Open Journal of Genetics
Vol.1 No.3(2011), Article ID:8892,5 pages DOI:10.4236/ojgen.2011.13011
Identification of the keratin-associated protein 13-3 (KAP13-3) gene in sheep
1Gene-Marker Laboratory, Faculty of Agriculture and Life Sciences, Lincoln University, Lincoln, New Zealand;
2Lincoln Research Centre, AgResearch Limited, Lincoln, New Zealand.
Email: *jon.hickford@lincoln.ac.nz
Received 10 September 2011; revised 26 October 2011; accepted 13 November 2011.
Keywords: Wool; KAP13-3 Gene (KRTAP13-3); Variation; PCR-SSCP
ABSTRACT
Keratin-associated proteins (KAPs) are a major structural component of hair and wool fibres, and play a critical role in determining the properties of the fibre. To date, forty functional high sulphur KAP genes from fourteen families have been identified in humans, but only seven functional high sulphur KAP genes have been identified in sheep. This led us to search for the ovine KAP13-3 gene, a gene encoding a high sulphur KAP. In this study, the notional KAP13- 3 gene (KRTAP13-3) was amplified using primers designed based on a reported bovine KRTAP13-3 sequence. PCR-single stranded conformational polymorphism (PCR-SSCP) analysis was used to screen amplicons derived from the gene in one hundred and forty seven New Zealand Romney cross-bred sheep. Five unique banding patterns were revealed. Either one PCR-SSCP pattern (homozygous) or a combination of two patterns (heterozygous) was observed for each sheep. Sequencing of PCR amplicons representtative of different SSCP patterns revealed five different DNA sequences. The sequences derived from the amplicons showed a low homology to other known ovine KRTAPs, but had a high homology with previous reported KRTAP13-n sequences from human and cattle, with the closest homology being with bovine KRTAP13-3, suggesting the sequences represent the ovine KRTAP13-3 locus. Among the five allele sequences, four nucleotide substitutions were identified within the coding region. Of these substitutions, three were non-synonymous and would result in amino acid changes (p.Arg79Cys, p.Arg81Gln and p.Tyr130His). This variation in the KAP13-3 gene may affect gene expression, the structure and assembly of the protein, and consequently influence wool traits, if KAP13-3 is of importance to wool fibre structure.
1. INTRODUCTION
Keratin-associated proteins (KAPs) are a key structural component of hair and wool fibres and cross-link with the keratin intermediate filament proteins via extensive disulfide bonding [1]. They are believed to play a critical role in defining the physico-mechanical properties of the hair and wool fibres. KAPs characteristically possess either a high cysteine or high glycine-tyrosine content. Generally, based on their amino acid composition, they can be divided into three broad groups: high sulphur (HS; ≤30 mol% cysteine), ultra-high sulphur (UHS; >30 mol% cysteine) and high glycine-tyrosine (HGT), although this classification is arbitrary and becoming inadequate [1] as increasing variation is found in these genes.
KAPs can be further subdivided into families based on nucleotide and amino acid sequence homology [1], and to date a total of 27 families have been identified across species [2,3]. Of these, KAP1-KAP3, KAP10-KAP16 and KAP23-KAP27 are HS families, KAP4, KAP5, KAP9 and KAP17 are UHS families, and KAP6-KAP8 and KAP18-KAP22 are HGT families [2,3].
While in humans there are fourteen HS-KAP families consisting of forty functional KAP genes [4], only seven functional ovine HS-KAP genes from three families have been reported to date. These ovine HS-KAP genes are KRTAP1-1 [5], KRTAP1-2 [6], KRTAP1-3 [5], KRTAP1-4 [5], KRTAP3-2 [7], KRTAP3-3 [7] and KRTAP11-1 [8]. Many homologues of the human HS-KAP genes remain unidentified in sheep.
In this study, we report the identification of an ovine homologue of the KAP13-3 gene (KRTAP13-3) and the search for genetic variation in the gene using PCR-single strand conformational polymorphism analysis.
2. MATERIALS AND METHODS
2.1. Sheep Investigated and DNA Isolation
One hundred and forty seven New Zealand Romney cross-bred sheep were investigated. Blood samples were collected onto FTA cards (Whatman BioScience, Middlesex, UK) and genomic DNA was purified using a two-step washing procedure as described in Zhou et al. [9].
2.2. PCR Primers and Amplification
Two PCR primers (5’-tacattcaaactcagaatcttc-3’ and 5’- tgaatttggctcttctacaag-3’) were designed to amplify the entire coding region of ovine KRTAP13-3, based on a reported bovine KRTAP13-3 sequence (ENSBTAG00000040032). Primers were synthesized by Integrated DNA Technologies (Coralville, IA, USA).
