peroxidase enzyme (H2O2 donor and consumer) contains many isoforms which partake in a variety of metabolic functions. In animals, peroxidase enzymes are involved in phagocytosis and immune cell function [19,20], cell adhesion [21], antioxidant function [22,23] and the oxidative polymerization of hydroquinones to melanin [24].
The present study aimed to investigate whether the studied isoenzymes differently expressed in the different tissues of the Nile tilapia Oreochromis niloticus cultivated in the Saudi Arabian farms.
2. Materials and Methods
Fish samples were freshly obtained from a fish Farm in Makkah city, immediately taken to the lab. Male samples weighing 241 - 276 g were used. The samples were then autopsied and the organs were taken and stored at –80˚C for further laboratory use. The targeted organs were the liver, the kidney, a muscle and the heart.
The Isozymes used were: malate dehydrogenase (MDH), acid phosphatase (Acph.) and peroxidase (Px). For isoenzyme extraction, approximately 0.5 g of tissue was homogenized in 2 mL saline solution NaCl (0.9%) using a manual Homogenizer. The homogenates were centrifuged at 5000 rpm for 10 minutes and the supernatants were kept at –20˚C until use. For electrophoresis, 30 μL of the extract was mixed with 10 μL of treatment buffer and 35 μL of this mixture was applied to the well. Isoenzymes were electrophorased in 10% native-polyacrylamide gel as described by Stegemann et al. [25]. After electrophoresis, the gels were stained according to their enzyme system with the appropriate substrate and chemical solutions then incubated at room temperature in dark for complete staining. In most cases the incubation for about 1 to 2 hours is enough.
In gels staining, protocols of Jonathan and Wendell [26] for MDH, Wendel and Weeden [27] for Acph and Heldt [28] for Px were used. Gels were washed two or three times with tap water; fixed in ethanol: 20% glacial acetic acid (9:11 v/v) for 24 hours; and photographed.
After the appearance of the enzyme bands, the reaction was stopped by washing the gel two or three times with tap water. This was followed by adding the fixative solution, which consists of ethanol and 20% glacial acetic acid (9:11 v/v). The gel was kept in the fixative solution for 24 hours and then was photographed.
All gels were scanned using Gel Doc-2001 Bio-Rad system. For isoenzymes, the bands of enzyme activity were designated using the known system of nomenclature [29]. An abbreviation which corresponds to the name of the enzyme designated each locus. When multiple loci were involved, the fastest anodal protein band was designated as locus one, the next as locus two and so on.
The data were expressed as means ± standard error of mean. The percentage amounts for isoforms were compared using the independent t-test. All analyses were performed using the Statistical Package for Social Sciences (SPSS) software v. 13 in a PC-compatible computer and the significance was set at P < 0.05.
3. Results and Discussion
Malate dehydrogenase isoenzyme showed three fractions in the electrophoretic pattern (Figure 1). The first two fractions (MDH-1 and MDH-2) were recorded in muscle and heart tissues only while the third fraction was recorded in all studied tissues. Table 1 showed the mean and standard error for the percentage amount of the studied isoenzymes in the different Tilapia tissues. The MDH-1 isoform that occurred only in muscle and heart didn’t show any variation in its activity between these tissues. On the other hand, the MDH-2 isoform that also occurred only in muscle and heart showed a significant increase (P < 0.05) in its activity in muscle than in the heart. MDH-3 was significantly higher in liver (P < 0.01) and kidney (P < 0.05) than in muscle. The percentage amount of the total enzyme was significantly higher in muscle tissue (P < 0.01, P < 0.05) than in liver and kidney tissues.
