Open Journal of Genetics
Vol.4 No.3(2014), Article
ID:47423,6
pages
DOI:10.4236/ojgen.2014.43024
QTL Positioning of Thousand Wheat Grain Weight in Qaidam Basin
Le Wei1*, Shenggui Bai2*, Jianmin Li1, Xiujuan Hou3, Xin Wang3, Hongqing Li3, Bo Zhang1,4, Wenjie Chen3, Dencai Liu3, Baolong Liu1,4#, Huaigang Zhang1,4#
1College of Biologic and Geographic Sciences, Qinghai Normal University, Xining, China
2Huangzhong County Grassland Station, Huangzhong, China
3Key Laboratory of Adaptation and Evolution of Plateau Biota (AEPB), Northwest Institute of Plateau Biology, Chinese Academy of Sciences, Xining, China
4Qinghai Province Key Laboratory of Crop Molecular Breeding, Xining, China
Email: #blliu@nwipb.cas.cn, #hgzhang@nwipb.cas.cn
Copyright © 2014 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 8 April 2014; revised 7 May 2014; accepted 6 June 2014
ABSTRACT
TGW is an important agronomic trait that influences the yield and milling quality of wheat. In this paper, after having investigated the phenotype of TGW from 114 ITMI recombinant inbred genealogies in 4 years in Qaidam Basin, a typical plateau oasis agricultural area, and having combined 1410 molecular markers, 7 major QTL loci of the thousand gain weight were screened out with the aid of QTL network software. These loci included qTgw1B (42.6 cm), qTgw2A (77.9 cm), qTgw2D1 (25.4 cm), qTgw2D2 (51.8 cm), qTgw6A1 (56.1 cm), qTgw6A2 (62.2 cm) and qTgw7A (75.7 cm) with their genetic contribution rates between 3.29% - 19.36%. There were two epistatic effect loci 2A-2D and 2A-6B with their genetic contribution rates as 2.3% and 5.3% respectively. The quantitative genetic locus positioning of thousand hexaploid wheat grain weight in Qaidam Basin can assist us in better understanding the genetic regulatory network formed by TGW, and can also provide a theoretical basis for improving thousand wheat grain weight in this ecological area.
Keywords:Triticum aestivum, TGW, QTL Analysis, Qaidam Basin

1. Introduction
Bread wheat (Common wheat) is one of the world’s most important food crops, providing 40% of the world’s food and heat. With the increasing number of the world’s population, the global demand for wheat will increase by 40% before 2020. Therefore, a method to solve this problem is to breed more productive varieties of wheat [1] . TGW is an important agronomic trait for wheat and is one of the three key elements that determine the wheat yield. As TGW increases, so does the wheat yield. Since 1940, TGW of Chinese wheat varieties has increased from the average 31.5 grams in the 1940s to 44.64 grams in 2000, with an addition of 2.19 grams every ten years [2] . At the same time, high TGW can improve the seed germination rate, promote the seedling growth and enhance the variety’s ability to resist natural disasters [3] . Furthermore, high TGW can also improve the flour yield and affect the milling quality of wheat grains [4] . Accordingly, high TGW is also regarded as a significant characteristic of common wheat domestication and modern wheat breeding [5] .
Thousand wheat grain weight is a complex quantitative genetic trait. The positioning of quantitative genetic loci is the preliminary work of molecular genetic mechanism to examine the size and trait of wheat grains; meanwhile, it benefits the molecular marker assistance selection that is carried out on the separating generation in the breeding process [6] . The statistics has already shown that by 2010 at least 116 QTL loci of thousand wheat grain weight were positioned and distributed on all the 21 pairs of wheat chromosomes [7] . Since the quantitative genetic trait is susceptible to natural conditions and environment, different QTL loci of thousand wheat grain weight usually play varied roles under different ecological conditions. Besides, due to differences in experimental materials and environments during different experiments, there exists no such an experiment that includes the whole QTL loci, nor does there exist a QTL locus that can be separated in all experiments [7] . Consequently, the excavation work of QTL loci should be implemented under more diversified ecological conditions so as to comprehensively analyze the genetic regulatory network QTL loci of thousand wheat grain weight.
