Open Journal of Animal Sciences
Vol.2 No.3(2012), Article ID:21369,9 pages DOI:10.4236/ojas.2012.23024
Association analyses of single nucleotide polymorphisms in the leptin and leptin receptor genes on milk and morphological traits in Holstein cows
![]()
1Agricultural Research Council-Animal Production Research Centre (CRA-PCM), Animal Production Research Centre, Via Salaria, Monterotondo, Roma, Italy; *Corresponding Author: giovanna.dematteis@entecra.it
2Agricultural Research Council-Research Centre for Fodder Crops and Dairy Productions (CRA-FLC), Research Centre for Fodder Crops and Dairy Productions, Via Porcellasco Cremona, Italy
Received 2 April 2012; revised 9 May 2012; accepted 17 May 2012
Keywords: Polymorphis; Leptin Gene; Leptin Receptor Gene and Dairy Traits
ABSTRACT
To date, only a few SNP in the leptin receptor gene have been detected and analyzed while the leptin gene has been extensively characterized and several polymorphisms were detected and associated to functional and productive traits in cattle. In this work, for the first time, the bovine leptin receptor gene was fully characterized by sequencing the whole coding region and part of the 5’ flanking region. A group of 95 Holstein cows was genotyped in order to search for SNP at both the LEPR and LEP genes, and to associate them with milk and morphological traits. Nine novel SNP in the leptin receptor gene and one novel SNP in the leptin gene were detected. Four SNP of the leptin receptor gene and one SNP of leptin gene showed a significant effect on one or more analyzed traits, and, in all cases, the greatest effect was observed on fat content. This study provided knowledge of the existence of further polymorphisms in the leptin receptor and leptin genes that have an influence on some important economic traits.
1. INTRODUCTION
The leptin receptor (LEPR) is a glycoprotein with a single transmembrane-spanning region; it consists of 1165 amino acids and is a member of the class I cytokine receptor family [1]. Reference [2] reviewed the activity of the LEPR gene, which is expressed in six isoforms, due to alternative RNA splicing. The isoforms are divided into three classes: long, short, and secretory. The long, fully active isoform (LEPR-b) is expressed mainly in the hypothalamus, where it takes part in the energy homeostasis and in the regulation of the activity of the secretory organs.
The bovine LEPR gene is located on chromosome 3q33 [3] and is composed of eighteen exons. It has only been partly sequenced and, until now, few SNP were detected. Reference [4] detected five SNP localized in exon 4 of the LEPR gene in some Chinese cattle breeds and, after evaluating their effects on growth traits, they suggested that this gene may be a potential candidate for growth in animals. More interesting were the findings of Liefers [5] who detected a C-T missense mutation that causes the Threonine-Methionine amino acid substitution in the intracellular domain of the LEPR-b isoform. These authors reported that this SNP is associated with leptin concentration during late pregnancy. The same mutation was further associated with fat and protein content in Jersey cattle, but not with milk yield [6]. Reference [7] reported only a weak association of this SNP with milk yield and days to first service, while [8] found no significant association either with milk production, or feed intake, or body energy traits, in UK dairy cows. Also reference [9] could not associate this SNP either with milk yield or composition.
The leptin hormone, a peptide of 146 amino acids, is synthesized and secreted primarily by adipose cells and by other organs and tissues, including stomach, muscle, placenta and fetal tissues [10]; it is involved in the feed intake, the energy partitioning and the metabolism of the cow [11,12]. The leptin gene (LEP) is located on chromosome 4q32 in the bovine [13] and is composed of three exons, spanning around 18.9 kb of the genome [14]. Several polymorphism of the LEP have been described [15,16] and associated with serum leptin concentrations [17,18], feed intake, milk yield, energy balance and fertility [16,19], milk energy output [8,20], milk composition [9], energy storage [21,22] and growth, fertility and milk production [7]. While the LEP gene has been extensively characterized, and several polymorphisms were associated to functional and productive traits in cattle, only few SNP in the LEPR gene have been detected and analyzed. The purpose of this work was therefore to fully characterize the bovine LEPR gene by sequencing the whole coding region and part of 5’ flanking region, in order to search for novel SNP and to evaluate the effect of the SNP of both LEP and LEPR genes on milk production and morphological traits of the same group of animals.
2. MATERIAL AND METHODS
2.1. Animals and Phenotypes
Official milk recording data, spanning over 10 years time, from 95 Holstein cows were used in this study. Cows belonged to four commercial neighboring farms located in Northern Italy, the most vocated Italian area for milk production. DNA was obtained from blood for each recorded animal. The following parameters were considered: fat and protein content at the test day; mature equivalent (ME) milk yield, fat and protein content. For 48 of these cows, the results of the morphological evaluation, performed by the Herdbook experts were available; therefore the following parameters were also included in the statistical analysis: stature, body depth, angularity, rump angle, rump width, rear legs, udder depth.
2.2. SNP Detection and Genotyping
Polymorphism detection and genotyping was performed for all samples by direct sequencing on the 3500 Genetic Analyzers (Applied Biosystems, Foster City, CA, USA). For the LEPR gene, 19 primer pairs were designed on the Bos taurus whole genome shotgun sequence (accession no. NW_001494806) and are summarized in Table 1. The amplified fragments include the whole coding sequence and part of the 5’ flanking region. For the LEP gene, all cows were genotyped at the already reported SNP by [15,16,23]. Primers used and position on the gene are summarized in Table 2. In order to sequence the amplicons Promoter 1 (1141 bp) and Promoter 2 (613 bp) that presented a very high CG content together with several deletions and insertion it was necessary to design four more forward (P1-for1, P1-for2, P1-for3, P2-for) and two more reverse primers (P1-rev, P2-rev), that fell inside the amplicon, so to obtain shorter sequences ensuring a greater analytical specificity, without affecting the efficiency.
