Advances in Biological Chemistry
Vol.3 No.2(2013), Article ID:30770,11 pages DOI:10.4236/abc.2013.32024
Molecular cloning and characterization of human age-related NADH oxidase (arNOX) proteins as members of the TM9 superfamily of transmembrane proteins
![]()
Mor-NuCo, Inc., Purdue Research Park, West Lafayette, USA
Email: *dj_morre@yahoo.com
Copyright © 2013 Xiaoyu Tang. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
Received 11 January 2013; revised 9 March 2013; accepted 12 April 2013
Keywords: Age-Related NADH Oxidase (arNOX), TM-9 Superfamily of Transmembrane Proteins; Molecular Cloning; Human Serum; Plasma and Body Fluids; Saliva
ABSTRACT
Age-related NADH oxidase (arNOX) proteins are superoxide-generating cell surface oxidases that increase in activity with age beginning at about 30 y. A soluble and truncated exfoliated form of the activity is present in blood and other body fluids. The activity was purified to apparent homogeneity from human urine and resolved by 2-D gel electrophoresis into a series of 24 to 32 kDa components of low isoelectric point. The purified proteins were resistant both to N-terminal sequencing and trypsin cleavage. Cleavage with pepsin revealed peptides corresponding to the TM9 family of transmembrane proteins. Peptide antisera raised to all five members of the human TM9 family sequentially blocked the arNOX activity of human saliva and sera. The soluble truncated Nterminus of the human homolog TM9SF4 was expressed in bacteria. The recombinant protein was characterized biochemically and exhibited arNOX activity. The findings identify five arNOX isoforms each of which correspond to one of the five known TM9 family members. The exfoliated soluble arNOX forms are derived from the 24 to 32 kDa N-termini exposed to the cell’s exterior at the cell surface. Each of the shed forms contain putative functional motifs characteristic of ECTO-NOX (ENOX) proteins despite only minimal sequence identity. Our findings identify arNOX as having functional characteristics of ENOX proteins and the TM9 superfamily of proteins as the genetic origins of the five known arNOX isoforms present in human sera, plasma and other body fluids.
1. INTRODUCTION
Our work has identified a family of superoxide-generating oxidases that increases in activity with age beginning at about age 30 [1]. Shed forms of the activity are present in blood and other body fluids where they have been shown to participate in a variety of action-at-a-distance phenomena associated with several age-related disorders such as skin aging [2] and coronary artery disease [3,4].
Cell associated arNOX is membrane anchored with its catalytic terminus directed toward the cells’ exterior [1]. A ca. 30 kDa fragment is shed and enters the blood and other body fluids or is internalized into endosomes [5]. The soluble fragments also generate superoxide as estimated from the superoxide dismutase-inhibited reduction of ferricytochrome c, a standard measure of superoxide production [6]. At peak production, which occurs on average at about age 65, approximately 0.06 nmoles superoxide are produced/107 cells (human buffy coats or epidermal epithelia). The value in sera is approximately 0.3 nmoles/ml/min which results in several millimoles of superoxide being produced on a daily basis in the proximity of circulating lipoproteins or matrix proteins of the skin just from the circulating form.
In this report, we identify the five circulating human arNOX isoforms as exfoliated N-termini of the five members of the TM9 family of transmembrane proteins (GenBank NP_006396, NP_001014842, NP_004791, NP_064508, NP_055557). The identification is based on cloned TM9SF4 and TM9SF2, two of the superfamily members and their expression in bacteria and inhibition by specific peptide antisera for the three additional superfamily members, TM9SF1a, TM9SF1b and TM9SF3.
2. METHODS
2.1. Materials
pET11b vector and BL21 (DE3) competent cells were purchased from Novagen (Madison, WI). I.M.A.G.E. Full length cDNA IRAUp969E0589D was from ima-Genes (Berlin, Germany).
2.2. Expression of Recombinant TM9SF4 and TM9SF2 (Full Length or M1 to E258)
Plasmids carrying TM9SF4 sequence were prepared by inserting the TM9SF4 sequence into the pET11b vector (between NheI and BamHI sites). The TM9SF4 sequence was amplified from cloned I.M.A.G.E. full length cDNA IRAUp969E9580D by PCR. The primers used were 5’-GATATACATATGGCTAGCATGGCGACGGCGATGGAT-3’ (forward) and 5’-TTGTTAGCAGCCGGATCCTCAGTCTATCTTCACAGC-3’ (reverse). The PCR products then were doubly digested with NheI and BamHI and were ligated into pET11b vector.
DNA sequences of the ligation products (pET11bTM9SF4) were confirmed by DNA sequencing. Then pET11b-TM9SF4 was transformed into BL21 (DE3) competent cells. A single colony was picked and inoculated into the 5 ml LB/AMP medium. The overnight culture (1 ml) was diluted into 100 ml LB/AMP media (1:100 dilution). The cells were grown with vigorous shaking (250 rpm) at 37˚C to an OD600 of 0.4 - 0.6 and IPTG (0.5 mM) was added for induction. Cultures were collected after 5 h shaking (250 rpm) at 37˚C. Expression of the TM9SF4 was confirmed by SDS-PAGE with silver staining.
