Natural Science
Vol.07 No.03(2015), Article ID:54637,7 pages
10.4236/ns.2015.73016
Phylogenetic Analysis and Taste Cell Expression of Calpain 9 in Catfish (Ictalurus punctatus)
Tetsuya Ookura1,2*, Eiki Koyama3,4, Anne Hansen5, John H. Teeter1, Yukio Kawamura2,6, Joseph G. Brand1,3,7*
1Monell Chemical Senses Center, Philadelphia, USA
2Division of Food Function, National Food Research Institute, Tsukuba, Japan
3Department of Biochemistry, School of Dental Medicine, University of Pennsylvania, Philadelphia, USA
4Division of Orthopedic Surgery, The Children’s Hospital of Philadelphia, Philadelphia, USA
5Cell and Developmental Biology, School of Medicine, University of Colorado, Denver, USA
6Department of Food and Nutrition, Kyoto Women’s University, Kyoto, Japan
7Veterans Affairs Medical Center, University of Pennsylvania, Philadelphia, USA
Email: *ookura@affrc.go.jp, *brand@monell.org
Copyright © 2015 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 20 February 2015; accepted 10 March 2015; published 13 March 2015
ABSTRACT
The calpains, calcium-activated neutral proteases, play important roles in calcium-regulated intra-cellular signal transduction cascades. Here we report the isolation and initial characterization of a cDNA encoding a calpain 9, digestive tract specific calpain, from catfish taste epithelium. This calpain 9 (Ip-CAPN9a) shares 61% identity with human calpain 9. Phylogenetic analysis provides evidence that catfish calpain 9 and the related enzymes from Oncorhynchus mykiss, Danio rerio, Xenopus laevis, Mus musculus, Rattus norvegicus and Homo sapiens make up a distinct clade within the tissue-specific calpain family. Northern blot analysis reveals that Ip-CAPN9a is predominantly expressed in barbell and digestive tract, but not expressed in brain. An antibody against the N- terminal segment of Ip-CAPN9a recognizes cells within the taste buds in catfish barbells.
Keywords:
Gene Evolution, Taste, Novel Type

1. Introduction
The calpains are cytoplasmic, calcium-activated neutral cysteine-proteases. They function in Ca2+ signaling by modulating biological activities of their substrates through limited proteolysis. Several structural types of calpains are known, many of these being tissue-specific. Calpain 3 (CAPN3) is specifically expressed in muscle [1] , and mutations of the CAPN3 gene were reported in limb girdle muscular dystrophy type 2A [2] . A splice variant of calpain 3, Lp82, is expressed in the eye lens [3] . Calpain 8, predominantly expressed in stomach, shows two alternative splice products, one with and one without a calcium binding domain [4] . Calpain 9 (CAPN9) is predominantly expressed in the digestive tract and plays a role in gastric mucosal defense in collaboration with calpain 8 [5] [6] . Down-regulation of calpain 9 has been observed in stomach cancer [7] and hypertension [8] . The physiological function of calpain 9 in taste cells is still unclear.
All (classical) calpains are heterodimeric. Of the two subunits that comprise the calpain heterodimer, the larger one can be divided into four domains: domain I has a regulatory function; domain II is the cysteine protease domain; domain III is C2-like β-sandwich and domain IV is responsible for Ca2+-binding with five EF-hand motifs [9] . X-ray crystallographic analysis of rat calpain 4 revealed that several amino acid pairs in EF-hand motifs specified calcium-binding properties [10] [11] . Recently, the Km values (125 mM) for Ca2+ ions for human recombinant CAPN9 have been reported [12] . The physiological substrates of calpain 9 remain to be elucidated.
Knowing that calpains are involved in intracellular calcium-mediated events, we reasoned that calpains may play a role in taste transduction, a process regulated by calcium signaling. In the visual system, calpains were thought to play roles in adaptation processes through arrestin degradation [13] . The calpains may also play limited roles in cleavage of the IP3 receptor [14] , PLC β3 [15] , G proteins [16] , and other members of the calcium signaling cascades [17] .
Chemical stimulation of taste receptor cells generally results in changes in activity of intracellular calcium, leading to either release of neurotransmitter from the stimulated cell or communication with a specialized synaptic cell [18] . This taste-mediated increase in calcium may be sufficient to activate calpains that stimulate the degradation of signaling molecules.
