Journal of Biomedical Science and Engineering
Vol.7 No.3(2014), Article ID:43086,11 pages DOI:10.4236/jbise.2014.73017
Extracting and smoothing contours in mammograms using Fourier descriptors
![]()
1LETS, GRETMAT, National Advanced School of Engineering, Yaoundé, Cameroon
2Department of Electrical and Electronic Engineering, College of Technology, University of Buéa, Buéa, Cameroon
3Centre Mère et Enfant, Fondation Chantal Biya, Yaoundé, Cameroun
4CTMED, Technical University of Cluj-Napoca, Cluj-Napoca, Romania
5Faculty of Science, University of Yaoundé I, Yaoundé, Cameroon
Email: alain_tiedeu@yahoo.fr
Copyright © 2014 Cyrille K. Feudjio et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In accordance of the Creative Commons Attribution License all Copyrights © 2014 are reserved for SCIRP and the owner of the intellectual property Cyrille K. Feudjio et al. All Copyright © 2014 are guarded by law and by SCIRP as a guardian.
Received 19 November 2013; revised 25 December 2013; accepted 3 January 2014
ABSTRACT
Contour is an important pattern descriptor in image processing and particularly in region description, registration and length estimation. In many applications where contour is used, a good segmentation and an efficient smoothing method are needed. In X-ray images, such as mammograms, where object edge is not clearly discernible, estimating the object’s contour may yield substantial shift along the boundary due to noise or segmentation drawbacks. An appropriate smoothing is therefore required to reduce these effects. In this paper, an approach based on local adaptive threshold segmentation to extract contour and a new smoothing approach founded on Fourier descriptors are introduced. The experimental results of extraction obtained from a set of mammograms and compared with the breast regions delineated by radiologists yielded a percent overlap area of 98.7% ± 0.9% with false positive and negative rates of 0.36 ± 0.74 and 0.93 ± 0.44 respectively. The proposed method was tested on a set of images and improved the accuracy, leading to an average error of less than one pixel.
KEYWORDS
Mammogram; Segmentation; Breast Contour; Smoothing; Fourier Descriptors
1. INTRODUCTION
Contour is one of the essential object features in image analysis. Despite its apparent simplicity, it is a feature difficult to accurately compute from a digital image. To accurately extract an object contour, a clear distinction between foreground and background is required. Furthermore, the shape of the contour extracted from digital image must be regular and smooth to ease object recognition or accurate length estimation.
Identifying and extracting breast contour in mammograms is of great importance due to the fact that it can improve computer-aided diagnosis (CAD) algorithms since it restricts the searching space for calcifications and lesions detection [1] and thus computation time [2]. Breast boundary delineating appears as a constraint in searching the reference point for registration between left and right mammograms in the process of asymmetry detection [3, 4]. Furthermore, nipple detection [5,6] and pectoral muscle [7,8] extraction which are some of the important preprocessing steps in CAD for breast cancer, critically depend on breast contour detection.
Unfortunately, in mammograms, breast edge is not clearly visible and the image background sometimes contains artifacts or is corrupted by noise. Developing a contour detection algorithm able to deal with different kinds of noise and artifacts present in the background of a mammogram is hard to achieve since it is difficult to distinguish the skin-air interface that separates the breast tissue from the background [9,10]. As a result, pixels are often shifted or misestimated after segmentation from the true contour. Smoothing the contour therefore appears as a solution to reduce these effects.
A wide variety of approaches have so far been developed to extract breast boundary in mammograms. Although some have shown promising results, they neither preserve the nipple—except in rare cases—nor produce a smooth contour. Smoothing the contour reduces shift of pixels along contour allowing easy estimation of length and removal of irrelevant objects present in the background while preserving skin and nipple. It is therefore of utmost importance in the event of breast deformation modeling.
1.1. Related Works
A literature review reveals that works aiming at segmenting the breast profile in mammograms can be categorized in methods based on histogram, gradients, polynomials modeling, region growing, active contours and wavelets [9-32].
One of the earliest attempts to segment breast region from background in mammograms was based on a simple threshold technique [11]. This approach consisted in setting a single threshold to separate non breast region from breast region [3,12,13]. However, as a single threshold is used for partitioning the image, it will inevitably result in misclassification of some background pixels as the breast region and vice-versa due to overlap between these regions. The use of local thresholding [9,14] had been shown to give better results as it thresholds each pixel using local information from its neighborhood. This method can efficiently deal with noise present on the breast edge area.
An interesting algorithm described by Lou et al. [15] considers that the traces of the intensity value from the breast region to the skin-air interface evolve as a monotonic decreasing function. It first searches an initial boundary set of pixels in the image, and then an estimated boundary is obtained by extrapolation. The contour is finally derived from the extrapolated points of the boundary after refinement and linking. The extrapolation finally introduces errors in the boundary detection Limin Yu et al. [16] used rational wavelet filtering to extract the breast contours in mammograms. Based on the observation that the low frequency images of a mammogram provide a better separation between breast region and background compared to gray level intensities (Heine et al. [17]), taking a reduced size of wavelet expansion image at low frequencies and applying a single threshold to eliminate noise, the largest object in the resulting image is identified as the breast region.
Semmlow et al. [18] applied spatial filters and Sobel operator edge detector to obtain breast boundary. Mendez et al. [19] used a gradient to identify the boundary after two-level histogram threshold techniques. Morton et al. [20] and Zhou et al. [21] developed similar methods where, after subscripting the background using an initial threshold, edge was found through line gradient analysis. Abdel-Mottaeb et al. [22] identified the breast edge through evaluation of different thresholds using edge gradient of resulting images and their union. Gradient-based methods present the advantage of using local information to extract the edge in image but show the same weakness as the histogram-based approaches as the final edge is to be selected through a threshold process.
Chandrasekhar et al. [23] applied a polynomial fitting model to approximate the breast region. A rough threshold segmentation was firstly done to approximate breast region. Then by fitting polynomials increasingly, several thresholds are generated. The selection of threshold and order of the polynomial was geared at minimizing the mean square error. Saha et al. [24] used a classifier where scale-based fuzzy technique connectivity was applied to segment breast region.
Wirth and Stapinski [25] and McLoughlin and Bones [26] used active contours to extract breast contour. They performed a global threshold and then modeled the background noise using Poisson approximation and the final binary mask obtained from the threshold of the model was used as seed for the snake algorithm. Ojala et al. [27] described a semi automated method where a boundary traced interactively was used to initialize the snake. Later, they described an active contour method for extracting and smoothing breast contours in mammograms [28]. Petroudi et al. [10] used a level set introduced by Chan and Vese [29] to extract the breast boundary. They firstly estimated the breast edge area by thresholding the image at the mean value and then enhanced the edge area. The binary mask was then used to initialize the active contour algorithm using the image with the enhanced breast edge. Ferrari et al. [30] used an active contour model especially designed to be locally adaptive to cope with noise, artifact and labels present near the boundary. Active contour methods are initialization dependent and sometimes a prior segmentation must be done to approximate the shape of the object in order to speed up the convergence. This technique is also complex and time consuming and can impose limitation on practical applications [9,10, 30].
One can clearly see that there is still no optimal segmentation method of breast region in digital mammograms. One inherent limitation of methods developed so far is the fact that very few of them preserve skin or nipple in profile. Generally, histogram intensity-based approaches are fast and not computationally demanding but suffer from reduced effectiveness in segmenting accurate breast regions and delineating a smooth contour. Useful results can nevertheless be obtained from global and local thresholding [10].
1.2. Proposed Method and Advantages
Extracting the breast contour should adequately model soft tissue/air interface and preserve nipple if in profile [2]. The use of local thresholding methods had earlier been said to be promising as they combine fairly good results and simplicity, unlike the algorithm described in [30] which can yield good results but is complex and has long execution time [9]. As breast contour detection is very often the first step in CAD systems, the segmentation method applied should be fast enough so as not to computationally overweight on the overall CAD system.
A global segmentation method like that of Otsu [31] tries to set a single threshold value for the whole image. This method is very fast and gives good results for discernible objects in images. However, when the boundary is not clearly visible or when the image has low contrast, this method tends to shift the edge or produce marginal noise along the edge.
Local binary segmentation methods [32,33] try to overcome these problems by computing a threshold for each pixel using information from the local neighborhood of the reference pixel. These methods are able to achieve good results for degraded images but they are often slow since the computation is conducted from the local neighborhood of each pixel in the image.
A good compromise would be therefore to devise tricks to speed up local binary segmentation methods. This is precisely what we do in this paper by using a segmentation approach based on local adaptive threshold using sum-tables implemented through the concept of integral images. By using the approach of sum-tables [34], local binary segmentation can be achieved with speed close to that of global segmentation but with better performance.
Commonly, after segmentation, the contour of an object is jagged or extended due to noise in the boundary or segmentation method shortcomings. In most of the previous mentioned works, care was not taken on smoothness the contour yielded. The authors rather focused on the accuracy of their algorithm in terms of breast area described by the detected contour. In this paper, we propose an approach to produce a smooth contour of breast in mammograms by the use of Fourier Descriptors (FDs). The main difficulty when using FDs for smoothing contour is to estimate the optimal number of FDs necessary for a proper smoothing. In Section 2, a method for automated detection of the adequate number of Fourier Descriptors (nFD) useful for a proper smoothing is introduced.
