Advances in Microbiology
Vol.07 No.04(2017), Article ID:75654,15 pages
10.4236/aim.2017.74022
Isolation and Identification of Potential Probiotic Bacteria from Cattle Farm Soil in Dibrugarh District
Nuredin Mohamedkassm Siraj1*, Kaushal Sood2, Raj Narayan Singh Yadav3
1Asmara College of Health Sciences, Asmara, Eritrea
2Centre for Biotechnology and Bioinformatics, Dibrugarh University, Assam, India
3Department of Life Sciences, Dibrugarh University, Assam, India

Copyright © 2017 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY 4.0).
http://creativecommons.org/licenses/by/4.0/



Received: February 23, 2017; Accepted: April 22, 2017; Published: April 26, 2017
ABSTRACT
Several studies have been done to isolate probiotic bacteria from different sources. In this present study, an attempt was made to isolate, screen and identify potential probiotic bacteria from cattle farm soil in Dibrugarh district, Assam, India. At the level of screening, the result showed the isolates designated as DUA4, DUD3 and DUE2 showed a percent survival rate of 75.36, 69.14 and 52.36 respectively at a pH of 2.5. Similarly survival rate of the same isolates in 0.5% bile salt condition was found to be 117.17%, 144.59% and 118.10% for the isolates DUA4, DUD3 and DUE2 respectively. Antimicrobial activity of the isolates towards the indicator organisms tested showed that DUA4 inhibited gram positive organisms while DUD3 showed activity against both gram positive and gram negative bacteria. All the three isolates showed activity against L. monocytogenes. Autoaggregation ability of the isolates DUA4, DUD3 and DUE2 was found to be 44.15%, 54.11% and 9.42% respectively. The adhesion ability of the isolates DUD3, DUA4 and DUE2 to xylene was 61.78%, 45.37% and 14.83% respectively. Antimicrobial susceptibility test of the isolates showed that the isolates are in general sensitive to antibiotics tested. Phenotypic and genotypic analysis based on the 16S rRNA gene sequence of the isolates DUA4, DUD3 and DUE2 resulted in the identification and designation of the isolate DUA4 as Bacillus spp., DUD3 as Enterococcus faecium and DUE2 as Enterobacter sp. In conclusion, the study has indicated the possibility of isolating potential probiotic bacterial strains from cattle farm soil.
Keywords:
Probiotic, Soil, Cattle Farm, Enterococcus faecium, Bacillus spp.

1. Introduction
Probiotics possess a number of unique phenotypic and genotypic characteristics, which are also exploited for the purpose of their isolation, screening, identification and characterization. Some of the main criteria for the selection of probiotics include the ability of the organisms to resist gastric acidity and pancreatic juice; their antagonistic property against food spoilage and pathogenic organisms and their stability via adherence to epithelial cells and food matrix [1] . Moreover, synthesis of essential metabolites, maintenance of homeostasis in the immune system and most importantly their safe nature for application are also characteristics that enable them to deliver health benefit [2] .
Probiotic bacteria are widely distributed in nature and their most common habitats include fermented and unfermented food items (like milk and milk products, meat, cereals and vegetables), intestines of animals, soil, water bodies, manure and sewage and these are commonly used as sources for their isolation [3] [4] . Isolation of probiotics from fermented foods of different kinds was reported in many publications but there are scarce reports on the isolation of probiotics from soil specimens. Cattle farm soil could be one of the important sources for the recovery of probiotic organisms among the sources other than food.
In this study, soil sample from cattle farm area was selected as a source for obtaining probiotic bacteria with the idea of increasing the possibility of isolating probiotic organisms as these samples presumably contain probiotic organisms originating both from the soil and the gastro intestinal tract of cattle. The objective of this study was to isolate, screen and identify potential probiotic bacterial strains from soil samples collected from cattle farm areas in Dibrugarh district, Assam, India.
2. Methods
2.1. Sample Collection and Processing
In this study, a total of 60 cattle farm soil specimens were collected from sites located in six sample collection zones in Dibrugarh district designated as A-F. Approximately 100 gm of soil specimen was collected from each site from the soil 5cm below the surface. Equal weight (10 gm) of all the ten samples collected from the same zone was properly mixed and blended to obtain a homogenous soil samples representative of the respective zone. The stock soil samples so prepared was stored at −20˚C and used for the isolation probiotic bacteria
2.2. Isolation and Screening of Bacteria from Cattle Farm Soil
Isolation of soil microbes was done using spread plate technique on de Mann Rogossa Sharpe (MRS) agar following overnight enrichment on MRS broth and appropriate tenfold serial dilutions. The preparation was incubated at 36°C aerobically under static condition. All the isolates with apparently different morphotypes were sub-cultured into MRS agar and subjected for cellular and cultural morphological studies.