The Qaidam Basin is an inland basin with its elevation ranging from 2600 meters to 3000 meters. This area is marked by high altitude, thin cloud cover, long insolation duration and perennial drought with little rain. It belongs to a typical plateau oasis agricultural area since spring wheat is produced by taking advantage of sufficient surface water and groundwater. Because of the large temperature difference between day and night, the wheat photosynthate accumulation is bettered and the wheat yield is heightened. So it is common to find the yield of spring wheat range between 11250 - 12750 kg/Ha. Moreover, high yield test and demonstration above the recorded 13500kg reached more than 30 times and there were 4 times to achieve the 15000 kg/Ha level, creating a record high of spring wheat production in our country and the world [8] . At present, domestic and foreign researchers have systematically and extensively studied both the physiological and ecological characteristics of spring wheat photosynthesis, panicle differentiation, grain filling and mineral nutrient absorption and utilization under the natural ecological conditions of the Qiadam Basin, and morphological selection, group features and physiological morphology of super-high yield wheat varieties in order to guide the new variety breeding and popularization of super-high yield wheat. However, no research on high yield trait in this area has been conducted from the perspective of genetics. The QTL analysis of thousand wheat grain weight in this area may lead to QTL loci different from those obtained in other ecological environments, thus explaining genetic regularities and characteristics of TGW.
2. Materials and Methods
2.1. Research Materials
The population used in this research is an ITMI population from 114 recombinant inbred genealogies, originally provided by Dr. R. P. Singh from CIMMYT in Mexico. Two parents and a population from 114 recombinant inbred genealogies were planted in Xiangride farm, Dulan County, Haixi Prefecture, Qinghai Province in 2010, 2011, 2012 and 2013. The length was 2 meters with a 20 cm spacing, which was repeated 3 times. The conventional field management was adopted. After harvest, 1000 wheat grains were chosen through counting machine to measure their weight.
2.2. Construction of Genetic Linkage Map
Frame map of ITMI population and separation data of 1410 molecular markers were retrieved from the GrainGenes database (http://wheat.pw.usda.gov). This map integrated 1410 molecular markers, including BARC, RFLP, AFLP, SSR and so on. It covered the 21 pairs of wheat chromosomes in genomes A, B and D with a total length of 3551 cm and the average distance between markers being 1.8 cm. On average, there were more than 60 marker loci on each chromosome.
2.3. Statistical Analysis and QTL Mapping
Phenotype data of thousand wheat grain weight were statistically analyzed through SPSS software and mapping was completed through SigmaPlot12 software. Genetic map was constructed by using 1410 SSR markers and QTL analysis was conducted by using QTLNetwork2.0 software based on mixed linear model. Taking P = 0.005 as the statistical detection threshold, when result comes out after the end of the software operation, it is considered that there will be 1 QTL related to traits on the marker should the marked P value be below the statistical detection threshold.
3. Research Results
3.1. Phenotype Differences of TGW
Table 1 showed the ITMI population performance values of TGW in Qaidam Basin. The maximum and minimum values of the average TGW in each year shared the same trend. The average TGW of Parent Opata and W7984 in 4 years were 43.27 and 38.27, within the range from the maximum value 60.31 to the minimum value 35.98, thus indicating a bidirectional transgressive segregation. The ITMI population phenotype of TGW in 4 years all showed a continuous distribution in line with a normal distribution in statistics with P < 0.05. There may be several quantitative trait locus controls (though data were not shown). Figure 1 is the frequency distribution diagram for the population in 4 years from 2010 to 2013.