This paper therefore puts forth a two-step algorithm of extracting and smoothing breast contours from mammograms. A local adaptive threshold method based on sums of table is used to extract breast profile followed by the use of FD to smooth the extracted contours. Furthermore, as the FDs are based on the Fourier transform, it makes our approach fast and easy to implement.
There are a few advantages pertaining to this work as compared to those in the literature. The first is the use of a speed-up version of local thresholding technique. The second is tackling of the problem of contours’ smoothness more or less neglected in a good number of previous works. This paper also has the merit of tackling this problem through the use of FDs which are derived from Fourier transforms which are fast and easy to use as they will be seen. Finally, we propose a method to automatically select the optimal number of FDs needed for a smoothing.
The rest of the paper is organized as follows: details of algorithms used in contour extraction and problems related to contour smoothing are the object of Section 2 while Section 3 presents results from test of our algorithm on a standard database. These results are commented and compared to those of previous works in Section 4. Finally Section 5 presents some conclusions and remarks.
2. CONTOUR EXTRACTION AND SMOOTHING
The segmentation approach used in this section is based on local adaptive threshold using sum-tables implemented through the concept of integral images. The following paragraph describes the method.
2.1. Local Adaptive Threshold for Segmentation Using Sum-Tables
Given an image in which the intensity of a pixel is at the position (i, j). In the local adaptive threshold technique, the goal is to compute a threshold T(i, j) for each pixel such that:
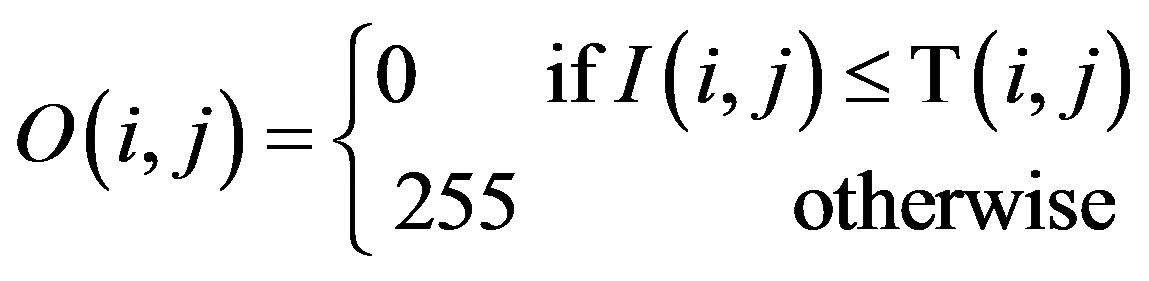 (1)
(1)
where O(i, j) is the output image.
The Sauvola’s binary segmentation technique [33] computes the local threshold T(i, j) of a pixel centered in a w × w window using the local mean m(i, j) and the local standard deviation s(i, j). The expression used in Sauvola’s binary segmentation method is of the form:
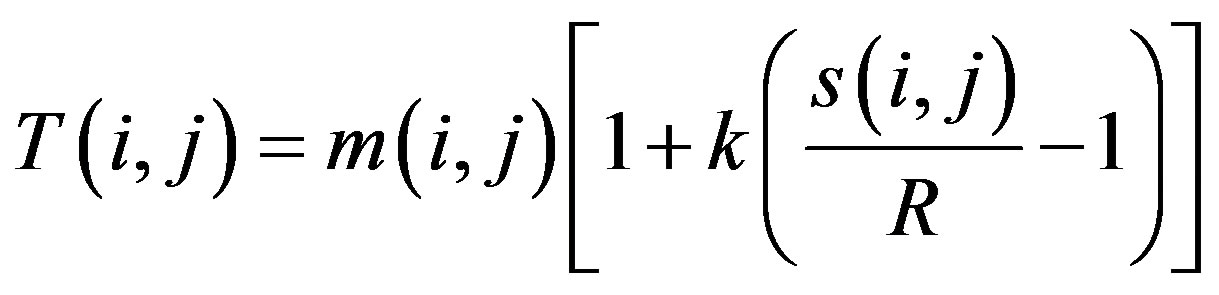 (2)
(2)
where R is the maximum value of the standard deviation and k is a parameter which takes positive values in the range [0.2, 0.5].
The use of the standard deviation in this formula is to adapt the threshold to pixel value in the window. When the contrast in the region is high, s(i, j) ≈ R which results in T(i, j) ≈ m(i, j). Otherwise, in the uniform region, the standard deviation is low and the threshold goes below the local mean therefore successfully removing the relatively dark region of the background.
Computing Equation (2) for an N × N image dimension results in computational complexity of O(W2N2) which makes the method time consuming. This complexity can be reduced to O(N2) by using the concept of integral image [35]. An integral image is defined as the image in which the intensity of a pixel at a given position is equal to the sum of all the pixels above the position and those left of the position. The expression of integral image is of the form:
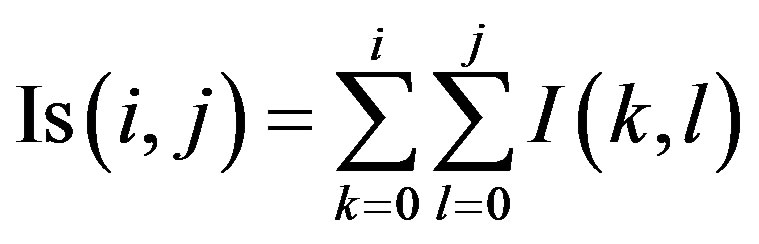 (3)
(3)
The local mean and the variance can be efficiently computed for any window size by using one addition and two subtractions instead of summing all pixels of the given window [33]. The local mean m(i, j) is defined as follows:
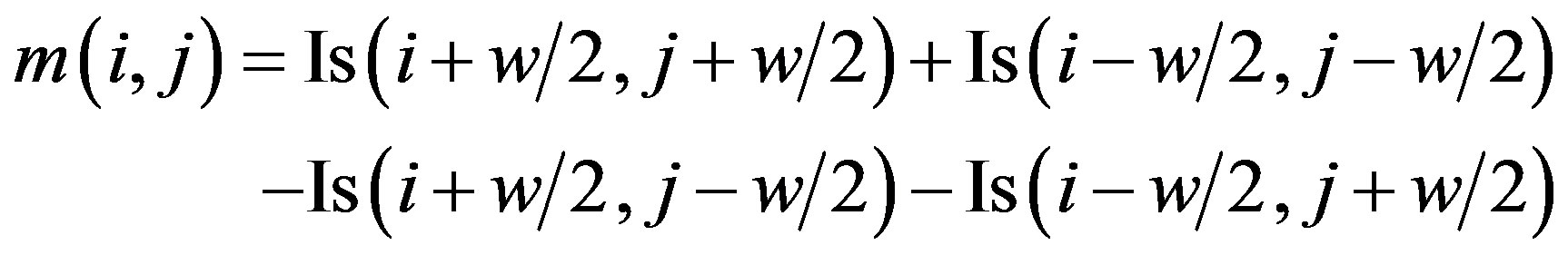 (4)
(4)
The standard deviation s(i, j) is defined as follows:
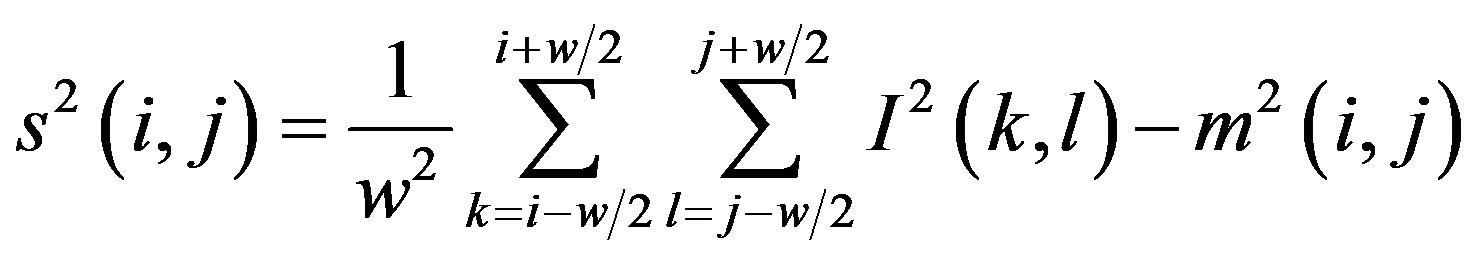 (5)
(5)
The result of the binary segmentation of mammograms using this approach is presented in Figure 1.
After segmentation, noisy points along contours are removed by performing a dilation followed by an erosion (Figure 1(c)). These two operations are carried out to avoid any displacement of the boundary.
Figure 2 shows some mammograms containing substantial noise in the background. After binary segmentation, the boundary of breast in mammogram is not entirely retrievable with the global thresholding method while the local thresholding method allows complete edge retrieval after morphological operations (erosion and dilatation) with a 7 × 7 structuring element.
These two cases of mammograms presented in Figure 2 are a good illustration that the local adaptive thresholding method using integral images can successfully extract boundary pixels when they are drowned in noise.