2.2.1. Acid and Bile Tolerance Tests
Preliminary selection of the potential probiotic bacteria was done by testing the isolates for their ability to grow on MRS agar adjusted to pH 5.5. Bacterial isolates that were resistant to such growth conditions were considered for acid and bile tolerance tests. A modified version of the previously developed method was used to test the ability of the isolates to survive in acidic condition of pH 2.5 for the specified period of 0, 1 and 2 hours [5] . The rate of survival of the different isolates upon exposure to acidic condition was done using viability testing following application of appropriate tenfold serial dilutions. Percentage of the viable cells was calculated in reference to the cell count at 0 hours of incubation. In this assay, L. casei ATCC 393 was used included for the sake of comparison.
In this study, previously described method was applied to test the bile salts tolerance ability of the isolates [6] . Freshly grown isolates were harvested and then subjected to MRS broth containing 0.5 % bile salts (HiMedia) and incubated for the duration of 0, 2 and 4 hours. Survival rate of the isolates was calculated using the same approach as that of acid tolerance test.
2.2.2. Antimicrobial Activity of Isolates
Five isolates selected based on their acid and bile tolerance were tested for their antimicrobial activity. Standard organisms, namely B. subtilis (ATCC 6051), E.coli (ATCC 25922), P.aeruginosa (MTCC 4673), S.epidermidis (MTCC 6810), L. monocytogenes (ATCC BAA-751), S. typhimurium (ATCC 49416), B. cereus (ATCC 11778) and S. aureus (MTCC 9542) were used as indicator organisms. Cell Free Culture Supernatant (CFCS) of the isolates neutralized using NaOH was used for testing antimicrobial activity of the isolates using well diffusion method as has been previously described [7] . Moreover, the culture supernatant was heated at 80 °C for 10 minutes to inactivate any available enzymes.
2.2.3. Autoaggregation and Hydrophobicity
Both autoaggregation and Microbial Adhesion to Hydrocarbon (MATH) was applied to the isolates DUA4, DUD3 and DUE2 which were selected based on their tolerance to acid and bile salt and antimicrobial activities. The reference strain L. casei ATCC 393 was included in the analysis for comparison sake. Cell suspension for autoaggregation test was prepared by mixing 0.2 ml of the bacterial suspension in Phosphate Buffer Solution (PBS) equivalent to 0.5 McFarland turbidity standard and 3.8 ml of PBS. Optical density reading at 600 nm was taken after incubation at room temperature for the specified time of 0, 1, 2, 3, 4 and 5 hours of incubation. Autoaggregation ability of the organisms was obtained by application of the following formula [8] :
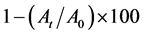
where At represents the absorbance at time t = 1, 2, 3, 4 or 5 hours and A0 is the absorbance at t = 0.
Microbial Adhesion to Hydrocarbons (MATH) test was applied to assess the hydrophobicity of the isolates. One ml of the cell suspension of the isolate in PBS equivalent to 1 McFarland turbidity standard was mixed with 3 ml of the hydrocarbon solvents namely Xylene, Chloroform and Ethyl acetate. Following proper mixing and incubation for 30 minutes, O.D. of the aqueous phase was measured at 600 nm and hydrophobicity of the isolates was calculated by applying the following formula [9] :
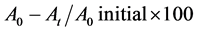
where A0, stands for the O.D. reading taken at the initial stage; At, an O.D. reading taken following phase separation time.
2.2.4. Antimicrobial Susceptibility of Isolates
Antimicrobial susceptibility test of the selected isolates was done using disc diffusion method [10] .
2.2.5. Phenotypic and Genotypic Study of Potential Probiotic Isolates
Phenotypic tests which include cellular, cultural, biochemical tests were performed using standard procedures for the identification of the bacterial isolates at different levels of the study. Molecular identification and phylogenetic analysis of the selected isolates was done based on the 16S rDNA gene sequence analysis. Genomic DNA isolation of the isolates was done by detergent lysis method [11] and Genomic DNA of the isolates were outsourced to Xcelris Labs Pvt. Ltd. (Ahmedabad, India) for 16S rDNA gene sequencing. Amplification of the isolated fragment of 16S rDNA gene was done by PCR using primers 8F and 1492R. Sequencing of the amplicon was carried out by automated genetic analyzer (BDT v3.1 Cycle sequencing kit on ABI 3730xl) using 704F and 907R primers. Sequence relatedness of the 16S rDNA gene with other organisms was done by the application of BLASTn alignment search tool on National Centre for Biotechnology Information (NCBI) genbank database. Phylogenetic analyses were conducted by using MEGA6 software and the evolutionary distances were computed using the Kimura 2-parameter method [12] [13] [14] . The evolutionary history was inferred using the Neighbor-Joining method [15] .