Acid phosphatase showed clear tissue specificity in its expression. The enzyme recorded two types (Acph-1 and Acph-2) in liver tissue while only one cathodal type appeared in muscle tissue. The fastest anodal type was expressed with significant increase in liver (P < 0.05) than in muscle tissues (Table 1). The electrophoretic pattern (Figure 2) clearly demonstrated high intensity (P < 0.05) for the bands recorded in liver tissues. No band was shown in kidney and heart tissues. Siddiqua et al. [30] and the references cited therein revealed that three types of acid phosphatase have been described in vertebrates based on molecular weight and its localization within the
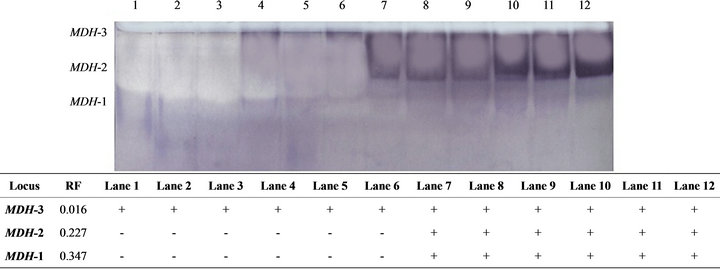
Figure 1. The electrophoretic profile (above) and the recorded isoforms with the relative mobility (RF) (below) of MDH isoenzymes in the studied tissue samples. Lanes are as follow: 1 - 3 (liver); 4 - 6 (kidney); 7 - 9 (muscle) and 10 - 12 (heart).
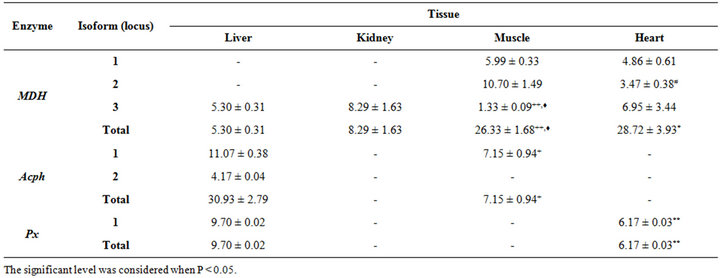
Table 1. Mean ± SE of the percentage amount for the studied isoenzymes in different tissues of Fish (Tilapia). The significance level was calculated by Student t-test. +significance level between liver and muscle; ♦significance level between kidney and muscle; *significance level between Liver and heart and #significance level between muscle and heart.
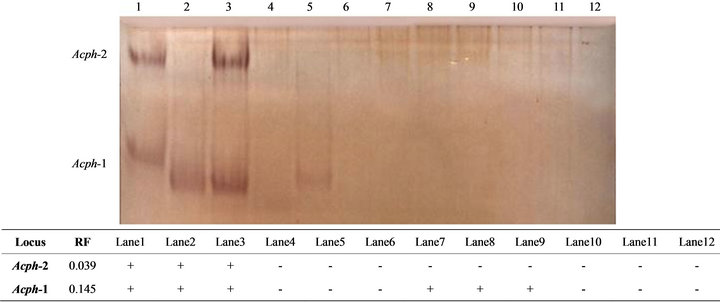

Figure 2. The electrophoretic profile (above) and the recorded isoforms with the relative mobility (RF) (below) of acid phosphatase (Acph) isoenzymes in the studied tissue samples. Lanes are as follow: 1 - 3 (liver); 4 - 6 (kidney); 7 - 9 (muscle) and 10 - 12 (heart).
cell organelle.
Regarding peroxidase isoenzymes, only one sharp fraction was recorded in liver and heart tissues (Figure 3). This enzyme could also be considered as a good indicator for tissue specificity where it was expressed as two fractions (Px-1 and Px-2) in liver and heart tissues with significantly higher activity in the liver (P < 0.01). In marine invertebrate [31], changes in peroxidase levels can signify immunomodulation due to contaminants and other environmental stressors. The enzyme processes an importance in the defenses of many different organisms [32]. Early roles of peroxidase induction in resistance may include the oxidation of substrates to produce cytotoxic molecules [33], production of reactive oxygen as cytotoxic molecules [34] and regulation of the oxidative state of the tissue [22]. We therefore, can assume that the less expression of peroxidase in the tilapia tissues may be attributed to the good environmental condition of the farm from which the samples collected. Many other factors can also be considered for this low expression such as the decreased quality of the chemicals used to investigate the enzyme, the bad storage of the tissues used or the bad handling of the samples obtained. These factors may also be considered for all isoenzymes studied herein.