Here we report on the identification of calpain 9 from catfish (designated Ip-CAPN9a) and on its expression in taste tissue. We first reported the molecular cloning of calpains from taste tissue in abstract form in 1997 [19] .
2. Materials and Methods
2.1. Calpain cDNA Isolation from Catfish Barbell Epithelium cDNA Library
A polyclonal anti-80 kDa antibody recognizing the taste cells of the catfish barbell taste buds by immunohistochemistry was used at 1:5000 dilution to isolate a cDNA clone from a λ-ZAP catfish barbell epithelium cDNA library using an immuno-screening kit (Stratagene, CA). DNA sequencing was performed by the dideoxy method using an ABI 373 DNA sequencer (Applied Biosystems, CA).
2.2. Phylogenetic Analysis of Calpain Family
The protein sequences of full-length calpain large subunits were obtained from the NCBI database. The accession numbers are shown in Table 1.
Amino acid sequences of calpains were aligned using MAFFT with G-insi options [20] . Two alignments were created: 1) one of the entire large subunit, 2) the other of only the calcium binding domain of the large subunits. With the alignments we ran maximum-likelihood analysis [21] with the RAxML software [22] over 10,000 rounds of heuristic search, using the JTT model of amino acid substitution [23] and the final tree was optimized using the JTT + Gamma model on the Cipres Portal v1.15. To derive branch support values, 300 rounds of nonparametric bootstrap analysis [21] were run with RAxML with the same substitution model as indicated above, with one heuristic search in each bootstrap replicate. The tree was rooted using the nematode CAPN5 as an outgroup. MEGA4 was used to visualize the trees [24] . Total RNA was isolated from barbell epithelium, olfactory epithelium, brain and intestine of catfish with Trizol reagent (Invitrogen, CA).
2.3. Northern Blot Analysis
Total RNA was isolated from barbell epithelium, olfactory epithelium, brain and intestine of catfish with Trizol reagent (Invitrogen, CA). From each tissue, 20 μg total RNA (except 10 μg RNA from olfactory rosette) was electrophoresed and blotted onto a Hybond-N plus membrane (Amersham Bioscience, NJ). Antisense RNA probes were synthesized using an RNA transcription kit (Promega, WI) and DIG-UTP (Roche Diagnostics, IN).
Table 1. List of the accession numbers cited in this report.
2.4. Antibody Preparation
Antibodies against the Ip-CAPN9a N-terminal segment were raised in rabbits injected with a synthetic peptide (amino acids 3-16: SASTLFSTSGNKIQ) conjugated to BSA, and purified by affinity chromatography with the immobilized synthetic peptide onto a Sepharose (QCB Inc., MA).
2.5. Immunohistochemistry
Catfish barbells were fixed overnight in 4% paraformaldehyde at 4˚C. Cryoprotection was carried out in 20% sucrose overnight. The tissue was then embedded in Tissue Tek OCT (Sakura Finetek, Torrance, CA). Cryosections (12 - 14 μm) were mounted on Superfrost Plus slides (VWR, West Chester, PA) and frozen at −80˚C until further use. Standard immunohistochemical procedures were used. Briefly, cryosections were rinsed in 0.1 M phosphate buffered saline (PBS), blocked in blocking solution containing 1% BSA, 3% normal donkey serum, and 0.3% Triton X-100 in PBS for 2 hours, and then incubated in the primary antiserum overnight (Ip-CAPN9a, 1:500 dilution in blocking serum). After 3 washes, 20 min each, the sections were incubated in secondary antibodies (FITC- or TRITC-conjugated, 1:400; Jackson ImmunoResearch, West Grove, PA) for 2 hours at room temperature. After incubation, sections were washed 3 times 20 min and coverslipped with Fluormount-G (Fisher Biotech, Birmingham, AL). Control slides were treated either without the primary antibody or with normal rabbit serum replacing the primary antiserum. The same protocol was used for whole mounts, i.e. whole barbells, however, the incubation times were longer: over 3 days for the primary and overnight for the secondary antibodies.
