Advances in Bioscience and Biotechnology
Vol.4 No.7A2(2013), Article ID:34657,8 pages DOI:10.4236/abb.2013.47A2005
The underground organ microbial complexes of moorland spotted orchid Dactylorhiza maculata (L.) Soó (Orchidaceae)
![]()
1Demidov Yaroslavl State University, Yaroslavl, Russia
2Academician Yu. Isakov Research Group, Bakulev Scientific Center of Cardiovascular Surgery, Moscow, Russia
Email: bioksusha@mail.ru, osipovga@mail.ru
Copyright © 2013 Nina V. Shekhovtsova et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received 8 May 2013; revised 8 June 2013; accepted 18 June 2013
Keywords: Dactylorhiza maculata; Orchidaceae; Tubers; Rhizosphere; Endophytic Bacteria; Associative Microorganisms
ABSTRACT
This paper is focused on the first study of microbiota of a moderate climate orchid from Northern hemisphere—Moorland Spotted Orchid Dactylorhiza maculata (L.) Soó growing in its natural habitat. There have been obtained detailed data concerning bacterial communities from rhizosphere and inner tissues of young and old tubers. It was done using the biomarker analysis method where the markers were detected by gas chromatography—mass-spectrometry. It is shown that the number of bacterial genera and the bacteria amount (105 - 108 CFU per gram of dry weight) in D. maculata microbial complexes decreases from rhizosphere to old tuber to young. At the same time all three bacterial cenoses closely resemble each other in biodiversity. Their constant members are gg. Hyphomicrobium, Methylococcus, Nitrobacter, Pseudomonas and Sphingomonas (Proteobacteria), Bacillus and Clostridium (Firmicutes), Rhodococcus and Streptomyces (Actinobacteria). There were found differences peculiar to the taxonomic structure of each microbial complex. The rhizosphere is dominated by Actinobacteria while in inner tissues Proteobacteria are prevalent. The bacteria of gg. Ochrobactrum, Xanthomonas, Butyrivibrio, Corynebacterium, Mycobacterium, Propionibacterium, Sphingobacterium and specific iron reducers were shown to belong to rhizosphere community. Genera of Agrobacterium and Bifidobacterium were found only in the endophytic microbiota of the young tuber while g. Aeromonas is specific for endophytic microbiocenosis of the old tuber. Endophytic bacteria complexes of the tubers differ in structure too: more numerous and diverse Actinobacteria mark the old tuber while the young organ has bigger share of gg. Bacillus and Clostridium.
The results suggest that the endophytic bacteria in tubers originate from rhizosphere. By means of plating on nutrient media there were isolated three phyla of bacteria—Proteobacteria, Firmicutes and Actinobacteria which made up 103 CFU per gram of fresh tissue weight. The phylum of Bacteroidetes, a minor part of the microbiocenosis was found only by biomarker method. The bacterial genera associated with D. maculata detected by us can positively influence the plant by means of their exometabolites. It can be used for strategic and tactical approaches to rare species conservation.
1. INTRODUCTION
The Orchid Family (Orchidaceae) is the largest family of monocotyledonous plants, spread all over the world [1]. They form symbiotic relationships with microorganisms essential for their life activity during all stages of onthogenesis [2,3]. The best studied are the relationships between orchids and mycosymbionts that enable germination of the seeds and accompany the grown plant through all life cycles [4,5]. At present the view of mycosymbiotic relationship has shifted in benefit of the multicomponent concept that would include other microorganisms along with mycorrhizal fungi [2]. It has been established that among those associate symbionts there are bacteria inhabiting the orchid underground organs [6,7]. The bacteria associated with subtropical and tropical orchids have been subjects of numerous researches, both in the plants natural habitat and under greenhouse conditions [6,8-10]. There are many studies of pathogenic bacteria affecting Orchidaceae cultivated in floriculture industry [11]. There are data concerning the biodiversity of associative bacteria obtained by their having been isolated and characterized using the plating method [9,10, 12]. However the results showing the domination of one or another group of microorganisms can be viewed as relative only because they depend strongly on the nutrient medium used in each particular case. The alternatives to plating method are molecular-biological and chemotaxonomic methods which allow to study the taxonomic structure of the microbial complexes without isolating them on nutrient media. The method of fatty acid biomarkers in particular is used to reconstruct both quantitative and qualitative structure of microbial complexes in situ [13].
It has been previously shown that the orchid symbiotic bacteria are able to synthesize substances (phytohormones, vitamins, ferments, amino acids, antibiotics) which control germination of seeds, the plant development and growth, forming of mycorrhiza and resistance towards phytopathogens [8,10,14,15].
There is definite lack of data concerning symbiotic bacteria associated with underground organs of moderate climate orchids from Northern hemisphere in their natural habitat [16,17]. The associative bacteria of Orchidaceae that might play a crucial part in controlling onthogenesis as well as in the expansion of the host plant ecological amplitude and symbioses, their forming and stability are practically unstudied [18]. The results of such studies could be used for the purposes of rare orchid species conservation, their introduction and reintroduction. They could be also implemented in optimization of the in vitro multiplication and cultivation of the orchids, and for decorative or medicinal purposes.
