American Journal of Plant Sciences
Vol.4 No.4(2013), Article ID:29824,11 pages DOI:10.4236/ajps.2013.44099
AtL1 a Non-LTR Retrotrasposon Fragment in the Genome of Arabidopsis thaliana with Homology to Plants and Animals
![]()
1Department of Life Sciences, University of Parma, Parma, Italy; 2Department of Physics and Earth Sciences “Macedonio Melloni”, University of Parma, Parma, Italy.
Email: nelson.marmiroli@unipr.it
Copyright © 2013 Giovanna Visioli et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received February 20th, 2013; revised March 25th, 2013; accepted April 1st, 2013
Keywords: Arabidopsis; Heat Shock; Plant Retrotransposons; LINE Retrotransposons; Horizontal Transposon Transfer (HTT)
ABSTRACT
We report the isolation of AtL1, a 249 bp non-LTR retrotransposon fragment from Arabidopsis thaliana by fingerprinting mRNAs extracted from A. thaliana plants, ecotype Columbia, in different heat stress conditions. Southern blot and PCR analysis suggested that AtL1 occurs as a singleor low-copy insert in the genome of A. thaliana ecotype Columbia. The presence of AtL1 in the genome of different Arabidopsis ecotypes was confirmed by PCR amplification and sequencing thus excluding all possible contamination. A preliminary scan of the AtL1 nucleotide sequence against the EMBL and NCBI databases revealed a high degree of similarity to a group of LINE type L1 retrotransposons of mammals and with a cDNA sequence of Artemisia annua. A phylogenetic analysis of LINE elements from animals and plants placed AtL1 and A. annua sequences in close proximity to some mammalian sequences but distant from the other plants LINE elements including those from Arabidopsis.
1. Introduction
Eukaryotic retrotransposons [1,2] have been grouped into two major subclasses: the long terminal repeat (LTR) elements and the non-LTR elements. The LTR retrotransposons are flanked by long terminal repeats and encode proteins similar to those of retroviruses; they have been found ubiquitous in the main taxonomic groups of plant species [3]. The non-LTR retrotransposons lack the terminal repeats and are usually classified into short interspersed nuclear elements (SINEs) and long interspersed nuclear elements (LINEs) [1].
All plant LINEs discovered so far belong to the L1 superfamily, which has been designated according to the human and mouse L1 LINEs [4,5]. The first LINE retrotransposon described in a plant species was the Cin4 element from Zea mays L., detected as an insertion in the 3’ untranslated region of the A1 gene [6]. Subsequently, LINEs were found in various plants such as Lilium speciosum Thunb. (del2), Arabidopsis thaliana (L.) Heynh. (Ta11-1, ATLN), Cannabis sativa L. (LINE-CS), Hordeum vulgare L. (BLIN), Oryza sativa L. (Karma), Ipomoea batatas (L.) Lam. (LIb) and in the unicellular green alga Chlorella vulgaris (Zepp) [7-14]. The presence of multiple LINE copies was documented also in Beta vulgaris L. [15,16] and in Gossypium spp. [17].
Sequence divergence and extreme heterogeneity are typical features of plant LINEs [18] and most of the LINE copies are heavily truncated and appear to be non-functional due to stop codons or to frame-shift mutations. Defective copies, often truncated at the 5’ end, were likely originated from incomplete reverse transcription/amplification, while sequence heterogeneity is mainly due to accumulations of nucleotide substitutions over long evolutionary time scales [19]. Of particular interest was the detection of a truncated LINE element in the mitochondrial genome of A. thaliana: its high similarity to nuclear retrotransposon sequences clearly suggests a transfer/insertion event from the nuclear to the mitochondrial compartment [20].
The mechanisms of transcription and mobilization of non-LTR retroelements is less well known than that of LTR retrotransposons in plants. It has been suggested that the activation of plant retrotransposons occurs under stress condition [21,22] and it is subjected to a developmental control [10]. In some cases, transcription of plant retrotransposons seems to be the result of a co-transcriptional event such as in the case of the S1 family of SINE in Brassica napus L. [23], or in the case of the LINE-like fragment (TSCL) isolated from A. thaliana inserted downstream of a tissue-specific promoter region [24].
In this paper, we report the identification and characterization of the AtL1 non-LTR retrotransposon sequence, from A. thaliana ecotypes Columbia, Landsberg and Wassilewskija with homology to LINE from animals and plants. An analysis of the phylogenetic relationships between AtL1 and an extensive set of LINE sequences belonging to animals and plants was performed and its evolutionary origin is discussed considering the possibility of an inter-kingdom horizontal transfer (HT) of retrotransposable elements (RTEs).
2. Materials and Methods
2.1. Plant Materials
Seeds of A. thaliana ecotypes Columbia, Landsberg and Wassilewskija were supplied by Nottingham Arabidopsis Stock Center, UK. Seeds of different batches were compared in parallel experiments. A. thaliana plants were grown in axenic conditions in Petri dishes on MS basal salt medium [25] supplemented with 1% (w/v) sucrose and 0.8% (w/v) agar, at 22˚C with a 16 h photoperiod. The fluorescence lamps provided an illumination of approximately 120 µEm−2∙s−1 at plant height. Particular attention was placed to the sterilisation of seeds and to the growth of the plants in extremely axenic conditions to prevent any type of contamination.
Ten days old plantlets were treated at different temperatures in a thermostatic chamber in the dark as previously described [26]: 1) 2 h at 22˚C, control temperature; 2) 2 h at 37˚C, a mild heat stress (HS) condition; 3) 2 h at 37˚C followed by 2 h at 45˚C, a HS treatment inducing thermotolerance; 4) 2 h at 45˚C, a severe and mostly lethal HS condition. Immediately after treatment plants were collected and frozen in liquid nitrogen for RNA and DNA analyses.