The contour extracted at this step (Figure 1(c) for instance) is jagged or may present some shifts of pixels on the edge due to the presence of noise along the boundary. Obtaining a real appearance of contour of the object requires some additional operations. This can be done by smoothing the extracted contour. The following section describes FDs in relation with contour smoothing.
2.2. Fourier Descriptors and Contours
Let’s consider a contour of an object made of N pixels.
 (a)
(a) (b)
(b) (c)
(c)
Figure 1. Binary segmentation of mammogram (mdb227) (a) initial image, (b) binary image, (c) cleaned edge in binary image.
 (a)
(a) (b)
(b) (c)
(c) (d)
(d) (e)
(e) (f)
(f) (g)
(g) (h)
(h)
Figure 2. Breast region segmentation in mammograms (mdb001 and mdb112) with Otsu technique (a), (e) and integral image method (c), (g). Images (b), (d), (f) and (h) are attempts of retrieving breast edge after segmentation.
Let Zk be the kth pixel in the contour and (xk, yk) its coordinate. The contour coordinate is therefore a vector Z of length N with individual elements Zk = 0,···, N − 1. Each contour element Zk can be represented as a complex Zk = xk + jyk. The advantage of this representation is that it reduces the problem from 2D to 1D. The discrete Fourier transform of Z is of the following form where Cl are the Fourier Descriptors coefficients of the contour:
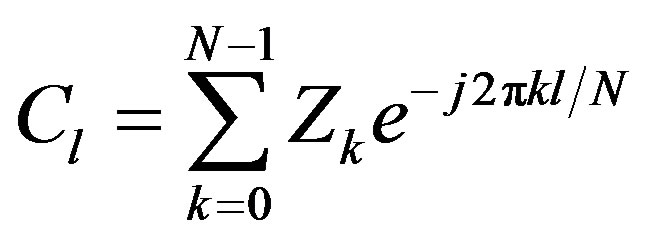 (3)
(3)
The key idea is that applying the Fourier transform to a closed contour (loop) is like computing the Fourier transform of a periodic signal. It represents the frequencies in the signal. In the case of an object contour, low frequency components of FDs contain information about the shape of the contour while the finer details along it are contained in the higher frequency components.
2.3. Contour Smoothing with FDs
Shape is the most important low level image feature to human perception. FDs have recently been applied to matching, retrieving or recognizing objects according to their shape [35-37]. They can also be used to smooth a contour. Considering a contour as a signal drowned in noise, FDs can be efficiently used for filtering the noise along the contour just as for any signal. Noise here is zizag or shift of the pixels along the contour. Smoothing tends to reduce that noise and produces a regular curve of contour as in its real state. The problem of smoothing a contour is simply the determination of the amount of FDs to be selected for reconstructing the contour curve.
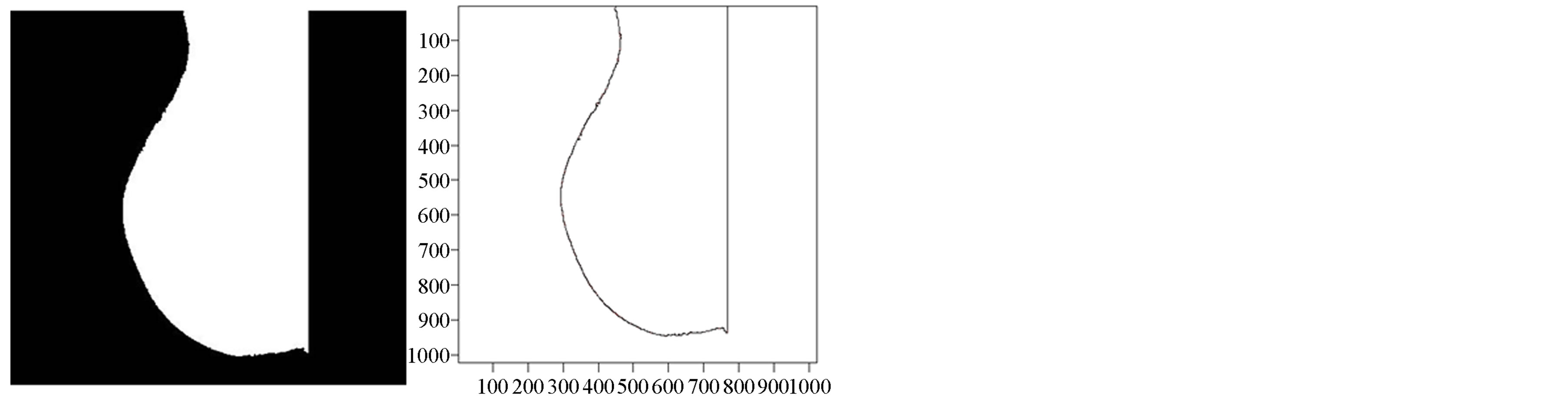
A contour with N points yields a set of N FDs coefficients. Figure 3 presents breast contour and its corresponding FDs. When more coefficients are selected, the smoothed curve gets closer to the segmented contour whereas considering very few coefficients will cause loss of significant corners or details with reduction of computation time. At this step, it is hard to determine the number of FDs (nFD) for a proper smoothing since there is no cut-off in FDs’ coefficients (see Figure 3(c)). Consequently, an optimal quantity of FDs’ coefficients cannot be straightly derived. A simple approach to cope with this problem is to study the effects of the variation of nFD on the smoothed curve. A comparison between segmented and smoothed contour is carried out to assess the similarity of both curves in order to derive the optimal nFD. This closeness is evaluated by computing Euclidean distances between two corresponding pixels in both curves which express gaps or errors. The contour will be smooth when errors are very low and remain under a limit value depending on the noise level in the contour of a given mammogram. Note that this limit value represents the maximal displacement done on a noisy pixel to smooth the contour curve. Next section explains how to derive the optimal nFD using errors produced with differrent nFD considered.
 (a)
(a)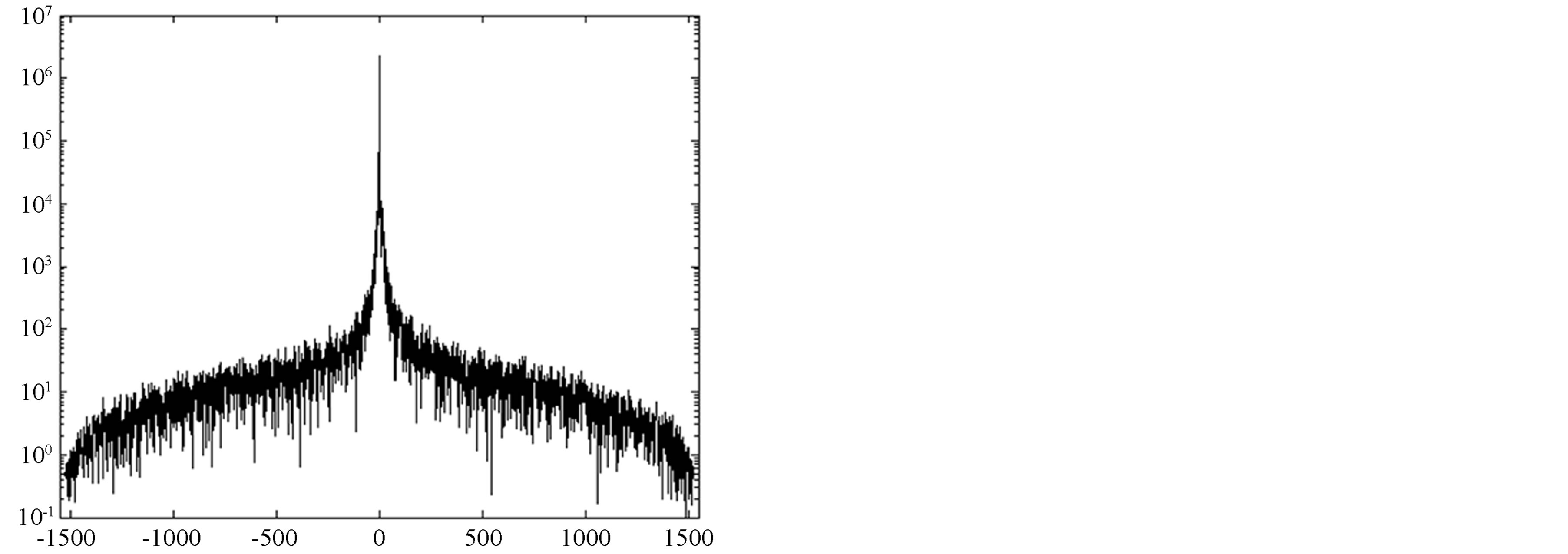 (b)
(b)
Figure 3. Extraction of mammogram contour (mdb227). (a) Breast region extracted, (b) Extracted contour, (c) Fourier descriptor coefficients.