3. Results
3.1. Molecular Cloning of a Catfish Tissue-Specific Calpain
We screened one million clones from a catfish (Ictalurus punctatus) barbell epithelium λ-Zap cDNA library with an anti-80 kDa antibody and isolated 2 positive clones. Blastp analysis revealed that one was a calpain, the other a MAGI-like protein. The calpain cDNA had 2458 nucleotides, encoding a polypeptide of 687 amino acids and was most closely related to human calpain 9 (digestive tract calpain) and fish calpain 9 (Figure 1(a)). We named this catfish calpain 9 “Ip-CAPN9a” (Ip = Ictalurus punctatus). Figure 1(a) shows the alignment of fish, frog, and mammalian calpain 9 and calpain 3. The black-boxed residues are those that are identical among calpain 9 and calpain 3. In calpains, the most diverse regions are at the N-terminus and at the calcium binding domain (Figure 1(a)). This calcium binding domain determines the calcium sensitivity of protease activity. In this domain, Ip-CAPN9a is also most closely related to human and fish calpain 9 (Figure 1(a)). X-ray crystallographic analysis of the m-calpain identified the Penta EF hand motifs and corresponding calcium-chelating amino acid residues (X, Y, Z, -Y, -X, -Z) within the calcium binding domain [10] [11] .
Figure 1(b) shows the Ca-chelating residues in each PEF motif for calpains and calmodulin. Unique amino acids were observed at PEF1-Y (A), PEF2-Y (M), PEF3-Y(R), PEF4-Y (E), PEF5 Z(R) and PEF5-Y (E) in Ip- CAPN9a. At (-Y) positions, some of these residues were identical among fish CAPN 9, while residues at the -Y position in frog CAPN 9 are identical to those of human CAPN9 as indicated by underline.
3.2. CAPN9 Clusters with CAPN3
The amino acid sequences from 648 to 821 in the large subunit were used to infer the phylogenetic relationships between the mammalian and fish calpain families. Phylogenic trees were built using RaxML. Five distinct monophyletic branches were observed: CAPN9, CAPN3, CAPN1, CAPN2 and CAPN11. Ip-CAPN9a is most homologous with rainbow trout CAPNg. Ip-CAPN9a is in the branch of the calpain 9 subfamily, joining the branch of the calpain 3 subfamily (Figure 2(a)), while rainbow trout CAPN1 and CAPN2 were in the branch of human CAPN1 and CAPN2, respectively (Figure 2(a)).
Figure 1. Comparison of catfish calpain 9 (Ip-CAPN9a) with other calpains. (a) Alignment of Ip-CAPN9a with other calpains, including human calpain 3, human calpain 9 and fish CAPN9. The Penta-EF hand motifs are indicated with arrows pointing out the calcium binding sites; (b) Alignment of calcium coordinating amino acids (X, Y, Z, -Y, -X, -Z) in Penta-EF hand motifs with calpain family and calmodulin.
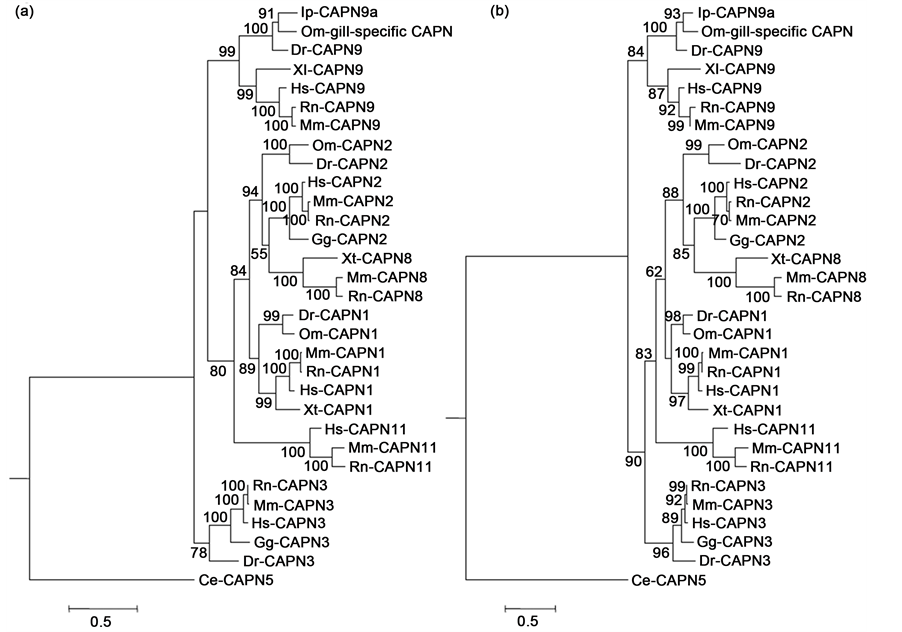
Figure 2. Phylogenetic trees of calpain large subunits (a) and penta EF-hand motifs (b). Phylogenetic trees estimated under the JTT + G model (RAxML with rapid bootstrap analysis with 300 bootstrapping runs). Nematode CAPN5 was used as an outgroup. Numbers above branches denote bootstrap values from 300 replicates (only values higher than 50% shown). The scale bar indicates an evolutionary distance of 0.5. The species abbreviations are as follows: Ce: Caenorhabditis elegans, Dr: Danio rerio, Gg: Gallus gallus, Hs: Homo sapiens, Ip: Ictarulus punctatus, Mm: Mus musculus, Om: Oncorhynchus mykiss, Rn: Rattus norvegicus, Xl: Xenopus laevis, Xt: Xenopus tropicalis.