This research aims to explore the rhizosphere and endophytic complexes of microorganisms in the young and old tubers of Moorland Spotted Orchid Dactylorhiza maculata (L.) Soó in its natural habitat of moderate climate Northern hemisphere, notably Central European Russia.
2. MATERIALS AND METHODS
2.1. Research Objects
The object of the research was the root layer in the soil, the inner tissues of tubers belonging to D. maculata grown in Central European Russia (Yaroslavl region) and microorganisms isolated from them. D. maculata is a permanent herbaceous plant with a young tuber forming yearly during the vegetation while the one formed in previous year dies off [19]. The D. maculata of the case study grew in a young birch forest with a touch of alder, mixed herbs and gramineous environment, in the loamy sod-podzolic soil. The humus content in the layer where the tubers were found (3 - 8 cm) was 1.4%, рН-4.2.
2.2. Sampling and the Preparation of Samples for Microbiological Analysis
The samples were gathered during the flowering period of D. maculata when the plants still had both their old and new tubers. The underground organs were carefully dug up and the samples of the rhizosphere were taken. Those included the 2 mm soil layer adjoining tubers. The soil samples were dried and cleaned from inclusions, crushed in sterile conditions and later used for marker fatty-acids detection.
The tubers have been washed three times in sterile tap water while shaken in shake-flask propagator for 15 min. Their surface has been sterilized in 6% chloramine solution for 6 min, then washed three times in sterile tap water and dried. Then the tubers were weighed and dried until they acquired constant weight, and used for detection of marker fatty-acids. Some of the tubers were left fresh so that they could be used to isolate the microorganisms from by cultivation method.
In all cases the average sample included materials from five plants.
2.3. Reconstruction of Bacterial Community Using Biomarker Analysis
Using gas chromatography-mass spectrometry method (GC-MS) specific chemical markers have been detected on the surface structures of microorganisms (fatty acids, hydroxy acids, aldehydes). Taxonomy structure of D. maculata rhizosphere microbiocenosis and that of the tuber inner tissues has been reconstructed through wellestablished algorithm [13]. The instrument used in this research was chromatograph mass spectrometer АТ 5973 D produced by Agilent Technologies (USA).
A melted sample (1.0 g) was subjected to acid methanolysis by treatment in 0.6 ml 1M HCl-methanol solution for 3 hrs at 80˚C. At this stage the compound lipids of microorganisms and other cells from the sample release fatty acids and aldehydes. Fatty acid methyl esters received in the solution were then extracted by applying 200 mkl of hexane, dried and treated by 20 mkl of N, O-bis(trimethylsilyl)trifluoroacetamide (BSTFA) for 15 min at 80˚C in order to produce trimethylsilyloxy acid esters as well as alcohol and sterols. For analysis purposes reagent solution (2 ml) was introduced into injector of the chromate-mass-spectrometer (GC-MS). This is usual procedure for lipid fraction investigation of biologic matter. GC-MS analysis was performed on chromatograph mass spectrometer GCMS-QP 2000 (Shimadzu, Japan). The quadrupole mass spectrometer has a resolution of 0.5 mass units over the whole mass range of 2 - 1000 amu. Ionization is performed by electrons at 70 eV. The sensitivity of GC-MS system is at 1 ng methyl stearate in constant scanning mode and the same in selected ion mode. Chromatographic probing was achieved by a fused silica capillary column HP-5MS 25 m long and 0.25 mm wide in internal diameter. Chromatography was performed in programmed temperature mode with the temperature range set from +130˚C to +320˚C, temperature rate 5 grad/min. Temperature in the injector was +280˚C, interface temperature +250˚C.
Mass-fragmentography method (MF) was used to analyze minor biogenic chemical components. A special program was designed to accumulate specific ion signals from microorganism marker compounds used for selective detection. The program sent periodic requests to five temporal groups each of which counted up to 20 ions. These groups that consisted of chemical substances ranged from decanoic acid to sitosterol, were timed during chromatography at 4 to 40 min intervals from the moment of the probe’s injection.
Measurement algorithm allows detecting about 140 microbial marker compounds if 3 - 4 parameters per compound are used: two specific ions, characteristic retention time as well as correlation of chromatographic peak areas.
The species structures of rhizosphere and tuber microbial communities were estimated using the database of fatty acid profiles and markers of microorganisms after the protocol described earlier. Identification of isolated microbial pure cultures and comparison of their residue profiles to library data was based on fatty acid structure. For this purpose both own identification program by image recognition and manuals of determinative bacteriology were used. On mass-fragmentograms the marker peak areas were integrated automatically by the task program using the internal standard. Then these data organized in Excel tables were processed by computational program. Calibration using tridecanoic acid trideuteromethyl ester was done and its results applied to the final computation.
Calculation of community structure and effective microbial cells number. The program designed for the analysis and the algorithm for identification and computation of effective microbial cells number were based on the data pertaining to chemical composition of individual and collective markers which characterized over 500 of microorganisms assumed to have formed the community.