2.2. RNA Isolation, DDRT-PCR and RT-PCR
Total RNA was extracted from axenically grown plants exposed to different thermal conditions using a phenolchloroform procedure, followed by LiCl precipitation and purification [27]. Differential Display (DDRT-PCR) was performed following the protocol of Liang and Pardee (1992) [28]. Reverse transcriptase (RT) reactions were performed using an oligonucleotide anchored primer (T12GA). Accidental amplification of contaminant DNA was excluded by subjecting the same samples to control reactions without reverse transcriptase. DNA-RNA hybrids were heat denatured at 95˚C, 2 min and PCR amplification was performed using the T12GA anchored primer combined with the arbitrary 10-mer primer AP5 (Table 1). The purified and amplified DDRT-1 fragment was cloned into the pGEM®-T vector (Promega, Madison, WI, USA) and sequenced following the protocol “CEQTM 2000 dye terminator cycle sequencing” (Beckman Coulter, Fullerton, CA, USA), by means of the automatic sequencer CEQ 2000 (Beckman Coulter). For semi-quantitative RT-PCR, total RNA isolated from plants subjected to different HS conditions was placed in a solution of 10 mM Tris-HCl pH 8.0, 10 mM MgCl2, containing 10 U of RNAse-free DNAse (Promega) for 30 min at 37˚C. RNA samples (1.5 μg) were retrotranscribed for 45 min at 37˚C with 100 U of M-MuLV Reverse Transcriptase RNase H Minus Point Mutant (Promega), in the presence of the specific primer LINE1-for (Table 1), priming DNA synthesis towards the 5’ end of AtL1 (Figure 1). To test for DNA contamination, control reactions were performed in the same conditions without RT. The amount of cDNA synthesized, as determined by spectrophotometer measurements, ranged from 300 - 600 ng per RT reaction. After cDNA synthesis, the RNADNA hybrids were heat denatured at 95˚C for 2 min. First strand cDNAs (500 pg) were then directly PCRamplified by using the same primer as for the RT reaction (AtL1-for) in combination with the LINE1-rev primer (Table 1), priming DNA synthesis towards the 3’
Table 1. Primer sequences.
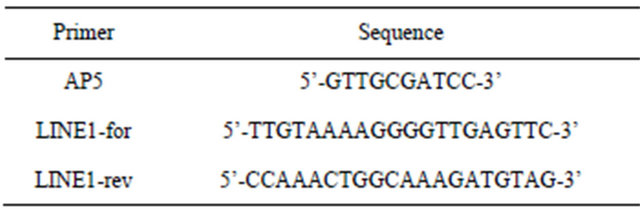
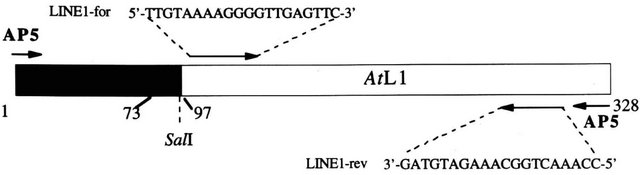
Figure 1. Drawing of DDRT-1 sequence. The region from position 1 to position 96 shows 100% identity to the 3’ end of the Athsp23.5 transcript (black box). The region from position 74 to position 322 shows a 78% similarity to the mammalian LINE-1 elements. Restriction map site SalI identifies AtL1 fragment. The primers utilized to amplify the retrotranscribed mRNA are indicated.
end of AtL1 (Figure 1). As internal control, the same PCR conditions were applied for the amplification of the Athsp23.5 cloned sequence [26]. PCR conditions were as follows: final volume 50 μl, 50 nM each primer, 0.5 U of Platinum® Pfx DNApolymerase (Invitrogen, Carlsbad, CA, USA), 100 mM each dNTP, 1.5 mM MgCl2, 50 mM KCl, 10 mM Tris-HCl pH 8.3, 20 cycles at 94˚C 30 sec, 50˚C 60 sec, 70˚C 60 sec followed by a final extension step at 70˚C for 5 min. The PCR products were separated by electrophoresis on 2% (w/v) agarose gel, blotted onto nylon membranes and hybridised with AtL1 probe.
2.3. DNA Isolation and Southern Blot Analysis
For Southern blot analysis, total DNA of ecotypes Columbia, Landsberg and Wassilewskija was isolated from axenically grown plants following a modified CTAB (hexadecyltrimethylammonium bromide) method as previously described [26]. DNA samples of Columbia ecotype were digested with each of the three restriction enzymes EcoRI, HindIII and EcoRV. Digestion, electrophoresis and blotting were performed following previously described methods [29]. Pre-hybridization and hybridization were performed at 65˚C in 5 × SSPE buffer, 5 × Denhardt’s solution and 0.5% (w/v) sodium dodecyl sulfate (SDS). Radiolabelled probes ([32P]-dCTP with a specific activity > 110 TBq/mmol) were prepared according to [30]. After hybridization, blots were washed twice in 2 × SSC buffer and 0.1% SDS for 5 min at room temperature, twice in 0.5 × SSC and 0.1% SDS for 15 min at 65˚C, and once in 0.1 × SSC and 0.1% SDS for 15 min at 65˚C. The consistency of these methods was confirmed by PCR analysis with primers LINE1-for and LINE1-rev, and by sequencing the amplicon obtained.
2.4. Bioinformatic Methods
The AtL1 nucleotide sequence was compared with the EMBL database by the FASTA program [31]. Other comparisons with databases were performed using the BLASTn program (http://www.ncbi.nlm.nih.gov) and the BLAST tool at TAIR (http://www.arabidopsis.org/). Multiple alignment of the amino acid sequences that were deduced from AtL1 and from an heterogeneous set of LINEs, spanning a broad spectrum of phylogenetic diversity, was built by the CLUSTALW2 program [32], using the BLOSUM matrix [33]. The alignment was visualised by using the Jalview program [34], in which the residues conservation, the quality of the alignment and the consensus sequence were also built. Phylogenetic and molecular evolutionary analyses were conducted using the MEGA version 4 software [35], available at http://www.megasoftware.net. The phylogenetic position of AtL1 was elucidated by the neighbor-joining (N-J) method [36]; reliability of the phylogenetic tree was assessed by a bootstrap test with 1000 data set resamplings.
3. Results and Discussion
3.1. Isolation and Characterization of DDRT-1 Fragment
A cDNA named DDRT-1 specifically transcribed in condition of expression of thermotolerance was isolated and cloned in our laboratory by fingerprinting mRNAs extracted from A. thaliana, ecotype Columbia. It was clearly excluded that the cDNA originated from accidental contamination by other forms of DNA (see Section 2).
Nucleotide sequence of DDRT-1 showed a size of 322 bp, the sequence of the arbitrary primer AP5 at the 5’ end, and the sequence complementary to AP5 at the 3’ end (Figure 1). DDRT-1 can be considered a product of the RNA retro-transcription obtained with amplification by arbitrary primers annealing at both ends in inverted fashion.
A search for sequence similarities against the EMBL and NCBI databases evidenced for DDRT-1 a perfect match, from nucleotide 1 to 96, to the 3’ region of the cDNA of the Athsp23.5 gene, which encodes for a low molecular weight mitochondrial heat shock protein [26]. In the Athsp23.5 gene the region of identity with DDRT- 1 spanned the end of the protein-coding region (63 bp) and the 33 bp located downstream of the stop codon.
The similarity ended there because the remaining portion of the DDRT-1 sequence exhibited a high degree of similarity with the LINE L1.1 retrotransposons of mammals. The region of similarity spans nucleotides 74 to 322 and this sequence has been named AtL1 (A. thaliana LINE-1; accession number AJ012311; Figure 1).
