Health
Vol.4 No.10A(2012), Article ID:24249,4 pages DOI:10.4236/health.2012.430145
Genome-wide comparison inferred the origin and evolution of B-cell epitopes on the proteins of human influenza A virus
![]()
1Centro de Biotecnología Genómica, Instituto Politécnico Nacional, Reynosa, México;
*Corresponding Author: gxianwu@yahoo.com
2Departamento de Inmunología, Escuela Nacional de Ciencias Biológicas, Unidad Profesional Lázaro Cárdenas, Instituto Politécnico Nacional, Reynosa, México
Received 14 September 2012; revised 12 October 2012; accepted 24 October 2012
Keywords: Influenza A Virus; H1N1; Pandemic; Epitopes; Genome Comparison
ABSTRACT
The novel strain H1N1 caused the outbreak of first pandemic influenza in 21 century. Now it is a common component of current seasonal influenza viruses. The recent transmission and plentiful genome sequences available provided a good opportunity to study the origin and evolution of epitopes on the proteins of human influenza virus. In the present study, the B-cell epitope compositions in the pandemic strains, circulating traditional seasonal strains, swine strains as well as highly virulent avian strain H5N1 were identified with the aid of the Immune Epitope DataBase (IEDB) and were compared at genomic level. A total of 14210 distinct sequences down-loaded from NCBI database were used for analysis. Some epitopes on proteins HA or NA, not conserved in recent seasonal strains, were found in 2009 pandemic strains but existed in the early human strains (1919-1935). The pandemic strain shared higher conserved epitopes with “bird flu” virus H5N1 than classic human seasonal strains. The epitopes that could exist at common antigenic regions of HA protein are needed to further identify. The genetic exchanges between human and swine population by transmission was very active but the princepal side of the transmission could be from swine to human. These results provided valuable information on influenza A virus evolution and transmission by means of epitope analysis at genomic level.
1. INTRODUCTION
In the interaction of influenza virus with human host, T cells and antibodies of immune system responses to the viruses are triggered by the recognition of viral epitopes by immune cell receptors, which is the key step of immune protection from infection. If the protein epitopes from different viral strains have identical sequences or even have subtle changes in sequence but keep the similar conformation and physicochemical properties in the interaction region, the immune recognition should occur in the same way. It is thus reasonable that the epitopes can be identified by the comparison of amino acid sequences of viral homologous proteins among serotype strains due to their similar function and conformation. In addition, the protective immunity is directly related to the composition of viral epitopes and the proportional pre-existing immunity of host, particularly the B-cellproduced neutralizing antibodies induced by the epitopes in new strains. Therefore, the identification of B-cell epitopes carried by the influenza viruses should be helpful for understanding the basis for viral infection and recombinant vaccine production.
During the post-pandemic period, 2009 H1N1 viruses, as a regular seasonal influenza virus, are co-circulating in many parts of the world along with classic seasonal influenza viruses. The data from Centers for Disease Control and Prevention (CDC) showed that in 2010-2011 influenza seasons, the pandemic strains still accounted for 26.5% of the influenza A infection cases in USA
(http://www.cdc.gov/flu/weekly/weeklyarchives2010-2011/weekly39.htm). The seasonal virus composition has been thus apparently changed due to the pandemic outbreak. This change should result in the pre-existing immunity alteration in human population. Thus, the comparison of epitope composition in the current seasonal viral strains (including the 2009 pandemic strains) with those of animal viral strains could give valuable reference data for the origin and evolution of epitopes on the proteins of human influenza A virus. In the present study, we focus on the B-cell epitope analysis at genomic level, comparing the epitope compositions of pandemic strain, classic seasonal strains, swine strains as well as highly virulent avian strain H5N1, and hope that the results could produce information to counteract new flu infection in the future.
2. MATERIALS AND METHODS
2.1. Searching the Amino Acid or Nucleotide Sequences of Each Gene in Influenza A Genomes from NCBI
The following parameters were used for searching the pandemic sequences of each viral protein in the influenza virus resource from the National Center for Biotechnology Information (NCBI) [1]:
Virus Species = Influenza A AND (Subtype = H1N1);
Host = Human;
Year from 2009 to 2010;
Complete sequences only;
INCLUDE = Pandemic only;
EXCLUDE = Lab strains;
EXCLUDE = Flu project;
Collapse similar sequences.
For searching the sequences of recent seasonal strains in the database, the following parameters were used:
Virus Species = Influenza A AND (Subtype = H1N1 or H3N2);
Host = Human;
Year from 1988 to 2008;
Complete sequences only;
EXCLUDE = Pandemic only;
INCLUDE = Lab strains;
INCLUDE = Flu project;
Collapse similar sequences.
For searching the sequences of swine strains and highly pathogenic avian strain H5N1, it is necessary to change the corresponding parameters. After eliminating the identical sequences, a total of 14210 distinct sequences were downloaded for analysis in the present research.
2.2. Searching the Epitopes in the Immune Epitope DataBase (IEDB)
The epitopes of influenza strains were extracted from the database IEDB [2]. The present study only focuses on B-cell epitopes for evaluating antibody-mediated protective immunity. Only the epitopes with 100% identity were considered. If the same protein of influenza A strains (H1, H3 and H5) has the same amino acid sequence of epitopes, they share the epitopes. If the same protein of the same serotype of influenza A strains has the same amino acids of discontinuous epitopes at the same relative sites, they share the epitopes. In the present study, discontinuous epitopes were only used for the conserved epitope analysis in pandemic H1N1 and seasonal influenza serotypes. In other words, discontinuous epitopes will not be taken account in the comparison with the strains from swine or strain H5N1. In order to include as much more information as possible, the searching of epitopes was based on each chromosomal fragment, not based on each complete genome. Considering that the structural constraints associated with their immune recognition could be similar cross species because B-cell epitopes were generally defined in different species [3], B-cell epitopes mapped in the context of any host organism were included. The epitopes with nested sequences were considered as separate epitopes because it could represent different epitopes. Even the nested epitopes represent the same individual epitope, the result of comparison with separate epitopes will be not affected because the test was performed under the same background. The shortest analyzed sequence of epitope was limited to be ≥5 residues for linear epitopes and ≥4 residues for discontinuous epitopes.