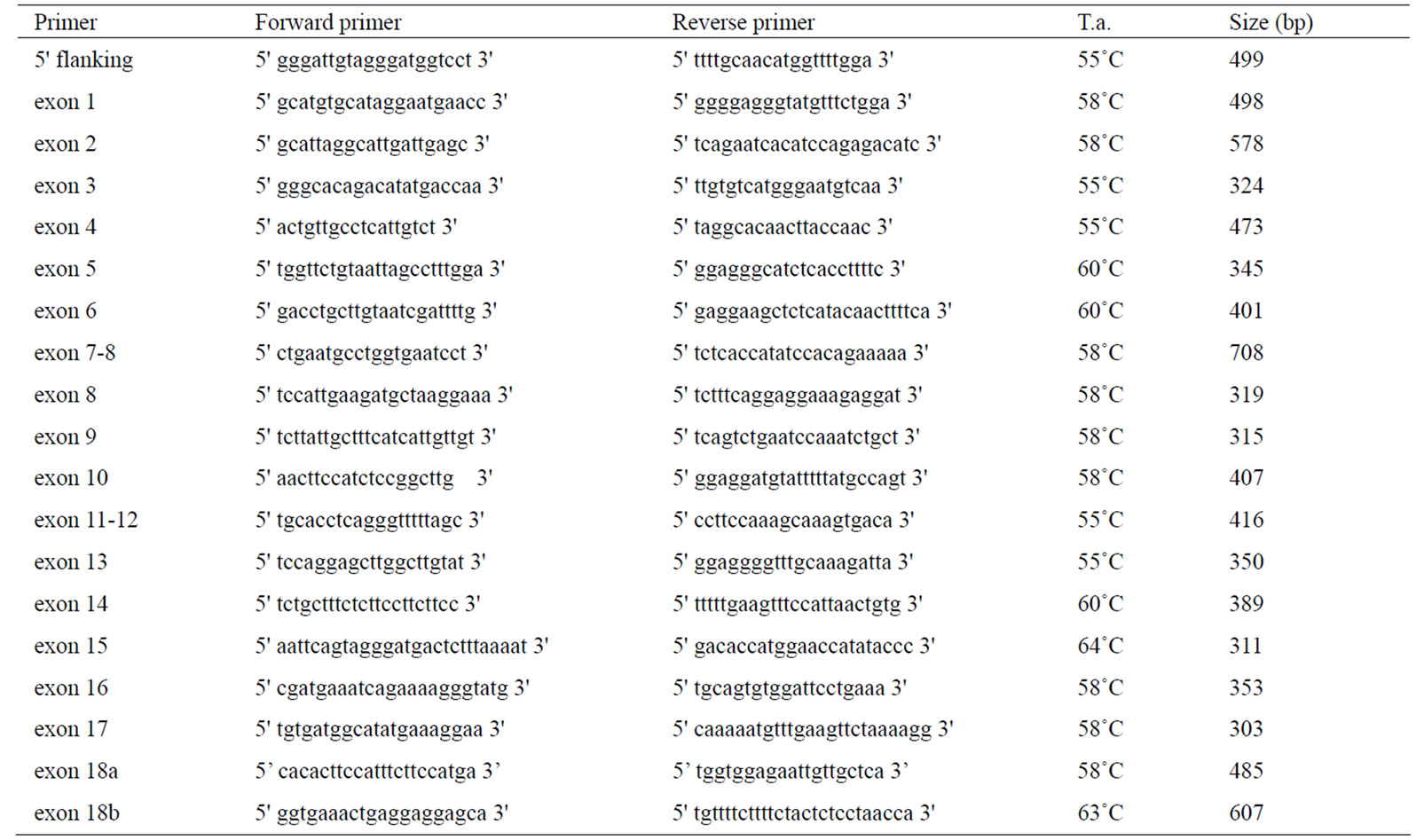
Table 1. Primers used for the amplification and sequencing of the bovine LEPR gene.
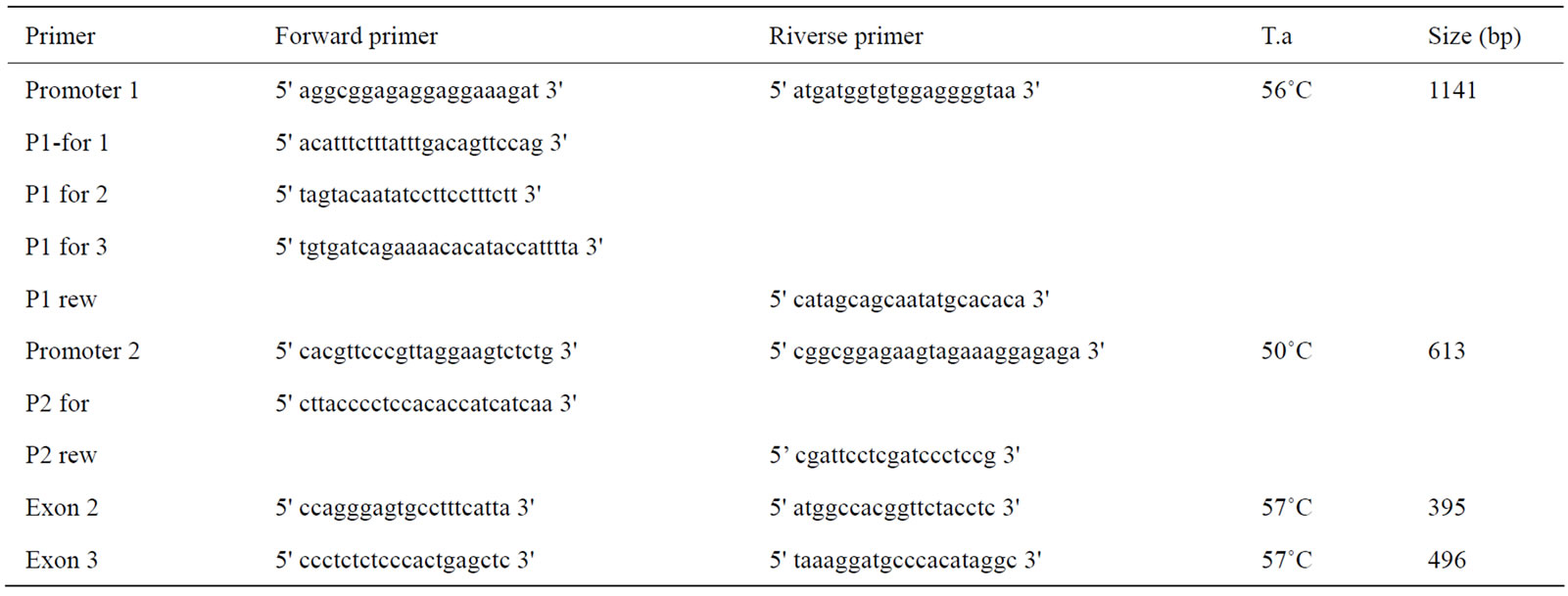
Table 2. Primers used for the amplification and sequencing of the bovine LEP gene.