Cultures were centrifuged at 6000 g for 20 min. Pellets were resuspended in 20 mM Tris-HCl, pH 8.0 (0.5 mM PMSF added). Cells were broken by French Press at 20,000 psi 3 times.
Plasmids carrying the TM9SF2 (S33-C250 of TM9SF2 with 6× His-tag at the N-terminus) sequence were prepared by inserting the TM9SF2 sequence into the pET11b vector (between NheI and BamHI sites). The TM9SF2 sequence was synthesized by GenScript USA, Inc. (Piscataway, NJ). DNA sequences of the ligation products (pET11b-TM9SF2) were confirmed by DNA sequencing. Expression of recombinant TM9SF2 was the same as that for TM9SF4.
2.3. Purification of Human arNOX and Peptide Analysis
Exfoliated arNOX was purified to apparent homogeneity from a 20% ammonium sulfate fraction from human urine (72-y male) followed by dialysis, YM-10 filtration and preparative SDS-PAGE. For sequencing, the sample was further purified on a Capto Q Imp Res HPLC column and a major fraction collected at 8 min with arNOX activity cleaved with 20 ng/µl trypsin, pH 7.8 or with 20 ng/µl pepsin, pH 3 (with HCl) at 37˚C for 2 h. For trypsin digestion, the sample was first suspended in 50 ml Tris-HCl, pH 8 containing 5 mM DTT and 8 M urea at 60˚C for 1 h. After cooling, 50 mM NH4HCO3 was added to reduce the urea concentration to 1 M before adding trypsin. Peptides were identified by Fourier Transform Ion Cyclotron Resonance Mass Spectrometry.
2.4. Cell Growth
Stock E. coli cells [BL21 (DE3)] containing plasmid pET11b-TM9SF4/-TM9SF2 were stored at −80˚C in glycerol stock solution. Small amount of cells were picked by using a sterile yellow pipette tip and added to 1 ml of LB medium containing ampicillin (100 µg/ml) (LB/amp medium) and mixed well. Twenty µl of the cell resuspension was plated on LB/amp. Cells were grown for more than 20 h until well-rounded colonies were formed. One colony was picked and mixed in 10 ml of LB/amp medium (inoculum). Cells were grown for 8 h and stored at 4˚C until use. The inoculum was centrifuged at 6000 rpm for 6 min. Supernatant was discarded and pellet was resuspended in 4 ml of LB/amp medium. Aliquots of 100 ml of LB/amp media were prepared in four 250 ml—Erlenmeyer flasks. One ml of cell resuspension was added to each flask and cells were grown for 8 h with shaking. IPTG (0.5 mM) was added for induction when the OD600 reached 0.4 to 0.6.
2.5. Cell Harvest and Protein Extraction
Cells were harvested by centrifugation at 6000 rpm for 6 min, washed with 20 mM Tris-HCI, pH 8.0 and resuspended in 20 mM Tris-HCI, pH 8.0 containing protease inhibitors (0.75 mM PMSF, 3 mM 6-aminocaproic acid and 1.5 mM benzamidine HCl).
Proteins were extracted by French Press (3 passages at 20,000 psi). The extracts were centrifuged at 10,000 rpm for 20 min. Supernatant was discarded and pellets (inclusion bodies) were resuspended in 20 ml of Tris buffer. Two ml of 20% Triton X-100 was added to each tube and sample volume was adjusted to 40 ml with Tris buffer. Tubes were incubated at room temperature for 1 h with shaking and centrifuged at 10,000 rpm for 20 min. Supernatants were discarded and pellets were washed two times with Tris buffer by resuspending in 25 ml of Tris buffer and centrifuged one time with 25 ml of deionized water.
2.6. Solubilization of the Inclusion Bodies
Pellets were resuspended in 20 ml of water and 4 ml of 0.5 M CAPS buffer, pH 11 (50 mM final concentration), 40 µl of 1 M DTT (1 mM final concentration) and 0.4 ml of 30% sodium lauroyl sarcosine (0.3 % final concentration) were added. Sample volumes were adjusted to 40 ml with water. Samples were incubated at room temperature for 17 h.