2.4. Optimal Choice of the nFD
Using all FDs coefficients leads to recovering the segmented contour (under smoothing) whereas considering only few of them causes the lost of contour (over smoothing). It is difficult to define either maximal error (ME) or nFD for a proper smoothing knowing that smoothing consists of shifting noisy pixels from their position on the segmented contour. Maximal error (ME) can be viewed as maximal shift to be done on a noisy pixel to align it on the contour curve. Mammograms being corrupted by different levels of noise, optimization aims at finding the best trade-off between noise reduction (pixels’ contour displacement) and nFD for proper smoothing. When computing the errors for different nFD, it is high when the nFD is low and decreases in amplitude as nFD increases and finally becomes null when all FD are considered. This is true for each pixel of the contour. The approach we propose to solve this optimization problem consists in studying the maximal error (ME) variation. Figure 4 shows the variation of maximal error (ME) for different nFD when smoothing breast contour in a mammogram.
Observing Figure 4, we notice that ME drops rapidly and becomes very small near zero and almost constant in a certain range of nFD. Another important remark we can
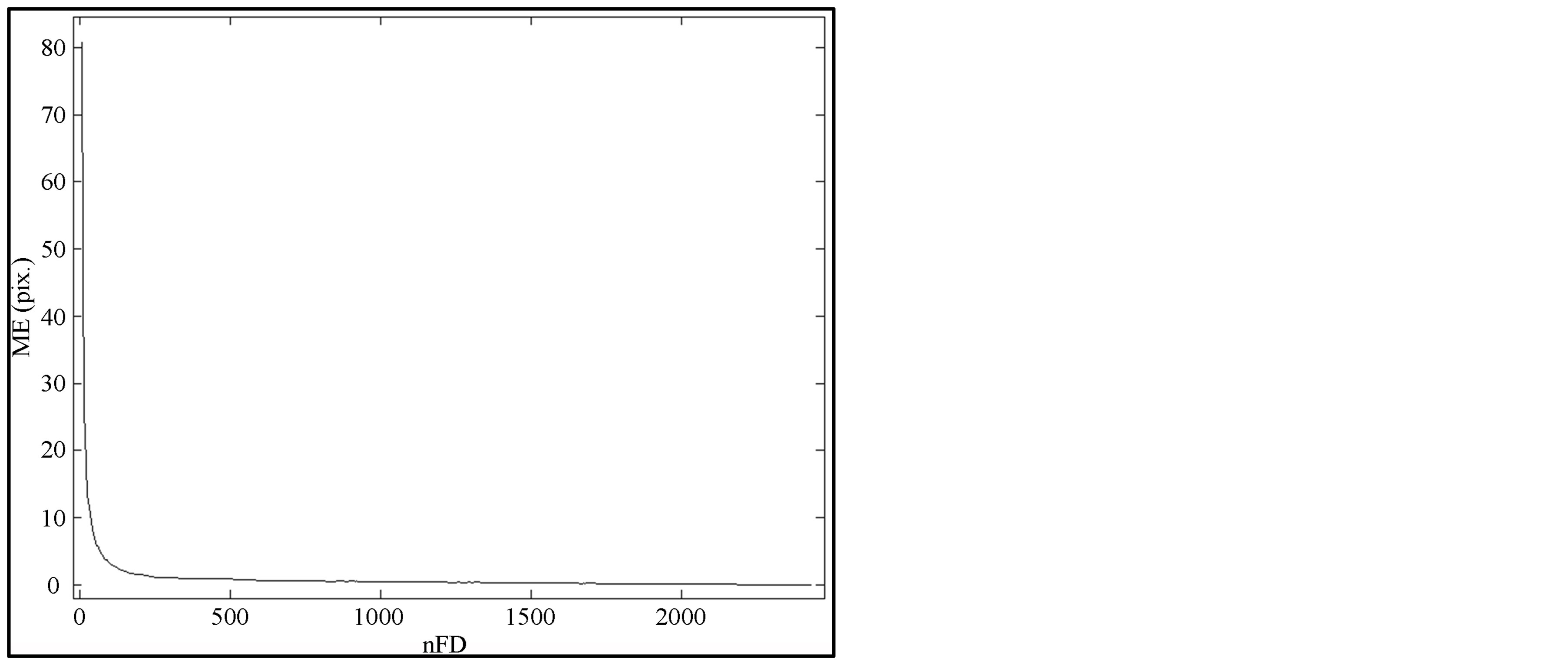
Figure 4. Maximal Error with respect to nFD used for contour smoothing (mdb015).
derive from this plot is that only few FDs are necessary for smoothing as ME is constant in a wide interval of values. The optimization process consists in searching for the nFD from which ME starts to be constant. This particular point is located on the elbow (knee) of the curve. The optimal nFD therefore corresponds to the amount of FDs for which ME is located at that elbow (knee) of the curve. It is a mathematical problem called “knee detection” that we address in the next subsection.
2.5. Knee Point Detection for Optimal Smoothing
Selecting the knee point—finding the good operating point—in a curve is difficult and even impossible to properly quantify since there exist neither a standard definition of the knee nor a general systematic approach for detecting one [37].
Recently, some accurate approaches have been developed to address the knee detection problem. Zhao et al [38] used an angle-based approach which is an extension of L-method for detecting knees in clustering applications. Their algorithm ignores the global trend of the curve and combines successive differences with the angles formed by y-axis to detect knees at local maxima of these angle values. Satopää et al. [37] attempted to give a standard definition of knee and discussed common methods used in knee detection. Their algorithm is based on threshold of local maxima of successive differences and can detect knee as well in online as offline data. While this algorithm is appealing, the choice of its sensitivity parameter is still questionable as it depends on the amount of data points considered. Although this algorithm yields knee detection results, it may computationally be expensive as we will show in the next section.
In our work, we rather used two basic criteria to detect the knee point for optimal smoothing. Our motivation in using these criteria is based on the fact that in each case of contour, the ME always exhibits the same behavior. The first criterion delimits the curve’s area where the knee is expected while the second checks if the slope of the curve is becoming null. The main advantages of this approach are 1) the limit values (maximal and/or minimal) of the data are generally known in advance and it is possible to define the convergence parameters, 2) the best operating point when smoothing lies at the end of knee’s curve and the method should give the possibility to tune up this value, 3) the knee is detected without delay. The knee detection algorithm works as follows:
1) Compute the error for different value of nFDi and select MEi
2) Obtain a knee by curve using the following equation on all computed MEi: yi = max(ME) − MEi
3) Check if the following criteria are satisfied
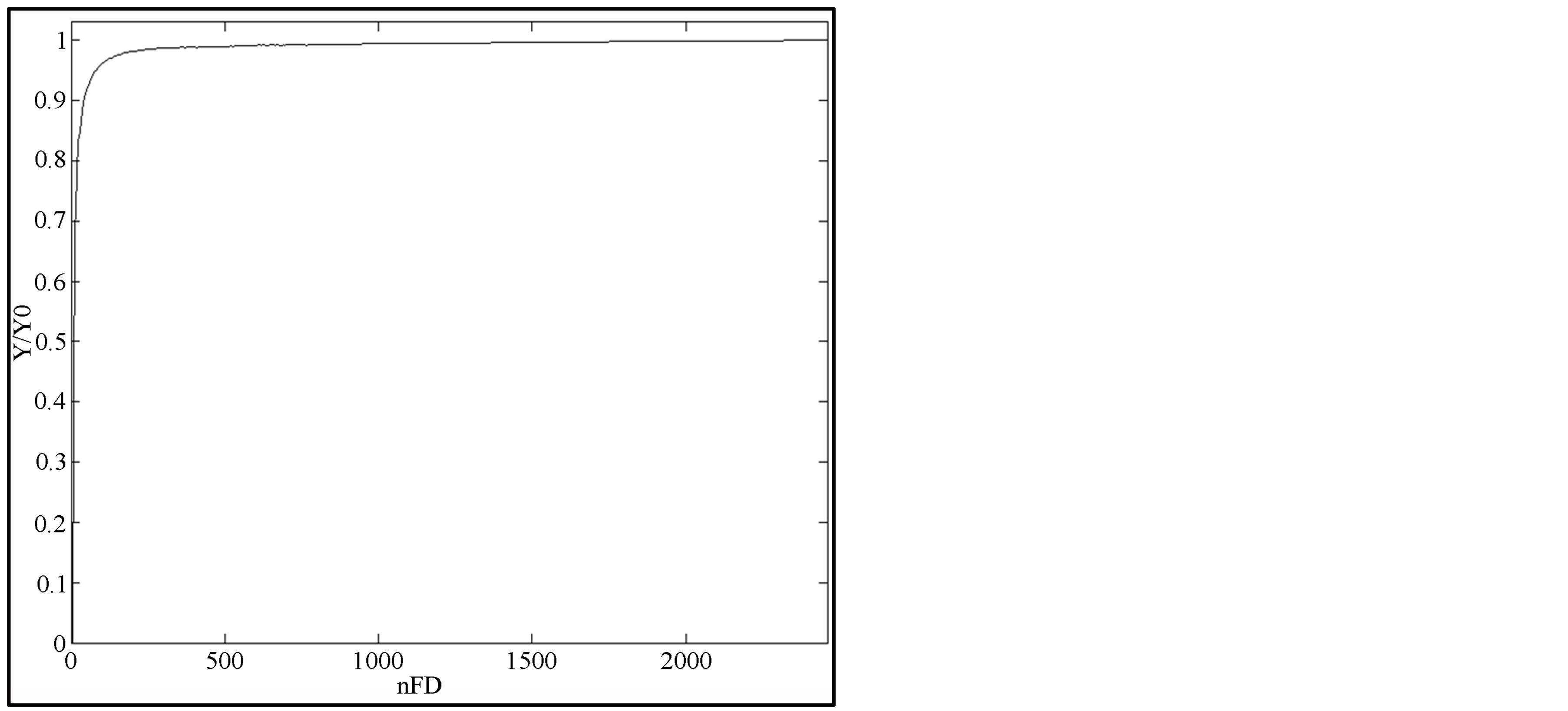
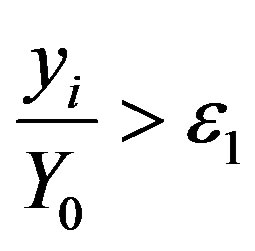 (7a)
(7a)
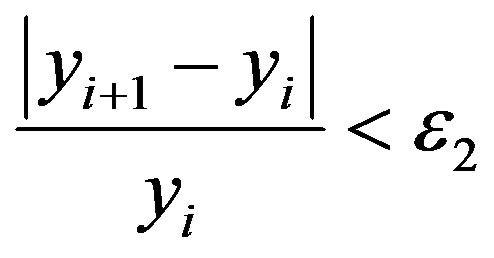 (7b)
(7b)
where Y0 is the maximal value of yi when ME is null and ε1, ε2 are the stopping criteria.
