Advances in Parkinson's Disease
Vol.2 No.1(2013), Article ID:27828,3 pages DOI:10.4236/apd.2013.21005
Analysis of known point mutations and SNPs in genes responsible for monogenic Parkinson’s disease in Russian patients
![]()
1Institute of Molecular Genetics, Russian Academy of Sciences, Moscow, Russia; *Corresponding Author: FilatovaEV@img.ras.ru
2Research Centre of Neurology, Russian Academy of Medical Sciences, Moscow, Russia
Received 28 January 2012; revised 11 January 2013; accepted 20 January 2013
Keywords: Parkinson’s Disease; Point Mutations; Single-Nucleotide Polymorphisms
ABSTRACT
Background: Parkinson’s disease (PD) is caused by complex interactions between genetic and environmental factors. Mendelian forms of PD rarely occur in practice, but respective genes may play some role in pathogenesis of a common sporadic form of the disease. Methods: We analyzed most frequent known point mutations (PMs) and single-nucleotide polymorphisms (SNPs) in genes responsible for monogenic PD in 408 Russian patients, using arrayed primer extension (APEX), real-time PCR, and restriction fragment length polymorphism analysis. Results: We detected only three heterozygous PMs in the PARK2 gene in three non-related patients with early-onset sporadic PD. No association between PD and the studied SNPs was identified. Conclusion: The examined PMs and SNPs in genes responsible for monogenic PD do not contribute significantly to the development of sporadic PD in Russia.
1. INTRODUCTION
Parkinson’s disease (PD) is a predominantly sporadic condition, caused by complex interactions between genetic and environmental factors. Mutations in, at least, five genes have been shown to cause multiple cases of monogenic PD in different populations: SNCA, PARK2, PINK1, PARK7 (DJ-1), and LRRK2 [1,2]. The roles of other genes (including UCHL1, ATP13A2, VPS35) are still contentious. Mutations in these genes may play a role in the pathogenesis of sporadic PD [3,4], including Russian population [5-7].
Consequently, we analyzed the frequency distribution of the genotypes of the most frequent point mutations (PMs) and single-nucleotide polymorphisms (SNPs) in the genes involved in the pathogenesis of monogenic PD in russian population to evaluate their role in the pathogenesis of sporadic form of PD in Russia. We recruited 408 Russian patients with various forms of PD. We analyzed the frequency distributions of the genotypes of the most frequent point mutations (PMs) and single nucleotide polymorphisms (SNPs) in several genes, involved in different monogenic parkinsonian syndromes, in this cohort.
2. METHODS/PATIENTS
We first studied two groups of patients and a small control group using an arrayed primer extension (APEX) technology: 21 patients with autosomal-recessive juvenile PD, 41 patients with early-onset sporadic PD with an age at onset of <45 years and 40 normal individuals. On the next stage, we analyzed an additional group of 346 patients with sporadic PD with an age at onset of 30 - 70 years and a control group of 224 normal individuals with an age of 30 - 70 years. All participants gave their informed consent, and the study was approved by the Ethics committee of the Research Center of Neurology RAMS. DNA was extracted from venous blood using the standard phenol-chloroform method.
During the first stage of the study, we performed an APEX analysis of the genotypes using the DNA array (AsperBioTech, Tartu, Estonia) according to the manufacturer’s recommendations. The DNA array is characterized in Table 1 (a complete list of all PMs and SNPs is available upon request). Most PMs present on the array are located within the PARK2 gene.
SNP rs1801582 in the PARK2 gene was further analyzed with the real-time PCR and SNP rs7966550 in LRRK2 was analyzed with the restriction-fragment-length polymorphism technique (Table 2). All reactions were
Table 1. Number of PMs and SNPs on the array.
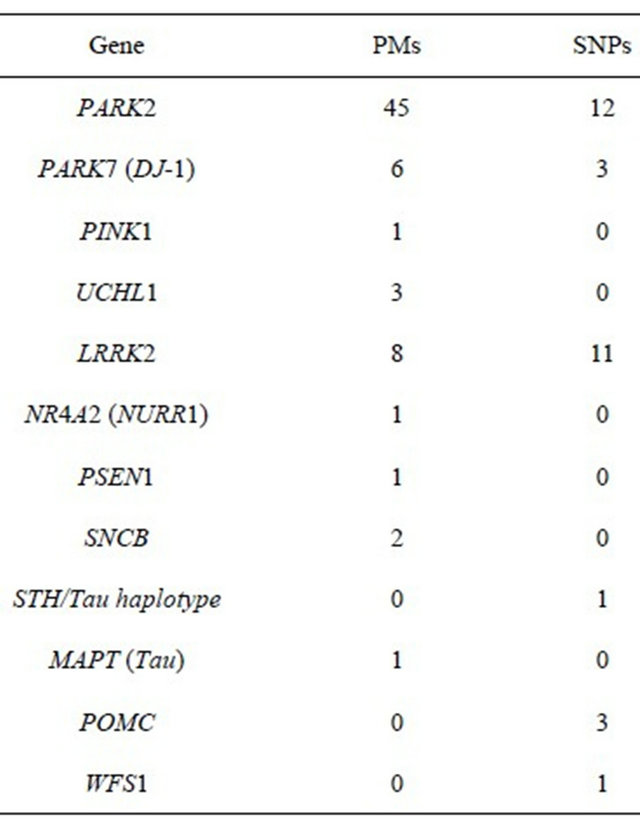
repeated twice.