3. Results
3.1. Primary Isolation of the Potential Probiotic Bacteria
During primary isolation, based on gram staining and morphological study, 26 of the isolates were found to be bacteria and the remaining 15 were detected as yeast cells. Among the bacterial isolates 21 were gram-positive while the remaining 5 were gram-negative. All bacterial isolates (n = 26) were then considered for further screening, identification and characterization for probiotic characteristics.
3.2. Acid and Bile Tolerance Tests
Result for acid tolerance test of the probiotic candidate isolates tested in this study is given in Table 1.
Table 1. Acid tolerance of potential probiotic isolates obtained from cattle farm soil in Dibrugarh district.
The result shows that out of eleven organisms tested only five isolates were found to survive acidic condition of pH 2.5. The level of survival in acidic condition of these isolates namely DUA4, DUC1, DUD3, DUE2 and DUF3 expressed in terms of per cent was found to be 75.36%, 46.72%, 69.14%, 52.36% and 40.76 % respectively. The reference strains L. casei ATCC 393 showed a survival rate of 66.64%.
Bile tolerance test was conducted only on isolates namely DUA4, DUD3, DUC1, DUE2 and DUF3 and the result is shown in Table 2. All the isolates tested with the exception of DUF3 showed an increase in their population size showing a survival rate of 117.17%, 116.72%, 144.59%, 118.10% and 94.98% for the isolates DUA4, DUC1, DUD3, DUE2 and DUF3 respectively when treated with 0.5% bile salts.
3.3. Antimicrobial Activity of Potential Isolates
Isolates with the characteristic feature of acid and bile tolerance were selected for antimicrobial activity test and the results are given in Table 3. Isolates DUA4, DUD3 and DUE2 showed inhibitory activities against the tested indicator organisms with different degrees of activities while no activity was observed with the isolates DUC1 and DUF3. In general DUA4 and DUD3 showed antimicrobial activity against most of the indicators organisms tested. However, difference was observed in the spectrum of activities of the isolates and as such DUD3 has shown broad spectrum of activity while DUA4 has shown activities against gram positive indicator organisms only.
Table 2. Bile tolerance of potential probiotic isolates obtained from cattle farm soil in Dibrugarh district.
Table 3. Antimicrobial activity of potential probiotic isolates obtained from cattle farm soil in Dibrugarh district.
3.4. Antimicrobial Susceptibility Test of Potential Probiotic Isolates
Antimicrobial Susceptibility Testing (AST) of the selected three isolates namely E. faecium DUD3, Bacillus sp DUA4 and Enterobacter sp DUE2 showed that the isolates were sensitive to the antimicrobial discs tested. However, E. faecium DUD3 showed intermediate level of sensitivity to ampicillin and tetracycline (Table 4).
Table 4. Antimicrobial susceptibility of the potential probiotic isolates.
3.5. Auto Aggregation and Microbial Adhesion to Hydrocarbon
Results for the autoaggregation test done in this study are given in Figure 1. The aggregation ability of the isolates tested after 5 hours of incubation was 44.15%, 54.11%, 33.55% and 9.42% for the isolates DUA4, DUD3, DUE2 and L. casei ATCC 393 respectively.
Result for the hydrophobicity assay of the isolates using the hydrocarbon compounds xylene, chloroform and ethyl acetate has shown contrasting values and the results are given in Table 5. Microbial adhesion to hydrocarbon using xylene showed adhesion values of 45.37%, 61.78%, 14.83% and 5.89% for the isolates DUA4, DUD3, DUE2 and L. casei ATCC 393 respectively. Results of Adhesion assay using chloroform showed percent adhesion values of 24.02%, 37.43%, 21.12% and 41.21% for the isolates DUA4, DUD3, DUE2 and L. casei ATCC 393 respectively. As for the adhesion assay using Ethyl acetate, no considerable adhesion ability was observed with all the four bacteria tested showing adhesion value of only 7.35%, 3.97%, 6.62% and 13.73% for the isolates DUA4, DUD3, DUE2 and L. casei ATCC 393 respectively.