2.3. Statistical Analysis
The allele substitution effect of each SNP was estimated, similarly to [24,25], by regressing the number of copies of each allele against each of the traits separately, using the following Linear Mixed Model in SAS [26]:
Yijklm = μ + Bi + + Cj + Dk + Gl + Am + eijklmn
where: Yijklm = Phenotype1; μ = overall mean; Bi = fixed effect of the herd; Cj = fixed effect of the season/year of calving; Dk = fixed effect of the age at calving; Gl = allele substitution effect; Am = random animal effect; eijklmn = residual.
A GLM that included only the allele substitution effect as covariate was used for the ME milk yield, fat and protein content, because these parameters were already corrected for the fixed effects. A GLM that included the year of birth and the age at the evaluation as fixed effects was used for the morphological indexes.
A false discovery rate (FDR) was also calculated using PROC MULTTEST in SAS [26] to account for multiple testing, whereby significance was achieved at an adjusted FDR of P < 0.10, as suggested in [7]. Linkage disequilibrium (LD) between loci was estimated using PROC ALLELE in SAS [26] in order to evaluate the pair-wise correlation coefficients for all SNP within each of the two genes, so to avoid false-positive effects and not to ascribe to a SNP the effect of another one, in case the alleles of the second were highly correlated with the first.
2.4. Transcription Factor Binding Sites
The MATCH program [27] in the TRANSFAC® Professional 10.2 http://www.biobaseinternational.com/ [28] was used to perform the profile search for the transcripttion factor binding sites for the two genes. The vertebrate binding matrices and the only high quality matrix option were used in the profile selection, and the cut-offs for core and matrix similarity were set to 0.999 and 0.7 respectively, with no further change in the other options.
3. RESULTS
In Tables 3-4, the position in the gene and the minor allele frequency (MAF) of the detected SNP in the analyzed population, for the LEPR and the LEP gene was reported. Sequence analysis of the DNA of the 95 Holstein cows, across the 7.8 kb region of the bovine LEPR gene, that includes part of the 5’ flanking region and the whole coding region, revealed a total of 16 SNP; 9 of them being novel detected SNP, not previously referred in the literature (Table 3). The novel SNP LEPR01 and LEPR02 are located in the 5' flanking region; LEPR04, LEPR10 and LEPR16 are synonymous mutations, located, respectively, in exons 7, 12, and 18; LEPR06 is a missense mutation in exon 8 that causes the amino acid substitution S460G at the position 460 of the protein; LEPR09 is also a missense mutation in exon 11, producing the amino acid substitution R615S at the position 615 of the protein. Because [29] showed that the region from 428 to 635 of the protein includes the binding sites for the leptin, the two missense mutations may be fundamental for binding and activate the leptin hormone. Finally, LEPR11 and LEPR14 are located in the introns 15 and 17.