Within the calcium binding domain (residues 648 to 821), Ip-CAPN9a clustered with rainbow trout CAPN9, zebrafish CAPN9, human CAPN9, mouse CAPN9, rat CAPN9 and Xenopus CAPN9. Rainbow trout CAPN1 and CAPN2 grouped with human CAPN1 and CAPN2, respectively (Figure 2(b)).
3.3. Expression of IP-CAPN9a in Catfish Barbell Epithelium
We examined the expression of Ip-CAPN9a mRNA in catfish tissues. The 2.4 kb band was detected by Northern blot analysis predominantly in barbell, weakly in intestine and the olfactory rosette, and not in brain (Figure 3(a)). A polyclonal antibody, raised against the peptide corresponding to the deduced amino acid sequence 3-16 of Ip-CAPN9a, recognized the taste pore region of taste buds in catfish barbell epithelium (Figure 3(b)) and was weakly expressed in the taste cells of taste buds (Figure 3(c)).
4. Discussion
In this report, we described the isolation of a tissue-specific calpain cDNA out of a catfish barbell epithelium cDNA library. Calpains have been reported in other non-mammalian species including CAPN1 & CAPN2 in carp muscle [25] , those in trout muscle [26] and gill-specific calpain of rainbow trout [27] . The calpain reported on here encodes a 687 amino acid protein with a predicted molecular weight of 78,345 Da. From amino acid sequence comparison, this catfish calpain (Ip-CAPN9a) is most closely related to the CAPN9 of rainbow trout, zebrafish, human, mouse, rat and Xenopus (Figure 1(a)).
In a maximum-likelihood analysis with the whole protein sequence, the value of alpha was 0.926. Figure 2(a)

Figure 3. Expression analysis of Ip-CAPN9 in catfish. (a) Northern blot analysis of IP- CAPN9a in catfish tissues. Total RNA from taste bud-containing maxillary barbell and from other tissues as indicated for catfish: (lane 1. barbell epithelia (20 μg), lane 2. Intestine (20 μg), lane 3. olfactory rosette (10 μg), lane 4. brain (20 μg)); (b) Whole mount preparation of a catfish barbel showing the taste pores (arrows) of taste buds. The small fluorescent dots depict solitary chemosensory cells which also seem to be Ip-CAPN9a-positive. Scale bar: 50 μm; (c) Ip-CAPN9a-like immuno-reactivity in taste cells of taste buds on catfish barbel. Scale bar: 20 μm.
shows that Ip-CAPN9a clustered within the same node next to the CAPN3 subfamily, while rainbow trout capn1 and capn2 grouped with human capn1 and capn2. The same result was observed in the phylogenetic analysis with the Penta-EF hand motifs (α = 1.692) in Figure 2(b). These observations would suggest that CAPN3 and CAPN9 diverged from CAPN1 and CAPN2 prior to teleost evolution [28] .
A feature worth further study is the unique amino acid residues at the calcium binding site of Ip-CAPN9a, especially at the Y site (Figure 1(b)) [10] [29] . Considering their position within the calcium binding sites, it is likely that these unique amino acids participate in the calcium modulation of this enzyme.