4) If both criteria are not satisfied increment nFDi and go to step 1, otherwise stop and declare the knee found at nFDi.
3. EXPERIMENTS AND RESULTS
Our algorithm was tested on images from the MIAS database (Mammographic Image Analysis Society) [39]. The MIAS images are distributed as 8 bit-per-pixel grayscale images at 50 μm per pixel spatial resolution and 1024 × 1024 pixels size. Sixty-nine images were used in this study. The sixty-nine images were those on which the radiologist could confidently mark the breast contour in order to assess the success rate of our algorithm.
3.1. Radiologist Assessment
A radiologist was invited to manually delineate the contours of breast in mammograms which were used as standard reference. To ease the radiologist’s task, breast edge visualization was improved using logarithmic contrast enhancement. The breast contours obtained by our algorithm were compared to the radiologist ones. Several criteria were chosen so that a fair comparison of our results with those of previous studies could be made. Those performance criteria were defined as follows:
1) Per cent Overlap Area (POA):
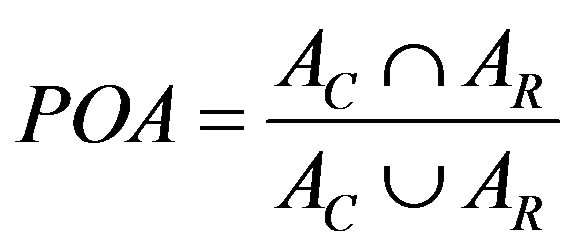
where AC and AR are the breast area delineated by the segmentation and the radiologist respectively.
2) False Positive (FP) pixels ratio
FP pixels are defined as pixels assigned to breast area by the algorithm but assigned outside the breast on the standard reference.
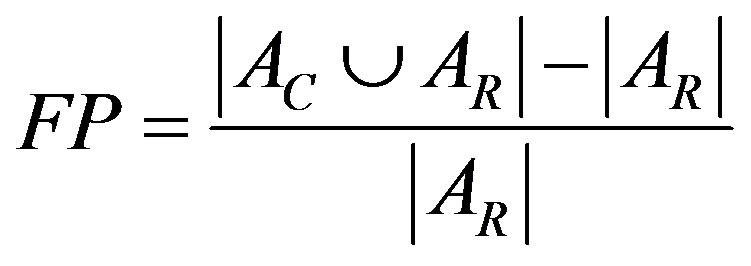
3) False Negative (FN) pixel ratio
FN pixels are defined as pixels assigned out of the breast by the algorithm but assigned inside the breast on the standard reference.
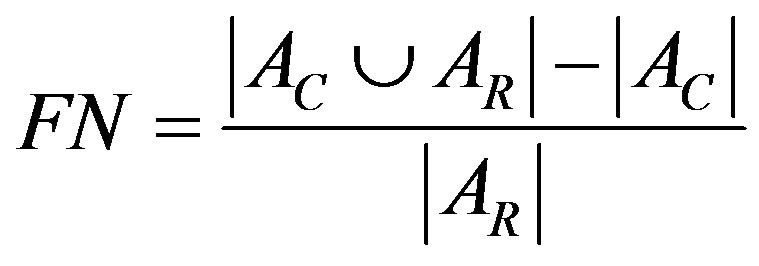
4) Hausdorff distance measure defined as:
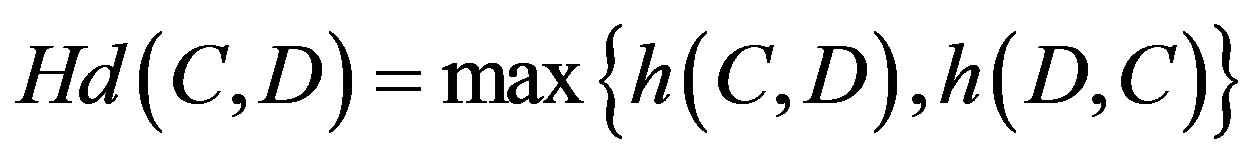
where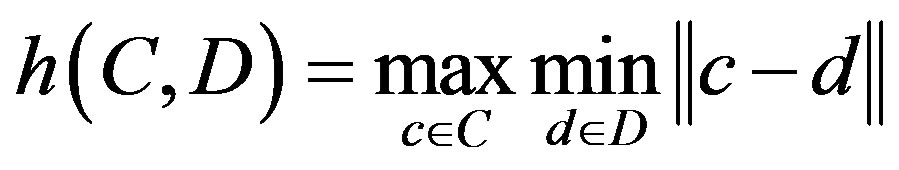 .
.
C is the set of points of the computer detected contour;
D is the set of points of the drawn contour by the radiologist.
The average proportion of these metrics and their corresponding standard deviation were computed for the 69 images used in this study. The average value and standard deviation of the percent overlap area POA are 98.7% ± 0.9%. The average value and standard deviation of false positive and false negative are 0.36% ± 0.74% and 0.93% ± 0.44% respectively. Figure 5 shows the initial mammogram with contour detected, drawn manually and superimposed on the same image.
3.2. Smoothing Contour
After segmentation, the computer detected contour is not always as smooth as the contour drawn manually (see Figure 5(c)). It usually appears jagged with some shifts of pixels in the noisy area of the boundary. Computing the length of the contour for instance with this rough detected contour may lead to over estimation. Smoothing is therefore applied to reduce noise along contour which is abnormal displacement. Figure 7 shows the result of smoothing of breast contour in a mammogram where smoothed contour and detected contour in a noisy area of the boundary are superimposed on the same mammogram.
3.3. Numerical Estimates and Error Rates
The smoothing is effective when the smoothed contour gets as closer as possible to the segmented contour. To determine how far smoothing should be done, ME, average value and standard deviation (STD) of distance between corresponding pixels in both contours were computed. To derive the optimal nFD, we execute the algo-
 (a)
(a) (b)
(b) (c)
(c)
Figure 5. Contour extraction (mdb108): (a) initial image with contour detected, (b) enhanced image with contour drawn manually, (c) detected (blue) and drawn (black) contour superimposed at a noisy area of the boundary.
 (a)
(a) (b)
(b) (c)
(c) (d)
(d)
Figure 6. Smoothing of breast contour (mdb108) (a) Contour obtained after segmentation, (b) contour smoothed, (c) and (d) comparison of smoothed contour (white) and extracted contour (magenta) at different spots of contour.
rithm described in Subsection 2.5 and plotted the curves of the two criteria defined. These plots are presented on Figure 7 and they help to easily determine the critical values of criteria used in the algorithm.
From the plots on Figure 7, we noticed that both curves become constant near 0.97 and 0.01 respectively. Taking into account that these curves are normalized and that all breast contours present the same behavior of ME, we set ε1 to 0.97 and ε2 to 0.01 in our algorithm in order to detect the knee of the ME curve.