Sequence analysis of the DNA of the 95 Holstein cows, across the 2.6 kb region of the bovine LEP gene, revealed a total of 26 SNP (24 in the promoter and 2 in the exons). Of the 26 detected SNP, 19 had already been reported by [16] (accession no. AJ571671); three SNP were reported in the dbSNP (rs29004171; rs29004172; rs29004173); one SNP (accession no. AB070368) had been previously reported by [23] and two SNP (rs29004488 and rs29004508) had been reported by [15]. Finally, a novel SNP, LEP05, was detected in the present study (g.2003435T>C; accession no. NW_001494939).

Table 3. Analyzed SNP in the LEPR gene.
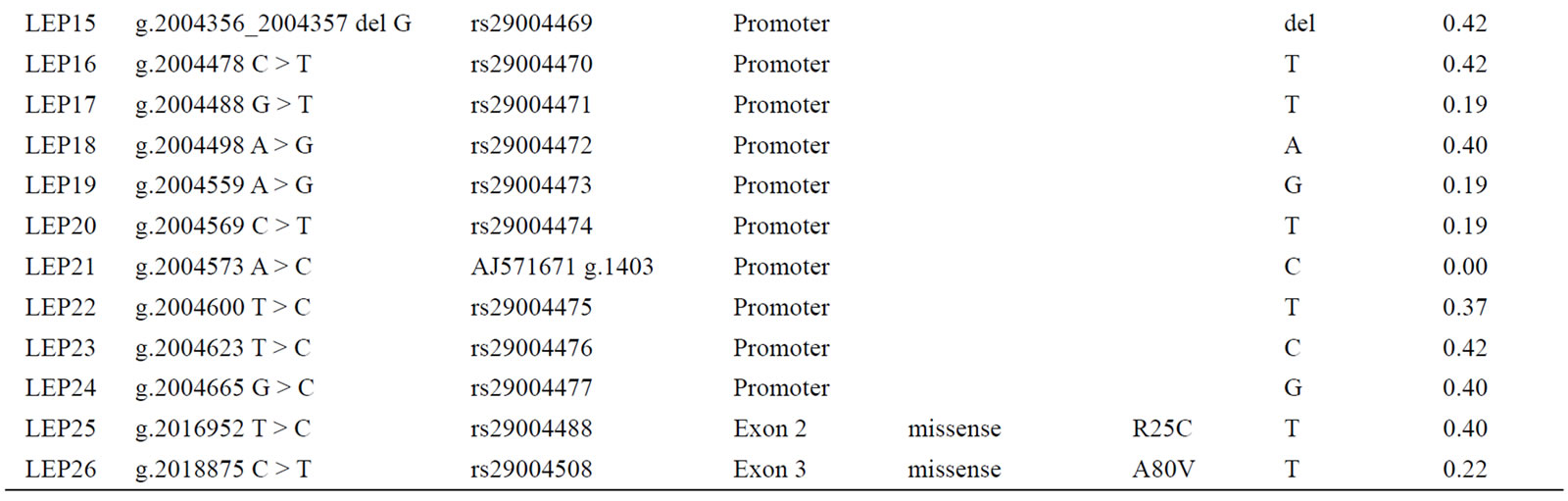
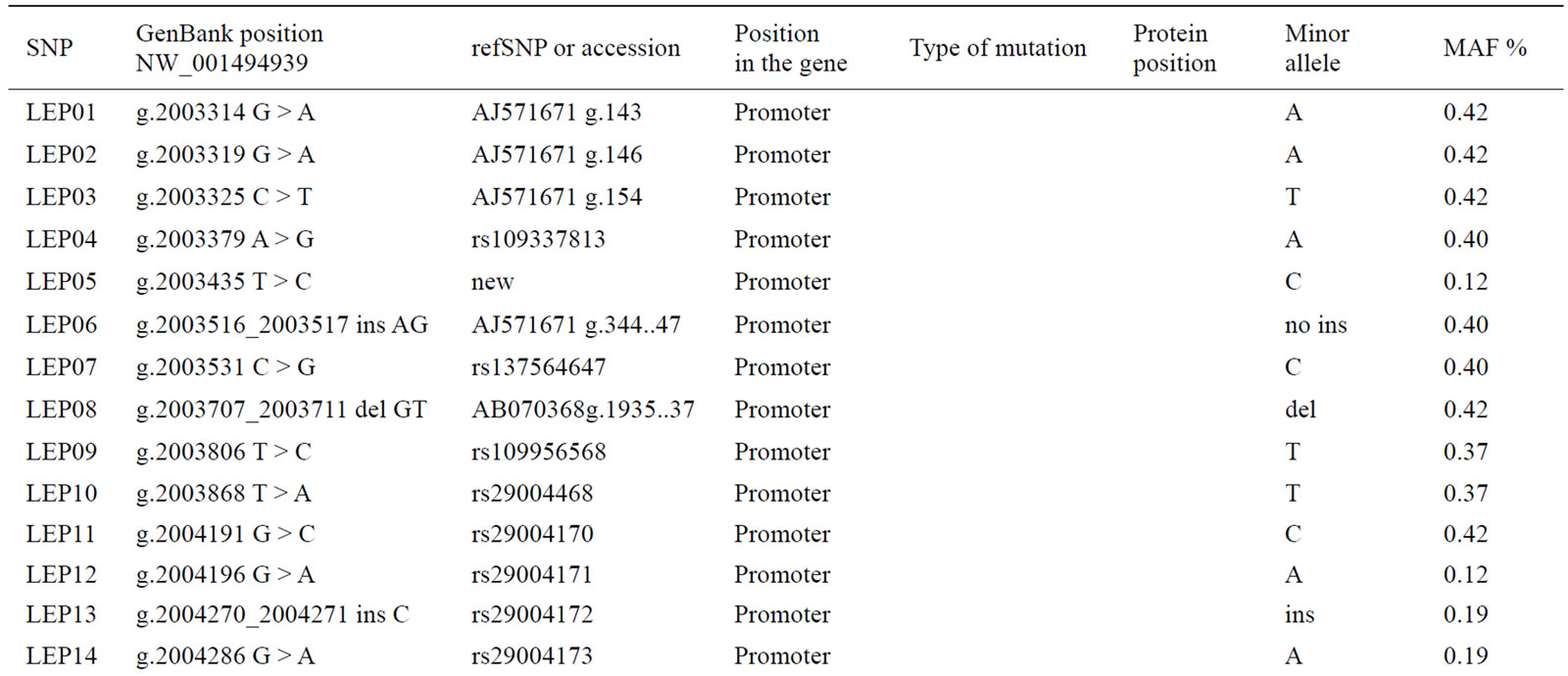
Table 4. Analyzed SNP in the LEP gene.
In this study, the in silico analysis with the MATCH software was performed to verify whether the SNP that resulted significantly associated to productive traits, were located in putative binding sites of transcription factors. The results of this analysis indicated that LEP14 is located at the putative binding site for v-Maf and NF-E2 transcription factors and that the A allele does not activate the binding sites of either of the two.
In the association analysis, of the 16 SNP of the LEPR gene, the following 6 were used: LEPR01 and LEPR02 because located in the promoter; LEPR03, LEPR09 and LEP15, because they encode a missense mutation; LEPR16 because of its short distance (30 bp) from the 3’UTR. Similarly, of the 26 SNP of the LEP gene, the following 4 were considered in the association analysis: LEP01 because of the significant associations reported by [7] on growth and fertility; LEP14, because of its position within the binding site of transcription factors (v-Maf and NF-E2), LEP25 and LEP26 because they encode a missense mutations.
In Table 5 and Table 6, the results of the association analysis are presented only for those SNP of LEP and LEPR that showed statistically significant effects, reporting also the effects of the same SNP on other traits, if trending to statistical significance (P < 0.09). Table 5 describes the association of 5 SNP with milk traits. Mean values and standard deviation for each trait are reported in the table, so to allow a direct evaluation of the extent of the allelic substitution effect. All these values fall in the range of the average production traits of Holstein cows in Italy.
Three SNP of the LEPR gene (LEPR03, LEPR09, and LEPR16) influenced ME fat content, with substitution effects ranging from 0.11 (P = 0.04) to 0.26 (P = 0.007); LEPR16 influenced also the test day fat content (0.38; P = 0.02). One SNP of LEP gene (LEP14) influenced the test day fat content (0.19; P = 0.02), and also showed a trend to influence ME fat content (0.10; P = 0.09). Furthermore, three SNP of the LEPR gene (LEPR01, LEPR09 and LEPR16) influenced protein content, with substitution effects ranging from 0.08 (P = 0.04) to 0.11 (P = 0.03) for ME protein content, and from 0.10 (P = 0.02) to 0.11 (P = 0.07) for the test day protein content.