Gene duplication [6] as well as post-translational processing mechanisms have been proposed as important factors in modulating tissue-specific enzymes. Removal of N-terminal amino acids [35], rate of translation of
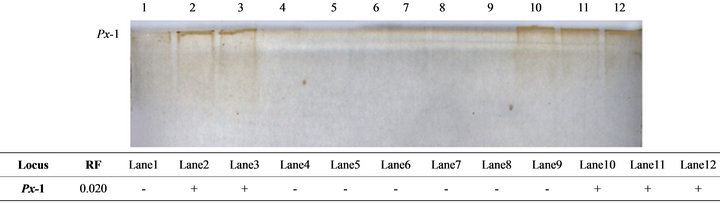

Figure 3. The electrophoretic profile (above) and the recorded isoforms with the relative mobility (RF) (below) of peroxidase (Px) isoenzymes in the studied tissue samples. Lanes are as follow: 1 - 3 (liver); 4 - 6 (kidney); 7 - 9 (muscle) and 10 - 12 (heart).
mRNA, or rate of protein degradation [36] have been suggested as post-translational processing mechanisms that lead to tissue-and organ-specific isoenzyme patterns in fishes. In the present study, it was obvious that the kidney was the least among the studied tissues in showing gene expression for the studied isoenzymes. The only expressed enzyme in kidney tissues was MDH. We may therefore conclude that liver, heart or muscle tissues are better applied in studying the isoenzymatic profiles for fish physiology and systematics. We also agreed with Shahjahan et al. [13] in that the findings in the present study may be extended to use genetic markers in various fields of physiology, taxonomy and toxicology in Nile tilapia (O. niloticus).
4. Conclusion
In this work it has been shown that liver, heart and muscle tissues are better applied in studying the isoenzymatic profiles for fish physiology and systematics. The present study may also be used as a clue for using genetic markers in various fields of physiology, taxonomy and toxicology. Application of more isoenzymes to freshly obtained tissues can give more obvious discrimination among the different fish organs.
5. Acknowledgements
We are grateful to Dr. Shawkat Ahmed at Ain Shams University of Egypt for his technical support in conducting the practical part of this work.
REFERENCES
- R. Trewavas, “Tilapiine Fishes of the Genera Sarotherodon, Oreochromis and Danakilia, ˮ Comstock Publishing Associates, London, 1983.
- S.-F. Li, Y. Zhao, W.-J. Fan, W.-Q. Cai and Y.-F. Xu, “Possible Genetic Reproductive Isolation between Two Tilapiine Genera and Species: Oreochromis niloticus and Sarotherodon melanotheron,ˮ Zoological Research, Vol. 32, No. 5, 2011, pp. 521-527.
- R. S. V. Pullin, A. E. Eknath, T. Gjedrum, M. M. Tayamen, J. M. Macaranas and T. A. Abella, “The Genetic Improvement of Farmed Tilapias (GIFT) Project, the Story So Far,ˮ NAGA. ICLARM Quarterly, Vol. 14, No. 2, 1991, pp. 7-9.
- S.-F. Li, J.-L. Zhao, M. Dey and R. Dunham, “Isozyme Variation of Nile Tilapia Oreochromis niloticus in China,ˮ Asian Fisheries Science, Vol. 14, 2001, pp. 411-416.
- Y. Mo, C. D. Young and R. W. Gracy, “Isolation and Characterization of Tissue-Specific Isozymes of Glucosephosphate Isomerase from Catfish and Conger,ˮ Journal of Biological Chemistry, Vol. 250, No. 17, 1975, pp. 6747- 6755.
- S. E. Fisher and G. S. Whitt, “Evolution of Isozyme Loci and Their Differential Tissue Expression. Creatine Kinase as a Model System,ˮ Journal of Molecular Evolution, Vol. 12, No. 1, 1978, pp. 25-55. doi:10.1007/BF01732544
- J. F. Leslie and P. J. Pontier, “Linkage Conservation of Homologous Esterase Loci in Fish (Cyprinodontoidei: Poeciliidae),ˮ Biochemical Genetics, Vol. 18, No. 1-2, 1980, pp. 103-105. doi:10.1007/BF00504363
- W. J. Berg and D. G. Buth, “Glucose Dehydrogenase in Teleosts: Tissue Distribution and Proposed Function,ˮ Comparative Biochemistry and Physiology, Vol. 77, No. 2, 1984, pp. 285-288.