3.2. AtL1 Sequence, Its Presence in the Genomes of the A. thaliana Ecotypes Columbia, Landsberg and Wassilewskija
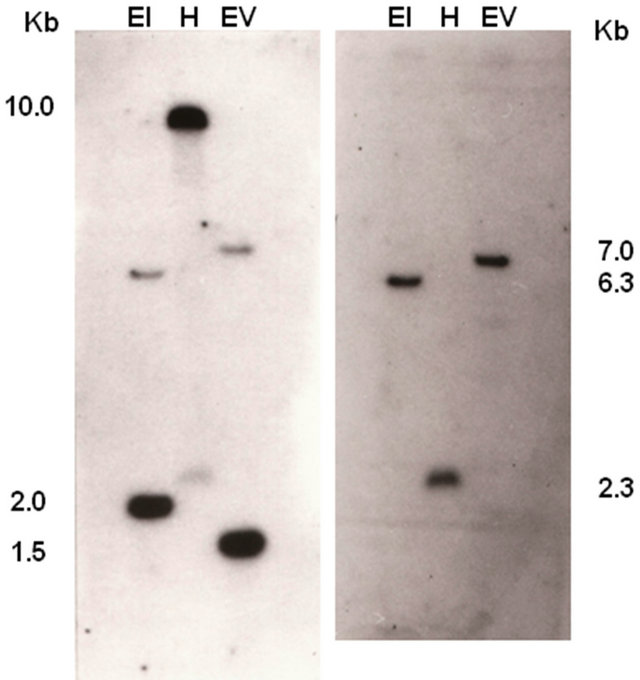
To confirm the presence of AtL1 sequence in the genome of A. thaliana ecotype Columbia, total genomic DNA was hybridized with the complete DDRT-1 sequence (322 bp), comprising both 96 bp from the 3’ end of the Athsp23.5 gene and the whole AtL1 sequence. Figure 2(a) shows the pattern of hybridization of DNA samples digested with restriction enzymes EcoRI, HindIII and EcoRV: hybridization bands corresponded to restriction fragments of 10.0, 2.0 and 1.5 kb respectively, in addition others bands were visible with less intensity (7.0, 6.3, and 2.3 kb). The same DNA blot was hybridized with a 204 bp probe whose sequence corresponds to the entire 3’ end of Athsp23.5 transcript (accession number Y11865),
 (a) (b)
(a) (b)
Figure 2. Southern blot analysis of total genomic DNA from A. thaliana ecotype Columbia. Restriction enzymes were: EcoRI (EI), HindIII (H), EcoRV (EV). Hybridization was performed as described in materials and methods using as probe: (a) The DDRT-1 fragment containing the AtL1 element; (b) A 204 nt fragment corresponding to the 3’ end of the Athsp23.5 transcript. The sizes of the genomic fragments are estimated by comparison with 1 Kb DNA ladder (New England Biolabs).
including the 96 bp present in DDRT-1, giving hybridization bands of 7.0, 2.3 and 6.3 kb (Figure 2(b)). The size of these restriction fragments are consistent with those reported for the hybridization of genomic DNA with the whole Athsp23.5 gene [26]. The results of the Southern blot were totally confirmed by PCR with primers LINE1- for and -rev, and by sequencing the amplicon obtained.
AtL1 was isolated from DDRT-1, in which it was linked, in opposite orientation, with a limited 3’ portion of the cDNA of a heat shock protein gene, Athsp23.5. A similar situation was found for the A1 gene in Z. mays and the Cin4 LINE element [6]. Two identical genomic clones isolated in our case from a library of the Columbia ecotype and containing the Athsp23.5 gene, did not reveal the presence of nearby or adjacent AtL1 sequence [26]. The two hybridization patterns found when using the AtL1cDNA or the 3’ end cDNA fragment of Athsp23.5, as probes (Figure 2), argue against a physical association of the two. AtL1 fragment therefore resides in a genomic location different from Athsp23.5 and the association between cDNA of AtL1 and of Athsp23.5 probably resulted during or after the processing of the relative transcripts. Southern blot analysis (Figure 2(a)), showed also that AtL1 did not occur at multiple locations in the genome of A. thaliana ecotype Columbia, a feature uncommon to other non-LTR transposons in plants [1116].
To confirm the presence of the AtL1 in the genome of different A. thaliana ecotypes with an independent approach, two specific primers were designed from the sequence of AtL1 (LINE1-for and LINE1-rev, Table 1 and Figure 1). These primers were used to amplify orthologous sequences out of genomic DNA extracted from A. thaliana ecotypes Columbia, Landsberg and Wassilewskija. Different amplicons were obtained, and among these a band of the expected dimension, about 200 bp, was present in all the ecotypes analyzed (Figure 3(a), lanes 1, 2, 3). Hybridization with DDRT-1 confirmed the homology of these 200 bp amplicons with AtL1 (Figure 3(b)). Sequencing revealed a complete nucleotide identity between the AtL1 fragments isolated from Columbia and Landsberg ecotypes, while two nucleotide substitutions, one A to G transition at nucleotide 52 and one G to T transversion at nucleotide 211, were found in the AtL1 orthologous from Wassilewskija.
To exclude possible accidental contamination during laboratory operation, amplification and cloning of the AtL1 sequences was confirmed in a different laboratory. During these controls an AtL1 sequence was identified also in the Arabidopsis ecotype Capo Verde Island (S. Gazzani personal communication). Repeated attempts of retrieving genomic clones spanning the homologous AtL1 sequences from A. thaliana genomic libraries in lambda phages were unyielding. A factor that can have played a negative role in the search for a genomic clone of AtL1 is the particular genomic locations of retrotransposons near heterocromatic, centromeric or telomeric regions. Of the 219 LINE homologues that have been discovered in A. thaliana by in silico analyses [11], only few were physically identified in 170 BAC and PAC clones.
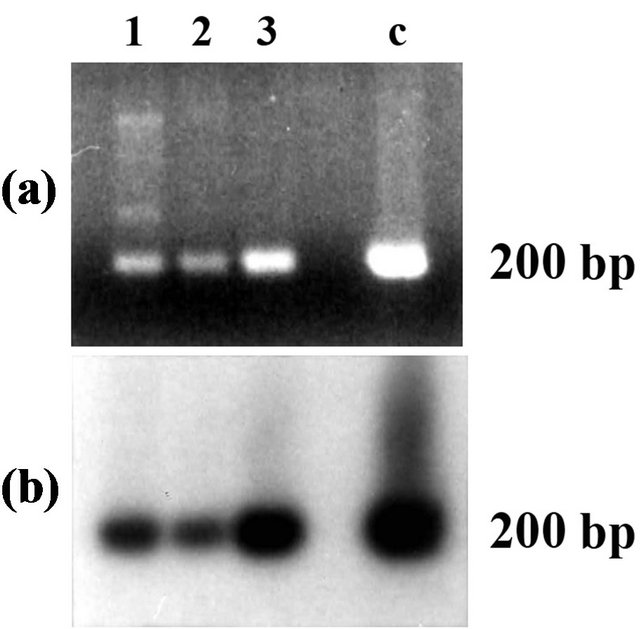
Figure 3. (a) Gel electrophoresis and (b) hybridization with the AtL1 probe of the PCR products obtained by amplification of genomic DNA of three different A. thaliana ecotypes, using LINE1-for and LINE1-rev primers. Lane labels correspond to the A. thaliana ecotypes Columbia (1), Landsberg (2) and Wassilewskija (3). Lane c corresponds to the positive control, amplified from the cloned AtL1 fragment. The size of the hybridizing fragments is estimated by comparison to a 100 bp DNA ladder.