Amplification was performed in a 20 μL reaction containing the genomic DNA on one 1.2 mm punch of FTA paper, 0.25 μM of each primer, 150 μM of each dNTP (Eppendorf, Hamburg, Germany), 2.5 mM of Mg2+, 0.5 U of Taq DNA polymerase (Qiagen, Hilden, Germany) and 1× reaction buffer supplied. The thermal profile consisted of 2 min at 94oC, followed by 35 cycles of 30 s at 94˚C, 30 s at 60˚C and 40 s at 72˚C, with a final extension of 5 min at 72˚C. Amplification was carried out in an iCycler (Bio-Rad, Hercules, CA, USA).
Amplicons were visualized by electrophoresis in 1% agarose (Quantum Scientific, Queensland, Australia) gels, using 1 × TBE buffer (89 mM Tris, 89 mM boric acid, 2 mM Na2EDTA) containing 200 ng/mL of ethidium bromide.
2.3. Single Strand Conformational Polymorphism Analysis
A 0.7-mL aliquot of each amplicon was mixed with 7 mL of loading dye (98% formamide, 10 mM EDTA, 0.025% bromophenol blue, 0.025% xylene-cyanol). After denaturation at 95˚C for 5 min, samples were rapidly cooled on wet ice and then loaded on 16 cm ´ 18 cm, 8% acrylamide:bisacrylamide (37.5:1) (Bio-Rad) gels. Electrophoresis was performed using Protean II xi cells (BioRad), at 280 V for 18 h at 15˚C in 0.5 ´ TBE buffer. Gels were silver-stained according to the method of Byun et al. [10].
2.4. Sequencing of Allelic Variants and Sequence Analysis
PCR amplicons representative of different SSCP patterns from sheep that appeared to be homozygous at KRTAP13-3 were directly sequenced at the Lincoln University DNA Sequencing Facility. For those alleles that were only found in heterozygous sheep, they were sequenced using a rapid approach described previously in Gong et al. [11]. In this approach, a band corresponding to the allele was excised as a gel slice from the polyacrylamide gel, macerated, and then used as a template for re-amplification with the original primers. This second amplicon was purified and then sequenced in both directions.
Sequence alignments, translations and comparisons were carried out using DNAMAN (version 5.2.10, Lynnon BioSoft, Vaudreuil, Canada). The BLAST algorithm was used to search the NCBI GenBank (http://www.ncbi. nlm.nih.gov/) databases for homologous sequences. Potential phosphorylation sites were predicted using the NetPhos 2.0 Server (http://www.cbs.dtu.dk/services/NetPhos/).
3. RESULTS
All the sheep blood samples generated PCR amplicons of the expected size (594 bp), and these amplicons exhibited different banding patterns upon SSCP analysis. Either one or a combination of two patterns was observed for each sheep, and in total, five different patterns were identified (Figure 1).
Sequencing of PCR amplicons representative of different SSCP patterns revealed five different DNA sequences. All of these sequences showed a low sequence homology to other known ovine KRTAPs, but were homologous to KRTAP13-n sequences in human and cattle, with the closest homologue being bovine KRTAP13-3. These ovine sequences were named KRTAP13-3 alleles A-E and deposited in the NCBI GenBank with the accession numbers JN377429-JN377433.
Among the sheep investigated, alleles A, B and C were common and occurred at frequencies of 28.9%, 35.0% and 32.0%, respectively. Alleles D and E were minor alleles, and both were detected at a frequency of 2.0%.

Figure 1. PCR-SSCP of the ovine KAP13-3 gene. Sheep carrying genotypes representative of the five unique PCR-SSCP patterns corresponding to five allele sequences A-E are shown.
Four nucleotide substitutions were identified within the ovine KRTAP13-3 coding region (Table 1). Of these substitutions, three were non-synonymous and would result in amino acid changes (p.Arg79Cys, p.Arg81Gln and p.Tyr130His).
The ovine KRTAP13-3 sequences all contained an open reading frame of 571 nucleotides, encoding a notional polypeptide of 156 amino acids. This polypeptide sequence was homologous with both human (accession numbers NM_181599.2, NM_181621.3, NM_181622.1 and NM_181600.1) and bovine KAP13-n sequences, but was homologous to bovine KAP13-3 (ENSBTAG000 00040032) than bovine KAP13-1 (ENSBTAG0000 0003613) (Figure 2). The ovine KAP13-3 polypeptide contained a high level of serine (22.44 mol%), cysteine (11.54 mol% - 12.18 mol%), arginine (10.26 mol% - 11.54 mol%) and glycine (9.62 mol%), and these were the predominant amino acids, making up 53.82 mol% - 55.78 mol% of the total amino acid content. Aspartic acid, glutamic acid, methionine and tryptophan were rare (accounting for 0.64 mol% each) and alanine was absent from the polypeptide. The notional polypeptide sequences had calculated isoelectric point (pI) values of 9.41, 9.26, 9.28, 9.04 and 9.41 for variants A to E, respectively. Between 18 to 21 serine and threonine resides in the notional ovine KAP13-3 polypeptide were predicted to be phosphorylated (Figure 2).