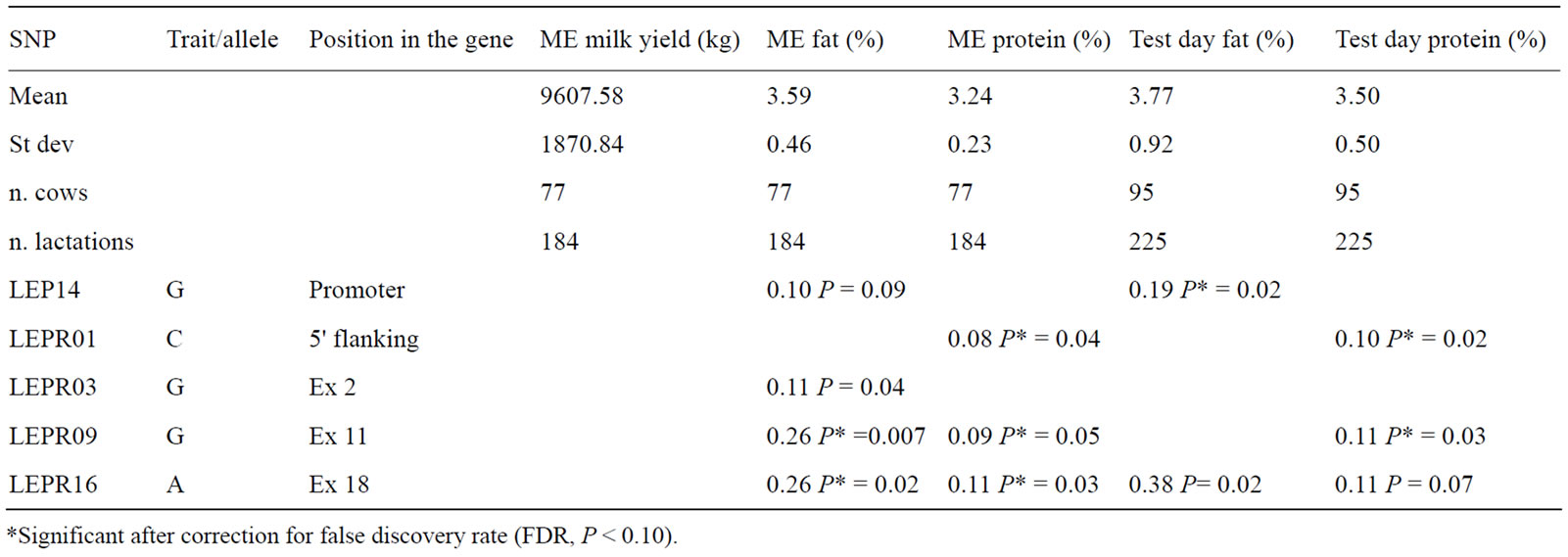
Table 5. Allele substitution effect of the SNP in the LEP and LEPR genes on milk production traits: mature equivalent (ME) and test day.
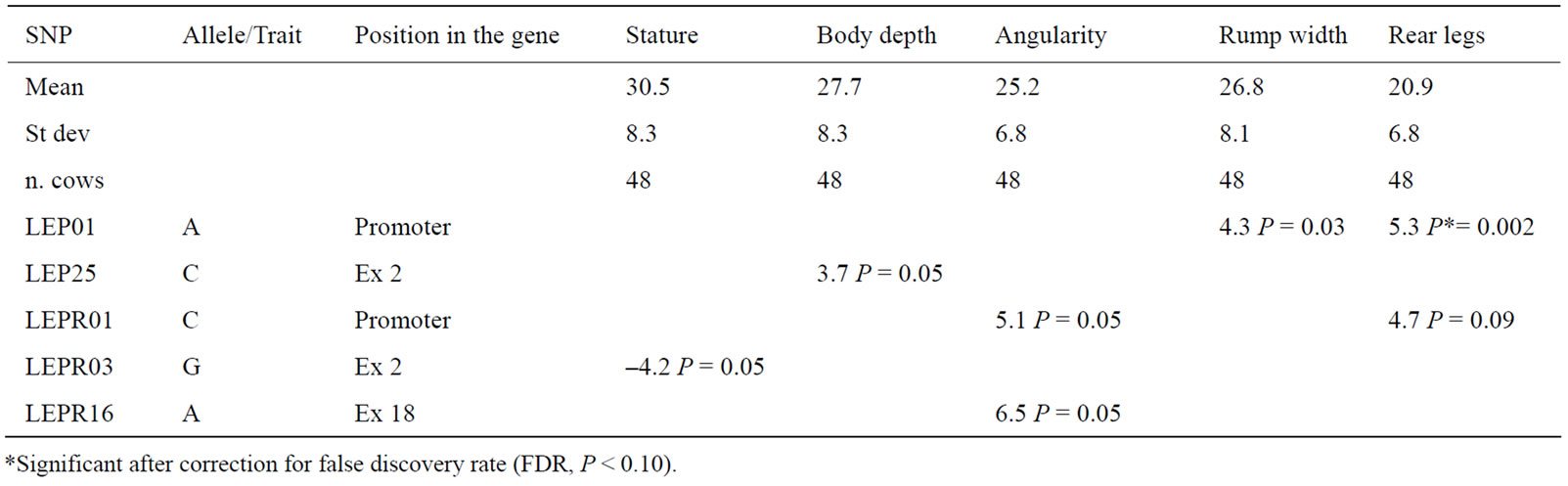
Table 6. Allele substitution effect of the SNP in the LEP and LEPR genes on morphological indexes expressed in points (pts). The effect refers to the allele of column 2.
In Table 6, the results of the association analysis between 5 SNP of the LEP and LEPR genes and morphological indexes, expressed in points (pts) are shown. These traits were included in this study under the hypothesis that the genes that regulate the fat storage physiology, like the LEP and LEPR genes, might affect some morphological traits; similarly, Waters [30] quantified the associations of some SNP in the genes of the somatotrophic axis with milk performance and conformation traits in Holstein cows. Three SNP of the LEPR gene (LEPR01, LEPR03, and LEPR16) showed a significant effect on either the angularity or the stature or the rear legs (Table 6). Only one SNP of the LEP gene (LEP01) affected rump width and rear legs with values respectively of 4.3 pts and 5.3 pts. None of the analyzed SNP affected significantly either the rump angle or the udder depth. In Table 7 and Table 8, separately for the SNP of the LEPR and the LEP genes, in order not to ascribe to a SNP the effect of another one, for the SNP that showed some effect on any of the traits, pair-wise allelic correlations were reported.
4. DISCUSSION
These results showed that fat content is the most affected trait by the LEPR and LEP gene polymorphisms, while protein content is significantly affected only by EPR gene polymorphisms. As regards to the LEPR gene, two SNP, LEPR09 and LEPR16, influence both fat and protein (Table 5). LEPR09 encodes a missense mutation in exon 11 causing the amino acids substitution R615S at residue 615. The Arginine is a polar and basic amino acid while the Serine is polar and neutral; moreover, the substitution falls in the extracellular domain, five residuals before the starting of the second W-S-X-W-S motif. Reference [31] reported that the W-S-X-W-S motifs are involved in the ligand binding. It is then possible to justify the role played by LEPR09 in the variation of fat and protein content. Also LEPR16 affects both parameters, but because it encodes a synonymous mutation, it is likely that such effect be due to the high pair-wise correlations with LEPR09 (0.87; P < 0.0001; Table 7). LEPR03 influenced only fat content; this SNP encodes the missense mutation, in exon 2, causing the amino acid substitution L35V at residue 35, this substitution falls in the extracellular region of LEPR molecule very near the putative signal peptide region. The evidence that the considered SNP affect particularly milk fat is corroborated by the function of the two genes in lipid regulation; however, because the considered animal sample was relatively small, the results should be regarded as an indication of the possible association of these SNP with the considered traits, that need to be further confirmed in independent dairy populations.