2.3. Searching the Epitope Sequences Conserved in Strains of 2009 Pandemic H1N1 Influenza against the Recent Human Seasonal Strains
The conserved epitopes of pandemic strains were shown by the number of epitopes in pandemic strain that are present in the seasonal strain, which generally indicates the reference of pre-existing immunity level to human host. After obtaining the required epitopes of strains at certain duration, the identification of conserved epitopes was manually implemented. In order to understand the pre-existing immunity before the pandemic, the epitopes conserved in the pandemic strains were obtained by comparison of epitopes from pandemic strains with those from recent seasonal influenza strains (H1N1 and H3N2) circulating in the previous 20 years (1988 to 2008) [4]. In regard to the post-pandemic case, the components of current seasonal strains have been changed due to the pandemic outbreak and should contain the pandemic strain. The comparisons were performed as the epitopes of current seasonal strains including pandemic strains with those of swine strains as well as the high virulent “bird flu” strain H5N1, which was more concerned about. The B-cell-related pre-existing immunity in general human population was considered by the epitopes that were 100% conserved in 30% or more of the strains of any given years [4]. In other words, the epitopes in a viral population represent the B-cell-related pre-existing immunity in human population when the viruses were isolated or sampled from human host. In present study, the surface proteins, in addition to HA, NA (neuraminidase), included the protein M2 (ion channel) due to the existence of ectodomain on the virus surface, which also could provide the epitopes for neutralization function.
2.4. Multiple Sequence Alignment
The MAFFT software [5] was employed to generate the sequence alignments that are necessary to create the phylogenetic trees and the graphical logos.
2.5. Phylogenetic Analysis
The multiple sequence alignment files generated on the basis of NA and HA protein sequences were processed with the SeaView software [6] to construct phylogenetic trees using the neighbor-joining method with bootstrap of 1000 replications.
2.6. The Production of Logo Map for Epitope 97748 (Accession Number in IEDB) by Historical Times
The web logo software was employed to describe graphically the similarity sequence of epitope 97748 using the alignments of NA protein through times [7].
2.7. Tridimensional Structure of HA Protein Epitopes in 2009 Pandemic Strains
The hemagglutinin from the H1N1 strain A/California/ 04/2009 [8] was used for describing the localization of B-cell epitopes. The coordinates of the crystal structure were obtained from the PDB database (http://www.pdb.org) (PDB ID: 3LZG). UCSF Chimera Software [9] were performed for visualizing, mapping the epitopes onto the protein and producing the images.
3. RESULTS
3.1. Epitope Composition Comparison between Pandemic Strains and Traditional Seasonal Influenza Strains (H1N1 and H3N2)
The pandemic strain was recently transmitted from swine and abundant sequences can be available. Thus we first compared this strain with traditional seasonal influenza viruses.
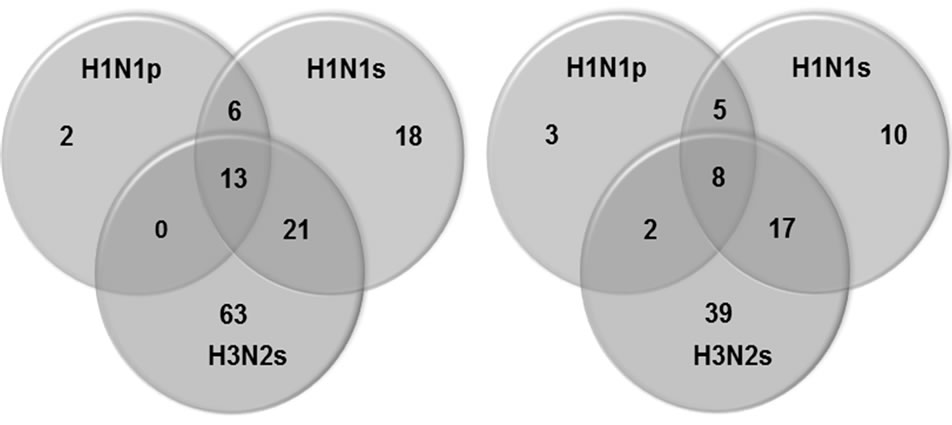
A total of 1148 distinct fragments of 2009 pandemic strains were applied to search the epitopes in the database. The 21 B-cell epitopes were found and distributed in the protein HA(7), NA(3), M2(1), M1(2), NP(5), PB1(2) and PB2(1). 16% (19/121) of the B-cell conserved epitopes in seasonal strains (seasonal H1N1 or H3N2) were shared in 2009 pandemic strains as shown in Figure 1(a), of which the neutralizing epitopes from HA, NA and M2 only consisted of 9% (11/121). There were 3 epitopes present with less than 4% in the population, indicating that the rare epitopes could be mutations occurred in transmission during the pandemic. If just considered the epitopes in the general population, the conserved epitopes kept 19% (15/81) (Figure 1(b)) and neutralizing conserved epitopes had 12% (10/81). The results indicated that B-cell epitopes in pandemic strain, particularly the epitopes in surface proteins HA, NA, and M2, only conserved a small part of epitopes in seasonal strains, similar but less to the previous report [4]. Crossreaction experiments have also confirmed it [10-14].