3.3. Expression of AtL1 in Plants Exposed to Different Thermal Conditions
AtL1 was isolated as a product of an in vitro cDNA synthesis starting from mRNA extracted from plants subjected to HS. To verify the presence of the AtL1 transcript in different HS conditions, RNAs were extracted from A. thaliana plants subjected to: 1) mild heat stress (37˚C); 2) HS treatment leading to expression of thermotolerance (37˚C to 45˚C); and 3) severe (lethal) HS (45˚C) and were analyzed by semi-quantitative RT-PCR with AtL1 specific primers. The RT-PCR amplification products were blotted and then hybridized with the AtL1 fragment. As internal control, the same RNA preparation was amplified by RT-PCR with primers designed for the transcript of Athsp23.5 [26]. Figure 4 shows the hybridization of RT-PCR products with AtL1: in all four lanes there is a signal corresponding to an amplified cDNA fragment of about 200 bp. Temperature treatments causing a severe HS or expression of thermotolerance led to the higher levels of the AtL1 transcription (Figure 4, lanes c and d). Expression of AtL1 was low but detectable at the temperature of 22˚C (Figure 4 lane a). The temperature dependent regulation of AtL1 could be either accidental, the gene contains an internal heat shock promoter, or instrumental in the regulation of Athsp23.5.
In the past years it was demonstrated that transcription and mobilization of active retrotransposons in plant species can be induced by various stress conditions [21,22, and references therein]. This led to the hypothesis that transposable elements are involved in host adaptation to environmental changes [37,38]. Transcription of the Tto1 element from tobacco, induced in response to a variety of environmental stresses, is mediated by a cis-regulatory element that was localized in the 5’ LTR [39]. In the same plant species, during microbial infections and wounding the expression of the Tnt1A retrotransposon is
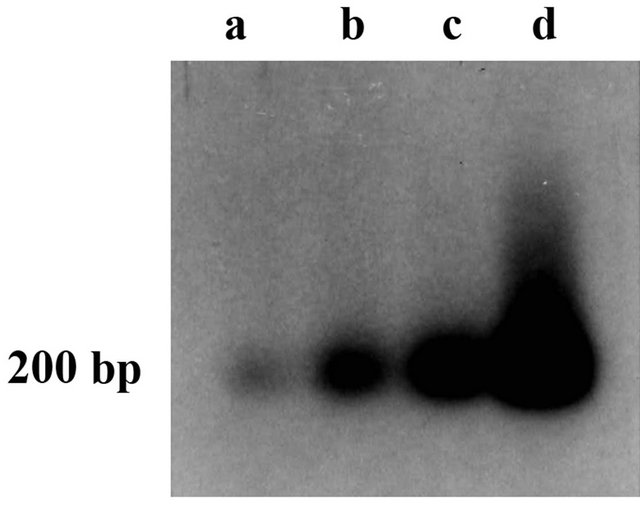
Figure 4. Reverse transcriptase-PCR analysis of AtL1. Hybridization with the AtL1 probe of the RT-PCR amplification products derived from RNA extracted from A. thaliana plants subjected to different HS conditions: (a) 22˚C; (b) 37˚C; (c) 37˚C to 45˚C; (d) 45˚C. The PCR reactions were performed with LINE1-for and LINE1-rev primers. The size of the hybridizing fragments is estimated by comparison to a 100 bp DNA ladder.
induced by tandemly repeated cis-acting regulatory elements that were detected in the 5’ LTR U3 region [40].
3.4. Phylogenetic Analysis
In silico comparison of the AtL1 sequence with the plant DNA library in the TAIR database show only a limited similarity with known nucleotide sequences, including many LINE elements so far characterized in the A. thaliana species [8,11]. An additional search (http://www. arabidopsis.org/Blast) within deposited plant cDNA sequences at TAIR revealed a high level of homology (81% identity) with a cDNA sequence isolated from Artemisia annua L. (EY057597).
A pairwise alignment of the inverted complementary sequence of AtL1 with one of the most similar sequences (accession number S65824), which corresponds to the region of a human LINE L1.1 encoding for reverse transcriptase domains V-VII, stressed the presence in AtL1 of three short deletion mismatches that alter its proper reading frame. A manual optimization of these frame shifts brought to the alignment in Figure 5 of the amino acid sequence deduced from AtL1 in comparison with LINE sequences: one from human (LINE L1.1 S65824) (48% identity) and one from A. thaliana (Ta27-AC007195) (19% identity) [8]. The inconsistent recognition, by the FASTA program or the BLASTn program, of a significant matching between AtL1 and Ta27 nucleotide sequences, as well as with the other LINE copies contained in the genome of A. thaliana, could be ascribed only to the remarkable difference existing at the level of base composition: AtL1 39% A, 23% T, 15% G and 23% C; Ta27 was 31% A, 31% T, 21% G and 17% C.
To better understand the degree of similarities between AtL1 and other retrotransposon sequences, additional LINEs belonging to animals and plants were considered. A scan of the AtL1 sequence against the various libraries of the EMBL database evidenced relevant similarities to LINEs of non-human mammals, such as capuchin monkey (Cebus albifrons) (M81409), cow (X59856), dog (AB01223), rabbit (M18818) and mouse (M13002). The set of plant LINEs was extended by including divergent members of the A. thaliana LINE family Ta11 (L47913); A. thaliana Ta12 (AF058826); A. thaliana Ta14 (AB 009056); A. thaliana Ta19 (AC006920); A. thaliana Ta21 (AC006570); A. thaliana Ta22 (AC002387); A. thaliana Ta23 (AC007069); A. thaliana Ta27 (AC 007195) [8,11] and the Cin4 element of Z. mays (Y00086) [6]. In addition by using the BLASTp programme at NCBI and the human LINE L1.1 (S65824) protein sequence as a query, a search for protein sequences in the Viridiplantae database was performed and additional plant putative non-LTR retroelements were identified and added to the analysis. In particular a putative non-LTR retroelement RT from O. sativa (ABF94719.1), a Sorghum bicolor (L.) Moench non-LTR RT sequence (XP002465860) and two Arabidopsis sequences annotated as non-LTR retroelement RT-like proteins (AAF 99785; BAB09379) were identified with an elevated score. In addition a search by using tBLASTn programme of LINE L1.1 protein (S65824) against the database of translated plant sequences found many high similarities with sequences of different plant species. Among them two A. thaliana (AL353814; AB028607), a sequence of Aegilops tauschii Coss. (CT009625) and a Brassica rapa L. translated sequences (AC189475.2) were fished out in our analyses.