2.7. Refolding of Recombinant TM9SF4/TM9SF2
After solubilization, samples were centrifuged at 10,000 rpm for 20 min and supernatants were collected. The supernatants were filtered through a 0.45 µm syringe filter. The filtrates were poured into two dialysis bags (3500 MWCO, flat width 45 mm and diameter 29 mm, Spectra Por) and dialyzed against cold dialysis buffer A (25 mM Tris-HCl, pH 8.5, 1 mM cysteamine, 0.1 mM cystamine, 1 mM 6-aminocaproic acid and 0.5 mM benzamidine HCl) with 3 changes, against cold dialysis buffer B (25 mM Tris-HCl, pH 8.0, 1 mM 6-aminocaproic acid and 0.5 mM benzamidine HCl) with one change and against dialysis buffer C (50 mM Tris-HCl, pH 8.0, 1 mM 6-aminocaproic acid and 0.5 mM benzamidine HCl) with one change. Dialysis was at least 17 h for each change. After dialysis, PMSF was added to a final concentration of 0.5 mM and samples were centrifuged at 10,000 rpm for 20 min. Supernatant was collected and concentrated to about 16 ml by using a Centriplus Concentrator (Amicon, MWCO 10,000, 470 rpm, 2800× g). Refolded TM9SF4/TM9SF2 was aliquoted to 0.5 ml into microcentrifuge tubes and stored at −80˚C. After refolding, the proteins were purified further by isoelectric focusing.
2.8. Isoelectric Focusing (IEF) Gel Purification
The refolded recombinant proteins were fucosed on Criterion IEF gels (Bio-Rad, Hercules, CA). The IEF gels were cut at pIs of 7.0, 6.0, and 4.5 based on IEF standards (Bio-Rad). The slices were soaked in 50 mM Tris-Mes buffer, pH 7 at 4˚C overnight to elute the proteins.
2.9. Measurement of arNOX Activity
Measurements of superoxide production were based on a standard method where reduction of ferricytochrome c by superoxide was monitored from the increase in absorbance at 550 nm with reference at 540 nm [6]. As a further check for the specificity of the arNOX activity, 60 units of superoxide dismutase (SOD) were added near the end of the assay to ascertain that the rate returned to base line. Rates were determined over 1 min at intervals of 1.5 min using a SLM Aminco DW 2000 spectrophotometer (Milton Roy, Rochester, NY) in the dual wavelength mode of operation. An extinction coefficient of 19.1 mM−1·cm−1 was used for reduced ferricytochrome c. Oxidation of NADH was determined spectrophotometrically from the disappearance of NADH measured at 340 nm in a reaction mixture containing 25 mM Tris-Mes (pH 7.2), 1 mM KCN to inhibit mitochondrial oxidase activity, 150 µM NADH, and the enzyme at 37˚C with temperature control (±0.5˚C) and stirring. Prior to assay, 1 µM reduced glutathione was added to reduce the protein in the presence of substrate. After 10 min, 0.03% hydrogen peroxide was added to reoxidize the protein under renaturing conditions in the presence of substrate. Measurements were initiated after an additional 10 min. A millimolar extinction coefficient of 0.805 was used to determine specific activity.
Oxidation of reduced coenzyme Q10 (CoQ10H2) was measured as the disappearance of reduced CoQ at both 290 and 410 nm [7]. A millimolar extinction coefficient of 0.805 was used to calculate the rate of CoQ10H2 oxidation.
Protein disulfide-thiol interchange was determined spectrophotometrically from the increase in absorbance at 340 nm resulting from the cleavage of dithiodipyridine (DTDP). Specific activities were calculated using a millimolar extinction coefficient of 6.21 [8].
Proteins were estimated by the bicinchoninic acid method [9] with bovine serum albumin as the standard.
2.10. Removal of Copper (II)
Purified recombinant TM9SF4 was concentrated to 0.7 mg/mL by using a Centricon concentrator (Millipore Corp., Danvers, MA) fitted with a 10,000 nominal molecular weight limit ultracel YM membrane. Samples (500 µL) were combined with 1 µL of trifluoroacetic acid (TFA) in the presence or absence of 15 µL of 10 mM bathocuproine and assayed for superoxide production and NADH oxidation.
2.11. Preparation of Antisera to Human arNOX
Rabbits were immunized with low molecular weight arNOX fragments purified from human urine (Figure 1(a)) to generate polyclonal antisera (Covance, Denver, PA). A spleen from one of the rabbits was used as a source of splenocytes for fusion with rabbit plastocytoma cells to generate a monoclonal antisera (Epitomics, Burlingame, Ca).
2.12. Preparation of Peptide Antibodies
Peptide antibodies were generated in rabbits to the N-terminal sequences of the exfoliated proteins. A cysteine residue was added to the N-terminus of each peptide to facilitate coupling to the carrier protein KLH.
