Open Journal of Statistics
Vol.05 No.07(2015), Article ID:62000,12 pages
10.4236/ojs.2015.57070
Robust Inference for Time-Varying Coefficient Models with Longitudinal Data
Zhaofeng Wang1, Jiancheng Jiang2, Qunyi Qiu3
1Jishou University, Jishou, China
2University of North Carolina, Charlotte, NC, USA
3Peking University, Beijing, China

Copyright © 2015 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).


Received 28 September 2015; accepted 14 December 2015; published 17 December 2015

ABSTRACT
Time-varying coefficient models are useful in longitudinal data analysis. Various efforts have been invested for the estimation of the coefficient functions, based on the least squares principle. Related work includes smoothing spline and kernel methods among others, but these methods suffer from the shortcoming of non-robustness. In this paper, we introduce a local M-estimation method for estimating the coefficient functions and develop a robustified generalized likelihood ratio (GLR) statistic to test if some of the coefficient functions are constants or of certain parametric forms. The robustified GLR test is robust against outliers and the error distribution. This provides a useful robust inference tool for the models with longitudinal data. The bandwidth selection issue is also addressed to facilitate the implementation in practice. Simulations show that the proposed testing method is more powerful in some situations than its counterpart based on the least squares principle. A real example is also given for illustration.
Keywords:
Local Polynomial Smoothing, Longitudinal Data, Local M-Estimators, Generalized Likelihood Ratio Tests

1. Introduction
The defining characteristic of a longitudinal data study is that individuals are measured repeatedly over a given time period, longitudinal studies are in contrast to cross-sectional studies, in which a single outcome is measured for each individual. The repeated measurements within each subject are generally correlated with each other, but different individuals can be assumed to be independent. The primary advantage of a longitudinal study is its effectiveness for studying changes over time. Statistical research in this field has been very active, and many parametric models have been developed. See Diggle et al. [1] , Davidian and Giltinan [2] , Vonesh and Chinchilli [3] , and the references therein.
As overwhelming longitudinal data exist in biomedical studies, there are increasing demands for a generally applicable inference tool for analysing these datasets. The parametric models are efficient for analysing longitudinal data but may suffer from mis-specification. To reduce possible modeling bias, different nonparametric and semiparametric methods have been studied in the literature, for example Zeger and Diggle [4] , Müller [5] , Eubank et al. [6] , He et al. [7] [8] , among others. In particular, a useful nonparametric model for analysing time-varying effects of covariates receives much attention. Examples include Hoover et al. [9] , Fan and Zhang [10] , Huang et al. [11] and others. These works focus on the least-squares based estimation. While they are useful in some applications, the shortcoming for lack of robustness is naturally raised. This motivates us to consider a robust inference tool for the time-varying coefficient models.
In this paper, we consider a local M-estimation approach based on local polynomial smoother and a robustified “generalized likelihood ratio (GLR)” statistic to test if parts of the coefficients are constants or of certain parametric forms. This in particular allows one to nonparametrically check the goodness-of-fit of the usual linear models widely used in practice (see for example Diggle et al. [1] ). We conduct extensive simulations to demonstrate that the proposed estimation method is robust against outliers and error distributions, and that the robustified GLR tests are powerful than its counterpart when the error deviates away from normality.
This paper is organized as follows. In section 2, the local M-estimation approach is introduced; a data-driven bandwidth selection rule is also given. In Section 3, we focus on the robustified GLR tests. Simulations are conducted in Section 4, where the performances of the different tests are compared, and a real example is also used to illustrate the proposed method. Finally, the paper is concluded by a discussion.
2. Model and Estimation
2.1. Local M-Estimation
Consider the following time-varying coefficient model,
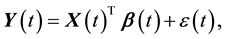 (1)
(1)
where 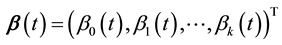 for
for 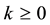 and
and 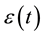 is a zero-mean correlated stochastic process that cannot be explained by the covariates
is a zero-mean correlated stochastic process that cannot be explained by the covariates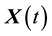 . As in Hoover et al. [9] we regard the repeated observations
. As in Hoover et al. [9] we regard the repeated observations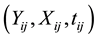 , for
, for 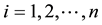 and
and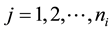 , as a random sample from model (1), that is,
, as a random sample from model (1), that is,
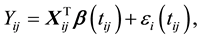 (2)
(2)
where 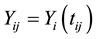 is the observed response,
is the observed response, 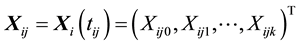 is the observed covariates for the ith subject at time
is the observed covariates for the ith subject at time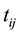 , and
, and 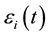 is a zero-mean stochastic process with covariance function
is a zero-mean stochastic process with covariance function 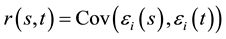 . For this model, we assume that the measurements on the responses for different subjects are independent, but
. For this model, we assume that the measurements on the responses for different subjects are independent, but 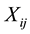 may be correlated at different time points within each subject,
may be correlated at different time points within each subject,
There are several methods for estimating the coefficient