The amino acidic sequence comprised between RT domains V and VII was deduced from all the sequences considered and a multiple alignment containing AtL1 was built by the CLUSTALW2 program (Figure 6). A visual inspection of the alignment showed a high conservation

Figure 5. Optimal alignment of the amino acid sequences corresponding to the reverse transcriptase domains V-VII. The sequences were deduced from AtL1, from a human LINE-1 L1.1 element (Acc. Number S65824) and from the A. thaliana Ta27 LINE (Acc. number AC007195). Asterisks denote perfectly conserved amino acid residues. The character “:” indicates highly similar amino acid residues. The character “.” indicates low similar amino acid residues. When positioned within the amino acid sequence (see AtL1) the character “*” denotes an internal stop codon.
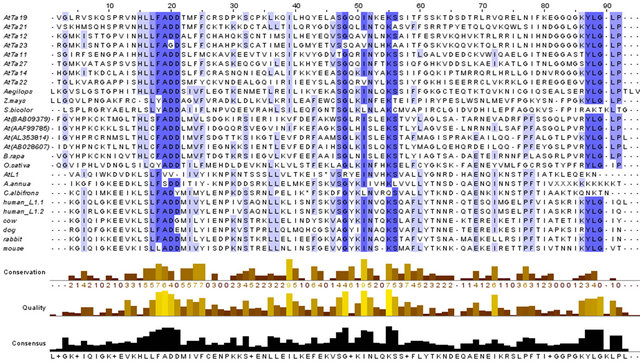
Figure 6. Multiple alignment of 25 amino acid sequences corresponding to the reverse transcriptase domains V-VII. They were deduced from the corresponding nucleotide sequences belonging to plant and vertebrate LINEs, whose accession numbers are as follows: AtL1 (AJ012311); Human L1.1 (S65824); Human L1.2 (M80343); C. albifrons (M81409); Cow (X59856); Dog (AB01223); Rabbit (M18818); Mouse (M13002); A. thaliana Ta11 (L47913); A. thaliana Ta12 (AF058826); A. thaliana Ta14 (AB009056); A. thaliana Ta19 (AC006920); A. thaliana Ta21 (AC006570); A. thaliana Ta22 (AC002387); A. thaliana Ta23 (AC007069); A. thaliana Ta27 (AC007195); Z. mays (Y00086); B. rapa (AC189475.2); O. sativa (ABF94719.1); A. thaliana (AAF99785; BAB09379; AL353814; AB028607) S. bicolor (XP002465860); Ae. tauschii (CT009625). Amino acid conservation, quality and consensus sequence are also given. Identical residues are coloured in a scale of blue according to the % of the residues in each column that agree with the consensus sequence.
of amino acid residues inside the group formed by AtL1, A. annua and LINE from vertebrates, different from that visualized in the other plant sequences. In particular a strong conservation of KLS residues at 14-16 positions is peculiar to this group of sequences.
A phylogenetic analysis was carried out by evaluating the alignments in Figure 6 which included LINEs from plants and animals. Using the N-J method, it was obtained a phylogenetic tree which summarized the evolutionary relationships among the 25 LINE homologues under examination (Figure 7). Robustness of the main branching nodes of the tree was evaluated by the bootstrap test, which emphasized the presence of three different clusters: 1) AtL1, Artemisia annua sequence, and mammal LINEs including human LINEs; 2) plant sequences isolated by homology to human LINE L1.1 (S65824); and 3) LINE homologues which were previously isolated in A. thaliana by homology with other plant LINEs. AtL1 sequence, as well as A. annua sequence, albeit of plant origin, was unambiguously positioned in the cluster of animal LINEs. An unexpected similarity between plant and animal retrotransposons has been found for a non-LTR element of the RTE clade: the closest relative to RTE elements of medaka fish clustered with a corresponding RTE of the plant kingdom [41]. For the same element, it has also been reported that brown algae and sea urchin share higher homology levels than expected.
Usually the most parsimonious hypothesis explaining these inconsistencies in phylogenetic grouping is horizontal transfer (HT): in this case, of transposable elements (HTT) [42-44]. Grandbastien et al. [45] first hypothesized a possible horizontal transfer of a LTR retroelement between insects and plants. They found, in fact, a much higher similarity between the elements Tnt1 of Nicotiana tabacum L. and copia of Drosophila melanogaster than might be expected between plants and Diptera. HTT was further supported by a comprehensive

Figure 7. Consensus phylogenetic tree obtained from N-J analysis of the reverse transcriptase nucleotide sequence from vertebrate and plant LINEs. The internal node numbers represent the bootstrap values, expressed as a percentage of all trees, obtained from 1000 replicates. Only nodes supported by bootstrap proportions higher than 60% are indicated.
study of Konieczny and co-workers [46], by comparing Ty1-copia-like elements from A. thaliana with those of other organisms including D. melanogaster, yeast and tobacco. The authors evidenced a clade showing a pattern consistent with horizontal inheritance. HTT including tRNA-related SINEs from animal to plants has also been proposed by Pozueta-Romero et al. [47]: tRNA-related region of plant SINEs, belonging to the Ts family, shows an 80% identity with hybrid sequences derived from the tRNALys and tRNAMet genes of the nematode Caenorhabditis elegans. A SINE element similar to the equivalent of a mammalian one has also been detected in a cDNA fragment isolated from Pisum sativum L. [48]. As regards the LINEs, the HTT of Bov-B LINEs from marsupials to Squamata to the ancestor of Ruminantia, likely mediated by ticks, is one of the most compelling examples of gene exchange between distantly related species [49,50].
4. Concluding Remarks
In this paper we present the identification and characterization of AtL1, a defective reverse transcriptase LINE-like element from A. thaliana, with high homology tomammalian LINEs. The existence of AtL1 has been demonstrated by three different types of evidences: 1) Southern blot analysis; 2) sequencing of PCR amplification products; 3) RT-PCR amplification with internal specific primers. Phylogenetic relationship with LINEs from animals and plants suggests a HT from animals to plants rather than within plant species.