- R. W. Holt and W. S. Leibel, “Coexpression of Distinct Eyeand Liver-Specific LDH Isozymes in Cichlid Fish,ˮ Journal of Experimental Zoology, Vol. 244, No. 2, 1987, pp. 337-343. doi:10.1002/jez.1402440219
- F. Basaglia, “Interspecific Gene Differences and Phylogeny of the Sparidae Family (Perciformes, Teleostei), Estimated from Electrophoretic Data on Enzymatic TissueExpression,ˮ Comparative Biochemistry and Physiology, B, Vol. 99, No. 3, 1991, pp. 495-508. doi:10.1016/0305-0491(91)90329-C
- D. Xia, T. Wu and H. Wang, “Differential Gene Expression for Lactate Dehydrogenase of Mandarian Fish (Sinaperca chuatsi),ˮ Aquaculture, Vol. 108, No. 3-4, 1992, pp. 207-214. doi:10.1016/0044-8486(92)90107-V
- M. Seimiya, T. Kusakabe and N. Suzuki, “Primary Structure and Differential Gene Expression of Three Membrane Forms of Guanylyl Cyclase Found in the Eye of the Teleost Oryzias latipes, ˮJournal of Biological Chemistry, Vol. 272, 1997, pp. 23407-23417. doi:10.1074/jbc.272.37.23407
- R. Shahjahan, A. Karim, R. A. Begum, M. S. Alam and A. Begum, “Tissue Specific Esterase Isozyme Banding Pattern in Nile Tilapia (Oreochromis niloticus), ˮUniversity Journal of Zoology, Vol. 27, 2008, pp. 1-5.
- C. R. Goward and D. J. Nicholls, “Malate Dehydrogenase: A Model for Structure, Evolution, and Catalysis,ˮ Protein Science, Vol. 3, No. 10, 1994, pp. 1883-1888. doi:10.1002/pro.5560031027
- R. Mishra and S. P. Shukla, “Endosulfan Negatively Modulates Mitochondrial Malate Dehydrogenase from the Freshwater Catfish, Clarias batrachus,ˮ Ecotoxicology and Environmental Safety, Vol. 56, No. 3, 2003, pp. 425-433. doi:10.1016/S0147-6513(03)00006-X
- A. Chaudhuri and G. Krishna, “Tissue Specifisty and Degree of Polymorphism of Five Enzyme Systems of L. rohita, from River Yamuna,ˮ Fish Genetics and Biodiversity Conservation, Vol. 5, 1998, pp. 358-360.
- S. Fujimoto, Y. Urata, T. Nakagawa and A. Ohara, “Characterization of Intermediate Molecular Weight Acid Phosphatase from Bovin Kidney Cortex, ˮJournal of Biochemistry, Vol. 96, No. 4, 1984, pp. 1079-1088.
- J. Shan, “Dissertation on: Transcriptional Regulation of the Human Prostatic Acid Phosphatase Gene,ˮ University of Oulu, Oulu, 2002, p. 15.
- A. Rodriguez, M. A. Esteban and J. Meseguer, “Phagocytosis and Peroxidase Release by Seabream (Sparus aurata L.) Leukocytes in Response to Yeast Cells, ˮ The Anatomical Record Part A: Discoveries in Molecular, Cellular, and Evolutionary Biology, Vol. 272, No. 1, 2003, pp. 415-423. doi:10.1002/ar.a.10048
- I. M. Soares-da-Silva, J. Ribeiro, C. Valongo, R. Pinto, M. Vilanova, R. Bleher and J. Machad, “Cytometric, Morphologic and Enzymatic Characterization of Haemocytes in Anodonta cygnea,ˮ Comparative Biochemistry and Physiology A: Molecular & Integrative Physiology, Vol. 132, No. 3, 2002, pp. 541-553. doi:10.1016/S1095-6433(02)00039-9
- T. Holmblad and K. Soderhall, “Cell Adhesion Molecules and Antioxidative Enzymes in a Crustacean, Possible Role in Immunity,ˮ Aquaculture, Vol. 172, No. 1-2, 1999, pp. 111-123. doi:10.1016/S0044-8486(98)00446-3
- S. C. Gamble, P. S. Goldfarb, C. Porte and D. R. Livingstone, “Glutathione Peroxidase and Other Antioxidant Enzyme Function in Marine Invertebrates (Mytifus edulis, Pecten maximus, Carcinus maenas and Asterias rubens),ˮ Marine Environmental Research, Vol. 39, No. 1-4, 1995, pp. 191-195. doi:10.1016/0141-1136(94)00031-J
- T. S. Galloway and M. H. Depledge, “Immunotoxicology in Invertebrates: Measurement and Ecotoxicological Review,ˮ Ecotoxicology, Vol. 10, No. 1, 2001, pp. 5-23. doi:10.1023/A:1008939520263
- M. D’Ischa, A. Napolitano and G. Prota, “Peroxidase as an Alternative to Tyrosinase in the Oxidative Polymerization of 5,6-Dihydroxyindoles Tomelanin(s), ˮBiochimica et Biophysica Acta, Vol. 1073, No. 2, 1991, pp. 423-430. doi:10.1016/0304-4165(91)90152-7
- H. Stegemann, A. M. R. Afify and K. R. F. Hussein, “Cultivar Identification of Dates (Phoenix dactylifera) by Protein Patterns,ˮ Second International Symposium of Biochemical Approaches to Identification of Cultivars, Braunschweing, West Germany, 1985, p. 44.