TM9SF1A&1B (aa 72 - 87) IRHKSKSLGEVLDGDR TM9SF2 (aa 89 - 104) GKESENLGQVLFGER TM9SF3 (aa 70 - 88) KKSISHYHETLGEALQGVE TM9SF4 (aa 69 - 84) ITYKAENLGEVLRGDR
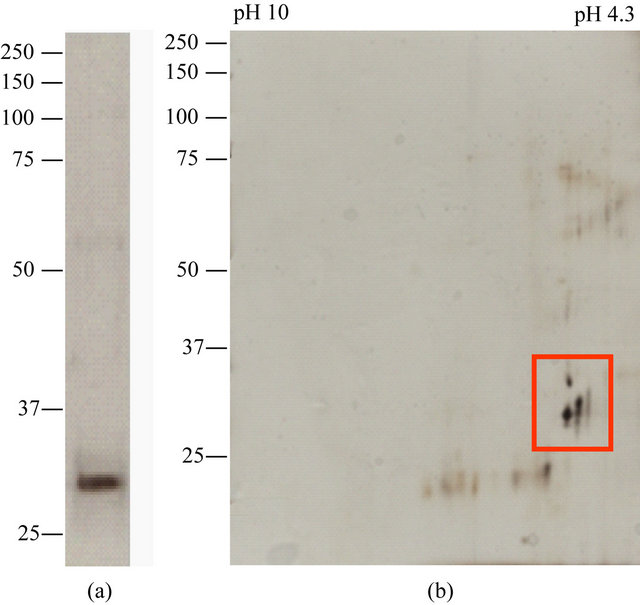

Figure 1. SDS-PAGE of arNOX purified from human urine (72-y M). (a). The protein was purified to apparent homogeneity by SDS-PAGE. (b). Two-dimensional gel separation of preparation of Figure 1(a) silver stained. Two sets of proteins are evident, one at about 24 kDa and one at about 32 kDa but with different isoelectric points.
For immunoassay, the arNOX source was coated on each of 5 replicated wells of a 96 well ELISA plate and after appropriate washing and blocking, the isoformspecific antibodies were added singly to each of the wells or as an equal mixture if the objective was to quantitate total arNOX. A peroxidase second antibody was added followed by a colorimetric substrate and the resultant absorbance was determined in an automated plate reader. The log of absorbance readings was proportional to the amount of arNOX present and quantitated by means of a standard curve using recombinant TM9SF4.
2.13. Site-Directed Mutagenesis
Amino acids indicated were replaced by alanines by site-directed mutagenesis according to Braman et al. [10]. Numbered amino acids and nucleotide positions of splice variant products refer to numbers assigned to amino acids of the full length transcript.
2.14. Statistical Analyses
Means and standard deviations were analyzed for statistical significance using a two-tailed test.
3. RESULTS
Based on specific activity measurements and SDS-PAGE, a soluble preparation of arNOX proteins was isolated to apparent homogeneity from human urine (Figure 1(a)). When resolved by 2-D gel electrophoresis, the material concentrated from urine revealed only a small number of proteins of similar molecular weights (30 to 32 kDa) but differing acidic isoelectric points (Figure 1(b)). The arNOX activity of unfractionated urine exhibited a pattern of five different maxima, each recurring every 26 min (Figure 2). The activity was inhibited by addition of superoxide dismutase. When subjected to analysis by N-terminal sequencing (Tufts Core Facility, Boston, MA), no sequence was obtained suggesting that the fragments were both N-terminally blocked and trypsin resistant as has been characteristic of ENOX proteins in general. In contrast, pepsin digests analyzed revealed with a high confidence level a peptide of human origin ATMDWLPWSLLLF… corresponding to the N-terminus of a member of the TM9 superfamily of transmembrane proteins, designated as SF4.
To determine if the human arNOX proteins might similarly derive from TM9 superfamily members [5,11], recombinant human TM9SF4 was generated from cDNA corresponding to full length TM9SF4. We were unable to express full length TM9SF4 in E. coli. Moreover, the bacteria transfected with cDNA to full length TM9SF4 grew at only 20% to 30% of the rate of non-transfected bacteria. Therefore, since we were primarily interested in the exfoliated form of the protein, cDNA corresponding to the ca. 30 kDa N-terminus of TM9SF4 was prepared and expressed. The truncated form exhibited the marked oscillating activity characteristic of other ENOX proteins family members [12; Figures 2-7].
Crude lysates of bacteria expressing the truncated TM9SF4 exhibited single maxima of arNOX activity separated at intervals of approximately 26 min (Figure 3) which is characteristic of the arNOX activities of the proteins purified from urine except that, with urine, there were at least five such maxima repeating every 26 min
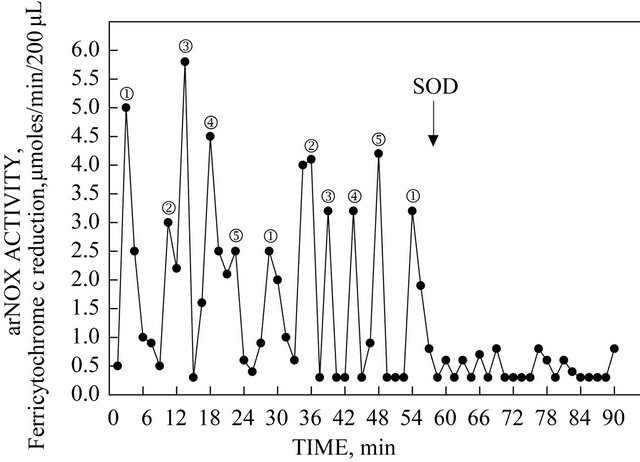
Figure 2. arNOX activity of the human urine fraction utilized in the generation of the data of Figure 1. The repeating pattern is that of five oscillating maxima that recur at intervals of 26 min. SOD = 60 units of superoxide dismutase added after 60 min.