3.6. Phenotypic and Genotypic Study of Isolates
Results of the selected isolates for the phenotypic analysis showed that DUA4 was gram positive spore forming bacillus giving positive results for oxidase, catalase and nitrate reductase tests. Moreover, results for sugar fermentation tests and other biochemical test show the relatedness of the isolate to Bacillus spp. (data not shown). The isolate DUD3 on the other side, was found to be gram positive cocci with negative results for catalase, oxidase and nitrate reductase tests and with the overall biochemical test results resembling E. faecium (data not shown). In a similar way application of the phenotypic tests applied on DUE2 also showed results that match with Enterobacter species.
Genotypic confirmation of the identification was done using 16S rDNA gene sequence analysis. Based on the maximum scores obtained by application of BLAST analysis on the 16S rDNA gene sequence of the isolates DUA4, DUD3
Figure 1. Autoaggregation of potential probiotic isolates obtained from cattle farm soil in Dibrugarh District.
Table 5. Microbial adhesion to Hydrocarbons of potential probiotic isolates obtained from cattle farm soil in Dibrugarh district.
and DUE2 using NCBI genbank database, the isolates were identified as Bacillus sp. DUA4 (maximum identification of 97% with B. cereus ATCC 14579 with query coverage of 94%), Enterococcus faecium DUD3 (maximum identification of 99% with E. faecium strain DSM 20477 with query coverage of 99%) and Enterobacter spp DUE2 (maximum identification of 97% with Enterobacter cancerogenus strain LMG with query coverage of 94%) respectively. The 16S rDNA gene sequence of the isolates was submitted to NCBI and accession number KT340076, KT340075 and KX097968 was obtained for the isolates Bacillus sp. DUA4, Enterococcus faecium DUD3 and Enterobacter spp DUE2 respectively. The finding was in agreement with the results obtained using phenotypic tests applied. Phylogenetic tree was constructed using MEGA6 program and the output for the two potential isolates namely Bacillus sp. DUA4 and Enterococcus faecium DUD3 is given in Figure 2, A and B respectively.
4. Discussion
A total of 41 organisms with apparently different morphotype were isolated out of which 15 were detected as yeast cells and the remaining were gram-positive (n = 11) and gram-negative (n = 16). Considerably high number of yeast cells were isolated during the primary isolation.The soil of Dibrugarh is acidic with documented pH value in the range of 4.6 to 5.57 [16] . Therefore, considering the fact that low pH favours the growth of yeasts, the isolation of yeasts in considerably


Figure 2. (a) and (b). Molecular Phylogenetic analysis of the isolate DUA4 (a) and DUD3 (b) with L. casei ATCC 393 as an out group. Phylogenetic analysis of the selected probiotic isolates was done using MEGA 6 software. Inference of the evolutionary history was done using the Maximum Likelihood method based on the Tamura et al., model [14] . The phylogenetic tree is drawn to scale with the length of the branch arm representing the number of base substitution and the values at the node indicating bootsrap values based on 1000 replications.
high number is justifiable [17] .
In this study, acid tolerance of DUD3 (Enterococcus faecium DUD3) was found to be relatively lower than that of DUA4 (Bacillus sp. DUA4) and this was found to be in agreement with the findings from other studies [18] . Bacillus spp. has shown excellent level of resistance to acidic pH value of 2.5% and 0.5% bile and one such previously conducted study indicated that all the 13 Bacillus isolates studied showed resistance to both acid and bile salts [19] [19] . In this study the isolate DUA4 has been found to form endospore which may increase the ability of tolerance to acid and bile salts. However, there is conflicting report about the role of spore formation and resistance to acid and bile. Most of the reports indicate that acid tolerance increases with the formation of spores while some others report that there is no difference in the susceptibility of the spores and vegetative forms to acid and bile salts [20] [21] .
In this study, higher resistance to acid and bile salts was in general shown by the isolates DUA4 (Bacillus sp. DUA4) and DUD3 (Enterococcus faecium DUD3). Both acid and bile tolerance characteristics of probiotic bacteria are required for their viability in the food, for the passage through the stomach and survival in the small intestine [22] [23] . Though the level of acid resistance of probiotic cultures is expected to be in the pH range of 2.5 - 3.0, the combination of the gastric acid with the food materials in the stomach provides more protection increasing their ability to survive in the gut [24] [25] [26] [27] . Moreover, acid tolerance of probiotic bacteria is known to depend on the type of organisms tested, growth medium and the incubation conditions and thus meaningful comparison could be done in consideration to these variables [28] . Different concentration of bile salt has been used for the same assay and accordingly, concentration of 0.3 % was found to be discriminatory and is also recommended for the screening of probiotic isolates [29] [30] [31] .