Cell envelope of different organisms is known to have up to 200 fatty acids in its lipid structure, depending on the type of the cell-bacterial, fungous, plant or animal. Some compounds are associated with one taxon only. Cell number or weight of such microorganisms were calculated by marker compound concentration using known data on fatty acid content in microbial cells and making allowances for preparation and device calibration conditions.
2.4. Similarity Estimation for Reconstructed Bacterial Cenoses
Similarities in bacterial genera that make up respective bacterial communities have been estimated. Sørensen’s Similarity Index was calculated as

where a is the number of shared bacterial genera, b— number of genera only in community 1; с—number of genera only in community 2. Cs ranges from 0 (no species overlap) to 1 (complete overlap) [20].
2.5. Isolation of Bacteria from the Tubers
Prepared tubers were sliced into 5 mm segments under sterile conditions, put into sterile mortar and ground into powder with a small amount of water. The resulted suspension was then serially diluted tenfold. From each dilution 0.1 ml was plated onto two selected media: (1) peptone yeast glucose agar (PYGA), containing per liter: glucose, 1 g; peptone, 2.0 g; casein hydrolyzate, 1.0 g; yeast extract, 0.5 g; agar, 17.0 g; and (2) starch-ammonium agar (SAA), containing per liter: starch, 10.0 g; (NH4)2SO4, 2.0 g; K2HPO4, 1.0 g; yeast extract, 0.5 g; MgSO4 × 7H2O, 1.0 g; NaCl, 1.0 g; CaCO3, 3.0 g. Nystatin (50 μg·ml−1) was added to all media to inhibit fungal growth. All plates were incubated at 28˚C until single bacterial colonies arose. Bacteria were isolated from young and old tubers separately during flowering when the orchid has both.
2.6. Determination of Bacterial Number
The number of bacteria inhabiting tubers was calculated by Koch plating method according to which the number of bacterial colonies cultivated on each plate was estimated. The bacterial number was calculated as:
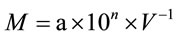
where M is the number of bacterial cells per 1 ml of initial organ suspension, a is an average of colonies from three replicates, V = 0.1 ml (aliquot, plated on the plate), 10n is dilution factor. Сolony-forming units (CFU) per gram of fresh tissue weight were counted after 3 weeks of incubation at 28˚C.
2.7. Determination of Bacteria Phenotypic Properties
Different types of bacterial colonies were counted. The colonies were further purified by three times streaking onto PYGA and SAA plates to obtain pure cultures. The bacterial cultures were classified into morphological groups based on their phenotypic properties as described in Bergey’s Manual of Systematic Bacteriology (2nd Edition, Springer, N.Y., 2005-2012). The appearance of colonies and morphology of cells was studied after bacteria have been plated in fresh medium for 24, 48 and 72 hrs respectively. The Gram stain was performed in order to divide bacteria into Gram-positive and Gram-negative. Morphological and tinctorial properties have been studied by means of light microscopy.
2.8. Statistical Analysis
Isolation of associate bacteria has been replicated three times. The data on bacterial number and strain content values have been subjected to statistical analysis using Excel Package. Bacterial population percentage (%) was estimated as follows: the number of microbial colonies of given bacterial strain divided by the total number of bacterial colonies × 100%. Values calculated are the mean of three replicates, standard error is shown as ± m in the text.
3. RESULTS AND DISCUSSION
GC-MS analysis of FA fractions in rhizosphere and tuber specimens revealed the 81 fatty acid biomarkers of bacteria. Among them the following chemicals are found: (1) unbranched hydroxy fatty acids (FA)—saturated 3-hydroxydecanoic (3hC10:0), 3hC11:0, 3hC12:0, 2hC12:0, 3hC13:0, 3hC14:0, 2hC14:0, 3hC15:0, 2hC15:0, hC16:0, 3hC17:0, 10hC18:0, 3hC18:0, 3hC20:0, 3hC22:0, 2-hydroxy-cyclo-nonadecanoic (2hC19cyc), and unsaturated 10hС18:1; (2) branched unsaturated hydroxy FA—hydroxy-iso-undecanoic (hiC11:0), 3-hydroxy-anteiso-tridecanoic (3haC13:0), 3hiC13:0, 3hiC14:0, 3hiC15:0, 3haC15:0, 2hiC15:0, hiC16:0, 3hiC17:0 and 3haC17:0; (3) unbranched saturated FA—from decanoic (C10:0) to nonadecanoic (C19:0), except of C11:0 и C16:0, as well as cyclopropane FA—C17:0cyc and C19:0cyc; (4) branched saturated FA—iso-tridecanoic (iC13:0), anteiso-tridecanoic (aC13:0), iC14:0, iC15:0, iC16:0, iC17:0, iC18:0, aC15:0, aC17:0, iC19:0, aC19:0, 10-methyl-tetradecanoic (10MeС14:0), 10MeС16:0, 10MeС17:0 and 10MeС18:0; (5) unbranched unsaturated FA—dodecenoic (C12:1), cis-11,12-tetradecenoic (C14:1d11c), C14:1d5, C15:1, C16:1d9c, C16:1d7c, trans-7,8-hexadecenoic (C16:1d7t), C16:1d5c, C17:1, C18:1d7c, C18:1d9c, C20:1d9c and 11 Me18:1; (6) branched unsaturated FA—iso-hexadecenoic (iC16:1), iC17:1 and 11Me18:1; aldehydes (7) unbranched saturated—C14a, C15a, C16a, C17a, (8) unbranched unsaturated—C18:1a, C18:1d7a, (9) branched saturated —iC14a, i15a, аC15a, i16a, i17a, a17a and (10) sterol —coprostanol.