Two epitopes were new to recent seasonal strains. One is discontinuous epitope (accession number in IEDB: 91060 with sequence N140, P152, K168, K169, corresponding the pandemic strain N142, P154, K170, K171) in HA protein, located at the antigenic region Sa (N142, K170, K171) and Ca2 region (P154). It seems that this epitope has only been found in the population of swine H1N1 strains, as present in the 1930 swine Iowa strains, swine strain in human infection at the 1976 outbreak in Fort Dix, New Jersey and the current swine population. Another is the epitope on NA protein with the accession number 97748 in IEDB as the sequence at sites 448 - 468 (novel pandemic strain as reference), VNSDTVGWSWPDGAELPFTID. It occupied 88% pandemic population. It is important to note that this percentage should be underestimated since this epitope is located at the end of the protein and some of the examined sequences were incomplete, missing few amino acids at this region (but the NCBI database identified the sequences as full-length proteins). If these samples are removed from the data, this epitope is present as 99% of pandemic genomes. Compared to early human H1N1 strains, this epitope existed in most strains isolated during 1919-1934, as shown in Figure 2. It is only one amino acid different from the strain 1918 (T467D). However, we cannot deny the existence of this epitope in 1918 H1N1 population
 (a) (b)
(a) (b)
Figure 1. Comparison of B-cell epitopes between novel pandemic strain (H1N1p) and recent seasonal strains H1N1(H1N1s) and H3N2(H3N2s) (1988-2008). (a) The epitopes present on all the sequenced genomes of each viral group; (b) The epitopes present on ≥30% of the sequenced genomes of each viral group.

Figure 2. The logo presentation of the epitope with accession number 97748 and its homolog on NA protein in human historical H1N1 strains. The numbers under the logos represent the corresponding amino acid position of 2009 pandemic strain. Each logo consists of stacks of letters, one stack for each position in the sequence. The overall height of each stack indicates the sequence conservation at that position (measured in bits), whereas the height of symbols within the stack reflects the relative frequency of the corresponding amino acid at that position. The right numbers represented the years of sample collection. The number on the figure bottom indicated the location of amino acids on the protein using the novel pandemic strain as a reference.
due to the unique genome data available from samples of this year. In a comparison of epitope sequences in the strains from 1935 to 2008, the evolutionary changes in this epitope can be inferred. N450 existed in all the 1918- 1934 strains but more and more strains fixed this mutation N450D as circulating in humans. The most obvious changes were the amino acids, V454 and G455, which have been replaced by A454 and N455, fixed in the later seasonal viral population. Therefore, this epitope existed or was similar to that in the strains before 1935. Another long epitope (DNSIRIGSKGDVFVIREPFISCSHLECRFFLTQGALLNDKH) existed in recent seasonal strains, “bird flu” H5N1 [15] and partial strains in the population between 1918-1934, such as the strain A/PuertoRico/8/ 1934, but only occurred in one 2009 pandemic strain ADH29478, while in general 2009 pandemic population, the corresponding sequences had only one amino acid difference (H126P).
Only a few epitopes are available on NA protein, which may be the reason why the previous researches did not compare this protein in more detail. Although the presented epitopes only showed the parts of the whole NA protein epitopes, higher similarity in protein sequences could indicate more epitopes that are similar present in the strains in general sense. We thus compared the whole sequence similarity of NA between strains by phylogenetic analysis, including the samples from Eurasian swine, American avian, Classical swine, and Human seasonal strains. The comparisons based on the sequences of protein showed that the NA of novel 2009 strain was more similar to early strains than recent human seasonal strains (Figure 3). Actually, the novel strain has 87% identity to the 1918 pandemic strain but only 81% to the vaccine seasonal strain ACA28845 A/Brisbane/59/2007/ 01. Therefore, the epitopes on NA in 2009 pandemic strains should be generally more similar to those in early strains of 20th century than in recent seasonal strains.
3.2. Epitope Composition Comparison between Current Seasonal Influenza Strains (H1N1, H3N2 and Pandemic) and Other Influenza A Groups
Since the novel pandemic strain is still an important composition of current seasonal influenza viruses, leading to the change in the pre-existing immunity in human population compared to the situation before the pandemic. Therefore, we compared the epitopes of current seasonal viruses, which included the pandemic strains, with those of swine strains and high virulent strain H5N1 in order to find some implications in immune protection on human. The 96% (112/117) current seasonal epitopes were shared with swine viruses, as shown in Figure 4(a), suggesting that the human viruses have close relation with swine viruses (inter-host transmission or transmission by reassortant strains). If only considering the epitopes in general population, the same result was clearer (Figure 4(b)). All the epitopes in current seasonal strains were conserved in swine population. H5N1 shared much more epitopes with swine viruses than with human viruses and there was no epitope that only shared with human viruses. There were 28 epitopes shared among swine, H5N1 and human viruses. However, the viral populations in H5N1 and swine viruses contained many new epitopes to human, which should be a threat to human population by reassortant strains.
The swine viruses had more novel epitopes than those from humans due to the existence of more serotypes of swine strains. We thus separately compared the serotypes H1N1 and H3N2 from swine viruses as well as current seasonal human viruses, as shown in Figure 5. Only a few epitopes are specific to swine or to human population. The further comparison showed that all the epitopes specific to human strains or to swine strains in serotypes H1N1, H3N2 belonged to small populations (less than 30%) (not shown). The results indicated that the epitopes specific to human strains or to swine strains could be the consequence of viral mutations in each host. Of course, it is also possible that small proportional epitopes come from reassortant strains by sporadic infection on human.