We here provide a comparison of knee point detection using the proposed criteria and Kneedle algorithm (which is the most recent work with best results dealing with the knee problem) [39]. Figure 8 shows the knee
 (a)
(a)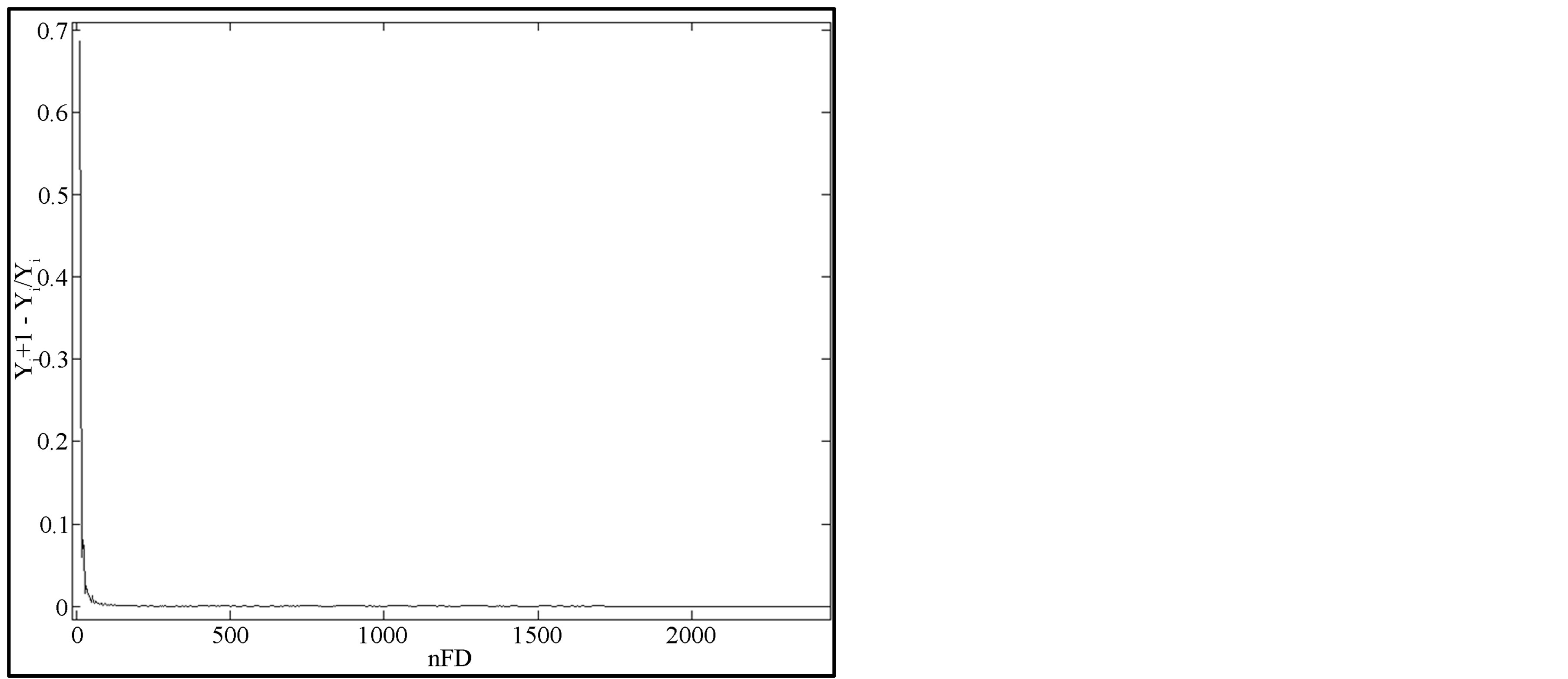 (b)
(b)
Figure 7. Variation of criteria for knee detection (mdb015).
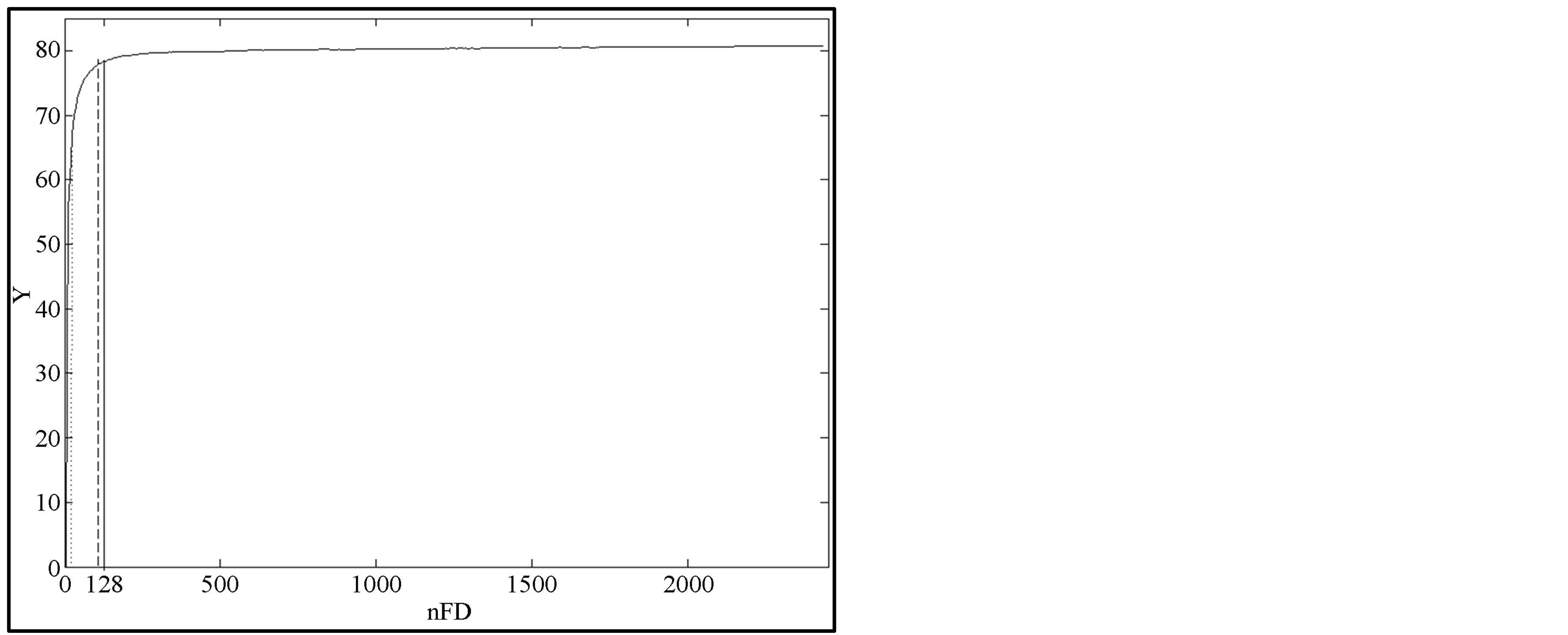
Figure 8. Knee detection (mdb015). Kneedle online detection (dot), Kneedle offline detection (dash), criteria method online and offline detection (solid).
point detected at nFD = 128 (solid) and nFD = 112 (dash) respectively with the proposed method and the Kneedle method for the case mdb015 offline. These values of nFD obtained by Kneedle algorithm and the proposed approach are quite close and they provide accurate smoothing (see Table 1 for estimates). Note that this result is obtained by computing ME for different amount of FD from 0 to the maximal value. Afterwards, the knee is detected by using the complete set of ME. This way of detecting the knee is called offline detection. This approach is computationally demanding since ME has to be computed iteratively for a huge amount of FDs.
To reduce the computation time, we attempt to detect the knee point at each iteration as nFD is increasing. This way of detecting knee is called online detection. From Table 1, we see that the same knee point is detected when using the proposed criteria online and offline. On the other hand, in the online cases, the Kneedle method detects the knee too much earlier (the word “earlier” refers to the knee position found. However, Kneedle detects the position of the knee in front of the knee curve rather than exactly on the knee) which leads to undersmoothing. Moreover the knee is detected with delay (computationally speaking as shown in Table 1, kneedle will declare the knee after receiving more points or more iterations and therefore with delay) which is globally linked to the parameter S. However, even if the value of parameter S can be tuned to detect nFD optimally, this induces an increase of delay. The delay or latency corresponds to the number of iterations that the algorithm has to perform after the point detected as knee point before declaring it. As the aim of optimization is to reduce computation by quickly detecting the best operating point in the curve, the proposed approach for knee detection appears more effective for this task. Note that the optimal nFD was obtained offline and online with the proposed method without any tuning of criteria.
Table 1 presents the knee point detected online and offline with Kneedle and our algorithm, the nFD derived and error estimates and parameters used in each case.
Table 2 presents average error rates obtained on a set of sixty-nine mammograms while smoothing breast contour. For each mammogram the following values were computed: the nFD found to be optimal (nFDo) for smoothing, the percentage of nFDo to nFD, the mean of error and the final ME (MEf) which represents the maximal displacement that occurred in a pixel. The cases were classified into three groups according to EMf less than 5 pixels, between 5 and 7 pixels and greater than 7 pixels. For each group of cases the table gives its percentage to the dataset and the averages of the ratio of nFD and mean error.