The minor microbial components have been used for calculation of reconstructed microbial complexes on the base of data about biomarkers of microorganisms which have been remarked in the rhizosphere and internal tissues of different plants including tropical orchids [13].
In the course of the research there have been reconstructed and described bacterial communities of the rhizosphere and inner tissues of both old and young tubers belonging to D. maculata (Table 1). The diversity of the communities included 35 genera of gram-positive and gram-negative bacteria that according to the phylogenetic system for procaryotic classification belong to four phyla—BXII. Proteobacteria, BXIII. Firmicutes, BXIV. Actinobacteria and BXX. Bacteroidetes.
The rhizosphere community of D. maculata contained 32 genera of which 15 belonged to Proteobacteria, 11— to Actinobacteria, and 4 genera to Firmicutes и Bacteroidetes respectively (Table 1). The most numerous were Actinobacteria (58%) with codominant Proteobacteria (35%). Firmicutes and Bacteroidetes were represented by 7% of all bacteria. Compared to tuber microbiota, the rhizosphere community had 8 bacterial groups—gg. Ochrobactrum, Xanthomonas, Butyrivibrio, Corynebacterium, Mycobacterium, Propionibacterium, Sphingobacterium and specific iron reducers. The only dominant group in the rhizosphere community was g. Streptomyces (15%) from the phylum of Actinobacteria.
The endophytic bacterial complex of D. maculata young tuber contained 20 genera, of which 10 genera belonged to Proteobacteria, 4 genera to Actinobacteria and Firmicutes each, 2 genera belonged to Bacteroidetes (Table 1). Groups arranged by number in decreasing order: Proteobacteria (56%)—Firmicutes (35%)—Actinobacteria (7%)—Bacteroidetes (2%) (Figure 1). Genera of Clostridium was the dominant one (made up 27% of community). Its codominants were proteobacteria gg. Sphingomonas, Methylococcus, Hyphomicrobium, Nitrobacter which made 10% of the community. Bacteria from genera Agrobacterium and Bifidobacterium were found in young tuber only.
The tissue microbiota in D. maculata old tuber consisted of 24 genera of which Proteobacteria 11 genera, Actinobacteria—7, Firmicutes—4 and Bacteroidetes—2 (Table 1). The biodiversity and numbers of bacteria per phyla in old tuber corresponded with the findings in the young tuber (Figure 1). The difference is the increasing of Actinobacteria in numbers and diversity and the decreasing of Firmicutes. Genera of Sphingomonas (17%), Methylococcus (17%) among Proteobacteria and g. Clostridium (14%) of Firmicutes dominate the group with gg. Hyphomicrobium, Methylococcus remaining co-dominants (no less than 11%). The endobacteria of g. Aeromonas are specific for the old tuber.