Northern blot analysis suggests that Ip-CAPN9a is predominantly expressed in barbell, slightly in olfactory rosettes and intestine, but not in brain (Figure 3(a)). Even though we were unable to detect Ip-CAPN9a mRNA expression in brain by northern analysis, we could not rule out the possibility that some specific cells within the brain may express Ip-CAPN9a as shown in Drosophila CalpA [30] . Salem et al. showed that rainbow trout gill-specific calpain, closely related to calpain 9, was not expressed in the small intestine. Different detecting probes used in the northern blot analysis may explain this difference [27] .
The immuno-histochemical studies demonstrated that polyclonal antibodies against a deduced peptide from the N-terminal segment of Ip-CANP9a recognized the taste pore region (Figure 3(b)) and taste bud cells (Figure 3(c)). Taken together with the northern analysis, these data suggest that Ip-CAPN9a is a chemosensory and digestive tract specific calpain in fish. The barbell of the catfish detects taste stimuli as does the tongue in mammals. According to the current views of the taste system, where the sense of taste is considered a “metabolic sense” acting both at the perceptual and the metabolic levels, it may not be surprising to find a digestive tract calpain within cells of the taste bud.
In spite of much effort, we could not detect a band on SDS-PAGE-western blot that would correspond in molecular weight (78 kDa) to Ip-CAPN9a (data not shown). Several explanations can be considered including 1) the likely high turnover rate for tissue-specific calpains [1] and 2) high calcium concentration around taste cells leading to calpain activation and autolysis. A similar anomaly is seen with human CAPN3. This calpain is difficult to detect by western blotting, most probably due to its high turnover rate and its lability after dissociation from connectin, to which CAPN3 binds through its IS2 region [31] .
Human CAPN9 and Ip-CAPN9a might be more fragile than human calpain 3. M- and μ-calpains cleave their N-terminal segments upon activation (autolysis). The antibody against Ip-CAPN9a was raised against its N- terminal segment (3 to 16 amino acids). The presumptive cleavage site on autolysis localizes near the end of this segment. We thus assume that the antibody recognition segment would be mostly released from Ip-CAPN9a upon its activation. Na+, Ca2+, Mg2+ ions are rich in catfish barbell mucin layer. This high calcium concentration could activate calpains during sample preparation.
The calpains have been implicated in degradation of the IP3 receptor [14] and of the enzyme, PLC beta-3 [15] . In a preliminary experiment [19] , we noted that the calpain inhibitor, the tripeptide, LLL, reduced the degradation of the IP3 receptor in catfish barbell upon stimulation of the preparation by the potent taste stimulus, L-ar- ginine. As there are no subtype specific calpain inhibitors, we cannot determine which calpain subtypes are responsible for the degradation of taste-related proteins.
Our discovery of Ip-CAPN9a should lead to the characterization of other teleost-related calpain subtypes. Studies of other tissue-specific type fish calpains will clarify how these calcium-dependent calpains are activated and inhibited under physiological conditions.
Acknowledgements
T.O thanks to the laboratory members in the Monell Chemical Senses Center and NFRI.
References
- Sorimachi, H., Toyama-Sorimachi, N., Saido, T.C., Kawasaki, H., Sugita, H., Miyasaka, M., Arahata, K., Ishiura, S. and Suzuki, K. (1993) Muscle-Specific Calpain, p94, Is Degraded by Autolysis Immediately after Translation, Resulting in Disappearance from Muscle. Journal of Biological Chemistry, 268, 10593-10605.
- Richard, I., Broux, O., Allamand, V., Fougerousse, F., Chiannilkulchai, N., Bourg, N., Brenguier, L., Devaud, C., Pasturaud, P., Roudaut, C., et al. (1995) Mutations in the Proteolytic Enzyme Calpain 3 Cause Limb-Girdle Muscular Dystrophy Type 2A. Cell, 81, 27-40. http://dx.doi.org/10.1016/0092-8674(95)90368-2
- Ma, H., Fukiage, C., Azuma, M. and Shearer, T.R. (1998) Cloning and Expression of mRNA for Calpain Lp82 from Rat Lens: Splice Variant of p94. Investigative Ophthalmology Visual Science, 39, 454-461.
- Sorimachi, H., Ishiura, S. and Suzuki, K. (1993) A Novel Tissue-Specific Calpain Species Expressed Predominantly in the Stomach Comprises Two Alternative Splicing Products with and without Ca(2+)-Binding Domain. Journal of Biological Chemistry, 268, 19476-19482.