3.2. Additive QTL Loci
Seven additive QTL loci were detected in the research during these 4 years, and they were qTgw1B (42.6 cm), qTgw2A (77.9 cm), qTgw2D1 (25.4 cm), qTgw2D2 (51.8 cm), qTgw6A1 (56.1 cm), qTgw6A2 (62.2 cm) and qTgw7A (75.7 cm) respectively ( Table 2). Among them, qTgw6A1 could be detected in 2010 and 2013 with its genetic contribution rates as 19.36% and 14.93% respectively. In 2012, the genetic contribution rates of qTgw6A2 Table 1. TGW of the RIL population. Figure 1. Frequency Distribution of TGW in ITMI Population. and qTgw7A were detected as 15.12% and 11.31% respectively. In 2013, the genetic contribution rate of qTgw2D was detected as 28.1%. When the data in 4 years were integrated together to carry out a QTL analysis, qTgw1B, qTgw2A, qTgw2D2 and qTgw6A2 could be detected with their genetic contribution rates as 3.49%, 6.91%, 3.29% and 11.24% respectively (Table 2). When the data collected in 2011 were used to carry out a QTL analysis, obvious QTL loci were not found. This probably was owing to the damage caused to the irrigation canals in planting area, which further caused widespread drought in wheat seedling and mature stages, thus affecting the wheat growth. Moreover, the additive alleles of qTgw2D1, qTgw2D2, qTgw6A1 and qTgw6A2 came from artificially synthesized wheat ITMI, while other three additive effects came from common wheat Opata. 3.3. Epistatic QTL During the period from 2010 to 2014, two significant epistatic effect loci 2A-2D and 2A-6B existed in ITMI wheat population with their genetic contribution rates as 2.3% and 5.3% respectively ( Table 3). No interaction effect between epistasis and environment was detected. The total contribution rate of epistatic effect reached 7.6%. 4. Discussions In this study, positioned qTgw6A1 (56.1 cm) and qTgw6A2 (62.2 cm) in genetic map belonged to the same section with a 6.1 spacing in between. This was probably due to annual differences of TGW effect. Without doubt, it might also because they were two different major QTL loci in themselves. A major gene GW2-6A of TGW has been positioned in wheat chromosomes [3] [9] . However, owing to parent differences between different populations and differences among the used molecular markers, it is hard for us to conform whether or not this gene was in the same locus with qTgw6A in this paper. This case can also be used to explain all QTL loci we have positioned. The quantitative genetic trait is characterized by its susceptibility to environmental factors. Generally speaking, positioned quantitative genetic trait loci often differ from each other resulting from varied sites and periods. Since researchers are confined by regions they belong to, QTL positioning experiments of TGW are more often than not conducted in rain-fed or saturated irrigation agricultural area with a lower altitude and better climate. In contrast, research work carried out in high altitude inland basin is relatively less. Qaidam Basin in this study belongs to an oasis agricultural area of plateau inland basin with its unique climatic conditions. During 1970-2000, the average annual temperature is 3.18˚C, the average rainfall is 193.8 mm with the annual evaporation amount reaching 2285.4 mm, the relative humidity is 40.2% and the sunshine duration is 3091.8 hours. With sowing period in April 15th and harvest period in September 15th, it enjoys a spring wheat growth period of 150 days. Once in an area of 0.26 Ha, the average amount of effective panicles arrived at 516300 per Ha, every panicle had 36.18 grains and TGW reached 56.2 grams. After being actually collected, the total output reached 3961.03 kilograms with the average output as 15195.75 kilograms per Ha. Therefore, QTL loci of TGW carried out in this area could possibly be found differ from those in other areas. Table 3. Estimated epistasis (AA) of QTLs for TGW. ITMI recombinant inbred population (W7984 × Opata85) was a mapping population used by International Triticeae Mapping Initiative to construct a wheat genetic linkage map. It had significant parental genetic differences of agronomic traits and 1410 molecular markers were established. On average, there were more than 60 marker loci on each chromosome, achieving a saturated degree. Thanks to the above mentioned advantages, scientists have already conducted QTL analyses on several wheat quantitative trait phenotypes by using this population. A major QTL with wheat macular disease was located in 1AS and a minor QTL in 4AL. Besides, there existed an epistatic effect between 1AS and 2DL [10] . Gene Pm15 with Blumeria graminis was positioned in 7D and 4B, gene Lr34 in 7D, and glutenin gene Gpc in 7A [11]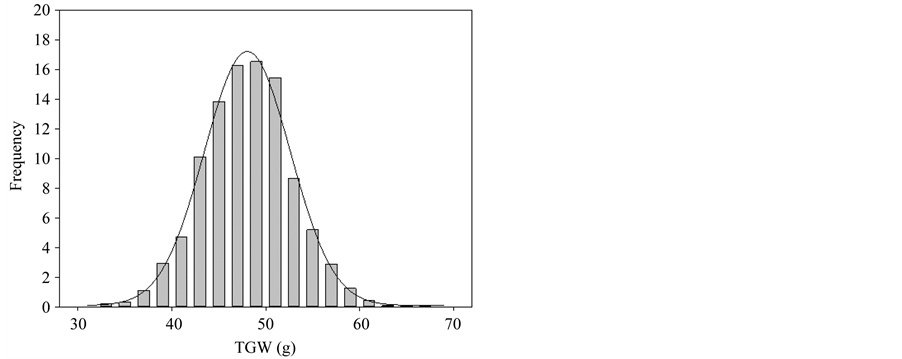
Artificially synthesized hexaploid wheat resulted from cross-breeding between tetraploid wheat (2n = 28) and diploid Aegilops (2n = 14) as well as from chromosome doubling. Because of selection effect, bottleneck effect and long-term selection, genetic basis of modern wheat is narrowing. On the other hand, two parents tetraploid wheat and diploid Aegilops of artificially synthesized wheat carried disease-resistance gene and gene increasing yield [14] . When gene of artificially synthesized hexaploid wheat was introduced in common wheat, wheat quality and quantity was improved [13] [15] -[18] . In this study, parent Opata55 of ITMI population was a common Mexican wheat variety bred by CIMMYT and parent W7984 (M6) was an artificially synthesized variety cross-bred between Altar 84 (T. durum, AABB) and TA 2465 (Aegilops tauschii, DD) with chromosome doubling [19] . In fact, additive alleles of positioned qTgw2D2, qTgw 2D1, qTgw6A1 and qTgw6A2 in Qiadam Basin came from artificially synthesized wheat W7984. Therefore, these QTL loci are probably excellent allelic variations that modern wheat lack at present. And these excellent allelic variations can be directly applied into new wheat variety breeding in a high-altitude oasis agricultural area.