Though the movement of genetic material between reproductively isolated species (HT) is known to play an important role in genome evolution in eukaryotes, the main obstacle to really support the possibility of HT between evolutionary distant kingdoms like animals and plants, is the difficulty of envisaging an efficient potential pathway for the exchange of the genetic material [51, 52]. On the other hand there could be, probably, no transposable elements (TEs) without HT, because this is an essential part of life cycle of TEs to avoid co-evolved suppression mechanisms with the purpose of limiting TEs mobility within lineages [53].
Viruses, bacteria, fungi, and also parasites and symbionts, metazoa and arthropods could be all considered potential vectors [43-45]. Often, viruses have been invoked as attractive candidates to transport DNA sequences between reproductively isolated species where they are carried by specific vectors, like insects [43,44, 54,55 and references therein], but they show a limited host range, thus a decreasing probability for cross-species infection with increasing evolutionary distance.
Transformation mediated by short free DNA molecules could be another possibility [43,44], and despite the fact that it is widely used to generate transgenic organisms, its importance is frequently ignored in the theoretical discussions concerning introgression of genetic material in evolutionary distant species. Degraded free DNA molecules arising from dead cells or organisms, or carried by a virus or bacteria can be assumed by an intermediate host vector, a parasitic or saprophytic metazoa or arthropod and injected or introduced into a susceptible recipient thereafter.
In Figure 8 is compared the amino acid sequences of AtL1, the human LINE L1.1 (S65824) and a putative reverse transcriptase sequence of the insect Bombyx mori (ADI61832.1) detected by search of homology of human LINE in BLASTp programme selecting the Arthropoda database. There is a sufficient similarity to suggest that Arthropoda could have played a role in HT between the two kingdoms. Degraded DNA fragments can be more resistant to cleavage by nucleolytic enzymes present in the digestive guts of the intermediate vectors and have a chance to survive, transferring first to gut lining cells and after that to cells of other tissues [43,44].
On the other hand, some kinds of virus are involved in facilitating the parasitism of lepidopterans by wasps (hymenopterans). Therefore at least one mechanism for TEs maintenance within insects through their viruses may exist. The viral particles encode for factors suppressing immunity of the lepidopterans, thus facilitating

Figure 8. Optimal alignment performed with CLUSTALW2 programme of AtL1 with the amino acid sequences corresponding to the reverse transcriptase domains V-VII from a human L1.1 element (Acc. Number S65824) and from the Bombyx mori endonuclease-reverse transcriptase sequence (Acc. Number ADI61832.1). Asterisks denote perfectly conserved amino acid residues. The character “:” indicates highly similar amino acid residues. The character “.” indicates lowly similar amino acid residues. When positioned within the amino acid sequence (see AtL1) the character “*” denotes an internal stop codon.
the growth of the wasp larvae. It has been suggested [56] that a close association between the parasitoid wasp and lepidopterans could facilitate HT for TEs. In addition to virus infection, some parasitic insects have also been implicated with HT because of their association with their host [57].
In conclusion, the HT of RTEs (LINE, AtL1) could be proposed only on the basis of theoretical considerations on previously reported cases and literature. But the advantages of HT are clear: 1) from genetic innovation without sex; to 2) diversification within lineages, with the general purpose for the evolution of increasing the plasticity of the genome.
5. Acknowledgements
The authors thank Dr. Silvia Gazzani (Environment and Natural Resources Area, Fondazione Edmund Mach, via Mach 1, San Michele all’Adige) for her great contribution to the experimental work and the preparation of the paper. We are grateful to the Nottingham Arabidopsis Stock Center for the kind gift of seeds. The work was supported by the contribution of FIL (University of Parma Local Funding for Research) to Prof. Nelson Marmiroli, Prof. Elena Maestri and Dr. Giovanna Visioli.
REFERENCES
- A. Kumar and J. L. Bennetzen, “Plant Retrotransposons,” Annual Review of Genetics, Vol. 33, 1999, pp. 479-532. doi:10.1146/annurev.genet.33.1.479
- H. L. Levin and J. V. Moran, “Dynamic Interactions between Transposable Elements and Their Hosts,” Nature Reviews Genetics, Vol. 12, No. 9, 2011, pp. 615-627. doi:10.1038/nrg3030
- D. F. Voytas, M. P. Cummings, A. Koniczny, F. M. Ausubel and S. R. Rodermel “Copia-Like Retrotransposons Are Ubiquitous among Plants,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 89, No. 15, 1992, pp. 7124-7128. doi:10.1073/pnas.89.15.7124
- B. A. Dombroski, S. L. Mathias, E. Nanthakumar, A. F. Scott and H. H. Kazazian Jr., “Isolation of an Active Human Transposable Element,” Science, Vol. 254, No. 5039, 1991, pp. 1805-1808. doi:10.1126/science.1662412
- D. D. Loeb, R. W. Padgett, S. C. Hardies, W. R. Shehee, M. B. Comer, M. H. Edgell and C. A. Hutchison III, “The Sequence of a Large L1Md Element Reveals a Tandemly Repeated 5’ End and Several Features Found in Retrotransposons,” Molecular and Cellular Biology, Vol. 6, No. 1, 1986, pp. 168-182. doi:10.1128/MCB.6.1.168
- Z. Schwarz-Sommer, L. Leclercq, E. Goebel and H. Saedler, “Cin4, an Insert Altering the Structure of the A1Gene in Zea mays, Exhibits Properties of Nonviral Retrotransposons,” The EMBO Journal, Vol. 6, No. 13, 1987, pp. 3873-3880.
- P. R. J. Leeton and D. R. Smyth, “An Abundant LINE-Like Element Amplified in the Genome of Lilium speciosum,” Molecular and General Genetics, Vol. 237, No. 1-2, 1993, pp. 97-104. doi:10.1007/BF00282789
- D. A. Wright, N. Ke, J. Smalle, B. M. Hauge, H. M. Goodman and D. F. Voytas, “Multiple Non-LTR Retrotransposons in the Genome of Arabidopsis thaliana,” Genetics, Vol. 142, No. 2, 1996, pp. 569-578.