The results of this work indicated that also LEPR01 affected significantly protein content but the effect, very similar to the effect of LEPR09, is likely due to the partial and highly significant correlation between the two variants (Table 7).
It is interesting to note that the majority of the authors, that have performed association studies on the LEPR gene, have analyzed only the SNP LEPR15 (rs133672995) which encodes the missense mutation Threonine to Metionine at the 944 residue of the protein sequence (accession no. NP_001012285). Reference [5] found that this SNP significantly influenced blood concentration of circulating leptin; however, no significant association of this SNP with milk-related traits was found [8,32], in agreement with our results.
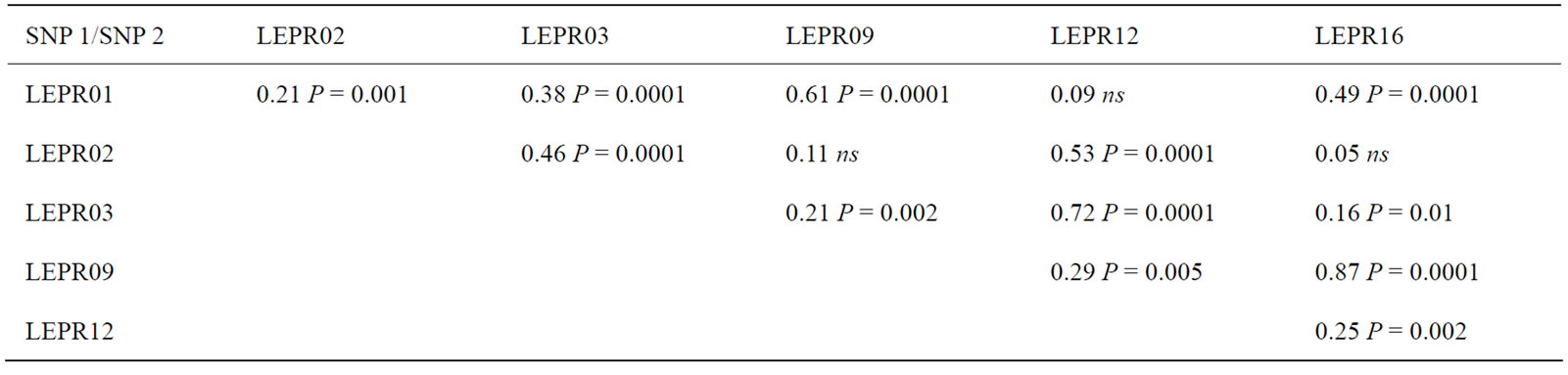
Table 7. Allelic correlations between the pairs of SNP of the LEPR gene that were included in the association analysis.
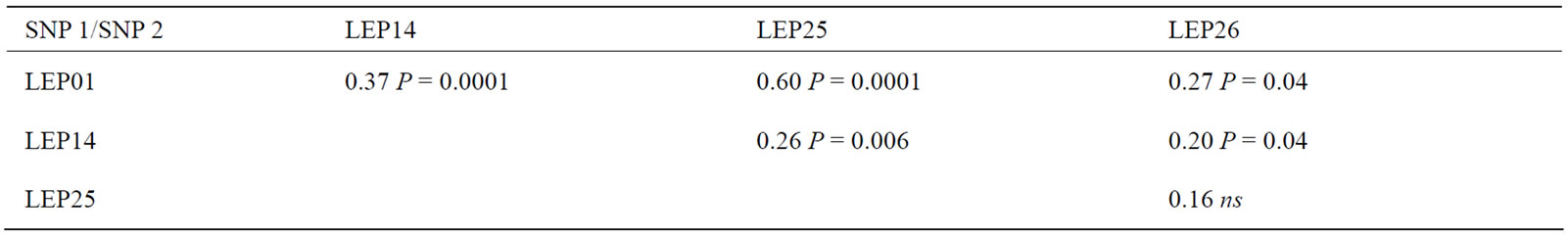
Table 8. Allelic correlations between the pairs of SNP of the LEP gene that were included in the association analysis.
Of the SNP of the LEP gene, only LEP14 showed significant effects on some of the considered traits. On the contrary, a significant effect of LEP25 on total milk yield was proposed by [19], while [16] found a significant effect of LEP09 on fat yield, and of LEP11 on protein content. Reference [33] showed that LEP09 and LEP25 were significantly associated with milk fat and protein content, while LEP26 was associated only with the survival in the herd. Reference [9] reported a significant effect of LEP01 and LEP09 with milk protein content. Finally, [7] investigated the associations between the SNP in the LEP and LEPR genes, with growth, milk production, and fertility traits and found that LEP01 was significantly associated with height at withers and with the total number of AI services in heifers, while LEP26 was significantly associated with crown rump length, age at first service, at daily milk yield and 305-d milk yield.
It is interesting to note that LEP14 falls in the core sequence of an important binding site of transcription factors: NF-E2 (gGCTGActttc). The NF-E2 is involved in the regulation of globin gene transcription [34] and shows a high degree of homology to the Nrf2 transcription factor [35] that plays a role in lipid metabolism [36, 37]. In fact, [38] reported that the expression of many genes, that are involved in the synthesis and catabolism of cholesterol and fatty acids, is inversely related to the amount of Nrf2. The Nrf2 might also modulate leptin expression, by interfering with lipogenic pathways, so corroborating the positive effect of allele G of LEP14 on milk fat content.
As regards to the association between the SNP and the morphology, some SNP of the LEPR gene seemed to influence both milk and morphological traits. In fact, the trait angularity, that summarizes the ideal dairy cow type, is positively influenced by the A allele of LEPR16, that showed a positive effect on both fat and protein content (Table 5), confirming that milk traits are positively correlated with the morphological parameters desired for the dairy cow type.
As regards to the LEP gene, we noted that the desired morphological parameters for the dairy cow (body depth, rump width and rear legs) are positively influenced by LEP01 and LEP25 (Table 6), and these SNP are also highly correlated (Table 8).