3. RESULTS
We detected only three heterozygous PMs in PARK2 in three unrelated patients with sporadic PD among 62 patients with juvenile and early-onset sporadic PD: M1L, A82G, and C253Y. These results were also confirmed by direct sequencing of the DNA samples. It has previously been shown that these three PMs lead to the development of autosomal-recessive PD [8-10], and there is a wide variety of other rare PMs in PARK2 [11-13].
We conducted a comparative analysis of the distribution of genotypes in our patients and in the small control group to evaluate the possible impact of the SNPs studied here on the development of PD in Russia. Two SNPs, rs7966550 in LRRK2 and rs1801582 in PARK2, occurred more frequently in the homozygous state in the Russian patients compared to the small control group. The frequency of the ТТ genotype of SNP rs7966550 was 0.952 in the Russian patients and 0.750 in the small control group. The GG genotype of SNP rs1801582 also occurred more frequently in the Russian patients than in the small control group (0.927 vs 0.700 respectively).
From the results of our APEX analysis, we inferred that these two SNPs may affect the development of PD in the Russian population. Therefore, we investigated genotypes distributions of these two SNPs in a more representative sample (346 patients) and a control population (224 normal subjects).
The results of this more detailed analysis of SNPs rs1801582 and rs7966550 are shown in Table 2. As can
Table 2. Genotype frequencies of SNPs in LRRK2 and PARK2.
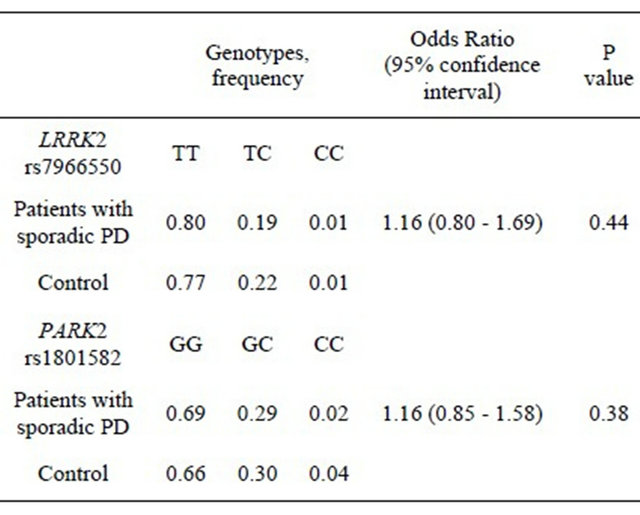
be seen from the data, this analysis did not confirm an association between either SNPs and the development of PD. Results were also consistent with Hardy-Weinberg equilibrium.
4. DISCUSSION
SNP rs7966550 in LRRK2 leads to a synonymous substitution (L953L). Therefore, it is not surprising that no association has yet been demonstrated between this SNP and PD. The differences in the genotype frequencies of our Russian patients and the small control group revealed with the APEX analysis are probably a consequence of the small size of our sample. The results of our detailed analysis of this SNP showed no association between rs7966550 and the development of PD.
Polymorphism rs1801582 in PARK2, which leads to the substitution V380L, was first described by Wang and co-workers [14]. The association of this SNP with PD has been shown in France and the USA [15,16]. However, several other studies have identified no such association [9,14,17]. We also identified no association between rs1801582 and PD (Table 2). Nevertheless, it has been suggested that this SNP, when combined with environmental factors, can affect the development of earlyonset PD [18].
Thus, we identified only three heterozygous PMs in three different patients with sporadic PD, indicating a high microheterogeneity of PMs in the Russian population. We also identified no association between the risk of PD and the SNPs studied. Therefore, known PMs in the genes associated with monogenic PD probably do not contribute significantly to the development of sporadic PD. However, we nave not yet screened for all the mutations that cause familial PD [3]. Therefore, all the key exons of the genes involved in the pathogenesis of PD should be resequenced in our sample.
5. ACKNOWLEDGEMENTS
This work was supported by the Russian Foundation for Basic Research (projects no. 10-04-00745-а, 12-04-31091, 12-04-01183-а), as well as by programs of the Russian Academy of Sciences (Molecular and Cellular Biology, Fundamental Sciences for Medicine), state contracts (no. 16.740.11.0630, 8605, and 8851).