where
The above local least squares based estimation is not robust against outliers and heavy tailed errors. To fix this problem, we propose to estimate the coefficient function by minimizing

where 







for 

The above method is in the same spirit of the local M-estimation studied for cross-sectional data in Fan and Jiang [19] and Jiang and Mack [20] . It can be shown that the estimator is 




The Newton algorithm can be used to find the solutions to the Equations (5). If the initial values of the parameters for iteration are good enough, for example, from the least squares estimation, then the iterative solutions can be found in a few steps, which is theoretically verified in Jiang and Mack [20] ) for nonparametrically modeling dependent data. In our simulations, the least squares based estimators in (2) will be employed as the initial values.
2.2. Bandwidth Selection
The performance of the estimator in (4) depends on the smoothing parameter h. There are several approaches to the selection of the bandwidth, such as the cross-validation (CV) and generalized cross-validation (GCV) criteria in Hastie and Tibshirani [24] . We here extend the CV method to the present situation for determining the bandwidth h. Specifically, denote by 


where
The minimization of (6) is time-consuming for simulations, even though it is not for real data analysis. We here suggest a pre-determination of 
1) Generate several samples (20 for instance) from (2), then minimize (6) to get the 
and compute the average 

2) Fix the bandwidth 
In step 1), the computational burden can be further reduced if one uses an easy-evaluated bandwidth as initial value, such as the one from the GCV criterion for the local least squares based estimator. Since the estimated bandwidths for the local least squares estimation and the local M-estimation are highly correlated in general, the minimization of 
3. Robustified GLR Tests
For simplicity, consider the following testing problem

where

where




Intuitively, large values of 








constructed if one replaces the 

Bootstrap Estimate of Null Distribution
To implement the robustified GLR tests, one needs to obtain the null distribution of 


1) Obtain the nonparametric estimate 

2) Compute the testing statistic 


3) For each given



4) Use the generated random sample 




5) Repeat steps 3 and 4 B times to obtain the bootstrap statistics
6) The P-value of the testing statistic 


Similar simulation approaches to determining the p-value of a testing statistic were given in Fan and Jiang [27] for additive models and Hui and Jiang [29] for DTARCH models. For the case that
Huang, Wu and Zhou, [11] ) to replace the bootstrap method above. Specifically, let 
where 


replacement from 




4. Numerical Studies
4.1. Robustness of the Estimation
In this section, we compare the performance of the local M-estimation with the local least squares estimation. For the outlier-resistant function, we opt for the Huber 
The rule of thumb in Section 2.1 for the choice of k will be employed.
Example 1. Consider the following model,

where










(A1)
(A2)
(A3)
(A4)
We simulated 200 samples of size 




The average of values of the above MADE over simulations was calculated and reported in Table 1 for the error models (A1) and (A3). The results show that the local LS estimator and the local M-estimators have similar accuracy if the error is normal, but the latter performs better than the former if the error deviates away from normality. We display in Figure 1 the estimators of the coefficient functions along with the widths of the envelopes formed by pointwise 2.5% and 97.5% quantiles of the estimators among simulations, where 