5. CONCLUSION
In this study, for the first time, the bovine LEPR gene was fully characterized by sequencing the whole coding region and part of 5’flanking region and several novel SNP were detected, some of which encode missense mutations; moreover, a novel SNP was detected in the promoter of the LEP gene. In this study, we described 26 SNP of the LEP gene and 16 SNP of the LEPR gene and we reported significant associations of some of them with important milk production and morphological traits. Previous authors had aimed to estimate the LEP and LEPR gene effects on milk production traits, but because only one or few SNP were considered in each study, the results were contrasting and difficult to be extended to other cow populations. To our knowledge, this is the first study that evaluated the effect of many SNP of the LEPR gene on dairy traits. Due to relatively small size of the considered animal sample, the detected effects need to be further confirmed in independent dairy populations; however, because the allele substitution effects of the SNP of both the genes were evident, in most cases, on fat content, the results of this study confirm the role that both genes play in influencing this trait.
6. ACKNOWLEDGEMENTS
This study is part of the GENZOOT research program, funded by the Italian Ministry of Agriculture (Rome, Italy).
REFERENCES
- Tartaglia, L.A. (1997) The leptin receptor. Journal of Biological Chemistry, 7, 6093-6096.
- Gorska, E., Popko, K., Stelmaszczyk-Emmel, A., Ciepiela, O. and Kucharska, A. (2010) Leptin recerptor. European Journal of Medical Research, 15, 50-54.
- Pfister-Genskow, M., Hayes, H., Eggen, A. and Bishop, M.D. (1997) The leptin receptor (LEPR) gene maps to bovine chromosome 3q33. Mammalian Genome, 8, 227. doi:10.1007/s003359900398
- Guo, Y., Chen, H., Lan, X., Zhang, B., Pan, C., Zhang, L., Zhang, C. and Zhao, M. (2008) Novel SNPs of bovine LEPR gene and their association with growth traits. Biochemical Genetics, 46, 828-834. doi:10.1007/s10528-008-9197-z
- Liefers, S.C., Verkamp, R.F., Te Pas, M. F.W., Delavaud, C., Chilliard, Y. and Van der Lende, T. (2004) A missense mutation in the leptin receptor gene is associated with leptin concentrations during late pregnancy. Animal Genetics, 35, 138-141. doi:10.1111/j.1365-2052.2004.01115.x
- Komisarek, J. and Dorynek, Z. (2006) The relationship between the T945M single nucleotide polymorphism in the leptin receptor gene (LEPR) and milk production traits in Jersey cows. Animal Science Papers and Reports, 4, 271-277.
- Clempson, A.M., Pollott, G.E., Brickell, J.S., Bourne, N.E., Munce, N. and Wathes, D.C. (2011) Evidance that leptin genotype is associated with fertility, growth, and milk production in Holstein cows. Journal of Dairy Science, 94, 3618-3628. doi:10.3168/jds.2010-3626
- Banos, G., Woolliams, J.A., Woodward, B.W., Forbes, A.B. and Coffey, M.P. (2008) Impact of single nucleotide polymorphisms in leptin, leptin receptor, growth hormone receptor, and diacylglycerol acyltransferase (DGAT1) gene loci on milk production, feed, and body energy traits of UK dairy cows. Journal of Dairy Science, 91, 3190- 3200. doi:10.3168/jds.2007-0930
- Glantz, M., Lindmark Månsson, H., Stålhammar, H. and Paulsson, M. (2011) Effect of polymorphisms in the leptin, leptin receptor and acyl-CoA:diacylglycerol acyltransferase 1 (DGAT1) genes and genetic polymorphism of milk proteins on bovine milk composition. Journal of Dairy Research, 30, 1-9.
- Forhead, A.J. and Fowden, A.L. (2009) The hungry fetus? Role of leptin as a nutritional signal before birth. Journal of Physiology, 587, 1145-1152. doi:10.1113/jphysiol.2008.167072
- Lagonigro, R., Wiener, P., Pilla, F., Woolliams, J.A. and Williams, J.L. (2003) A new mutation in the coding region of the bovine leptin gene associated with feed intake. Animal Genetics, 34, 371-374. doi:10.1046/j.1365-2052.2003.01028.x
- Liefers, S.C., Te Pas, M.F.W., Veerkamp, R.F. and van der Lende, T. (2002) Associations between leptin gene polymorphisms and production, live weight, energy balance, feed intake, and fertility in Holstein heifers. Journal of Dairy Science, 85, 1633-1638. doi:10.3168/jds.S0022-0302
- Stone, R.T., Kappes, S.M. and Beattie, C.W. (1996) The bovine homolog of the obese gene maps to chromosome 4. Mammalian Genome, 7, 399-400. doi:10.1007/s003359900119
- Taniguchi, Y., Itoh, T., Yamada, T. and Sasaki, Y. (2002) Genomic structure and promoter analysis of the bovine leptin gene. IUBMB Life, 53, 131-135. doi:10.1080/15216540211465
- Konfortov, B.A., Licence, V.E. and Miller, J.R. (1999) Re-sequencing of DNA from a diverse panel of cattle reveals a high level of polymorphism in both intron and exon. Mammalian Genome, 10, 1142-1145. doi:10.1007/s003359901180
- Liefers, S.C, Verkamp, R.F., Te Pas, M. F.W., Delavaud, C., Chilliard, Y., Platje, M. and Van der Lende, T. (2005) Leptin promoter mutations affect leptin levels and performance traits in dairy cows. Animal Genetics, 36, 111- 118. doi:10.1111/j.1365-2052.2005.01246.x
- Liefers, S.C., Te Pas, M.F.W., Veerkamp, R. F., Chilliard, Y., Delavaud, C., Gerritsen, R. and Van der Lende, T. (2003) Association of leptin gene polymorphisms with serum leptin concentration in dairy cows. Mammalian Genome, 14, 657-663. doi:10.1007/s00335-003-2275-y
- Nkrumah, J. D., Li, C., Yu, J., Hansen, C., Keisler, D.H. and Moore, S.S. (2005) Polymorphisms in the bovine leptin promoter associated with serum leptin concentration, growth, feed intake, feeding behavior, and measures of carcass merit. Journal of Animal Science, 83, 20-28.