- T. Higashiyama, Y. Noutoshi, M. Fujie and T. Yamada, “Zepp, a LINE-Like Retrotransposon Accumulated in the Chlorella Telomeric Region,” The EMBO Journal, Vol. 16, No. 12, 1997, pp. 3715-3723. doi:10.1093/emboj/16.12.3715
- M. Komatsu, K. Shimamoto and J. Kyozuka, “Two-Step Regulation and Continuous Retrotransposition of the Rice LINE-Type Retrotransposon Karma,” The Plant Cell, Vol. 15, No. 8, 2003, pp. 1934-1944.doi:10.1105/tpc.011809
- K. Noma, H. Ohtsubo and E. Ohtsubo, “ATLN Elements, LINEs from Arabidopsis thaliana: Identification and Characterisation,” DNA Research, Vol. 7, No. 5, 2000, pp. 291-303.doi:10.1093/dnares/7.5.291
- K. Sakamoto, N. Ohmido, K. Fukui, H. Kamada and S. Satoh, “Site Specific Accumulation of a LINE-Like Retrotransposon in a Sex Chromosome of the Dioecious Plant Cannabis sativa,” Plant Molecular Biology, Vol. 44, No. 6, 2000, pp. 723-732.doi:10.1023/A:1026574405717
- V. Vershinin, A. Druka, A. G. Alkhimova, A. Kleinhofs and J. S. Heslop-Harrison, “LINEs and gypsy-Like Retrotransposons in HordeumSpecies,” Plant Molecular Biology, Vol. 49, No. 1, 2002, pp. 1-14. doi:10.1023/A:1014469830680
- H. Yamashita and M. Tahara, “A LINE-Type Retrotransposon Active in Meristem Stem Cells Causes Heritable Transpositions in the Sweet Potato Genome,” Plant Molecular Biology, Vol. 61, No. 1-2, 2006, pp. 79-94. doi:10.1007/s11103-005-6002-9
- T. Schmidt, S. Kubis and J. S. Heslop-Harrison, “Analysis and Chromosomal Localization of Retrotransposons in Sugar Beet (Beta vulgaris L.): LINEs and Ty1-Copia-Like Elements as Major Components of the Genome,” Chromosome Research, Vol. 3, No. 6, 1995, pp. 335-345. doi:10.1007/BF00710014
- T. Wenke, D. Holtgraewe, A. V. Horn, B. Weisshaar and T. Schmidt, “An Abundant and Heavily Truncated NonLTR Retrotransposon (LINE) Family in Beta vulgaris,” Plant Molecular Biology, Vol. 71, No. 6, 2009, pp. 585- 597. doi:10.1007/s11103-009-9542-6
- J. S. Hawkins, G. Hu, R. A. Rapp, J. L. Grafenberg and J. F. Wendel, “Phylogenetic Determination of the Pace of Transposable Element Proliferation in Plants: copia and LINE-Like Elements in Gossypium,” Genome, Vol. 51, No. 1, 2008, pp. 11-18. doi:10.1139/G07-099
- S. E. Kubis, J. S. Heslop-Harrison, C. Desel and T. Schmidt, “The Genomic Organization of non-LTR Retransposons (LINEs) from Three Beta Species and Five Other Angiosperms,” Plant Molecular Biology, Vol. 36, No. 6, 1998, pp. 821-831. doi:10.1023/A:1005973932556
- C. Feschotte, N. Jiang and S. R. Wessler, “Plant Transposable Elements: Where Genetics Meets Genomics,” Nature Reviews Genetics, Vol. 3, No. 5, 2002, pp. 329- 341. doi:10.1038/nrg793
- V. Knoop, U. Unseld, J. Marienfeld, P. Brandt, S. Sunkel, H. Ullrich and A. Brennicke, “Copia-, gypsyand LINELike Retrotransposon Fragments in the Mitochondrial Genome of Arabidopsis thaliana,” Genetics, Vol. 142, No. 2, 1996, pp. 579-585.
- H. Ito, H. Gaubert, E. Bucher, M. Mirouze, I. Vaillant and J. Paszkowski, “An siRNA Pathway Prevents Transgenerational Retrotransposition in Plants Subjected to Stress,” Nature, Vol. 472, No. 7341, 2011, pp. 115-119. doi:10.1038/nature09861
- M. A. Grandbastien, “Activation of Plant Retrotransposons under Stress Conditions,” Trends in Plant Science, Vol. 3, No. 5, 1998, pp. 181-187. doi:10.1016/S1360-1385(98)01232-1
- J. M. Deragon, N. Gilbert, L. Rouquet, A. Lenoir, P. Arnaud and G. Picard, “A Transcriptional Analysis of the S1Bn (Brassica napus) Family of SINE Retroposons,” Plant Molecular Biology, Vol. 32, No. 5, 1996, pp. 869- 878. doi:10.1007/BF00020484
- M. L. Chye, K. Y. Cheung and J. Xu, “Characterization of TSCL, a Nonviral Retroposon from Arabidopsis thaliana,” Plant Molecular Biology, Vol. 35, No. 6, 1997, pp. 893-904. doi:10.1023/A:1005947804227
- T. Murashige and F. Skoog, “A Revised Medium for Rapid Growth and Bio Assays with Tobacco Tissue Cultures,” Physiologia Plantarum, Vol. 15, No. 3, 1962, pp. 473-497. doi:10.1111/j.1399-3054.1962.tb08052.x
- G. Visioli, E. Maestri and N. Marmiroli, “Differential Display-Mediated Isolation of a Genomic Sequence for a Putative Mitochondrial LMW HSP Specifically Expressed in Condition of Induced Thermotolerance in Arabidopsis thaliana (L.) Heynh,” Plant Molecular Biology, Vol. 34, No. 3, 1997, pp. 517-527. doi:10.1023/A:1005824314022
- A. J. van Tunen, R. E. Koes, C. E. Spelt, A. R. van der Krol, A. R. Stuitje and J. N. M. Mol, “Cloning of the Two Chalcone Flavanone Isomerase Genes from Petunia hybrida: Coordinate, Light-Regulated and Differential Expression of Flavonoid Genes,” The EMBO Journal, Vol. 7, No. 5, 1988, pp. 1257-1263.
- P. Liang and A. B. Pardee, “Differential Display of Eukaryotic Messenger RNA by Means of the Polymerase Chain Reaction,” Science, Vol. 257, No. 5072, 1992, pp. 967-971. doi:10.1126/science.1354393
- J. Sambrook, E. F. Fritsch and T. Maniatis, “Molecular Cloning: A Laboratory Manual,” 2nd Edition, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1989.
- A. P. Feinberg and B. Vogelstein, “A Technique for Radiolabelling DNA Restriction Endonuclease Fragments to High Specific Activity,” Analytical Biochemistry, Vol. 132, No. 1, 1983, pp. 6-13. doi:10.1016/0003-2697(83)90418-9
- W. R. Pearson and D. J. Lipman, “Improved Tools for Biological Sequence Comparisons,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 85, No. 8, 1988, pp. 2444-2448. doi:10.1073/pnas.85.8.2444
- M. A. Larkin, G. Blackshields, N. P. Brown, et al., “Clustal W and Clustal X Version 2.0,” Bioinformatics, Vol. 23, No. 21, 2007, pp. 2947-2948. doi:10.1093/bioinformatics/btm404
- S. Henikoff and J. G. Henikoff, “Amino Acid Substitution Matrices from Protein Blocks,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 89, No. 22, 1992, pp. 10915-10919. doi:10.1073/pnas.89.22.10915
- M. Waterhouse, J. B. Procter, D. M. A Martin, M. Clamp and G. J. Barton, “Jalview Version 2—A Multiple Sequence Alignment Editor and Analysis Workbench,” Bioinformatics, Vol. 25, No. 9, 2009, pp. 1189-1191. doi:10.1093/bioinformatics/btp033
- K. Tamura, J. Dudley, M. Nei and S. Kumar, “MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0,” Molecular Biology and Evolution, Vol. 24, No. 8, 2007, pp. 1596-1599. doi:10.1093/molbev/msm092
- N. Saitou and M. Nei, “The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees,” Molecular Biology and Evolution, Vol. 4, No. 4, 1987, pp. 406-425.