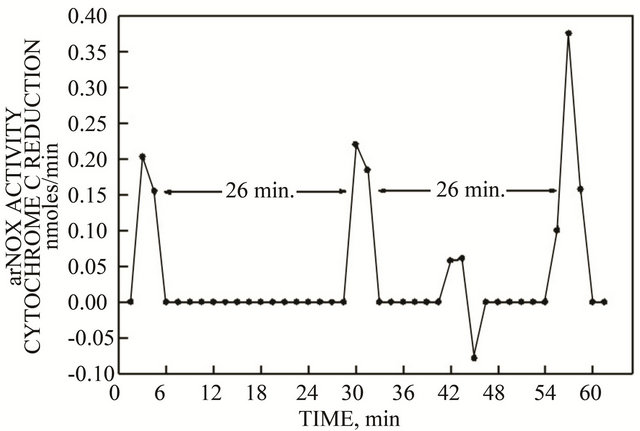
Figure 3. arNOX activity of the soluble fraction of a crude bacterial lysate containing recombinant TM9SF4 showed only bursts of superoxide generation as measured by reduction of ferricytochrome c. The bursts were separated by intervals of 26 min.
with a burst of superoxide suggestive of not one but as many as five arNOX proteins being present each contributing to a single superoxide burst every 26 min.
The activity pattern exhibited by the expressed truncated TM9SF4 proteins purified by isoelectric focusing exhibited five activity maxima within a time span of 26 min based on NADH oxidation where two of the maxima were separated by about 6 min and the remaining three maxima were separated from each other and from the two maxima separated by 6 min by intervals of about 5 min to generate the patterns shown in Figures 4(a) and (c). What is most characteristic of the arNOX activity patterns is that the maxima labeled ƒ exhibited a burst of superoxide production 2 to 4 times that of the other maxima (Figures 4(b) and (d)) and is the predominant activity observed in diluted preparations of the protein (Figure 3). The recombinant TM9SF2 and TM9SF4 oxidized reduced coenzyme Q in a standard assay (Figure 5) with activity measured either at A410 (Figures 5(a) and (c)) or at A290 (Figures 5(b) and (d)).
When protein disulfide thiol interchange activity, a second enzymatic activity associated with ENOX pro-
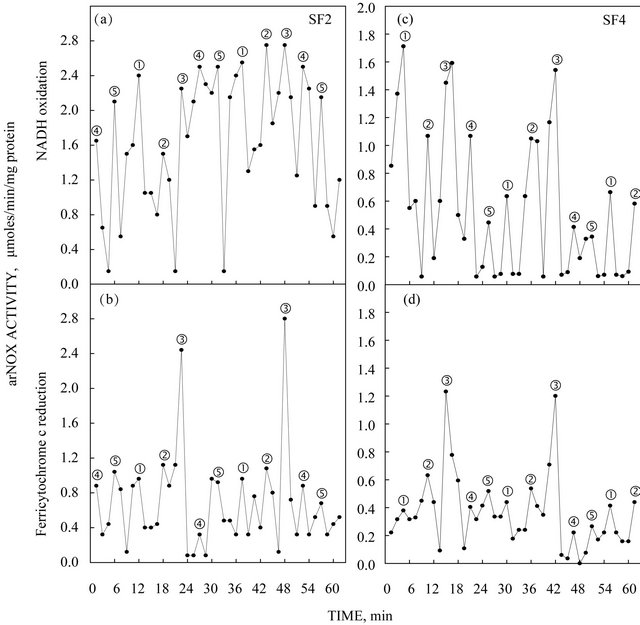
Figure 4. arNOX activity of recombinant TM9SF2 (a, b) and TM9SF4 (c, d) purified by isoelectric focusing showing the typical 5-peak pattern of activity characteristic of ENOX proteins in general. (a, c). NADH oxidation. (b, d). Superoxide generation. Superoxide generation is intensified with maximum 3 of the 5 maxima oscillatory pattern in keeping with results of Figure 3.