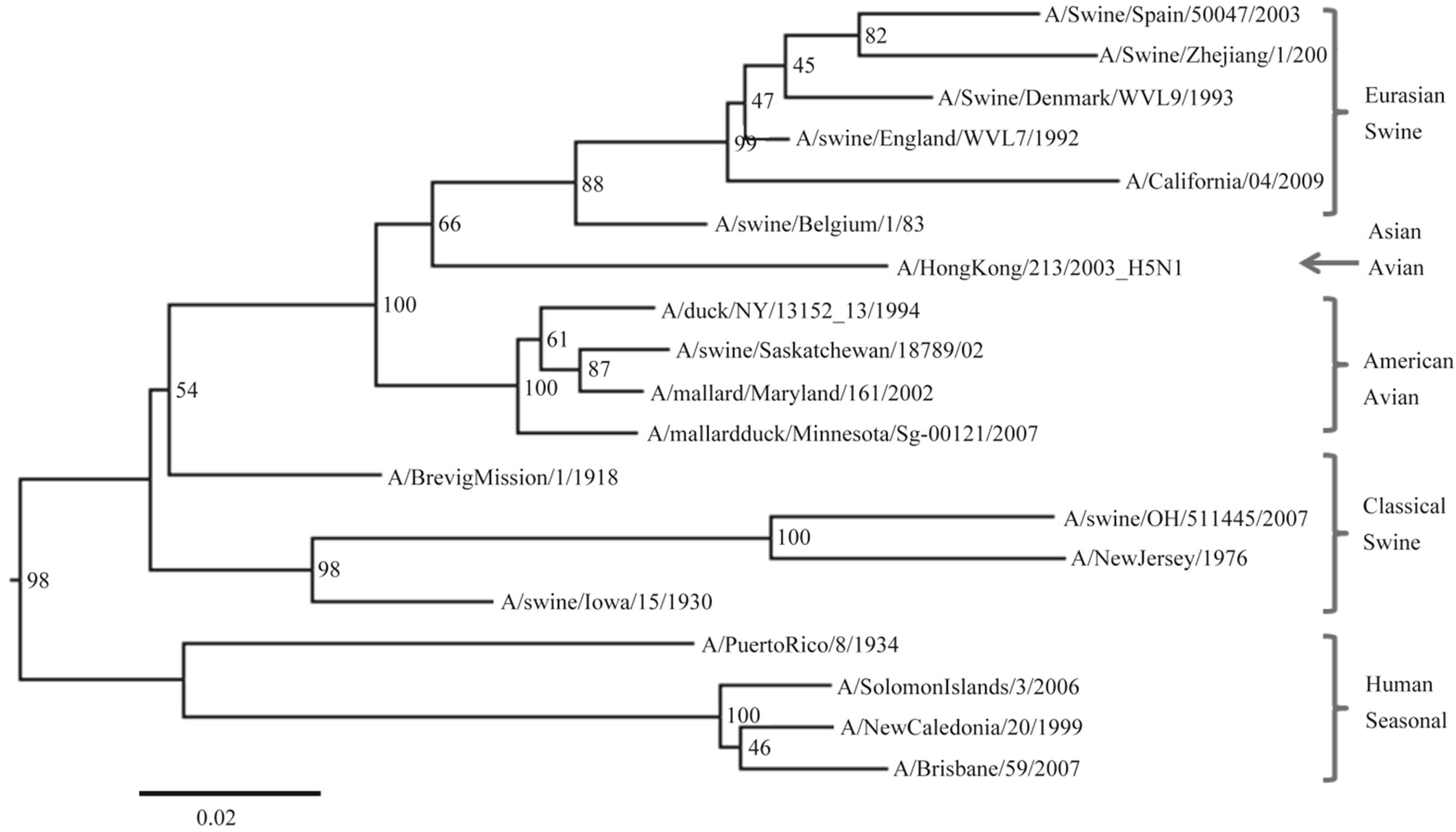
Figure 3. Phylogeny analysis of 2009 pandemic H1N1 strain with the relation to other groups of strains based on the sequences of NA protein. The strain groups have been indicated in the figure.
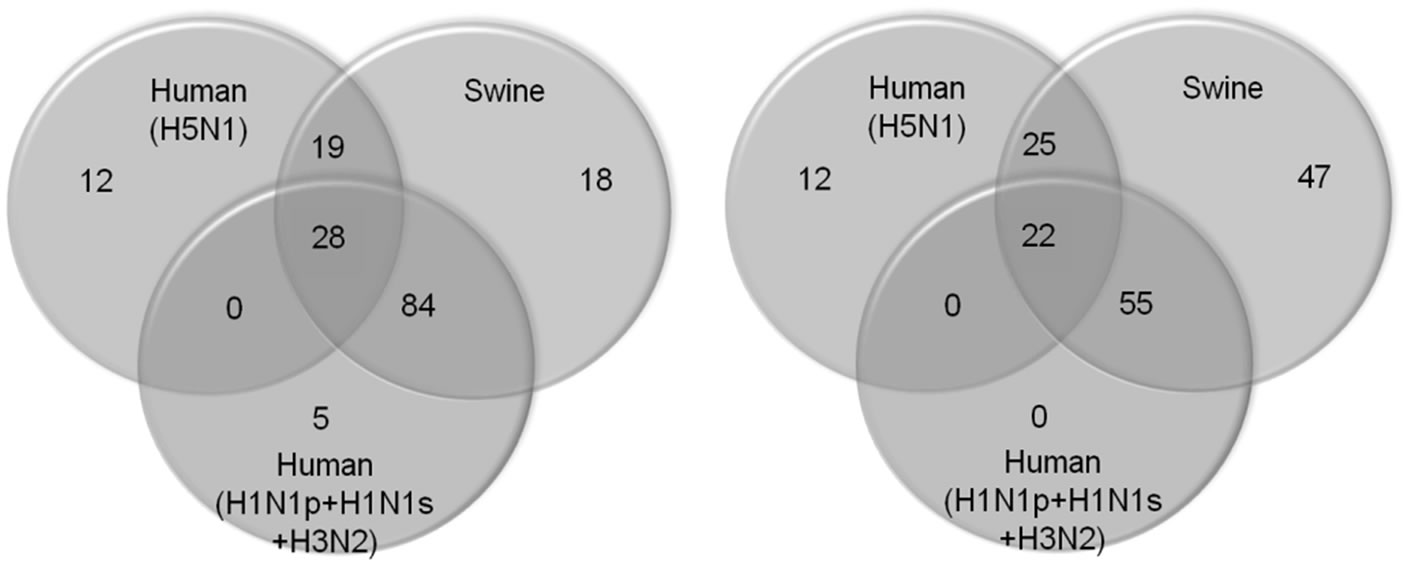
Figure 4. Comparison of B-cell epitopes among current seasonal strains (including the novel pandemic strain), the swine strains and the highly virulent strain H5N1 since 1988. (a) present on all the sequenced genomes of each viral group (discontinuous epitopes not included); (b) The epitopes present ≥30 percentage of the sequenced genomes of human seasonal viral groups.