Microbial complexes of D. maculata young and old tubers have their differences as well. While still forming, a tuber is colonized by bacteria of gg. Agrobacterium, Bifidobacterium, Riemerella that are no longer found in the old dying tuber. On the other hand the old tuber has bacteria specific for it—gg. Aeromonas, Caulobacter,
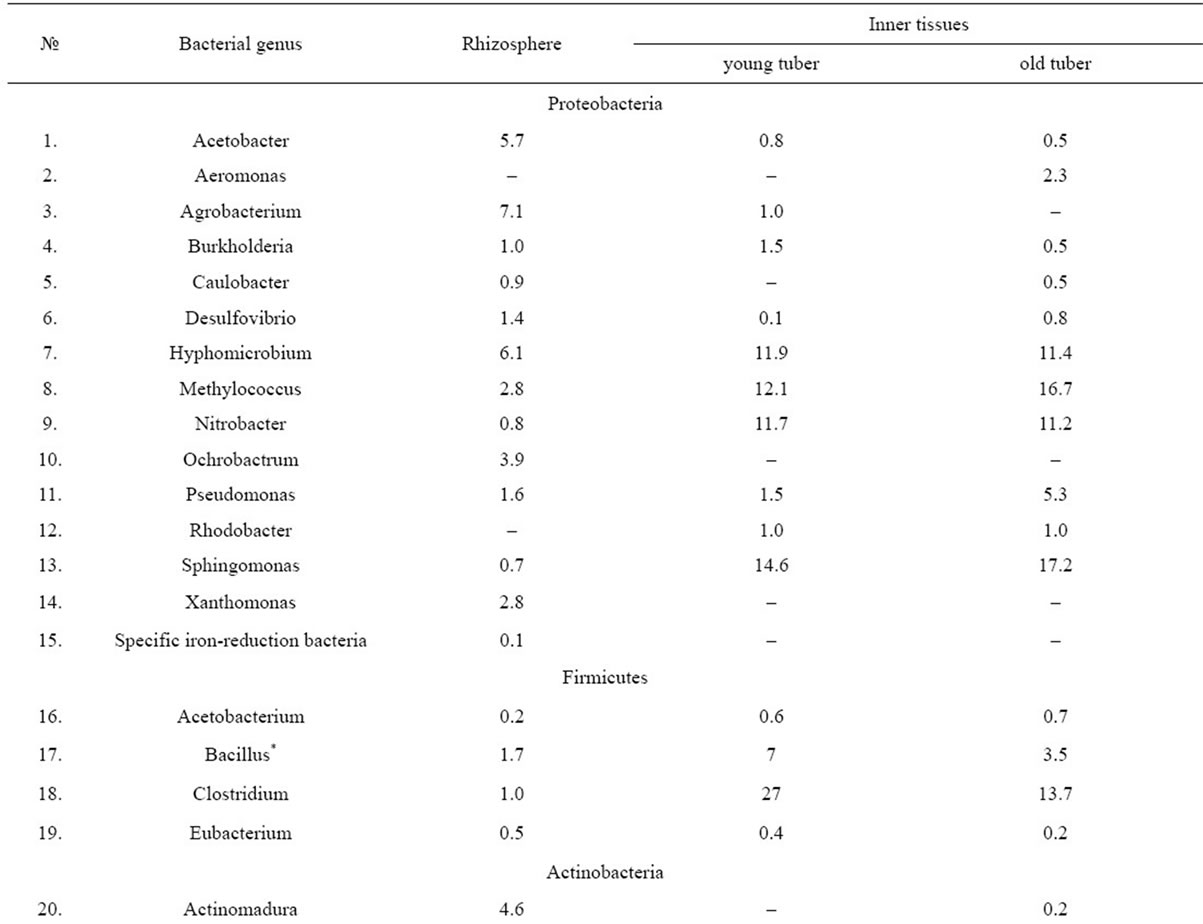
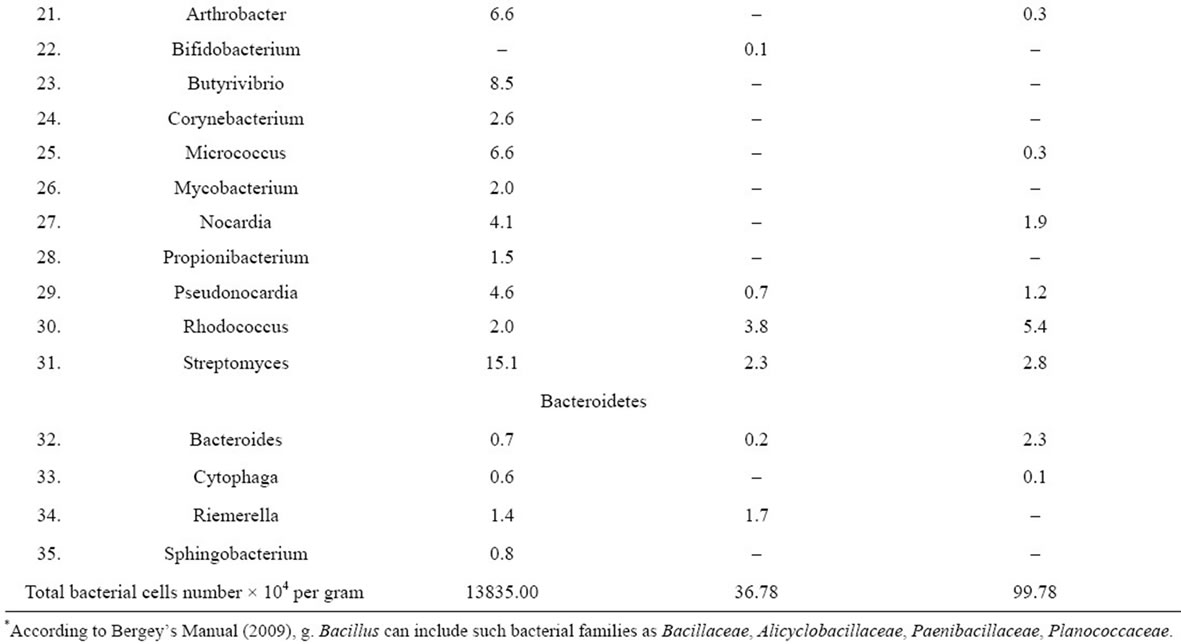
Table 1. Taxonomic structure of reconstructed microbial communities associated with underground organs of D. maculata (%).