- Lee, H.J., Sorimachi, H., Jeong, S.Y., Ishiura, S. and Suzuki, K. (1998) Molecular Cloning and Characterization of a Novel Tissue-Specific Calpain Predominantly Expressed in the Digestive Tract. Biological Chemistry, 379, 175-183. http://dx.doi.org/10.1515/bchm.1998.379.2.175
- Hata, S., Abe, M., Suzuki, H., Kitamura, F., Toyama-Sorimachi, N., Abe, K., Sakimura, K. and Sorimachi, H. (2010) Calpain 8/nCL-2 and Calpain 9/nCL-4 Constitute an Active Protease Complex, G-Calpain, Involved in Gastric Mucosal Defense. PLoS Genetics, 6, e1001040. http://dx.doi.org/10.1371/journal.pgen.1001040
- Yoshikawa, Y., Mukai, H., Hino, F., Asada, K. and Kato, I. (2000) Isolation of Two Novel Genes, Down-Regulated in Gastric Cancer. Japanese Journal of Cancer Research, 91, 459-463. http://dx.doi.org/10.1111/j.1349-7006.2000.tb00967.x
- Markmann, A., Schafer, S., Linz, W., Lohn, M., Busch, A.E. and Wohlfart, P. (2005) Down-Regulation of Calpain 9 Is Linked to Hypertensive Heart and Kidney. Cellular Physiology and Biochemistry, 15, 109-116. http://dx.doi.org/10.1159/000083643
- Sorimachi, H., Ishiura, S. and Suzuki, K. (1997) Structure and Physiological Function of Calpains. Biochemical Journal, 328, 721-732.
- Lin, G.D., Chattopadhyay, D., Maki, M., Wang, K.K., Carson, M., Jin, L., Yuen, P.W., Takano, E., Hatanaka, M., DeLucas, L.J. and Narayana, S.V. (1997) Crystal Structure of Calcium Bound Domain VI of Calpain at 1.9 A Resolution and Its Role in Enzyme Assembly, Regulation, and Inhibitor Binding. Nature Structural Molecular Biology, 4, 539- 547. http://dx.doi.org/10.1038/nsb0797-539
- Blanchard, H., Grochulski, P., Li, Y., Arthur, J.S., Davies, P.L., Elce, J.S. and Cygler, M. (1997) Structure of a Calpain Ca2+-Binding Domain Reveals a Novel EF-Hand and Ca2+-Induced Conformational Changes. Nature Structural Biology, 4, 532-538. http://dx.doi.org/10.1038/nsb0797-532
- Lee, H.J., Tomioka, S., Kinbara, K., Masumoto, H., Jeong, S.Y., Sorimachi, H., Ishiura, S. and Suzuki, K. (1999) Characterization of a Human Digestive Tract-Specific Calpain, nCL-4, Expressed in the Baculovirus System. Archives of Biochemistry and Biophysics, 362, 22-31. http://dx.doi.org/10.1006/abbi.1998.1021
- Azarian, S.M., King, A.J., Hallett, M.A. and Williams, D.S. (1995) Selective Proteolysis of Arrestin by Calpain. Molecular and Its Effect on Rhodopsin Dephosphorylation. The Journal of Biological Chemistry, 270, 24375-24384. http://dx.doi.org/10.1074/jbc.270.41.24375,
- Magnusson, A., Haug, L.S., Walaas, S.I. and Ostvolt, A.C. (1993) Calcium-Induced Degradation of the Inositol(1,4,5)- Trisphosphate Receptor/Ca2+-Channel. FEBS Letters, 323, 229-232. http://dx.doi.org/10.1016/0014-5793(93)81345-Z
- Banno, Y., Nakashima, S., Hachiya, T. and Nozawa, Y. (1995) Endogenous Cleavage of Phospholipase C-beta3 by Agonist-Induced Activation of Calpain in Human Platelets. The Journal of Biological Chemistry, 270, 4318-4324. http://dx.doi.org/10.1074/jbc.270.9.4318
- Greenwood, A.F. and Jope, R.S. (1994) Brain G-Protein Proteolysis by Calpain: Enhancement by Lithium. Brain Research, 636, 320-326. http://dx.doi.org/10.1016/0006-8993(94)91031-6
- Shinozaki, K., Maruyama, K., Kume, H., Tomita, T., Saido, T.C., Iwatsubo, T. and Obata, K. (1998) The Presenilin 2 Loop Domain Interacts with the Mu-Calpain C-Terminal Region. International Journal of Molecular Medicine, 1, 797- 799.