4. DISCUSSION
The epitopes borne on the 44 genomes of 2009 H1N1 strains have been examined at the early stage of pandemic outbreak [4] with the help of this database Immune Epitope Database (IEDB) [16]. No epitopes different from the recent seasonal human influenza virus have been found in the comparison. The subsequent related researches commonly focus on the comparison between novel pandemic strains with seasonal strains [4,17,18], or only on hemagglutinin (HA) protein [19,20] or T-cell epitopes [18,21,22]. The highest proportion of epitopes in the database IEDB is from HA protein, showing that HA protein is the most important B-cell epitope resource in influenza A virus. HA protein contain the regions rich in epitopes that are located at the globular cap of homotrimer and have been divided into four structural sites, Sa and Sb, proximal to the receptor-binding pocket, Ca (Ca1 and Ca2) at the subunit surface and Cb within the vestigial esterase domain [23]. The structural comparison between the 2009 pandemic strain and its closely related 1918 pandemic strain showed that there were 10

Figure 5. Comparison of B-cell epitopes between current seasonal strains H1N1 and H3N2 with the swine corresponding serotypes of strains since 1988. The arrows indicated intertransmission between them.
amino acid alternation in these antigenic regions [24]. We found two epitopes in novel pandemic strains that were not conserved in recent seasonal viruses, one of which was discontinuous epitope in HA protein and located at the antigenic regions Sa and Ca2. However, the six linear epitopes in HA protein were sited outside of those regions, epitope ID 62335 (SVSSFERFEIFPK) (123 - 136, the novel pandemic strain as a reference) close to the region Sa at the cap and others located at stem or close to membrane-based domain (Figure 6). Some epitopes at the cap of HA were not definite in novel pandemic strain but their antibody binding sites have been determined at Sa or close to Sa or Sb regions [14,25]. Therefore, the epitopes at the common antigenic zones of HA globular cap principally did not yet confirmed by experiments in pandemic strain. Recently, a novel epitope between the receptor-binding pocket and the Ca2 antigenic site was found [26]. The epitopes in the globular head typically elicit strain-specific responses because of the hypervariability of this region. It could be predicted that more epitopes will be discovered in the future for this pandemic strain.
Our comparison on NA protein in present study is consistent with that obtained from the research on HA [8]. These authors compared the crystal structures of HA1 and showed that the antigenic region, particularly within the Sa antigenic site, is similar to those of human H1N1 viruses circulating early in the 20th century but much more different from those of recent human seasonal strains. Similarly, our analysis also showed that the epi-
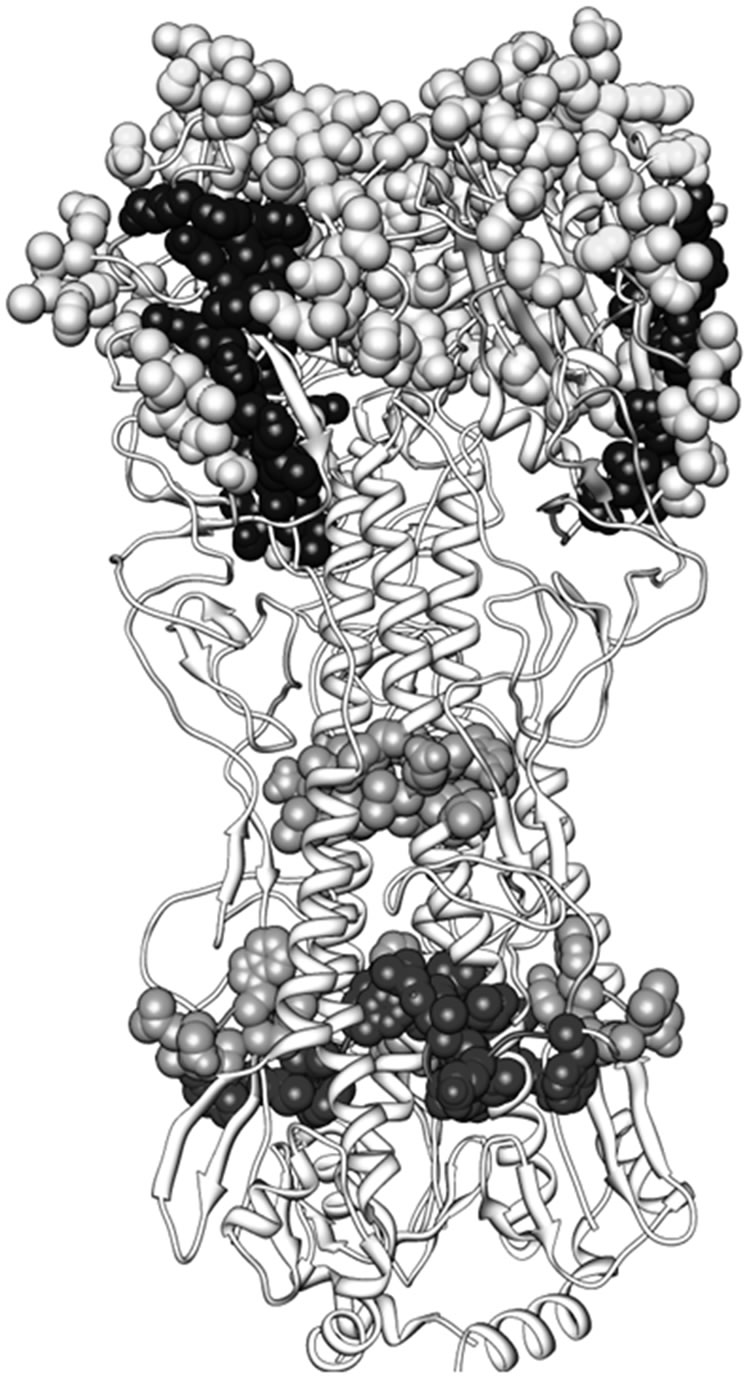
Figure 6. Tridimensional structure of homotrimer HA, indicating the location of common antigenic regions of HA (light blue) and the B-cell epitopes from 2009 pandemic strains (other colors). Only one discontinuous epitope (accession number in IEDB: 91060) (yellow) was located at the common antigenic regions.
topes on NA in 2009 pandemic strains share more similarities to H1N1 strains circulating in 1918-1934 than those in recent seasonal strains. Thus, both proteins (HA, NA) of the most importantly immune effect of the novel H1N1 strains provided the same evidence on human preexisting immunity that can explain why the aged people have less clinical symptom than young ones.