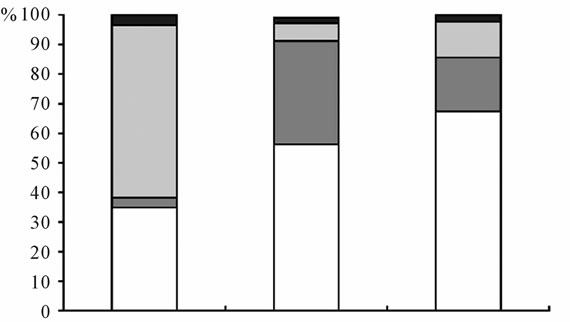 (a)
(a) (b)
(b)
Figure 1. Distribution of number (a) and diversity (b) in the composition of rhizosphere and endophytic microbial complexes of the D. maculata.
Actinomadura, Arthrobacter, Micrococcus, Nocardia and Cytophaga. This variability in old and young tuber microbial complexes corroborates the previous conclusion that the endophytic bacteria composition in one and the same orchid species can vary depending on different factors [6,7].
14 genera of the 35 identified by us were earlier described in the researches of subtropical and tropical orchids from greenhouses and their natural habitat. From the surface and inner tissues of epiphyte and terrestrial orchid species there have been isolated following bacteria—gg. Acetobacter, Azospirillum, Burkholderia, Pseudomonas, Arthrobacter, Streptomyces, Bacillus [7,9, 12,14]. The species of such genera as Agrobacterium, Sphingomonas, Micrococcus, Mycobacterium, Nocardia, Rhodococcus were found to be specific for root surface [7,8,12]. K.G. Wilkinson et al. (1994) showed g. Xanthomonas to be an endophytic habitant only. The genera of Pseudomonas, Sphingomonas, Xanthomonas, Micrococcus, Nocardia, Rhodococcus, Streptomyces, Bacillus were also identified in association with epiphyte orchid air-roots [7,8]. The most commonly mentioned among bacteria found with tropical and subtropical orchids are gg. Pseudomonas, Bacillus, Burkholderia, Xanthomonas, Arthrobacter, Rhodococcus that have also been identified by us, and gg. Enterobacter, Erwinia, Flavobacterium and Kurthia, not found in association with Moorland Spotted Orchid [7,9,10,12,14].
In bacterial complexes of D. maculata the number of genera and the bacteria amount decreases from rhizosphere to old tuber to young. However, the biodiversity similarities in all three bacteriocenoses according to Sø- rensen’s index are rather close. The similarity coefficient for rhizosphere and old tuber communities is 0.80, for rhizosphere and young tuber it is 0.71 and for old and young tuber it is 0.77. Among constant components of D. maculata rhizosphere and tubers both are bacteria gg. Hyphomicrobium, Methylococcus, Nitrobacter, Pseudomonas and Sphingomonas (Proteobacteria), Rhodococcus and Streptomyces (Actinobacteria), Bacillus and Clostridium (Firmicutes) (Table 1). Taking into account the high degree of similarity between communities (more than 0.6) it is reasonable to assume that endophytic bacteria in the tubers originate from rhizosphere. Rhizobacteria outnumber endophytic bacteria in old and young tuber by 10 - 100 times. Young tuber has the least number of microorganisms (37% of their number in old tuber) which is probably explained by the late start of bacterial colonization in storage organs.
As the starch-ammonium agar (SAA) plating shows, the number of hydrolytics participating in decomposition of the storage organ biopolymers is higher in old tuber compared to the young. It is 4583 ± 290 and 1100 ± 90 CFU/g of fresh tissue weight respectively (Figure 2). At the same time the proportion is quite reverse for copiotrophes consuming low molecular compounds. Using peptone yeast glucose agar (PYGA) it was possible to extract only 3967 ± 220 CFU/g of fresh tissue weight from young tuber and 2083 ± 110 CFU/g from old. Independent of media used for plating, the bacterial groups cul-
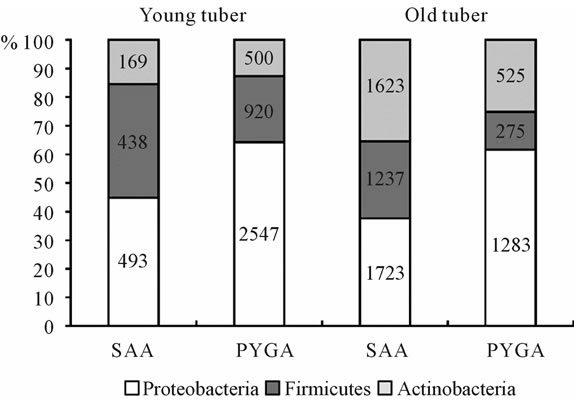
Figure 2. Structure of microbial complexes in young and old tubers of D. maculata, isolated on starch-ammonium agar (SAA) and peptone yeast glucose agar (PYGA). The number shows the absolute amount of bacteria, CFU per gram of fresh tissue weight.
tivated range in descending order like this—Proteobacteria—Firmicutes—Actinobacteria for young tuber and Proteobacteria—Actinobacteria—Firmicutes for old tuber. This mostly corroborates the data obtained for reconstructed microbial communities of storage organs as shown above.