We did simulations with the error distributions in (A2) and (A4). The results are similar and omitted for saving space.
Table 1. MADEs under the errors (A1) and (A3) for Example 1.
Figure 1. Estimated curves and widths of envelopes. Upper panel: results for the error in (A1); lower panel: results for the error in (A2). (a) and (c): the average of estimated curves; solid―true curves, dash-dotted―local LS estimator, dashed―local M-estimator. (b) and (d): pointwise widths of envelopes for
Example 2. In this example, we consider the model (9) with much more complex structures for the covariates, the errors, and the coefficient functions than those in Example 1. Different from Example 1, the two co-
variates are now correlated, and the error 




where 



(B1)
(B2)

(B3)

We conducted 400 simulations. In each simulation, a samples of size 

Since 





4.2. Null Distribution and Power of Test
In this section, we conduct simulations to show that our conditional bootstrap method gives a good estimate of the null distribution, and to compare the powers of different nonparametric tests.
Example 3. To compare the powers of different tests, we consider the testing problem (7) for the model (9). The null model is the constant-coefficient model with




(C1)
(C2)
(C3)
We use a sequence of alternative models, 
We conducted 400 simulations. The sample size is the same as in Example 1. The bootstrap replicates number is
Table 2. MADEs for Example 2.
Figure 2. Estimated curves and envelopes. Upper panel: estimated curves for 
Figure 3. Histograms and estimated probability densities of TN’s. Left panel: based on the local LS-estimator; right panel: based on the local M-estimator with Huber 
imately equal type I errors. Figure 4 shows the powers of the tests based on the local LS and local M-estimators. It demonstrates that the proposed test is more powerful than that based on local LS-estimator when the error deviates away from the normal distribution. Both tests are approximately the same powerful under the normal error.
4.3. A Real Example
We here illustrate how to use the proposed method in practice. Consider the body-weight of male Wistar rats dataset in Brunner et al. ([30] , Table A11). For this dataset, the objective of the experience is to assess the toxicity of a drug on the body-weight evolution of Wistar rats. A group of ten rats was given a placebo, while a second group of ten was given a high dose of the drug. For each rat in the study, its body-weight was observed once a week over a period of 22 weeks. Figure 5(a) and Figure 5(b) displays the data curves for the two groups.
To check if the body-weights of the two test groups differ in their evolution over time, Brunner et al. [2] compared the time curves of the mean body-weights for both groups, and evaluated the ANOVA-type statistics. Their results do not support the conjecture of different body-weight evolutions in the two groups.
We are interested in investigating the conjecture. Since Figure 5(a) and Figure 5(b) exhibit the time-varying feature of the body-weights of rats, it seems reasonable to model the dataset via the following time-varying coefficient model:

where 


Since in the beginning the average body-weight of the rats in the placebo group is bigger than that in the
Figure 4. Powers of TN’s. Left panel: powers under significance level 95%; right panel: powers under significance level 90%. (a) (b): results under the error in (C1); (c) (d): results under the error in (C2); (e) (f): results under the error in (C3). Solid - power based on the local LS-estimator; dash-dotted―power based on the local M-estimator with Huber’s 
Figure 5. Data curves and the estimated coefficient functions. Left panel: (a) body-weights for the placebo group; (b) body-weights for treatment group. Right panel: (c) estimated curve for

treatment group, we subtract the average body-weight in each group from the body-weight of each subject. This will make the estimators of the coefficient functions be approximately zeros at the beginning. Figure 5(c) and Figure 5(d) report the estimators of 



Now we consider the following two hypothesis testing problems:
1)