Note that while smoothing breast contour in mammograms with the proposed method, the maximal displace-
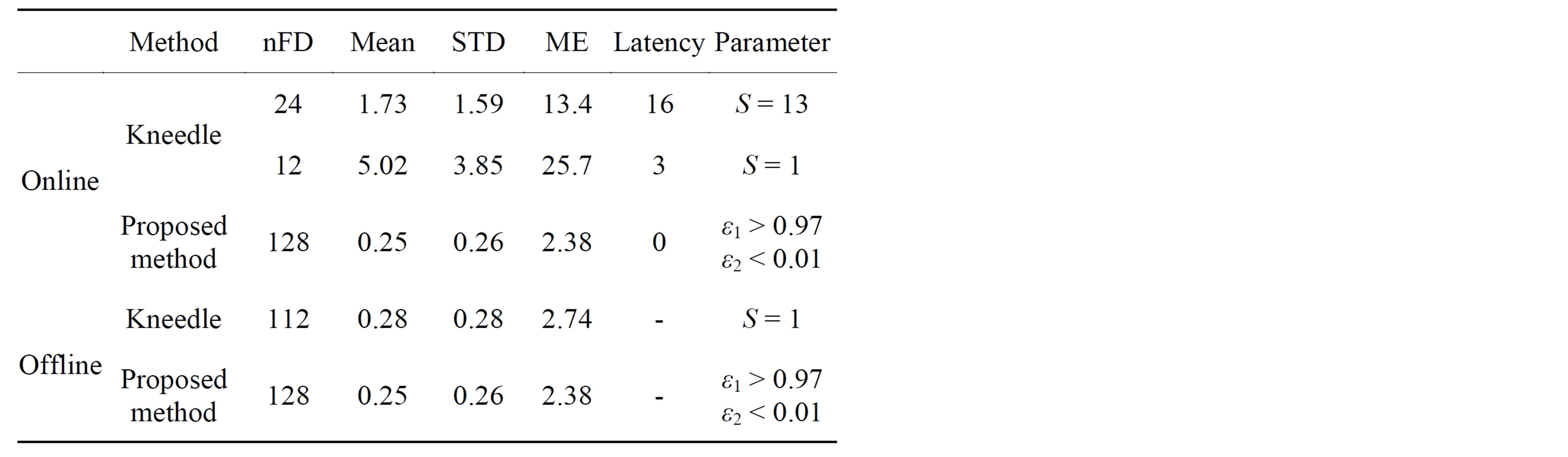
Table 1. Online and offline knee point detection and latency.
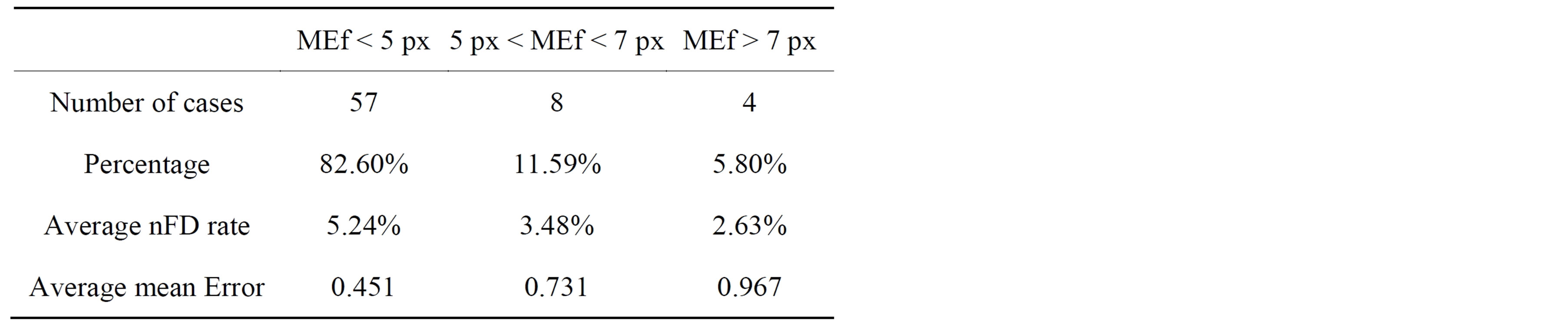

Table 2. Estimates of smoothing with the proposed method.
ment of pixel or MEf found for the whole dataset was less than 10 pixels which represents a shift in distance less than 2 mm.
4. DISCUSSION
The segmentation results obtained with the proposed method demonstrate that the algorithm can work well with images of breast containing some noise in the boundary (see Figure 2). The main advantages of this method are that 1) it preserves nipple if it is in profile, 2) is non iterative and 3) the computation time was set to be constant and non-dependent on the local window size as in global segmentation threshold while 4) providing more accurate results.
The local adaptive threshold approach was compared favorably to active contour models and it was shown in [9] that it does not conserve nipple in profile. Moreover, it was found to cause high rate of false negative by not detecting the nipple region. Also, large relative background misclassification pixels results from snake being unable to identify gradient edge and is therefore attracted towards high-density region inside breast instead of growing outwards in the direction of the true boundary [30].
Table 3 shows the comparison between the segmentation method presented in this work and others reported methods of breast profile extraction in mammograms and the performance metrics provided in their publications: The mean and standard deviation of POA, FP and FN in comparison to radiologist’s manually drawn breast profile. The three published methods are based on Level set [10], active contour [30] and breast border enhanced al
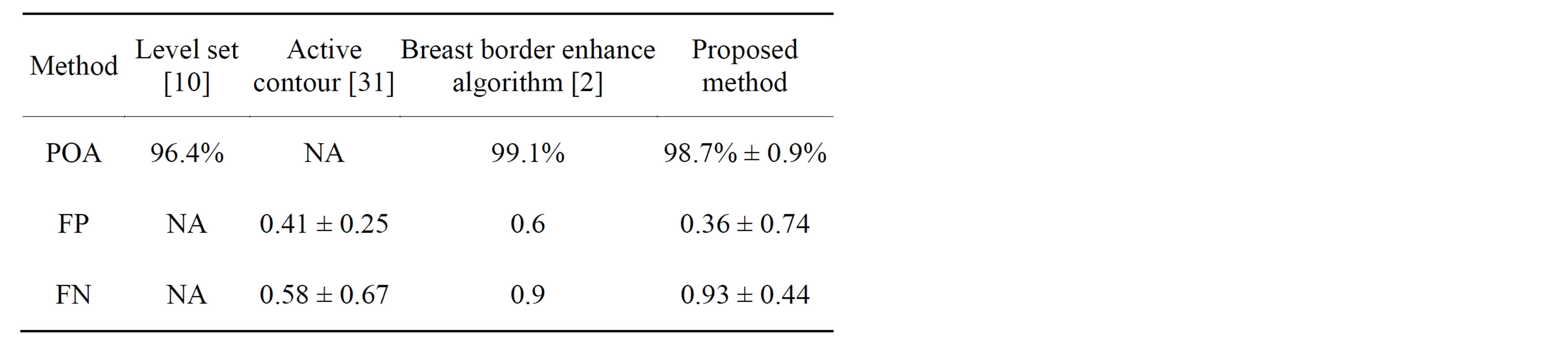
Table 3. Comparison with reported studies using performance metrics. Mean value and standard deviation of POA, FP and FN. Data not provided in the publications are marked as NA.
gorithm [2]. The Hausdorff distance obtained by the proposed approach was 3.07 ± 2.83 mm while the other reported studies did not evaluate this performance metric.
This comparison shows that the approach presented can perform as well or better that those reported. However, the major difference between these approaches lies on computation time. It was reported that extracting breast contour on a typical image of the dataset used in this work by active contour may required up to 3 min whereas only few seconds (less than 3 s) are needed with our approach.
We observed that in the noisy area of the breast boundary, the extracted contour was not smooth but rather jagged. A smoothing approach based on Fourier transform was introduced. Quantitative study on maximal error allowed us to establish two criteria which enable us to derive optimal nFD for smoothing. We show that using those criteria, the knee can be detected efficiently without delay as compared to the kneedle method. Globally the kneedle method tends to quickly detect the knee point on a curve particularly for online case (see Figure 8) and its accuracy depends both on S parameter and the number of data point received. It is possible for both methods to adjust their parameters to detect the knee point faster (front of the knee) or more precisely (end of the knee) but it is obvious that the kneedle method may be more conservative to produce accurate detection [37].
5. CONCLUSION
In this paper, a novel approach for accurate extraction of breast profile in mammograms is introduced. The proposed method relied on a two-step approach. The breast region is firstly estimated by the means of local threshold technique built up with sum table principle and the breast contour is then smoothed using FD technique. The proposed approach of extracting and smoothing contour is quite easy to implement as it is based on Fourier Transform. The optimal nFD was found by tackling the problem of knee detection. The result of knee detection problem showed that the proposed method found good values of the optimal nFD and was computationally less consuming when compared to other knee detection methods. The use of optimal nFD therefore yields accurate smoothing of contour with respect to noise in the boundary. The method was applied to a set of mammograms and yielded accurate extraction of breast contour. Moreover, the average error obtained was less than one pixel size for all the images processed. This method can be useful for preprocessing breast in application dealing with breast tissues characterization and breast cancer detection systems as it accurately delineated the area where further analyses had to be carried out.
ACKNOWLEDGEMENTS
This work was partially supported both by the Romanian research government funding “Eugen Ionescu” No. CE/DG/99/2010, grant established by AUF (Agence Universitaire de la Francophonie) and French research government funding.
REFERENCES
- Chandrasekhar, R. and Attikiouzel, Y. (2001) Mammogram-attribute database: A tool for mammogram segmentation and analysis. Proceedings of IASTED International Conference SPPRA, Rhodes, 3-6 July 2001, 143-148.
- Maitra, I.K., Nag, S. and Bandoypadhyay, S.K. (2011) Accurate breast contour detection algorithms in digital mammogram. International Journal of Computation and Applications, 25, 1-13.