- S. R. Wessler, “Plant Retrotransposons: Turned on by Stress,” Current Biology, Vol. 6, No. 8, 1996, pp. 959- 961. doi:10.1016/S0960-9822(02)00638-3
- D. Lisch, “How Important Are Transposons for Plant Evolution?” Nature Reviews Genetics, Vol. 14, No. 1, 2013, pp. 49-61.doi:10.1038/nrg3374
- S. Takeda, K. Sugimoto, H. Otsuki and H. Hirochika, “A 13-bp cis-Regulatory Element in the LTR Promoter of the Tobacco Retrotransposon Tto1 is Involved in Responsiveness to Tissue Culture, Wounding, Methyl Jasmonate and Fungal Elicitors,” The Plant Journal, Vol. 18, No. 4, 1999, pp. 383-393. doi:10.1046/j.1365-313X.1999.00460.x
- S. Vernhettes, M. A. Grandbastien and J. M. Casacuberta, “In Vivo Characterization of Transcriptional Regulatory Sequences Involved in the Defence-Associated Expression of the Tobacco Retrotransposon Tnt1,” Plant Molecular Biology, Vol. 35, No. 5, 1997, pp. 673-679. doi:10.1023/A:1005826605598
- V. Zupunski, F. Gubensek and D. Kordis, “Evolutionary Dynamics and Evolutionary History in the RTE Clade of Non-LTR Retrotransposons,” Molecular Biology and Evolution, Vol. 18, No. 10, 2001, pp. 1849-1863. doi:10.1093/oxfordjournals.molbev.a003727
- H. S. Malik, W. D. Burke and T. H. Eickbush, “The Age and Evolution of Non-LTR Retrotransposable Elements,” Molecular Biology and Evolution, Vol. 16, No. 6, 1999, pp. 793-805. doi:10.1093/oxfordjournals.molbev.a026164
- F. Bushman, “Lateral DNA Transfer: Mechanisms and Consequences,” Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 2002.
- S. Schaack, C. Gilbert and C. Feschotte, “Promiscuous DNA: Horizontal Transfer of Transposable Elements and Why It Matters for Eukaryotic Evolution,” Trends in Ecology and Evolution, Vol. 25, No. 9, 2010, pp. 537-546. doi:10.1016/j.tree.2010.06.001
- M. A. Grandbastien, A. Spielmann and M. Caboche, “Tnt1, a Mobile Retroviral-Like Transposable Element of Tobacco Isolated by Plant Cell Genetics,” Nature, Vol. 337, No. 6205, 1989, pp. 376-380.doi:10.1038/337376a0
- A. Konieczny, D. F. Voytas, M. P. Cummings and F. M. Ausubel, “A Superfamily of Arabidopsis thaliana Retrotransposons,” Genetics, Vol. 127, No. 4, 1991, pp. 801- 809.
- J. Pozueta-Romero, G. Houlné, and R. Schantz, “Identification of a Short Interspersed Repetitive Element in Partially Spliced Transcripts of the Bell Pepper (Capsicum annuum) PAPGene: New Evolutionary and Regulatory Aspects on Plant tRNA-Related SINEs,” Gene, Vol. 214, No. 1-2, 1998, pp. 51-58. doi:10.1016/S0378-1119(98)00217-0
- F. Martin-Laurent, D. van Tuinen, E. Dumas-Gaudot, V. Gianinazzi-Pearson, S. Gianinazzi and P. Franken, “Differential Display Analysis of RNA Accumulation in Arbuscular Mycorrhiza of Pea and Isolation of a Novel Symbiosis-Regulated Plant Gene,” Molecular and General Genetics, Vol. 256, No. 1, 1997, pp. 37-44. doi:10.1007/s004380050543
- D. Kordis and F. Gubensek, “Unusual Horizontal Transfer of a Long Interspersed Nuclear Element between Distant Vertebrate Classes,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 95, No. 18, 1998, pp. 10704-10709. doi:10.1073/pnas.95.18.10704
- A. Morton Walsh, R. D. Kortschak, M. G. Gardner, T. Bertozzi and D. L. Adelson, “Widespread Horizontal Transfer of Retrotransposons,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 110, No. 3, 2013, pp. 1012-1016. doi:10.1073/pnas.1205856110
- A. Roulin, B. Piegu, P. M. Fortune, et al., “Whole Genome Surveys of Rice, Maize and Sorghum Reveal Multiple Horizontal Transfers of the LTR-Retrotransposon Route 66 in Poaceae,” BMC Evolutionary Biology, Vol. 9, 2009, Article 58. doi:10.1186/1471-2148-9-58
- K. R. Oliver and W. K. Greene, “Transposable Elements: Powerful Facilitators of Evolution,” BioEssays, Vol. 31, No. 7, 2009, pp. 703-714. doi:10.1002/bies.200800219
- J. C. Silva, E. L. Loreto and J. B. Clark, “Factors That Affect the Horizontal Transfer of Transposable Elements,” Current Issues in Molecular Biology, Vol. 6, No. 1, 2004, pp. 57-71.
- M. G. Kidwell, “Lateral Transfer in Natural Populations of Eukaryotes,” Annual Review of Genetics, Vol. 27, 1993, pp. 235-256. doi:10.1146/annurev.genet.27.1.235
- P. J. Keeling and J. D. Palmer, “Horizontal Gene Transfer in Eukaryotic Evolution,” Nature Reviews Genetics, Vol. 9, No. 8, 2008, pp. 605-618. doi:10.1038/nrg2386
- M. Yoshiyama, Z. Tu, Y. Kainoh, H. Honda, T. Shono and K. Kimura, “Possible Horizontal Transfer of a Transposable Element from Host to Parasitoid,” Molecular Biology and Evolution, Vol. 18, No. 10, 2001, pp. 1952- 1958. doi:10.1093/oxfordjournals.molbev.a003735
- C. Gilbert, S. Schaack, J. K. Pace, P. J. Brindley and C. Feschotte, “A Role for Host-Parasite Interactions in the Horizontal Transfer of Transposons across Phyla,” Nature, Vol. 464, No. 7293, 2010, pp. 1347-1350. doi:10.1038/nature08939