The result of antimicrobial activity test of CFCS of the Isolates Bacillus sp DUA4 (DUA4) and E. faecium DUD3 (DUD3) showed broad spectrum activity. The presence of strains of bacteriocinogenic enterococci that are capable of showing broad spectrum antimicrobial activity has also been reported in other studies [32] . Moreover, antimicrobial activity of bacteria could be attributed to a number of factors other than bacteriocin. In this study in order to rule out the antagonistic effect of acidity of the culture supernatant, neutralization of CFCS was done using NaOH. All the three isolates tested in this study showed activity against L. monocytogenes, food spoilage bacteria which is relevant in the food and dairy industries [33] [34] .
In this study autoaggregation and MATH test was used to assess the adhesion capacity of the isolates. The ideal method for testing adhesion capacity of organisms is the application of in vitro adhesion assay using cell lines. Autoaggregation and cell hydrophobicity tests which were designed as alternative tests to adhesion assay were proven as reasonably reliable screening method when compared to the use of cell lines which is technically and economically demanding [8] [35] [36] . In this study, the aggregation ability of the isolates tested was found to be 44.15%, 54.11%, 9.42% and 33.55% for the isolates DUA4, DUD3, DUE2 and L. casei ATCC 393 respectively. In a similar study, an autoaggregation value of 40.2% was reported for potential probiotic Bacillus sp. Isolated from human faeces [37] .
Result for the hydrophobicity assay of the isolates using the hydrocarbon compounds xylene, chloroform and ethyl acetate in the current study has shown contrasting values for the isolates tested. In performing hydrophobicity test either xylene or n-hexadecane has been used in different researches but no significant difference in result was observed with the two solvents [38] . The MATH like other probiotic traits is strain specific. Analysis of 8 strains of bacillus species for MATH has shown a variation of results with a hydrophobicity value ranging from 14.1% to 83.6% [39] . In the absence of standard guideline for the interpretation of hydrophobicity assay, hydrophobicity value of more than 40% is considered to be significant value for hydrophobic nature [40] . A hydrophobicity value of 58%, 82% and 12.4% was obtained for Bacillus subtilis JK-04 isolated from human faeces using hydrocarbons xylene, ethyl acetate and chloroform respectively [37] .
The antimicrobial susceptibility study of the isolates in this study showed that all the isolates showed sensitivity to the antibiotics tested except E. faecium DUD3 against ampicillin and tetracycline.However, it is also important to consider the transferability of the resistant trait to other microbes. Among the different mechanisms of gene exchange between bacteria, conjugation which involves the transfer of mobile genetic elements such as plasmids is the most common mechanism for the horizontal transfer of antibiotic resistance gene [41] . Enterococci are also known to be very well receptive for conjugation [42] , but are also successful donor organisms for the transfer of antibiotic resistance genes to unrelated enterococci, lactobacilli, other Gram-positives including Bacillus subtilis [43] [44] [45] .
The resolution power of the application of the microscopic, growth characteristics and biochemical tests for the identification of the potential probiotic bacteria was not adequate. Therefore, molecular methods based on 16S rDNA sequence were used as a supplementary test for the identification of the potential probiotic isolates. Moreover, Consensus sequence of the 16S rDNA gene sequence was also used to construct phylogenetic tree of the isolates. Construction of phylogenetic tree based on 16S rDNA gene sequence has been found to be reasonably reliable in comparison to trees constructed using whole genome sequences [46] .
5. Conclusion
Three bacterial isolates with different level of probiotic capacity were identified and E. faecium DUD3 was found to be the most potential of all. From this study, it can be concluded that there is a potential to isolate probiotic bacteria from cattle farm soil.
Acknowledgements
Authors would like to acknowledge the support received from the Indian Council for Cultural Relations and the Government of Eritrea. Authors would also like to acknowledge the Department of Biotechnology, Government of India and Dibrugarh University for providing the essential facilities for research at the Centre for Biotechnology and Bioinformatics. Authors also thank Debajit Borah, Assistant professor in Centre for Biotechnology and Bioinformatics in Dibrugarh University, for his assistance during the study.