5. Conclusions
The phenotype of TGW of ITMI population in the typical plateau oasis agricultural area, a quantitative genetic trait, was conformed to a normal distribution. In this area, 7 major QTL loci of the thousand gain weight were positioned, including qTgw1B (42.6 cm), qTgw2A (77.9 cm), qTgw2D1 (25.4 cm), qTgw2D2 (51.8 cm), qTgw6A1 (56.1 cm), qTgw6A2 (62.2 cm) and qTgw7A (75.7 cm) with their genetic contribution rates between 3.29% - 19.36%. There were two epistatic effect loci 2A-2D and 2A-6B with their genetic contribution rates as 2.3% and 5.3% respectively.
Acknowledgements
This research is financially supported by the National Natural Science Foundation of China (no. 31071417, 31260322), the West Light Foundation of Chinese Academy of Sciences, State Key Laboratory Breeding Base for Innovation and Utilization of Plateau Crop Germplasm and Pilot Projects of Designer Breeding by Molecular Module.
References
- Dixon, J.M. (2009) Wheat Facts and Futures 2009. CIMMYT (International Maize and Wheat Improvement Center).
- Wang, L.F., Ge, H.M., Hao, C.Y., Dong, Y.S. and Zhang, X.Y. (2012) Identifying Loci Influencing 1,000-Kernel Weight in Wheat by Microsatellite Screening for Evidence of Selection during Breeding. PLoS ONE, 7, e29432.
- Zhang, K.P., Wang, J.J., Zhang, L.Y., Rong, C.W., Zhao, F.W., Peng, T., Li, H.M., Cheng, D.M., Liu, X., Qin, H.J., et al. (2013) Association Analysis of Genomic Loci Important for Grain Weight Control in Elite Common Wheat Varieties Cultivated with Variable Water and Fertiliser Supply. PLoS ONE, 8, e57853.
- Campbell, K.G., Bergman, C.J., Gualberto, D.G., Anderson, J.A., Giroux, M.J., Hareland, G., Fulcher, R.G., Sorrells, M.E. and Finney, P.L. (1999) Quantitative Trait Loci Associated with Kernel Traits in a Soft x Hard Wheat Cross. Crop Science, 39, 1184-1195. http://dx.doi.org/10.2135/cropsci1999.0011183X003900040039x
- Peng, J.H., Ronin, Y., Fahima, T., Roder, M.S., Li, Y.C., Nevo, E. and Korol, A. (2003) Domestication Quantitative Trait Loci in Triticum dicoccoides, the Progenitor of Wheat. Proceedings of the National Academy of Sciences of the United States of America, 100, 2489-2494. http://dx.doi.org/10.1073/pnas.252763199
- Kumar, N., Kulwal, P.L., Gaur, A., Tyagi, A.K., Khurana, J.P., Khurana, P., Balyan, H.S. and Gupta, P.K. (2006) QTL Analysis for Grain Weight in Common Wheat. Euphytica, 151, 135-144. http://dx.doi.org/10.1007/s10681-006-9133-4
- Zhang, L.Y., Liu, D.C., Guo, X.L., Yang, W.L., Sun, J.Z., Wang, D.W. and Zhang, A.M. (2010) Genomic Distribution of Quantitative Trait Loci for Yield and Yield-related Traits in Common Wheat. Journal of Integrative Plant Biology, 52, 996-1007. http://dx.doi.org/10.1111/j.1744-7909.2010.00967.x
- Chen, D. (1993) Eco-Physiological Characteristics and Cultivation Techniques of High Yield of Spring Wheat in Qaidam Basin. In: Su, L., Ed., New Progress in China’s Cultivation of Wheat, China Agriculture Press, Beijing, 327-340.