- Yin, F.F., Giger, M.L., Doi, K., Metz, C.E., Vyborny, C.J. and Schmidt, R.A. (1991) Computerized detection of masses in digital mammograms: Analysis of bilateral subtraction images. Medical Physics, 18, 955-963.
- Ferrari, R.J., Rangayyan, R.M., Desautels, J.E.L. and Frère, A.F. (2001) Analysis of asymmetry in mammograms via directional filtering with Gabor wavelets. IEEE Transactions on Medical Imaging, 20, 953-964. http://dx.doi.org/10.1109/42.952732
- Olsen, C. and Georgsson, F. (2005) Problem related to automatic nipple extraction. Lecture Notes in Computer Science, 3540, 470-480. http://dx.doi.org/10.1007/11499145_48
- Chandrasekhar, R. and Attikiouzel, Y. (1997) A simple method for automatically locating the nipple on mammograms. IEEE Transactions on Medical Imaging, 16, 483- 494. http://dx.doi.org/10.1109/42.640738
- Raba, D., Oliver, A., Marti, J., Peracaula, M. and Espunya, J. (2005) Breast segmentation with pectoral muscle suppression on digital mammograms. Lecture Notes on Computer Science, 3523, 471-478. http://dx.doi.org/10.1007/11492542_58
- Kwok, S.M., Chandrasekhar, R., Attikiouzel, Y. and Rickard, M.T. (2004) Automatic pectoral muscle segmentation on mediolateral oblique view mammograms. IEEE Transactions on Medical Imaging, 23, 232-246. http://dx.doi.org/10.1109/TMI.2004.830529
- Shahedi, B.K.M., Amirfattahi, R., Azar, F.T. and Sadri, S. (2007) Accurate breast region detection in digital mammograms using local adaptive thresholding method. Proceedings of International WIAMIS, Santorini, 6-8 June 2007, 26-29.
- Liasis, G. and Petroudi, S. (2011) Estimation of the breast boundary in mammograms using level sets. Journal of Medical Imaging and Health Informatics, 1, 199-206. http://dx.doi.org/10.1166/jmihi.2011.1029
- Hoyer, A. and Spiesberg, W. (1979) Computerized mammogram processing. Phillips Technical Review, 38, 347- 355.
- Feudjio, C., Tiedeu, A., Gordan, M., Domngang, S. and Vlaicu, A. (2012) Computerized detection and smoothing contour in mammograms. Proceedings of 11th African Conference on Research in Computer Science and Applied Mathematics, Algiers, 13-16 October 2012, 299-306.
- Bick, U., Giger, M.L., Schmidt, R.A., Nishikawa, R.M., Wolverton, D.E. and Doi, K. (1995) Automated segmentation of digitized mammograms. Academic Radiology, 2, 1-9. http://dx.doi.org/10.1016/S1076-6332(05)80239-9
- Masek, M., Attikiouzel, Y. and deSilva, C.J.S. (2000) Skinair interface extraction from mammograms using local thresholding algorithm. Proceedings ICB, Brno, June 2000, 204-206.
- Lou, S.L., Lin, H.D., Lin, K.P. and Hoogstrate, D. (2000) Automatic breast region extraction from digital mammograms for PACS and telemammography applications. Computerized Medical Imaging and Graphics, 24, 205-220. http://dx.doi.org/10.1016/S0895-6111(00)00009-4
- Yu, L., Ma, F., Jayasuriya, A., Sigelle, M. and Perreau, S. (2007) A new contour detection approach in mammogram using rational wavelet filtering and MRF smoothing. Proceedings of 9th Biennial Conference of the Australian Pattern Recognition Society on Digital Image Computing Techniques and Applications, Glenelg, 3-5 December 2007, 106-111.
- Heine, J. and Kallergi, M. (1998) Multiresolution wavelet approach for separating the breast region from the background in high resolution digital mammography. Digital Mammography, Kluwer Academic Publishers, Nijmegen, 295-298.
- Semmlow, J.L., Shadagopappan, A., Ackerman, L.V., Hand, W. and Alcron, F.S. (1980) A fully automated system for screening xeromammograms. Computerized and Biomedical Research, 13, 350-362. http://dx.doi.org/10.1016/0010-4809(80)90027-0
- Mendez, A.J., Tahoces, P.J., Lado, M.J., Souto, M., Correa, J.L. and Vidal, J.J. (1996) Automatic detection of breast border and nipple in digital mammograms. Computer Methods and Programs in Biomedicine, 49, 253- 262. http://dx.doi.org/10.1016/0169-2607(96)01724-5
- Morton, A., Chan, H. and Goodsitt, M. (1996) Automated model guided segmentation algorithm. Medical Physics, 23, 1107-1108.
- Zhou, C. and Chan, H. (2001) Computerized image analysis: Estimation of breast density on mammograms. Medical Physics, 28, 1056-1069. http://dx.doi.org/10.1118/1.1376640
- Abdel-Mottaleb, M., Carman, C.S., Hill, C.R. and Vafai, S. (1996) Locating the boundary between the breast skin edge and the background in digitized mammograms. In: Doi, K., Giger, M.L., Nishikawa, R.M. and Schmidt, R.A. (Eds.), Digital Mammography, Elsevier, Amsterdam, 1996, 467-470.
- Chandrasekhar, R. and Attikiouzel, Y. (1996) Gross segmentation of mammograms using a polynomial model. Proceedings of International Conference of the IEEE EMBS, Amsterdam, 31 October-3 November 1996, 1056- 1058.
- Saha, P.K., Udupa, J.K., Conant, E.F., Chakraborty, D.P. and Sullivan, D. (2001) Breast tissue density quantification via digitized mammograms. IEEE Transactions on Medical Imaging, 20, 792-803. http://dx.doi.org/10.1109/42.938247
- Wirth, M. and Stapinski, A. (2004) A segmentation of the breast region in mammograms using snakes. Proceedings of the 1st Canadian Conference on Computer and Robot Vision, Ontario, 17-19 May 2004, 385-392.
- McLoughlin, K.J. and Bones, P.J. (2000) Location of the breast-air boundary for a digital mammogram image. Proceedings of Image Vision Computing Conference, Hamilton, November 2000.
- Ojala, T. and Liang, J. (1999) Interactive segmentation of the breast from digitized mammograms with united snakes. Technical Report 315, Turkey Centre for Computer Science, Turkey.
- Ojala, T., Nappi, J. and Nevalainen, O. (2001) Accurate segmentation of the breast region from digitized mammograms. Computerized Medical Imaging and Graphics, 25, 47-59. http://dx.doi.org/10.1016/S0895-6111(00)00036-7
- Chan, F.T. and Vese, L. (2001) Active contours without edge. IEEE Transactions on Image Processing, 10, 266- 277. http://dx.doi.org/10.1109/83.902291
- Ferrari, R.J., Frère, A.F., Rangayyan, R.M., Desautels, J.E.L. and Borges, R.A. (2004) Identification of the breast boundary in mammograms using active contour models. Medical and Biological Engineering and Computing, 42, 201-208. http://dx.doi.org/10.1007/BF02344632
- Otsu, N. (1979) A Threshold selection method from gray level histograms. IEEE Transactions on Systems Man and Cybernetics, 9, 62-66. http://dx.doi.org/10.1109/TSMC.1979.4310076
- Shafait, F., Keysers, D. and Breuel, T.M. (2008) Efficient implementation of local adaptive thresholding techniques using integral images. Proceedings of Conference DRR, San Jose, 26-31 January 2008, 681510.
- Sauvola, J. and Pietikainen, M. (2000) Adaptative document image binarization. Pattern Recognition, 33, 225- 236. http://dx.doi.org/10.1016/S0031-3203(99)00055-2
- Ekombo, P.L.E., Ennahnahi, N., Oumsis, M. and Meknassi, M. (2009) Application of affine invariant Fourier descriptors to shape-based image retrieval. International Journal of Computer Science and Network Security, 9, 240-247.
- Crow, F.C. (1984) Summed-area tables for texture mapping. Proceedings of SIGGRAPH, Minneapolis, 22-27 July 1984, 207-212.
- Larsson, F., Felsberg, M. and Forssén, P.-E. (2009) Patch contour matching by correlating Fourier descriptors. Proceedings of DICTA, Melbourne, 1-3 December 2009, 40- 46.
- Satopää, V., Albrecht, J., Irwin, D. and Raghavan, B. (2011) Finding a kneedle in a haystack: Knee detection points in system behavior. Proceedings of 31st ICDCSW, Minneapolis, 20-24 June 2011, 166-171.
- Zhao, Q., Xu, M. and Fränti, P. (2008) Knee point detection on bayesian information criterion. Proceedings 20th IEEE International Conference on Tools with Artificial Intelligence, Dayton, 3-5 November 2008, 43-48.
- Suckling, J., Parker, J., Dance, D.R., Astley, S., Hutt, I., Boggis, C.R.M., Ricketts, I., Stamatakis, E., Cerneaz, N., Kok, S.-L., Taylor, P., Betal, D. and Savage, J. (1994) The mammographic image analysis society digital mammogram database. Proceedings of the 2nd International Workshop on Digital Mammography, New York, Elsevier Science 1994, 375-378.