We’ve calculated the number of endophytic bacteria in D. maculata tubers as 103 CFU per gram of fresh tissue weight (Figure 2) which is keeping within the 103 and 105 CFU/g of plant fresh tissue range, common popularity size of endophytic bacteria associated with the most of investigated plant species [21,22]. However it is considerably less than the number of bacteria extracted from roots of greenhouse orchids—106 CFU per gram of dry tissue weight [8,12].
As for biodiversity, from young tuber there were isolated 18 strains of bacteria, of which 7 belong to Proteobacteria, 7—Actinobacteria и 5—Firmicutes. Actinobacteria were at their most numerous and diverse in old tuber. Steady dominance of Proteobacteria was probably due to the fact that tubers contained considerable amount of low molecular compounds all through their growth and development. The change in dominant status within the hydrolytic group can be caused by the lower content of available substrate in old tuber while the amount of intercellular spaces within it has grown. In young tuber its active growing doesn’t give enough time for intercellular spaces to form which leads to poorer aeration of the organ.
By analyzing reconstructions of microbial communities and the results of bacterial plating on nutrient media it becomes possible to highlight the processes of microbial succession in tubers. As shown above, the shift in dominance within the hydrolytic group can be explained by changes in storage organ physiology. In its initial stage the forming tuber is colonized by fast-growing Firmicutes while towards the final stage Actinobacteria become the dominant hydrolytics.
Regarding potential properties of associated bacteria it is important to note that gg. Agrobacterium, Burkholderia, Pseudomonas, Sphingomonas, Xanthomonas, Arthrobacter, Micrococcus, Mycobacterium, Rhodococcus and Bacillus, associated with D. maculata and previously identified in tropic orchids are able to produce auxins [8,10,12,14]. Bacteria from gg. Bacillus, Pseudomonas и Xanthomonas found in substrate roots of Australian orchids, promote germination of their seeds together with mycosymbiont [14]. Bacteria Bacillus, Sphingomonas and Mycobacterium isolated from aerial roots of epiphyte greenhouse orchids also promote germination in absence of mycosymbiont [14]. The rhizobacteria identified as Bacillus and Burkholderia, isolated from Cattleya walkeriana orchid were shown to promote survival and growth of in vitro germinated plantlets during ex vitro acclimatization [10]. Similar properties of bacteria identified in rhizosphere and tubers of D. maculatа can be subject of future research.
4. CONCLUSIONS
This paper describes the first comprehensive study of microbiota of a moderate climate orchid from Northern hemisphere in its natural habitat, case Moorland Spotted Orchid Dactylorhiza maculata. There have been obtained detailed data concerning bacterial communities from rhizosphere and inner tissues of its young and old tubers. For the first time it was done using the biomarker analysis method where the markers were detected by gas chromatography—mass-spectrometry.
There have been identified 32 genera of bacteria in the tuber rhizosphere, 20 genera among the young tuber endophytes and 20 genera of endophytes in the old tuber. The number of rhizobacteria was 108 CFU per gram of dry soil, varying in endophyte communities from 105 to 106 CFU per gram of dry tissue weight and 103 CFU per gram of fresh tissue weight. The number of endobacteria in the young tuber was less than in the old.
There are differences peculiar to the taxonomic structure of each microbial complex. By means of marker analysis it was found that the microbial complexes of bacteria associated with D. maculata contained four phyla —Proteobacteria, Firmicutes, Actinobacteria and Bacteroidetes. The first three were later detected by means of plating on nutrient media. The results suggest that the endophytic bacteria in tubers originate from rhizosphere.
The rhizosphere is dominated by Actinobacteria while in inner tissues Proteobacteria are prevalent. Endophyte bacterial complexes of the tubers differ in structure too: more numerous and diverse Actinobacteria mark the old tuber while the young organ has bigger share of the bacteria of gg. Bacillus and Clostridium.
The bacteria associated with D. maculata identified by us have been earlier shown to play a positive role in the life of other plant species by means of their exometabolites. This can be used for strategic and tactical approaches to rare species conservation.
5. ACKNOWLEDGEMENTS
The authors express their gratitude to T. G. Dobrovolskaya, chief science officer at the Lomonosov State University (Moscow) for her help in identifying bacteria, and also deep appreciation to the students and aspirants at the Botany and Microbiology Department of Demidov State University (Yaroslavl).
![]()
![]()
REFERENCES
- Dressler, R.L. (1981) The orchids: Natural history and classification. Harvard University Press, Cambridge and London, 332.
- Bayman, P. and Otero, J.T. (2006) Microbial endophytes of orchid roots. Soil Biology, 9, 153-177. doi:10.1007/3-540-33526-9_9
- Smith, S.E. and Read, D.J. (2008) Mycorrhizal symbiosis. Elsevier, Amsterdam, 804.