Cite this paper
Siraj, N.M., Sood, K. and Yadav, R.N.S. (2017) Isolation and Identification of Potential Probiotic Bacteria from Cattle Farm Soil in Dibrugarh Dis- trict. Advances in Microbiology, 7, 265- 279. https://doi.org/10.4236/aim.2017.74022
References
- 1. Havenaar, R. and Huis in't Veld, J.H.J. (1992) Probiotics; A General Review' in the Lactic Acid Bacteria in Health and Disease. In: Wood, B., Ed., Elsevier, London, 151 ± 170.
http://dx.doi.org/10.1007/978-1-4615-3522-5_6 - 2. Fuller, R. (1989) Probiotics in Man and Animals. Journal of Applied Microbiology, 66, 365-378.
http://dx.doi.org/10.1111/j.1365-2672.1989.tb05105.x - 3. Holzapfel, W.H., Haberer, P., Geisen, R., Bjorkroth, J. and Schillinger, U. (2001) Taxonomy and Important Features of Probiotic Microorganisms in food and Nutrition. American Journal of Clinical Nutrition. 73, 365-373.
www.ncbi.nlm.nih.gov/pubmed/11157343 - 4. Caplice, E. and Fitzgerald, G.F. (1999) Food Fermentation: Role of Microorganisms in Food Production and Preservation. International Journal of Food Microbiology, 50, 131-149.
http://dx.doi.org/10.1016/s0168-1605(99)00082-3 - 5. Pereira, D.I.A. and Gibson, G.R. (2002) Cholesterol Assimilation by Lactic Acid Bacteria and Bifidobacteria Isolated from the Human Gut. Applied and Environmental Microbiology, 68, 4689-4693.
http://dx.doi.org/10.1128/aem.68.9.4689-4693.2002 - 6. Klingberg, T.D., Axelsson, L., Naterstad, K., Elsser, D. and Budde, B.B. (2005) Identification of Potential Probiotic Starter Cultures for Scandinavian-Type Fermented Sausaes. International Journal of Food Microbiology, 105, 419-431.
http://dx.doi.org/10.1016/j.ijfoodmicro.2005.03.020 - 7. Mishra, V. and Prasad, D. (2005) Application of in Vitro Methods for Selection of Lactobacillus Casei Strains as Potential Probiotics. International Journal of Food Microbiology, 103, 109-115.
http://dx.doi.org/10.1016/j.ijfoodmicro.2004.10.047 - 8. Del Re, B., Sgorbati, B., Miglioli, M. and Palenzona, D. (2000) Adhesion, Autoaggregation and Hydrophobicity of 13 Strains of Bifidobacterium Longum. Letters in Applied Microbiology, 31, 438-442.
http://dx.doi.org/10.1046/j.1365-2672.2000.00845.x - 9. Rosenberg, M., Gutnik, D. and Rosenberg, E. (1980) Adherence of Bacteria to Hydrocarbons: A Simple Method for Measuring Cell-Surface Hydrophobicity. FEMS Microbiology Letters, 9, 29-33.
http://dx.doi.org/10.1111/j.1574-6968.1980.tb05599.x - 10. CLSI (Clinical and Laboratory Standards Institute) (2007) Performance Standards for Antimicrobial Susceptibility Testing; Seventeenth Informational Supplement. Document M100-S17, 27, 52-55.
http://dx.doi.org/10.1201/9781420014495.ch1 - 11. Harisha, S. (2007) Biotechnology Procedures and Experiments Handbook. Bio-technology Laboratory Manuals.
- 12. Felsenstein, J. (1985) Phylogenies and the Comparative Method. The American Naturalist, 125, 1-15.
https://doi.org/10.1086/284325 - 13. Kimura, M. (1980) A Simple Method for Estimating Evolutionary Rates of Base Substitution through Comparative Studies of Nucleotide Sequences. Journal of Molecular Evolution, 16, 111-120.
https://doi.org/10.1007/BF01731581 - 14. Tamura, K., Stecher, G., Peterson, D., Filipski, A. and Kumar, S. (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Molecular Biology and Evolution, 30, 2725-2729.
https://doi.org/10.1093/molbev/mst197 - 15. Saitou, N. and Nei, M. (1987) The Neighbor-Joining Method: A New Method for Reconstructing Phylogenetic Trees. Molecular Biology and Evolution, 4, 406-425.
- 16. Nath, T.N. (2014) Soil Bulk Density and Its Impact on Soil Texture, Organic Mattter Content and Available Macronutrients Acronutrients of Tea Cultivated Soil in Dibrugarh District of Assam, India. International Journal of Development Research, 4, 343-346.
http://www.journalijdr.com - 17. Phaff, H.J., Miller, M.W. and Mrak, E.M. (1966) Ecology. Life of Yeasts: Their Nature, Activity, Ecology and Relation to Mankind. Harvard University Press, Cambridge.