The fact that all the epitopes in the current seasonal human viruses in general population were shared with swine viruses implies that the genetic exchange existed between them. This genetic exchange could be caused by inter-host transmission of strains after chromosomal reassortment or direct inter-host transmission of strains, e.g. the novel pandemic strain. The 2009 pandemic strain H1N1 occurred in April in Mexico and USA in 2009 [27] and sooner it can be found the infection in swine population in Canada and other places of the world later [28- 31]. In 1918, both human and swine populations showed the similar symptoms caused by 1918 pandemic H1N1 virus at similar time so that 1918 pandemic strain resulted from the swine to human or vice verse is still blurred, except the inter-transmission between human and pigs [32-35]. The present epitope analysis showed the coincidence with inter-host genetic exchange, sometimes direct inter-host transmission, of human seasonal influenza viruses with the swine population. However, considering the lack of new human viral epitopes to the general swine virus population (Figure 4(b)), the genetic exchange could occur mostly from swine to human (Figure 5). Multiple reassortment between novel pandemic strains and endemic influenza viruses in pigs has been reported [36,37]. A new swine-origin H3N2 strain that contains M2 gene from 2009 pandemic strain has been recently isolated from several children in Pennsylvania and Indiana [38]. Therefore, such genetic exchange is very active along the evolutionary history.
It is necessary to note that all the data in the IEDB have been confirmed by experiments in the literature. As for the identified epitopes in this study, we only consider the 100% identity of epitopes for comparison. Because of the existence of epitope’s conformational plasticity that allows the alternative amino acids, the actual pre-existing immunity could be higher than our estimate. However, as regard to effectual immunity protection, people still need to face two main problems. One is the new epitopes that could be brought to human by new strains. These new epitopes may come from the genetic drift in animal host by the inter-transmission strains or the gene-reassortant strains. The other is that the virulence related to certain proteins or immunity of epitopes depends on the antibody titers in the infected individuals. As the case of 2009 pandemic influenza, the causing virus contained new epitopes and the infected individuals dominantly produced low titers of the broadly cross-reactive antibodies, which were unable to effectively provide protection from infection. This phenomenon represents a challenge for future research.
One limiting factor for this epitope analysis, based on the comparison of protein amino acid sequences, is not to take into account the glycosylation sites, which could affect the antibody binding or recognition because of the special barrier of glycans. HA and NA both are antigenic glycoproteins and have different glycosylation patterns in different serotypes of strains. Although the potential glycosylation sites could be predicted somehow on the proteins but the actual interaction on the immune recognition needs to be tested by specific experiments.
5. ACKNOWLEDGEMENTS
This work was supported by Secretaría de Investigación y Posgrado del Instituto Politécnico Nacional, México (Grant numbers: 20101372 and 20110896) and by the Consejo Nacional de Ciencia y TecnologíaMéxico (Fondo Mixto de Fomento a la Investigación Científica y Tecnológica, CONACYTGobierno del Estado de Oaxaca. Grant number. 122497). Xianwu Guo, Mario A. Rodríguez-Pérez and Ma. Isabel Salazar hold the scholarships from Comisión de Operación y Fomento de Actividades and Estímulos al Desempeño de los Investigadores/Instituto Politécnico Nacional.