- Hadley, G. (1982) Orchid mycorrhiza. In: Arditti, J., Ed., Orchid biology—Reviews and perspectives. Cornell Universtiy Press, Ithaca, 83-118.
- Rasmussen, H.N. (1995) Terrestrial orchids: From seed to mycotrophic plant. Cambridge University Press, Cambridge, 433.
- Wilkinson, K.G., Dixon, K.W. and Sivasithamparam, K. (1989) Interaction of soil bacteria, mycorrhizal fungi and orchid seeds in relation to germination of Australian orchids. New Phytologist, 112, 429-435. doi:10.1111/j.1469-8137.1989.tb00334.x
- Tsavkelova, E.A., Cherdyntseva, T.A, Botina, S.G. and Netrusov, A.I. (2004) Bacteria associated with the roots of epiphytic orchids. Mikrobiologiia, 73, 825-831.
- Tsavkelova, E.A, Cherdyntseva, T.A. and Netrusov, A.I. (2005) Auxin production by bacteria associated with orchid roots. Mikrobiologiia, 74, 55-62.
- Zambrano, E.R., Salgado, T.J. and Hernández, A.T. (2007) Estudio de bacterias asociadas orquídeas (Orchidaceae). Lankesteriana, 7, 322-325.
- Galdiano, J.R.F., Pedrinho, E.A.N., Castellane, T.C.L. and de Macedo Lemos, E.G. (2011) Auxin-producing bacteria isolated from the roots of Cattleya walkeriana, an endangered Brazilian orchid, and their role in acclimatization. Revista Brasileira de Ciência do Solo, 35, 83-92.
- Wu, Z.-Y., Fang, Y., Chen, X., Zhang, H.-L., Zhang, M.-Z. and Li B. (2010) Pathogen identification of bacterial soft rot in moth orchid in Zhejiang Province. Journal of Zhejiang Forestry College, 27, 635-639.
- Tsavkelova, E.A., Cherdyntseva, T.A. and Netrusov, A.I. (2007) Bacteria, associated orchid roots and microbial production of auxin. Microbiological Research, 162, 69- 76. doi:10.1016/j.micres.2006.07.014
- Osipov, G.A. and Turova, E.S. (1997) Studying species composition of microbial communities with the use of gas chromatography-mass spectrometry. Microbial community of kaolin. FEMS Microbiology Reviews, 2, 437- 446. doi:10.1111/j.1574-6976.1997.tb00328.x
- Wilkinson, K.G., Dixon, K.W., Sivasithamparam, K. and Ghisalberti, E.L. (1994) Effect of IAA оn symbiotic germination of аn Australian orchid and its production bу orchid-associated bacteria. Plant Soil, 159, 291-295.
- Tsavkelova, E.A., Cherdyntseva, T.A., Klimova, S.Yu., Shestakov, A.I., Botina, S.G. and Netrusov, A.I. (2007) Orchid-associated bacteria produce indole-3-acetic acid, promote seed germination, and increase their microbial yield in response to exogenous auxin. Archives of Microbiology, 188, 655-664.
- Shеkhovtsova, N.V., Pervushina, K.А., Маrakaev, О.А., Kholmogorov, S.V. and Osipov, G.А. (2010) Microorganisms associated with underground organs of central Russia Orchidaceae. Problemy Agrokhimii i Ekologii, 4, 30-36. doi:10.1007/s00203-007-0286-x
- Mineeva, T.I. and Voronina, E.Yu. (2011) Qualitative and numerical characteristics of some temperate orchid mycorrhizospheres. Proceedings of the 9th International Conference “Orchid Conservation and Cultivation”, SaintPetersburg, 26-30 September 2011, 300-305.
- Hofflich, G., Glante, F., Liste, H.H., Weise, J., Ruppel, S. and Schlozseidel, C. (1993) Phytoeffective combination effects of symbiotic and associative microorganisms оn Lеgumes. Symbiosis, 14, 427-438.
- Vakhrameeva, M.G., Tatarenko, I.V., Varlygina, T.I., Torosyan, G.K. and Zagulski, M.N. (2008) Orchids of Russia and adjacent countries (within the borders of the former USSR). A.R.G. Gantner Verlag K.G., Konigstein, 690.
- Magurran, A. (2004) Measuring biological diversity. Blackwell Publishing, Oxford, 215.
- Zinniel, D.K., Lambrecht, P., Harris, N.B., Feng, Zh., Kuczmarski, D., Higley, Ph., Ishimaru, C.A., Arunakumari, A., Barletta, R.G. and Vidaver A.K. (2002) Isolation and characterization of endophytic colonizing bacteria from agronomic crops and prairie plants. Applied and Environmental Microbiology, 68, 2198-2208. doi:10.1128/AEM.68.5.2198-2208.2002
- Lodewyckx, C., Vangronsveld, J., Porteous, F., Moore, E.R.B., Taghavi, S., Mezgeay, M. and van der Lelie, D. (2010) Bacteria and their potential applications. Critical Reviews in Plant Sciences, 21, 583-606. doi:10.1080/0735-260291044377