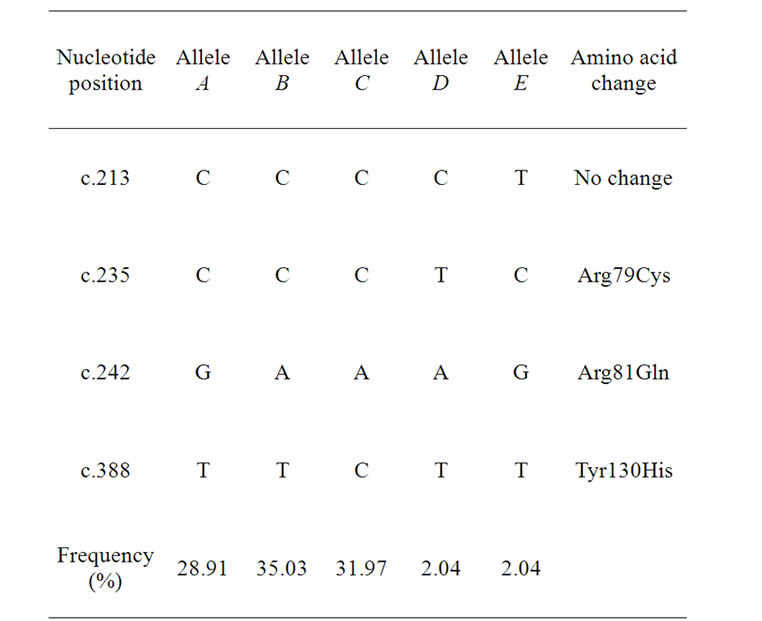
Table 1. Sequence variation and allele frequencies identified in the ovine KAP13-3 gene.

Figure 2. Amino acid sequence alignment of the ovine KAP13-3 sequences with known KAP13-n sequences from cattle and human. The amino acid sequences are predicted from nucleic acid sequences and are presented as a one-letter code. Amino acids identical to the top sequence are presented by dashes, and dots have been introduced to improve the alignment. Predicted phosphorylated residues in the ovine KAP13-3 sequences are shaded. The ovine sequences (accession numbers JN377429-JN377433 for alleles A-E, respectively) are indicated with a prefix “o”, the bovine sequences (ENSBTAG00000003613 and ENSBTAG00000040032 for KAP13-1 and KAP13-3, respectively) are indicated with a prefix “b”, and the human sequences (accession numbers NM_181599.2, NM_181621.3, NM_181622.1and NM_181600.1 for KAP13-1, KAP13-2, KAP13-3 and KAP13-4, respectively) are indicated with a prefix “h”.
4. DISCUSSION
This study has identified a notional KAP13-3 gene in sheep and investigated the variation in its coding region. The newly identified gene was intronless and contained an open reading frame encoding a cysteine-rich (11.54 mol% - 12.18 mol%) polypeptide. These characteristics are consistent with other HS-KAPs.
Some unique features were observed with the notional ovine KAP13-3 gene. Firstly, the putative polypeptide sequence had a high content (over 28 mol%) of serine and threonine collectively, with many of these residues (41% - 45%) notionally being able to be phosphorylated. This pattern is similar to ovine KRTAP11-1 of Gong et al. [8], but different from other known HS-KAP genes. While a comparable level of serine and threonine is also found in the KAP1 family members, but with the exception of KAP1-4, the proportion of these two amino acids that could potentially be phosphorylated in the putative KAP13-3 is two to three times higher than those KAP1 family members. While phosphorylation has not yet been investigated in KAPs, Ku et al. [12] reported that phosphorylation in keratins can affect keratin assembly and organization.
The putative ovine KAP13-3 appeared to be a basic protein. It contained many more (14.3 mol% - 15.5 mol%) positively charged amino acids (arginine, lysine and histidine) than negatively charged ones (aspartic acid and glutamic acid; 1.2 mol% in total), resulting in a high (9.0 - 9.3) calculated pI value. Such a high pI value has not been observed for any other known HS-KAP where pIs are typically less than 7.