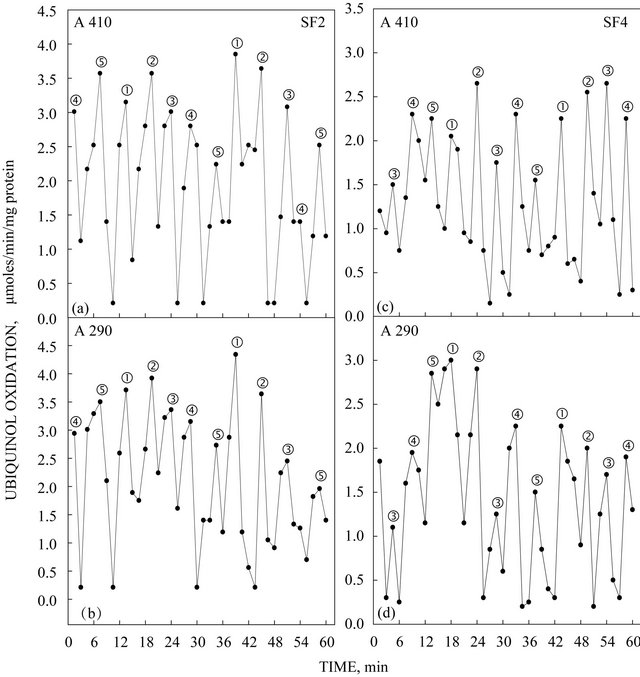
Figure 5. Ability of recombinant TM9SF2 (a, b) and TM9SF4 (c, d) purified by isoelectric focusing to oxidize hydroquinone (reduced coenzyme Q10) either by an increase in A410 (a, c) or by a decrease in A290 (b, d). As with NADH oxidation of Figure 4, the activity oscillates with prominent maxima separated by 6 min plus three additional maxima ƒ, „ and … separated from each other and from maximum ‚ by about 5 min to create a 26 min period created by the five recurring maxima.
teins, was measured, a five-maxima pattern similar to that generated for NADH oxidation was obtained (Figure 6). All three activities were resistant to a specific inhibitor of ENOX1, simalikalactone D, and inhibitors of ENOX2, phenoxodiol and capsaicin. Nor were the arNOX activity patterns phased by addition of melatonin as is characteristic of ENOX1 [13]. The period length, however, was increased to about 30 min by assay in D2O in place of water (Figure 7), a characteristic of ENOX proteins generally.
Results with recombinant TM9SF2 and TM9SF4 proteins expressed as N-terminal (ca. 15 kDa or 30 kDa) fragments were nearly identical.
Superoxide dismutase was added at the end of each assay to ascertain that activity based on superoxide production returned to base line. Both a specific arNOX inhibitor mixture of dormin + Schizandra + salicin [to 2.5 ml of assay volume were added 60 µl of an aqueous mixture of 4 mg/ml Schizandra (Schizandra chinensis extract, 9% schizandrins, Draco, San Jose, CA) plus 1 mg/ml salicin (Sigma, St. Louis, MO) and 20 µL of IBR Dormin (Israli Biotechnology Research, Ramat-Gan, Israel) = AgeLoc (NuSkin Enterprises, Provo, UT)] and an equal mixture of peptide antibodies to each of the 5 arNOX superfamily members at a titer of 1:10,000 inhibited the activity by >90%. Also inhibitory were gallic acid (EC50 = 2 µM), tyrosol (EC50 = 1 µM) and coenzyme Q10 (EC50 = 80 nM).
Peptide antibodies to each of the isoforms along with corresponding DNA probes to each of the isoforms were developed for the soluble forms of each of the five isoforms. The antibodies were used to systematically identify the 5 isoforms in human sera and saliva and to verify that they corresponded to the known sequences of the TM9 superfamily members. DNA sequence information was used to generate RT-PCR probes for each of the

Figure 6. Protein disulfide thiol interchange activity of recombinant TM9SF2 (a) and TM9SF4 (b) measured from the cleavage of dithiodipyridine (DTDP) substrate.
isoforms and demonstrate their expression in both human lymphocytes and human skin explants (not shown) to confirm the genetic origins of the five known arNOX isoforms of human sera, plasma and other body fluids as TM9 superfamily proteins.
Sequential addition of peptide antibodies to sera to associate specific maxima unaffected in the pre-bleed with specific TM9 family members SF1 to SF4 (Figure 8). After addition of SF1-specific antibody, two of the five maxima were no longer present. After addition of antibody specific to SF2 only 3 activity maxima remained. After TM9SF3-specific antibody (antisera) only one isoform remained and following addition of TM9- SF4 antibodies, the arNOX activity was completely inhibited. Because of N-terminus sequence similarity, antibodies to TM9SF1a and TM9SF1b cross-reacted and blocked both activities.
The ca. 30 kDa N-terminal regions of the TM9SF proteins, which are exposed at the external surface of the plasma membrane are shed into the blood and other body fluids (saliva, perspiration, urine). They are present in sera and plasma and were measured collectively as arNOX (Table 1). Each of the shed forms contains putative functional motifs required of an ENOX protein (Figure 9, Table 2). The functional motifs are located in the shed forms of the TM9 proteins but, despite the presence of some form in each of the superfamily members, sequence identity among the shed fragments with arNOX activity of different family members was minimal.
Table 1. Relative arNOX levels in skin, saliva and serum minus blank of four subjects ages 51- to 74-y. The antibody source was a mixture of equal amounts of each of the 4 peptide antisera of Figure 8.