- 18. Kabore, D., Sawadogo-Lingani, H., Dicko, M.H., Diawara, B. and Jakobsen, M. (2012) Acid Resistance, Bile Tolerance and Antimicrobial Properties of Dominant Lactic Acid Bacteria Isolated from Traditional “Maari” Baobab Seeds Fermented Condiment. African Journal of Biotechnology, 11, 1197-1206.
https://doi.org/10.5897/AJB11.2667 - 19. Compaoré, C.S., Jensen, L.B., Diawara, B., Ouédraogo, G.A., Jakobsen, M. and Ouoba, L. (2013) Resistance to Antimicrobials and Acid and Bile Tolerance of Bacillus spp. Isolated from Bikalga, Fermented Seeds of Hibiscus sabdariffa. African Journal of Food Science, 7, 408-414.
https://doi.org/10.5897/AJFS2013.1018 - 20. Duc, L.H., Hong, H.A., Barbosa, T.M., Henriques, A.O. and Cutting, S.M. (2004) Characterization of Bacillus Probiotics Available for Human Use. Applied and Environmental Microbiology, 70, 2161-2171.
https://doi.org/10.1128/AEM.70.4.2161-2171.2004 - 21. Guo, X., Li, D., Lu, W., Piao, X. and Chen, X. (2006) Screening of Bacillus Strains as Potential Probiotics and Subsequent Confirmation of the in Vivo Effectiveness of Bacillus subtilis MA139 in Pigs. Antonie van Leeuwenhoek, 90, 139-146.
https://doi.org/10.1007/s10482-006-9067-9 - 22. Lee, Y.K. and Salminen, S. (1995) The Coming of Age of Probiotics. Trends in Food Science & Technology, 6, 241-245.
- 23. Hemiksson, B.R., Wageinus, P., Franzen, G.L. and Lewin, F. (1999) Aspects of Reducing Gastrointestinal Adverse Effects Associated with Radiotherapy. Acta Oncologica, 38, 159-164.
https://doi.org/10.1080/028418699431564 - 24. Charteris, W.P., Kelly, P.M., Morelli, L. and Collins, J.K. (1998) Development and Application of an in Vivo Methodology to Determine the Transit Tolerance of Potentially Probiotic Lactobacillus and Bifidobacterium Species in the Upper Human Gastrointestinal Tract. Journal of Applied Microbiology, 84, 759-768.
https://doi.org/10.1046/j.1365-2672.1998.00407.x - 25. Wang, X., Brown, I.L., Evans, A.J., Conway, P.L. (1999) The Protective Effects of High Amylose Maize (Amylomaize) Starch Granules on the Survival of Bifidobacterium spp. in the Mouse Intestinal Tract. Journal of Applied Microbiology, 87, 631-639.
https://doi.org/10.1046/j.1365-2672.1999.00836.x - 26. Zárate, G., Chaia, A.P., González, S. and Oliver G. (2000) Viability and Beta-Galactosidase Activity of Dairy Propionibacteria Subjected to Digestion by Artificial Gastric and Intestinal Fluids. Journal of Food Protection, 63, 1214-1221.
https://doi.org/10.4315/0362-028X-63.9.1214 - 27. Huang, Y. and Adams, M.C. (2004) In Vitro Assessment of the Upper Gastrointestinal Tolerance of Potential Probiotic Dairy Propionibacteria. International Journal of Food Microbiology, 91, 253-260.
- 28. Hood, S.K. and Zoitola, E.A. (1988) Effect of Low pH on the Ability of Lactobacillus acidophilus to Survive and Adhere to Human Intestinal Cells. Journal of Food Science, 53, 1514-1516.
https://doi.org/10.1111/j.1365-2621.1988.tb09312.x - 29. Goldin, B.R., Gorbach, S.L., Saxelin, M., Barakat, S., Gualtieri, L. and Salminen, S. (1992) Survival of Lactobacillus Species (Strain GG) in Human Gastrointestinal Tract. Digestive Diseases and Sciences, 37, 121-128.
https://doi.org/10.1007/BF01308354 - 30. Chateau, N., Deschamps, A.M and Sassi, A.H (1994) Heterogeneity of Bile Salts Resistance in the Lactobacillus Isolates of a Probiotic Consortium. Letters in Applied Microbiology, 18, 42-44.
https://doi.org/10.1111/j.1472-765X.1994.tb00796.x - 31. Papamanoli, E., Tzanetakis, N., Litopoulou-Tzanetaki, E. and Kotzekidou, P. (2003) Characterization of Lactic Acid Bacteria Isolated from a Greek Dry-Fermented Sausage in Respect of Their Technological and Probiotic Properties. Meat Science, 65, 859-867.