2)
We used 


5. Discussion
We have introduced a robust inference method based on the local M-estimation method and the robustified GLR test. It is demonstrated that the local M-estimators are robust against outliers and error distributions, and the proposed robustified GLR test is more powerful than its counterpart (the GLR test) under certain situations with heavy tailed errors, while both of them perform well under the normal error. The proposed inference approach seems appealing in robustly modeling longitudinal data.
Our method is also applicable to other estimating methods, such as the global smoothing one in Huang et al. [11] , but will not be discussed further.
Acknowledgements
We thank the Editor and the referee for their comments. Supported in part by the NSFC grant 71361010 and by funds provided by the University of North Carolina at Charlotte. Correspondence should be addressed to Jiancheng Jiang at University of North Carolina at Charlotte, USA (E-mail: jjiang1@unc.edu).
Cite this paper
ZhaofengWang,JianchengJiang,QunyiQiu, (2015) Robust Inference for Time-Varying Coefficient Models with Longitudinal Data. Open Journal of Statistics,05,702-713. doi: 10.4236/ojs.2015.57070
References
- 1. Diggle, P.J., Liang, K.-Y. and Zeger, S.L. (1994) Analysis of Longitudinal Data. Clarendon Press, Oxford.
- 2. Davidian, M. and Giltinan, D.M. (1995) Nonlinear Models for Repeated Measurement Data. Chapman and Hall, London.
- 3. Vonesh, E.F. and Chinchilli, V.M. (1997) Linear and Nonlinear Models for Analysis of Repeated Measurements. Marcel Dekker, New York.
- 4. Zeger, S.L. and Diggle, P.L. (1994) Semiparametric Models for Longitudinal Data with Application to CD4 Cell Numbers in HIV Seroconverters. Biometrics, 50, 689-699.
http://dx.doi.org/10.2307/2532783 - 5. Müller, H.-G. (1998) Nonparametric Regression Analysis of Longitudinal Data. Lecture Notes in Statistics, Vol. 46, Springer-Verlag, Berlin.
- 6. Eubank, R.L., Huang, C., Maldonado, Y.M., Wang, N., Wang, S. and Buchanan, R.J. (2004) Smoothing Spline Estimation in Varying-Coefficient Models. Journal of the Royal Statistical Society, 66, 653-667.
http://dx.doi.org/10.1111/j.1467-9868.2004.B5595.x - 7. He, X., Fung, W.K. and Zhu, Z.Y. (2005) Robust Estimation in Generalized Partial Linear Models for Clustered Data. Journal of the American Statistical Association, 100, 1176-1184.
http://dx.doi.org/10.1198/016214505000000277 - 8. He, X., Zhu, Z.Y. and Fung, W.K. (2002) Estimation in a Semiparametric Model for Longitudinal Data with Unspecified Dependence Structure. Biometrika, 89, 579-590.
http://dx.doi.org/10.1093/biomet/89.3.579 - 9. Hoover, D.R., Rice, J.A., Wu, C.O. and Yang, L.-P. (1998) Nonparametric Smoothing Estimates of Time-Varing Coefficient Models with Longitudinal Data. Biometrika, 85, 809-822.
http://dx.doi.org/10.1093/biomet/85.4.809 - 10. Fan, J. and Zhang, J.T. (2000) Two-Step Estimation of Functional Linear Models with Applications to Longitudinal Data. Journal of the Royal Statistical Society: Series B (Statistical Methodology), 62, 303-322.
http://dx.doi.org/10.1111/1467-9868.00233 - 11. Huang, J., Wu, C. and Zhou, L. (2002) Vary-Coefficient Models and Basis Function Approximations for the Analysis of Repeated Measurements. Biometrika, 89, 111-128.