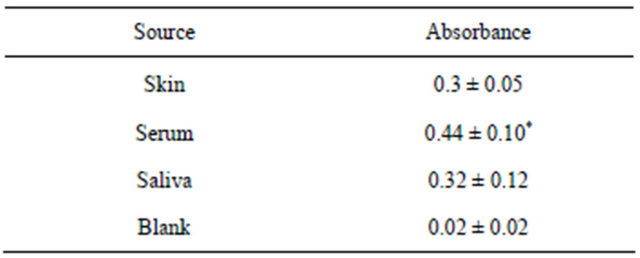
*By comparison, values minus blank for a younger volunteer (21-y) were 0.01 ± 0.01.
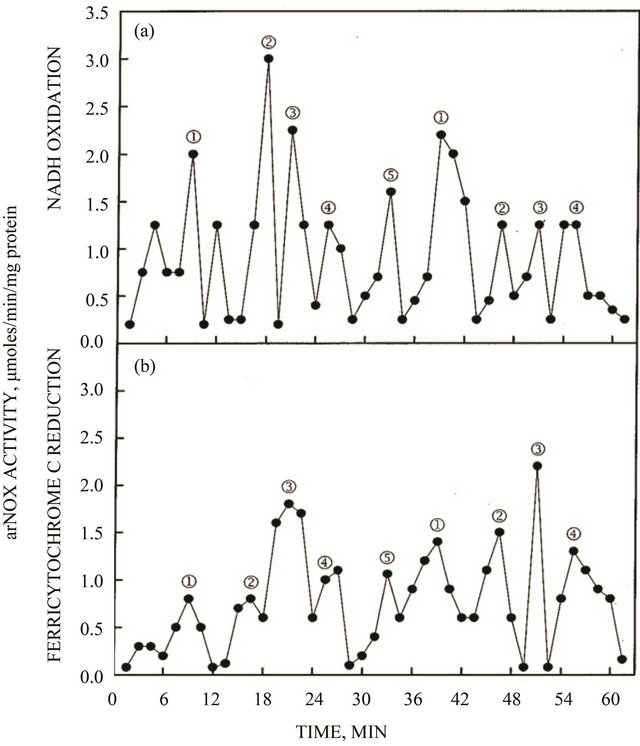
Figure 7. As in Figures 4(c) and (d) except with water replaced by D2O in the assay. As with other ENOX proteins, assay in the presence of heavy water increases the period length by about 25% from 26 to 30 min.
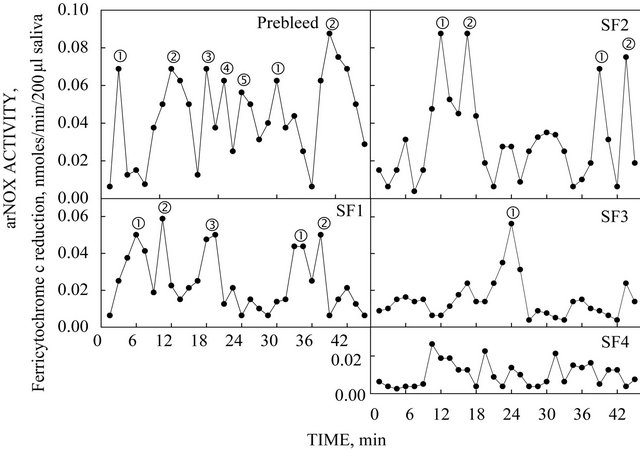
Figure 8. Results obtained with the sequential addition of peptide antisera to sera to show association of the five specific maxima unaffected by addition of the prebleed along with specific isoforms TM9SF1 to TM9SF4. After addition of TM9SF1-specific antisera, two maxima corresponding to SF1a and SF1b were lost. However, after addition of antisera specific to TM9SF2, only two maxima remained. After addition of TM9SF3-specific antisera, one maximum remained and after addition of TM9SF4 specific antisera, no maxima were observed.
| 1 MATAMDWLPWSLLLFSLMCETSAFYVPGVAPINFHQNDPVEIKAVKLTSSRTQLPYEYYS 61 LPFCQPSKITYKAENLGEVLRGDRIVNTPFQVLMNSEKKCEVLCSQSNKPVTLTVEQSRL 121 VAERITEDYYVHLIADNLPVATRLELYSNRDSDDKKKEKDVQFEHGYRLGFTDVNKIYLH 181 NHLSFILYYHREDMEEDQEHTYRVVRFEVIPQSIRLEDLKADEKSSCTLPEGTNSSPQEI 241 DPTKENOLYFTYSVHWEE Adenine nucleotide binding site (GXVXXG) Putative copper sites (YVH, HGY) Putative protein disulfide interchange site (CXXXC) Conserved CQ/CE |
Figure 9. The identities and locations of putative functional arNOX motifs of TM9SF4.
Table 2. Putative functional motifs of human SF9 superfamily members.