- Roper, S.D. (2006) Signaling in the Chemosensory Systems. Cellular and Molecular Life Sciences, 63, 1494-1500. http://dx.doi.org/10.1007/s00018-006-6112-9
- Ookura, T., Koyama, E., Puchalski, R.B., Teeter, J.H., Bayley, D.L. and Brand, J.G. (1997) Molecular Cloning of Calpain from Taste Epithelium of the Channel Catfish, Ictalurus punctatus. Chemical Senses, 22,764-765.
- Katoh, K., Kuma, K., Toh, H. and Miyata, T. (2005) MAFFT Version 5: Improvement in Accuracy of Multiple Sequence Alignment. Nucleic Acids Research, 33, 511-518. http://dx.doi.org/10.1093/nar/gki198
- Felsenstein, J. (1981) Evolutionary Trees from DNA Sequences: A Maximum Likelihood Approach. Journal of Molecular Evolution, 17, 368-376. http://dx.doi.org/10.1007/BF01734359
- Stamatakis, A. (2006) RAxML-VI-HPC: Maximum Likelihood-Based Phylogenetic Analyses with Thousands of Taxa and Mixed Models. Bioinformatics, 22, 2688-2690. http://dx.doi.org/10.1093/bioinformatics/btl446
- Jones, D.T., Taylor, W.R. and Thornton, J.M. (1992) The Rapid Generation of Mutation Data Matrices from Protein Sequences. Computer Applications in the Biosciences, 8, 275-282.
- Tamura, K., Dudley, J., Nei, M. and Kumar, S. (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Molecular Biology and Evolution, 24, 1596-1599. http://dx.doi.org/10.1093/molbev/msm092
- Toyohara, H. and Makinodan, Y. (1989) Comparison of Calpain I and Calpain II from Carp Muscle. Comparative Biochemistry and Physiology Part B: Comparative Biochemistry, 92, 577-581. http://dx.doi.org/10.1016/0305-0491(89)90134-X
- Salem, M., Nath, J., Rexroad, C.E., Killefer, J. and Yao, J. (2005) Identification and Molecular Characterization of the Rainbow Trout Calpains (Capn1 and Capn2): Their Expression in Muscle Wasting during Starvation. Comparative Biochemistry and Physiology Part B: Biochemistry and Molecular Biology, 140, 63-71. http://dx.doi.org/10.1016/j.cbpc.2004.09.007
- Salem, M., Rexroad, C.E. and Yao, J. (2006) Identification of a Novel Gill-Specific Calpain from Rainbow Trout (Oncorhynchus mykiss). Fish Physiology and Biochemistry, 32, 1-6. http://dx.doi.org/10.1007/s10695-005-0560-2
- Jekely, G. and Friedrich, P. (1999) The Evolution of the Calpain Family as Reflected in Paralogous Chromosome Regions. Journal of Molecular Evolution, 49, 272-281. http://dx.doi.org/10.1007/PL00006549
- Maki, M., Narayana, S.V.L. and Hitomi, K. (1997) A Growing Family of the Ca2+-Binding Proteins with Five EF- Hand Motifs. Biochemical Journal, 328, 718-720.
- Theopold, U., Pinter, M., Daffre, S., Tryselius, Y., Friedrich, P., Nassel, D.R. and Hultmark, D. (1995) CalpA, a Drosophila Calpain Homologue Specifically Expressed in a Small Set of Nerve, Midgut, and Blood Cells. Molecular and Cellular Biology, 15, 824-834.
- Sorimachi, H., Tsukahara, T., Okada-Ban, M., Sugita, H., Ishiura, S. and Suzuki, K. (1995) Identification of a Third Ubiquitous Calpain Species-Chicken Muscle Expresses Four Distinct Calpains. Biochimica et Biophysica Acta (BBA)- Gene Structure and Expression, 1261, 381-393. http://dx.doi.org/10.1016/0167-4781(95)00027-E
NOTES

*Corresponding authors.