REFERENCES
- Bao, Y., Bolotov, P., Dernovoy, D., Kiryutin, B., Zaslavsky, L., Tatusova, T., et al. (2008) The influenza virus resource at the National Center for Biotechnology Information. Journal of Virology, 82, 596-601. doi:10.1128/JVI.02005-07
- Peters, B., Sidney, J., Bourne, P., Bui, H.H., Buus, S., Doh, G., et al. (2005) The design and implementation of the immune epitope database and analysis resource. Immunogenetics, 57, 326-336. doi:10.1007/s00251-005-0803-5
- Nguyen, H.H., Zemlin, M., Ivanov, I.I., Andrasi, J., Zemlin, C., Vu, H.L., et al. (2007) Heterosubtypic immunity to influenza A virus infection requires a properly diversified antibody repertoire. Journal of Virology, 81, 9331- 9338. doi:10.1128/JVI.00751-07
- Greenbaum, J.A., Kotturi, M.F., Kim, Y., Oseroff, C., Vaughan, K., Salimi, N., et al. (2009) Pre-existing immunity against swine-origin H1N1 influenza viruses in the general human population. Proceedings of the National Academy of Sciences of USA, 106, 20365-20370. doi:10.1073/pnas.0911580106
- Katoh, K. and Toh, H. (2008) Recent developments in the MAFFT multiple sequence alignment program. Brief in Bioinformatics, 9, 286-298. doi:10.1093/bib/bbn013
- Gouy, M., Guindon, S. and Gascuel, O. (2010) SeaView version 4: A multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Molecular Biology and Evolution, 27, 221-224. doi:10.1093/molbev/msp259
- Crooks, G.E., Hon, G., Chandonia, J.M. and Brenner, S.E. (2004) WebLogo: A sequence logo generator. Genome Research, 14, 1188-1190. doi:10.1101/gr.849004
- Xu, R., Ekiert, D.C., Krause, J.C., Hai, R., Crowe Jr., J.E. and Wilson, I.A. (2010) Structural basis of preexisting immunity to the 2009 H1N1 pandemic influenza virus. Science, 328, 357-360. doi:10.1126/science.1186430
- Pettersen, E.F., Goddard, T.D., Huang, C.C., Couch, G.S., Greenblatt, D.M., Meng, E.C., et al. (2004) UCSF Chimera: A visualization system for exploratory research and analysis. Journal of Computational Chemistry, 25, 1605- 1612. doi:10.1002/jcc.20084
- Hancock, K., Veguilla, V., Lu, X., Zhong, W., Butler, E.N., Sun, H., et al. (2009) Cross-reactive antibody responses to the 2009 pandemic H1N1 influenza virus. The New England Journal of Medicine, 361, 1945-1952. doi:10.1056/NEJMoa0906453
- Rizzo, C., Rota, M.C., Bella, A., Alfonsi, V., Declich, S., Caporali, M.G., et al. (2010) Cross-reactive antibody responses to the 2009 A/H1N1v influenza virus in the Italian population in the pre-pandemic period. Vaccine, 28, 3558-3562. doi:10.1016/j.vaccine.2010.03.006
- Itoh, Y., Shinya, K., Kiso, M., Watanabe, T., Sakoda, Y., Hatta, M., et al. (2009) In vitro and in vivo characterization of new swine-origin H1N1 influenza viruses. Nature, 460, 1021-1025.
- Huang, D.T., Shao, P.L., Huang, K.C., Lu, C.Y., Wang, J.R., Shih, S.R., et al. (2011) Serologic status for pandemic (H1N1) 2009 virus, Taiwan. Emerging Infectious Diseases, 17, 76-78. doi:10.3201/eid1701.100014
- Li, G.M., Chiu, C., Wrammert, J., McCausland, M., Andrews, S.F., Zheng, N.Y., et al. (2012) Pandemic H1N1 influenza vaccine induces a recall response in humans that favors broadly cross-reactive memory B cells. Proceedings of the National Academy of Sciences of USA, 109, 9047-9052. doi:10.1073/pnas.1118979109
- Khurana, S., Suguitan Jr., A.L., Rivera, Y., Simmons, C.P., Lanzavecchia, A., Sallusto, F., et al. (2009) Antigenic fingerprinting of H5N1 avian influenza using convalescent sera and monoclonal antibodies reveals potential vaccine and diagnostic targets. PLoS Medicine, 6, e1000049. doi:10.1371/journal.pmed.1000049
- Peters, B., Sidney, J., Bourne, P., Bui, H.H., Buus, S., Doh, G., et al. (2005) The immune epitope database and analysis resource: From vision to blueprint. PLOS Biology, 3, e91. doi:10.1371/journal.pbio.0030091
- Zhou, J.J., Tian, J., Fang, D.Y., Liang, Y., Yan, H.J., Zhou, J.M., et al. (2011) Analysis of antigen epitopes and molecular pathogenic characteristics of the 2009 H1N1 pandemic influenza A virus in China. Acta Virologica, 55, 195-202. doi:10.4149/av_2011_03_195
- De Groot, A.S., Ardito, M., McClaine, E.M., Moise, L. and Martin, W.D. (2009) Immunoinformatic comparison of T-cell epitopes contained in novel swine-origin influenza A (H1N1) virus with epitopes in 2008-2009 conventional influenza vaccine. Vaccine, 27, 5740-5747. doi:10.1016/j.vaccine.2009.07.040
- Rodriguez-Alvarez, M., Velasco-Velasco, A.M., AlvarezAnell, N.J., Jimenez-Corona, M.E. and de Leon-Rosales, S.P. (2009) Identification of seasonal vaccine hemagglutinin subtype 1 (H1) epitopes in Mexican isolates of the new influenza A (H1N1) 2009 virus. Archives of Medical Research, 40, 687-692. doi:10.1016/j.arcmed.2009.12.002