http://dx.doi.org/10.1093/biomet/89.1.111 - 12. Lindsey, J.K. (1993) Models for Repeated Measurements. Oxford University Press, Oxford.
- 13. Jones, R.M. (1993) Longitudinal Data with Serial Correlation: A State-Space Approach. Chapman and Hall, London.
http://dx.doi.org/10.1007/978-1-4899-4489-4 - 14. Hand, D. and Crower, M. (1996) Practical Longitudinal Data Analysis. Chapman and Hall, London.
http://dx.doi.org/10.1007/978-1-4899-3033-0 - 15. Brumback, B. and Rice, J.A. (1988) Smoothing Spline Models for the Analysis of Nested and Crossed Samples of Curves (with Discussion). Journal of the American Statistical Association, 93, 961-994.
http://dx.doi.org/10.1080/01621459.1998.10473755 - 16. Chiang, C.T., Rice, J.A. and Wu, C.O. (2001) Smoothing Spline Estimation for Varying Coefficient Models with Repeatedly Measured Dependent Variables. Journal of the American Statistical Association, 96, 605-619.
http://dx.doi.org/10.1198/016214501753168280 - 17. Wu, C.O., Chiang, C.T. and Hoover, D.R. (1998) Asymptotic Confidence Regions for Kernal Smoothing of a Varying Coefficient Model with Longitudinal Data. Journal of the American Statistical Association, 93, 1388-1402.
http://dx.doi.org/10.1080/01621459.1998.10473800 - 18. Wu, C.O. and Chiang, C.T. (2000) Kernal Smoothing on Varying Coefficient Models with Longitudinal Dependent Variable. Statistica Sinica, 10, 433-456.
- 19. Fan, J. and Jiang, J. (2000) Variable Bandwidth and One-Step Local M-Estimator. Science in China Series A, 43, 65-81.
http://dx.doi.org/10.1007/BF02903849 - 20. Jiang, J. and Mack, Y.P. (2001) Robust Local Polynomial Regression for Dependent Data. Statistica Sinica, 11, 705-722.
- 21. Huber, P.J. (1964) Robust Estimation of a Location Parameter. The Annals of Mathematical Statistics, 35, 73-101.
http://dx.doi.org/10.1214/aoms/1177703732 - 22. Serfling, R.J. (1980) Approximation Theorems of Mathematical Statistics. Wiley, New York.
http://dx.doi.org/10.1002/9780470316481 - 23. Jiang, J., Jiang, X. and Song, X. (2014) Weighted Composite Quantile Regression Estimation of DTARCH Models. Econometrics Journal, 17, 1-23.
http://dx.doi.org/10.1111/ectj.12023 - 24. Hastie, T.J. and Tibshirani, R.J. (1990) Generalized Additive Models. Chapman and Hall, New York.
- 25. Fan, J., Zhang, C.-M. and Zhang, J. (2001) Generalized Likelihood Ratio Statistics and Wilks Phenomenon. The Annals of Statistics, 29, 153-193.
http://dx.doi.org/10.1214/aos/996986505 - 26. Fan, J. and Huang, T. (2005) Profile Likelihood Inferences on Semiparametric Varying-Coefficient Partially Linear Models. Bernoulli, to Appear.
http://dx.doi.org/10.3150/bj/1137421639 - 27. Fan, J. and Jiang, J. (2005) Nonparametric Inferences for Additive Models. Journal of the American Statistical Association, 100, 890-907.
http://dx.doi.org/10.1198/016214504000001439 - 28. Fan, J. and Jiang, J. (2007) Nonparametric Inference with Generalized Likelihood Ratio Tests (with Discussion). Test, 16, 409-444.
http://dx.doi.org/10.1007/s11749-007-0080-8 - 29. Hui, Y.V. and Jiang, J. (2005) Robust Modelling of DTARCH Models. The Econometrics Journal, 8, 143-158.
http://dx.doi.org/10.1111/j.1368-423X.2005.00157.x - 30. Brunner, E., Domhof, S. and Langer, F. (2002) Nonparametric Analysis of Longitudinal Data in Factorial Experiments. Wiley, New York.