- Buchanan, F.C., Van Kessel, A.G., Waldner, C., Christensen, D.A., Laarveld, B.S. and Schmutz, M. (2003) Hot topic: An association between a leptin single nucletotide polymorphism and milk and protein yield. Journal of Dairy Science, 86, 3164-3166. doi:10.3168/jds.S0022-0302begin_of_the_skype_highlighting
- Madeja, Z, Adamowicz, T., Chmurzynska, A., Jankowski, T., Melonek, J., Switonski, M. and Strabel, T. (2004) Short communication: Effect of leptin gene polymorphisms on breeding value for milk production traits. Journal of Dairy Science, 87, 3925-3927. doi:10.3168/jds.S0022-0302
- Corva, P.M., Fernandez Macedo, G.V., Soria, L.A., Papaleo Mazzucco, J., Motter, M., Villarreal Schor, A., Mezzadra, C.A., Melucci, L.M. and Miquel, M.C. (2009) Effect of leptin gene polymorphisms on growth, slaughter and meat quality traits of grazing Brangus steers. Genetics and Molecular Research, 8, 105-116. doi:10.4238/vol8-1gmr556
- Yang, D., Chen, H., Wang, X., Tian, Z., Tang, L., Zhang, Z., Lei, C., Zhang, L. and Wang, Y. (2007) Association of polymorphisms of leptin gene with body weight and body sizes indexes in Chinese indigenous cattle. Journal of Genetics and Genomics, 34, 400-405. doi:10.1016/S1673-8527
- Orrù, L., Abeni, F., Cavillo, G., Grandoni, F., Crisà, A. De Matteis, G., Scatà, M.C., Napoletano, F. and Mo’ioli, B. (2012) Leptin gene haplotypes are associated with change in immunological and hematological parameters in dairy cow during the peripartum period. Journal of Animal Science, 90, 16-26. doi:10.2527/jas.2010-3706
- Barendse, W., Harrison, B.E., Bunch, R.J. and Thomas, M.B. (2008) Variation at the Calpain 3 gene is associated with meat tenderness in zebu and composite breeds of cattle. BMC Genetics, 9, 41. doi:10.1186/1471-2156-9-41
- Sherman, E.L., Nkrumah, J.D., Murdoch, B.M, Li, C., Wang, Z. and Fu, A. (2008) Polymorphisms and haplotypes in the bovine NPY, GHR, GHRL, IGF2, UCP2, and UCP3 genes and their associations with measures of growth, performance, feed efficiency and carcass merit in beef cattle. Journal of Animal Science, 86, 1-16. doi:10.2527/jas.2006-799
- SAS (2007) Applied Statistics and the SAS Programming Language. Release 9.3, SAS Institute Inc., Cary.
- Kel, A.E., Gossling, E., Reuter, I., Cheremushkin, E., Kel-Margoulis, O.V. and Wingender E. (2003) MATCH: A tool for searching transcription factor binding sites in DNA sequences. Nucleic Acids Research, 31, 3576-3579. doi:10.1093/nar/gkg585
- Matys, V., Kel-Margoulis, O., Fricke, E., Liebich, I., Land, S., Barre-Dirrie, A., Reuter, I., Chekmenev, D., Krull, M., Hornischer, K., Voss, N., Stegmaier, P., Lewicki-Potapov, B., Saxel, H., Kel, A. E. and Wingender, E. (2006) TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Research, 34, 108-110. doi:10.1093/nar/gkj143
- Fong. T.M., Huang, R.R., Tota, M.R., Mao, C., Smith, T., Varnerin, J., Karpitskiy, V.V., Krause, J.E. and Van der Ploeg, L.H. (1998) Localization of leptin binding domain in the leptin receptor. Molecular Pharmacology, 53, 234- 240.
- Waters, S.M., Berry, D.P. and Mullen, M.P. (2012) Polymorphisms in genes of the somatotrophic axis are independently associated with milk production, udder health, survival and animal size in Holstein-Friesian dairy cattle. Journal of Animal Breeding and Genetics, 129, 70-78. doi:10.1111/j.1439-0388.2011.00938.x
- Kawachi H., Yang, S. H., Hamano, A., Matsui, T., Smith, S. B. and Yano, H. (2007) Molecular cloning and expression of bovine (Bos taurus) leptin receptor isoform mRNAs. Comparative Biochemistry and Physiology Part B: Biochemistry and Molecular Biology, 148, 167-173. doi:10.1016/j.cbpb.2007.05.007
- Liefers, S.C., Veerkamp, R.F., te Pas, M.F.W., Chilliard, Y. and Van der Leed, T. (2005) Genetics and physiology of leptin in periparturient dairy cows. Domestic Animal Endocrinology, 29, 227-238. doi:10.1016/j.domaniend.2005.02.009
- Giblin, L., Butler, S.T., Kearney, B.M., Waters, S.M., Callanan, M.J. and Berry, D.P. (2010) Association of bovine leptin polymorphisms with energy output and energy storage traits in progeny tested Holstein-Friesian dairy cattle sires. BMC Genetics, 29, 11-73.
- Andrews, N.C. (1998) Molecules in focus. The NFE2 transcription factor. The International Journal of Biochemistry & Cell Biology, 30, 429-432. doi:10.1016/S1357-2725(97)00135-0
- Moi, P., Chan, K., Asunis, I., Cao, A. and Kann, Y.W. (1994) Isolation of NF-E2-related factor 2 (Nrf2), a NF-E2-like basic leucine zipper transcriptional activator that binds to the tandem NF-E2/AP1 repeat of the betaglobin locus control region. The Proceedings of the National Academy of Sciences USA, 91, 9926-9930. doi:10.1073/pnas.91.21.9926
- Dyck, P.A., Hoda, F., Osmer, E.S. and Green, R.M. (2003) Microarray analysis of hepatic gene expression in gallstone-susceptible and gallstone-resistant mice. Mammalian Genome, 14, 601-610. doi:10.1007/s00335-003-2269-9
- Gao, L., Wang, J., Sekhar, K.R., Yin, H., Yared, N.F., Schneider, S.N., Sasi, S., Dalton, T.P., Anderson, M.E. and Chan, J.Y. (2007) Novel n-3 fatty acid oxidation products activate Nrf2 by destabilizing the association between Keap1 and Cullin 3. Journal of Biological Chemistry, 282, 2529-2537. doi:10.1074/jbc.M607622200
- Tanaka, Y., Aleksunes, L.M.,Yeager, R.L., Gyamfi, M.A., Esterly, N., Guo, G.L. and Klaassen, C.D. (2008) NF-E2- related factor 2 inhibits lipid accumulation and oxidative stress in mice fed a high-fat diet. Journal of Pharmacology and Experimental Therapeutics, 325, 655-664. doi:10.1124/jpet.107.135822
NOTES
1Test day milk fat percentage, test day milk protein percentage.