- 32. Belguesmia, Y., Choiset, Y., Prévost, H., Dalgalarrondo, M., Chobert, J.M. and Drider, D. (2010) Partial Purification and Characterization of the Mode of Action of Enterocin S37: A Bacteriocin Produced by Enterococcus faecalis S37 Isolated from Poultry Feces. Journal of Environmental and Public Health, 2010, Article ID: 986460.
https://doi.org/10.1155/2010/986460 - 33. Rodriguez, E.G., Gaya, B.P., Nanez, M. and Medina, M. (2000) Diversity of Bacteriocins Produced by Lactic Acid Bacteria Isolated from Raw Milk. International Dairy Journal, 10, 7-15.
- 34. De Vuyst, L., Foulquie Moreno, M.R. and Revets, H. (2003) Screening for Enterocins and Detection of Hemolysin and Vancomycin Resistance in Enterococci of Different Origins. International Journal of Food Microbiology, 84, 299-318.
- 35. Bernet, M.F., Brassart, D., Neeser, J.R. and Servin, A.L. (1993) Adhesion of Human Bifidobacterial Strains to Cultured Human Intestinal Epithelial Cells and Inhibition of Enteropathogen Cell Interactions. Applied and Environmental Microbiology, 59, 4121-4128.
- 36. Crociani, J., Grill, J.P., Huppert, M. and Ballongue, J. (1995) Adhesion of Different Bifidobacteria Strains to Human Entero-Cyte-Like Caco-2 Cells and Comparison with in Vivo Study. Letters in Applied Microbiology, 21, 146-148.
https://doi.org/10.1111/j.1472-765X.1995.tb01027.x - 37. Naidu, K.S., Govender, B.P. and Adam, J.K. (2014) Identification and Characterization of Bacillus sp. for Probiotic Properties Isolated from Human Faeces. Journal of Pure and Applied Microbiology, 8, 2855-2862.
- 38. Rao, K.P., Chennappa, G., Suraj, U., Nagaraja, H., Raj, A.C. and Sreenivasa, M.Y. (2015) Probiotic Potential of Lactobacillus Strains Isolated from Sorghum-Based Traditional Fermented Food. Probiotics and Antimicrobial Proteins, 7, 146-156.
https://doi.org/10.1007/s12602-015-9186-6 - 39. Simmonds, P., Mossel, B.L., Intaraphan, T. and Deeth H.C. (2003) Heat Resistance of Bacillus Spores When Adhered to Stainless Steel and Its Relationship to Spore Hydrophobicity. Journal of Food Protection, 66, 2070-2075.
https://doi.org/10.4315/0362-028X-66.11.2070 - 40. Boris, S., Sua′rez, J.E., Va′zquez, F. and Barbe`s, C. (1998) Adherence of Human Vaginal Lactobacilli to Vaginal Epithelial Cells and Interaction with Uropathogens. Infection and Immunity, 66, 1985-1989. www.ncbi.nlm.nih.gov/pubmed/9573080
- 41. Salyers, A.A. (1995) Antibiotic Resistance Transfer in the Mammalian Intestinal Tract: Implications for Human Health. Food Safety and Biotechnology, Springer-Verlag, New York, Heidelberg.
- 42. Clewell, D.B. and Weaver, K.E. (1989) Sex Pheromones and Plasmid Transfer in Enterococcus faecalis. Plasmid, 21, 175-184.
- 43. Christie, P.J., Korman, R.Z., Zahler, S.A., Adsit, J.C. and Dunny, G.M. (1987) Two Conjugation Systems Associated with Streptococcus faecalis Plasmid pCF10: Identification of a Conjugative Transposon That Transfers between S. facealis and Bacillus subtilis. Journal of Bacteriology, 169, 2529-2536.
https://doi.org/10.1128/jb.169.6.2529-2536.1987 - 44. Rice, L.B., Carias, L.L., Donskey, C.L. and Rudin, S.D. (1998) Transferable, Plasmid-Mediated VanB-Type Glycopeptide Resistance in Enterococcus faecium. Antimicrobial Agents and Chemotherapy, 42, 963-964.
- 45. Shrago, A.W. and Dobrogosz, W.J. (1988) Conjugal Transfer of Group-B Streptococcal Plasmids and Comobilization of Escherichia coli-Streptococcus Shuttle Plasmids to Lactobacillus plantarum. Applied and Environmental Microbiology, 54, 824-826.
- 46. Bansal, A.K. and Meyer, T.E. (2002) Evolutionary Analysis by Whole-Genome Comparisons. Journal of Bacteriology, 184, 2260-2272.
https://doi.org/10.1128/JB.184.8.2260-2272.2002