There are two types of keratins that cross-link with KAPs, and of these, the type I keratins are typically more acidic (pI 4.5 - 6.0), while the type II keratins tend to be more basic (pI 6.5 - 8.5) [13]. The high pI value of KAP13-3 may accordingly affect its interaction with keratins, and on a charge basis alone it would appear to have a greater affinity for the type I (acidic) keratins.
The ovine sequences identified were more homologous to bovine KRTAP13-3 than to bovine KRTAP13-1 (the only other known family member in cattle), suggesting that these sequences represent the ovine KRTAP13-3 locus. However, while ovine KRTAP13-3 showed a reasonable degree of homology to human KRTAP13-n sequences, it did not cluster to any particular human KRTAP13-n gene. This suggests that genes of the KAP13 family emerged before the divergence of sheep and cattle, but after the divergence of primate and the artiodactyle mammals. Investigation of this gene and homologues from diverse species would shed further light on the evolution of KRTAPs.
Four nucleotide substitutions were identified in the ovine KRTAP13-3 coding region, and three of them would result in amino acid changes. It is interesting to note that some amino acid changes may result in the gain or loss of potential phosphorylation sites and also result in variation in the net charge of the protein. This variation in ovine KRTAP13-3 may affect the expression, structure and assembly of the protein and consequently influence key wool traits that underpin its quality and value.
5. ACKNOWLEDGEMENTS
This work was financially supported by FRST (C10X0710) and the Lincoln University Gene-Marker Laboratory. The Wool Research Organisation of New Zealand Inc. and New Zealand Wool Industry Charitable Trust Postgraduate Scholarship to H. Gong is acknowledged. The authors thank Q. Fang, S. Byun and G. Yang for technical assistance.
REFERENCES
- Powell, B.C. and Rogers, G.E. (1997) The role of keratin proteins and their genes in the growth, structure and properties of hair. In: Jolles, P., Zahn, H. and Hocker, E. Eds., Formation and Structure of Human Hair, Birkhauser Verlag, Basel, 59-148.
- Rogers, G.E. (2006) Biology of the wool follicle: An excursion into a unique tissue interaction system waiting to be re-discovered. Experimental Dermatology, 15, 931-949.
- Rogers, M.A., Winter, H., Langbein, L., et al. (2007) Characterization of human KAP24.1, a cuticular hair keratin-associated protein with unusual amino-acid composition and repeat structure. Journal of Investigative Dermatology, 127, 1197-204.
- Rogers, M.A. and Schweizer, J. (2005) Human KAP genes, only half of it? Extensive size polymorphisms in hair keratin associated genes. Journal of Investigative Dermatology, 124, vii-ix.
- Powell, B.C., Sleigh, M.J., Ward, K.A., et al. (1983) Mammalian keratin gene families: Organisation of genes coding for the B2 high-sulphur proteins of sheep wool. Nucleic Acids Research, 11, 5327-5346.
- Gong, H., Zhou, H., Yu, Z., et al. (2011) Identification of the ovine keratin-associated protein KAP1-2 gene (KRTAP1-2). Experimental Dermatology, In press. doi:10.1111/j.1600-0625.2011.01333.x
- Frenkel, M.J., Powell, B.C., Ward, K.A., et al. (1989) The keratin BIIIB gene family: Isolation of cDNA clones and structure of a gene and a related pseudogene. Genomics, 4, 182-191.
- Gong, H., Zhou, H., Dyer J.M., et al. (2011) Identification of the ovine KAP11-1 gene (KRTAP11-1) and genetic variation in its coding sequence. Molecular Biology Reports, 38, 5429-5433. doi:10.1007/s11033-011-0697-2
- Zhou, H., Hickford, J.G.H. and Fang, Q. (2006) A twostep procedure for extracting genomic DNA from dried blood spots on filter paper for polymerase chain reaction amplification. Analytical Biochemistry, 354, 159-161.
- Byun, S.O., Fang, Q., Zhou, H., et al. (2009) An effective method for silver-staining DNA in large numbers of polyacrylamide gels. Analytical Biochemistry, 385, 174- 175.
- Gong, H., Zhou, H. and Hickford, J.G.H. (2011) Diversity of the glycine/tyrosine-rich keratin-associated protein 6 gene (KAP6) family in sheep. Molecular Biology Report, 38, 31-35.
- Ku, N.O., Liao, J., Chou, C.F., et al. (1996) Implications of intermediate filament protein phosphorylation. Cancer and Metastasis Reviews, 15, 429-444.
- Bowden, P.E., Stark, H.J., Breitkreutz, D., et al. (1987) Expression and modification of keratins during terminal differentiation of mammalian epidermis. Current Topics in Developmental Biology, 22, 35-68.