- Sun, Y., Shi, Y., Zhang, W., Li, Q., Liu, D., Vavricka, C., et al. (2010) In silico characterization of the functional and structural modules of the hemagglutinin protein from the swine-origin influenza virus A (H1N1)-2009. Science China Life Sciences, 53, 633-642. doi:10.1007/s11427-010-4010-8
- Tu, W., Mao, H., Zheng, J., Liu, Y., Chiu, S.S., Qin, G., et al. (2010) Cytotoxic T lymphocytes established by seasonal human influenza cross-react against 2009 pandemic H1N1 influenza virus. Journal of Virology, 84, 6527-6235. doi:10.1128/JVI.00519-10
- Gras, S., Kedzierski, L., Valkenburg, S.A., Laurie, K., Liu, Y.C., Denholm, J.T., et al. (2010) Cross-reactive CD8+ T-cell immunity between the pandemic H1N1-2009 and H1N1-1918 influenza A viruses. Proceedings of the National Academy of Sciences of USA, 107, 12599-12604. doi:10.1073/pnas.1007270107
- Brownlee, G.G. and Fodor, E. (2001) The predicted antigenicity of the haemagglutinin of the 1918 Spanish influenza pandemic suggests an avian origin. Philosophical Transactions of the Royal Society B: Biological Science, 356, 1871-1876. doi:10.1098/rstb.2001.1001
- Zhang, W., Qi, J., Shi, Y., Li, Q., Gao, F., Sun, Y., et al. (2010) Crystal structure of the swine-origin A (H1N1)- 2009 influenza A virus hemagglutinin (HA) reveals similar antigenicity to that of the 1918 pandemic virus. Protein Cell, 1, 459-467. doi:10.1007/s13238-010-0059-1
- Wrammert, J., Koutsonanos, D., Li, G.M., Edupuganti, S., Sui, J., Morrissey, M., et al. (2011) Broadly cross-reactive antibodies dominate the human B cell response against 2009 pandemic H1N1 influenza virus infection. Journal of Experimental Medicine, 208, 181-193. doi:10.1084/jem.20101352
- Krause, J.C., Tsibane, T., Tumpey, T.M., Huffman, C.J., Basler, C.F., Crowe Jr., J.E. (2011) A broadly neutralizing human monoclonal antibody that recognizes a conserved, novel epitope on the globular head of the influenza H1N1 virus hemagglutinin. Journal of Virology, 85, 10905-10908. doi:10.1128/JVI.00700-11
- Pourbohloul, B., Ahued, A., Davoudi, B., Meza, R., Meyers, L.A., Skowronski, D.M., et al. (2009) Initial human transmission dynamics of the pandemic (H1N1) 2009 virus in North America. Influenza and Other Respiratory Viruses, 3, 215-222. doi:10.1111/j.1750-2659.2009.00100.x
- Howden, K.J., Brockhoff, E.J., Caya, F.D., McLeod, L.J., Lavoie, M., Ing, J.D., et al. (2009) An investigation into human pandemic influenza virus (H1N1) 2009 on an Alberta swine farm. Canadian Veterinary Journal, 50, 1153- 1161.
- Weingartl, H.M., Berhane, Y., Hisanaga, T., Neufeld, J., Kehler, H., Emburry-Hyatt, C., et al. (2010) Genetic and pathobiologic characterization of pandemic H1N1 2009 influenza viruses from a naturally infected swine herd. Journal of Virology, 84, 2245-2256. doi:10.1128/JVI.02118-09
- Pasma, T. and Joseph, T. (2010) Pandemic (H1N1) 2009 infection in swine herds. Emerging Infectious Diseases of Canada, 16, 706-708. doi:10.3201/eid1604.091636
- Zhou, H., Wang, C., Yang, Y., Guo, X., Kang, C., Chen, H., et al. (2011) Pandemic (H1N1) 2009 virus in swine herds. Emerging Infectious Diseases of China, 17, 1757- 1759. doi:10.3201/eid1709.101916
- Anhlan, D., Grundmann, N., Makalowski, W., Ludwig, S. and Scholtissek, C. (2011) Origin of the 1918 pandemic H1N1 influenza A virus as studied by codon usage patterns and phylogenetic analysis. RNA, 17, 64-73. doi:10.1261/rna.2395211
- Taubenberger, J.K., Reid, A.H., Janczewski, T.A. and Fanning, T.G. (2001) Integrating historical, clinical and molecular genetic data in order to explain the origin and virulence of the 1918 Spanish influenza virus. Philosophical Transactions of the Royal Society B: Biological Science, 356, 1829-1839. doi:10.1098/rstb.2001.1020
- Taubenberger, J.K., Reid, A.H., Lourens, R.M., Wang, R., Jin, G. and Fanning, T.G. (2005) Characterization of the 1918 influenza virus polymerase genes. Nature, 437, 889- 893. doi:10.1038/nature04230
- Antonovics, J., Hood, M.E. and Baker, C.H. (2006) Molecular virology: Was the 1918 flu avian in origin? Nature, 440, E9.
- Vijaykrishna, D., Poon, L.L., Zhu, H.C., Ma, S.K., Li, O.T., Cheung, C.L., et al. (2010) Reassortment of pandemic H1N1/2009 influenza A virus in swine. Science, 328, 1529. doi:10.1126/science.1189132
- Ducatez, M.F., Hause, B., Stigger-Rosser, E., Darnell, D., Corzo, C., Juleen, K., et al. (2011) Multiple reassortment between pandemic (H1N1) 2009 and endemic influenza viruses in pigs. Emerging Infectious Diseases of US, 17, 1624-1629. doi:10.3201/eid1709.110338
- MMWR (2011) Swine-origin influenza A (H3N2) virus infection in two children—Indiana and Pennsylvania, July-August 2011. Morbidity and Mortality Weekly Report, 60, 1213-1215.