REFERENCES
- Xiromerisiou, G., Dardiotis, E., Tsimourtou, V., et al. (2010) Genetic basis of Parkinson disease. Neurosurgical Focus, 28, E7. doi:10.3171/2009.10.FOCUS09220
- Lesage, S. and Brice, A. (2009) Parkinson’s disease: From monogenic forms to genetic susceptibility factors. Human Molecular Genetics, 18, R48-R59. doi:10.1093/hmg/ddp012
- Lesage, S. and Brice, A. (2012) Role of Mendelian genes in “sporadic” Parkinson’s disease. Parkinsonism and Related Disorders, 18, S66-S70. doi:10.1016/S1353-8020(11)70022-0
- Sutherland, G.T., Halliday, G.M., Silburn, P.A., et al. (2009) Do polymorphisms in the familial Parkinsonism genes contribute to risk for sporadic Parkinson’s disease? Movement Disorders, 24, 833-838. doi:10.1002/mds.22214
- Illarioshkin, S.N., Ivanova-Smolenskaya, I.A., Markova, E.D., et al. (2000) Lack of alpha-synuclein gene mutations in families with autosomal dominant Parkinson’s disease in Russia. Journal of Neurology, 247, 968-969. doi:10.1007/s004150070056
- Illarioshkin, S.N., Shadrina, M.I., Slominsky, P.A., et al. (2007) A common leucine-rich repeat kinase 2 gene mutation in familial and sporadic Parkinson’s disease in Russia. European Journal of Neurology, 14, 413-417. doi:10.1111/j.1468-1331.2007.01685.x
- Semenova, E.V., Shadrina, M.I., Slominsky, P.A., et al. (2012) Analysis of PARK2 gene exon rearrangements in Russian patients with sporadic Parkinson’s disease. Movement Disorders, 27, 139-142. doi:10.1002/mds.23901
- Hedrich, K., Kann, M., Lanthaler, A.J., et al. (2001) The importance of gene dosage studies: Mutational analysis of the parkin gene in early-onset parkinsonism. Human Molecular Genetics, 10, 1649-1656. doi:10.1093/hmg/10.16.1649
- Oliveira, S.A., Scott, W.K., Martin, E.R., et al. (2003) Parkin mutations and susceptibility alleles in late-onset Parkinson’s disease. Annals of Neurology, 53, 624-629. doi:10.1002/ana.10524
- Rawal, N., Periquet, M., Lohmann, E., et al. (2003) New parkin mutations and atypical phenotypes in families with autosomal recessive Parkinsonism. Neurology, 60, 1378- 1381. doi:10.1212/01.WNL.0000056167.89221.BE
- Abbas, N., Lücking, C.B., Ricard, S., et al. (1999) A wide variety of mutations in the parkin gene are responsible for autosomal recessive Parkinsonism in Europe. Human Molecular Genetics, 8, 567-574. doi:10.1093/hmg/8.4.567
- Koziorowski, D., Hoffman-Zacharska, D., Sławek, J., et al. (2010) Low frequency of the PARK2 gene mutations in Polish patients with the early-onset form of Parkinson disease. Parkinsonism and Related Disorders, 16, 136- 138. doi:10.1016/j.parkreldis.2009.06.010
- Vinish, M., Prabhakar, S., Khullar, M., et al. (2010) Genetic screening reveals high frequency of PARK2 mutations and reduced Parkin expression conferring risk for Parkinsonism in North West India. Journal of neurology, neurosurgery, and psychiatry, 81, 166-170. doi:10.1136/jnnp.2008.157255
- Wang, M., Hattori, N., Matsumine, H., et al. (1999) Polymorphism in the parkin gene in sporadic Parkinson’s disease. Annals of Neurology, 45, 655-658. doi:10.1002/1531-8249(199905)45:5<655::AID-ANA15>3.0.CO;2-G
- Lincoln, S.J., Maraganore, D.M., Lesnick, T.G., et al. (2003) Parkin variants in North American Parkinson’s disease: cases and controls. Movement Disorders, 18, 1306- 1311. doi:10.1002/mds.10601
- Lücking, C.B., Chesneau, V., Lohmann, E., et al. (2003) Coding polymorphisms in the parkin gene and susceptibility to Parkinson disease. Archives of neurology, 60, 1253-1256. doi:10.1001/archneur.60.9.1253
- Martinez, H.R., Gonzalez-Gonzalez, H., Cantu-Martinez, L., et al. (2010) PARKIN-coding polymorphisms are not associated with Parkinson’s disease in a population from northeastern Mexico. Neuroscience Letters, 468, 264-266. doi:10.1016/j.neulet.2009.11.009
- Aguiar, P.D.C., Lessa, P.S., Godeiro, C.J., et al. (2008) Genetic and environmental findings in early-onset Parkinson’s disease Brazilian patients. Movement Disorders, 23, 1228-1233. doi:10.1002/mds.22032