- J. F. Jonathan and N. F. Wendell, “Visualization and Interpretation of Plant Isozymes,ˮ In: D. E. Soltis and P. S. Soltis, Ed., Isozymes in Plant Biology, Champan and Hall, London, 1990, pp. 5-45.
- J. F. Wendel and N. F. Weeden, “Visualization and Interpretation of Plant Isozymes,ˮ In: D. E. Soltis and P. S. Soltis, Eds., Isozymes in Plant Biology, Dioscorides Press, Portland, 1989, pp. 5-45. doi:10.1007/978-94-009-1840-5_2
- W. H. Heldt, “A Leaf Cell Consists of Several Metabolic Compartments,ˮ Plant Biochemistry and Molecular Biology, Institute of Plant Biochemistry, Gottingen with the Collaboration of Fiona, 1997.
- W. Allendorff and F. M. Utter, “Population Genetics,ˮ In: W. S. Hoaran and D. J. Randalal, Eds., Fish Physiology, Academic Press, New York, 1978, pp. 407-454.
- A. Siddiqua, A. Saeed, R. Naz, M. Sherazi, S. Abbas and A. Saeed, “Purification and Biochemical Properties of Acid Phosphatase from Rohu Fish Liver,ˮ International Journal of Agriculture and Biology, Vol. 14, No. 2, 2012, pp. 223-228.
- E. A. Dyrynda, R. K. Pipe, G. R. Burt and N. A. Ratcliffe, “Modulations in the immune Defenses of Mussels (Mytilus edulis) from Contaminated Sites in the UK,ˮ Aquatic Toxicology, Vol. 42, No. 3, 1998, pp. 169-185. doi:10.1016/S0166-445X(97)00095-7
- L. D. Mydlarz and C. D. Harvel, “Peroxidase Activity and Inducibility in the Sea Fan Coral Exposed to a Fungal Pathogen,ˮ Comparative Biochemistry and Physiology A: Molecular and Integrative Physiology, Vol. 146, No. 1, 2007, pp. 54-62. doi:10.1016/j.cbpa.2006.09.005
- A. J. Nappi and B. M. Christensen, “Melanogenesis and Associated Cytotoxic Reactions: Applications to Insect Innate Immunity,ˮ Insect Biochemistry and Molecular Biology, Vol. 35, No. 5, 2005, pp. 443-459. doi:10.1016/j.ibmb.2005.01.014
- G. P. Bolwell, D. R. Davies, C. Gerrish, C.-K. Auh, T. M. Murphy, “Comparative Biochemistry of the Oxidative Burst Produced by Rose and French Bean Cells Reveals Two Distinct Mechanisms, ˮPlant Physiology, Vol. 116, No. 4, 1998, pp. 1379-1385. doi:10.1104/pp.116.4.1379
- P. R. Laud and J. W. Campbell, “Genetic Basis for Tissue Isozyme of Glutamine Synthetase in Elasmobranchs,ˮ Journal of Molecular Evolution, Vol. 39, No. 1, 1994, pp. 93- 100. doi:10.1007/BF00178254
- T. H. Yang and G. N. Somero, “Activity of Lactate Dehydrogenase but Not Its Concentration of Messenger RNA Increases with Body Size in Barred Sand Bass, Paralabrax nebulifer (Teleostei),ˮ The Biological Bulletin, Vol. 19, No. 2, 1996, pp. 155-158.
NOTES
*Corresponding author.