- Su, Z.Q., Hao, C.Y., Wang, L.F., Dong, Y.C. and Zhang, X.Y. (2011) Identification and Development of a Functional Marker of TaGW2 Associated with Grain Weight in Bread Wheat (Triticum aestivum L.). Theoretical and Applied Genetics, 122, 211-223. http://dx.doi.org/10.1007/s00122-010-1437-z
- Faris, J.D., Anderson, J.A., Francl, L.J. and Jordahl, J.G. (1997). RFLP Mapping of Resistance to Chlorosis Induction by Pyrenophora Tritici-Repentis in Wheat. Theoretical and Applied Genetics, 94, 98-103. http://dx.doi.org/10.1007/s001220050387
- Borner, A., Schumann, E., Furste, A., Coster, H., Leithold, B., Roder, M.S. and Weber, W.E. (2002) Mapping of Quantitative Trait Loci Determining Agronomic Important Characters in Hexaploid Wheat (Triticum aestivum L.). Theoretical and Applied Genetics, 105, 921-936. http://dx.doi.org/10.1007/s00122-002-0994-1
- Nelson, J.C., Singh, R.P., Autrique, J.E. and Sorrells, M.E. (1997) Mapping Genes Conferring and Suppressing Leaf Rust Resistance in Wheat. Crop Science, 37, 1928-1935. http://dx.doi.org/10.2135/cropsci1997.0011183X003700060043x
- Huang, X., Kempf, H., Ganal, M. and Röder, M. (2004) Advanced Backcross QTL Analysis in Progenies Derived from a Cross between a German Elite Winter Wheat Variety and a Synthetic Wheat (Triticum aestivum L.). Theoretical and Applied Genetics, 109, 933-943. http://dx.doi.org/10.1007/s00122-004-1708-7
- Del Blanco, I., Rajaram, S. and Kronstad, W. (2001) Agronomic Potential of Synthetic Hexaploid Wheat-Derived Populations. Crop Science, 41, 670-676. http://dx.doi.org/10.2135/cropsci2001.413670x
- Del Blanco, I., Rajaram, S., Kronstad, W. and Reynolds, M. (2000) Physiological Performance of Synthetic Hexaploid Wheat-Derived Populations. Crop Science, 40, 1257-1263. http://dx.doi.org/10.2135/cropsci2000.4051257x
- Kunert, A., Naz, A.A., Dedeck, O., Pillen, K. and Léon, J. (2007) AB-QTL Analysis in Winter Wheat: I. Synthetic Hexaploid Wheat (T. turgidum ssp. Dicoccoides × T. tauschii) as a Source of Favourable Alleles for Milling and Baking Quality Traits. Theoretical and Applied Genetics, 115, 683-695. http://dx.doi.org/10.1007/s00122-007-0600-7
- Liu, S., Zhou, R., Dong, Y., Li, P. and Jia, J. (2006) Development, Utilization of Introgression Lines Using a Synthetic Wheat as Donor. Theoretical and Applied Genetics, 112, 1360-1373. http://dx.doi.org/10.1007/s00122-006-0238-x
- Narasimhamoorthy, B., Gill, B., Fritz, A., Nelson, J. and Brown-Guedira, G. (2006) Advanced Backcross QTL Analysis of a Hard Winter Wheat× Synthetic Wheat Population. Theoretical and Applied Genetics, 112, 787-796. http://dx.doi.org/10.1007/s00122-005-0159-0
- Zwart, R., Thompson, J., Sheedy, J. and Nelson, J. (2006) Mapping Quantitative Trait Loci for Resistance to Pratylenchus thornei from Synthetic Hexaploid Wheat in the International Triticeae Mapping Initiative (ITMI) Population. Crop and Pasture Science, 57, 525-530. http://dx.doi.org/10.1071/AR05177
NOTES

*These persons contribute to this manuscript equally.
#Corresponding authors.