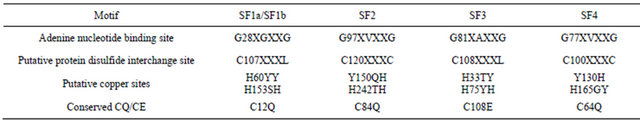
Confirmation of functional assignments of motifs common to arNOX proteins of Table 2 is provided for the specific functional motifs of arNOX TM9SF2 by site directed mutagenesis (Table 3). Within the CQ motif common to arNOX proteins, superoxide production was reduced substantially by the double mutation of C84A and Q85A and was reduced to background in one of the three repetitions. The C84A and Q85A replacements individually were without effect on specific activities. However the C84A, but not the Q85A, replacement resulted in a shortened period length decreased from 26 min to 22 min. A change in period length following one or more cysteine replacements has been observed with other ENOX proteins [13,14]. The G77V replacement in the putative adenine nucleotide binding motif largely eliminated NADH oxidation, was stimulatory to a superoxide generation and was without effect on disulfide-thiol interchange. A response to the C120A replacement was most evident with protein disulfide interchange. Putative copper site replacements, H153A and H242A, reduced activities in general. All three activities, NADH oxidation, superoxide generation and disulfidethiol interchange, were reduced to background levels with the triple mutation H152A, H153A and H242A.
Melatonin (100 µM) did not shift the phase of the period (not shown). However, after 1.5 periods, 39 min after melatonin addition, a maximum was skipped.
To determine if the TM9SF forms required copper for activity, recombinant TM9SF4 was assayed in the presence of trifluoroacetic acid (to unfold the protein) with or without the copper chelator, bathocuproine, to remove the copper. The activity, measured either based on NADH oxidation or superoxide production, was diminished when bathocuproine was present (not shown). TFA alone did not reduce activity and activity could be reTable 3. Confirmation of functional motifs of arNOX TM9SF2 by site-directed mutagenesis.
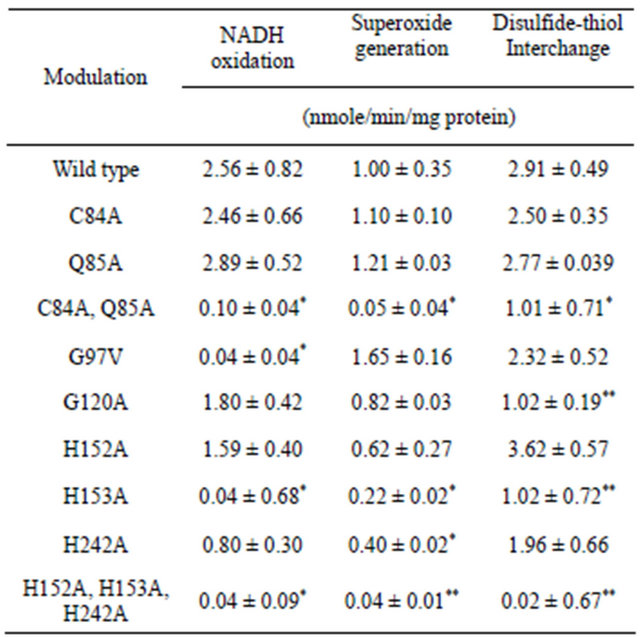 *Significant (p = 0.02 - 0.04); **Very Significant (p = 0.003 - 0.004).
*Significant (p = 0.02 - 0.04); **Very Significant (p = 0.003 - 0.004).
stored to TFA and bathocuproine treated preparations by additions of copper.
4. DISCUSSION
The age-related NADH oxidase of cells and body fluids [1] was difficult to clone in order to determine its genetic origins. In retrospect, the fact that human arNOX is not a single protein but a complex of at least five related proteins but with diverse N-terminal sequences contributed to the difficulty along with their derivation from a membrane associated precursor having 9 transmembrane regions [5]. As is characteristic of ENOX proteins generally, proteins with arNOX activity tend to form aggregates when concentrated and are blocked to N-terminal sequencing. In order to be cleaved enzymatically or otherwise, the protein must be unfolded. Based on studies with release of bound copper, low pH (0.2% trifluoroacitic acid) was required such that a combination of low pH and pepsin resulted in cleavage of identifiable peptide fragments, including one derived from the N-terminus of SF4 of the TM9 superfamily of membrane proteins.
Based on DNA sequence, TM9SF4 protein was expressed in E. coli and shown to have functional characteristics of an ENOX protein. The pattern of activity oscillations consisting of 5 unequally spaced maxima was the characteristic, yet unique, periodicity (26 min period instead of the 24 min period for ENOX1 or 22 min period for ENOX2). A further defining characteristic of the arNOX proteins is their ability to generate superoxide. Superoxide generation is evidenced by the superoxide dismutase-inhibited reduction of ferricytochrome c. ENOX1 and ENOX2 proteins carry out four electron transfers from NAD(P)H to molecular oxygen. Superoxide is not a reaction product of either ENOX1 or ENOX2 as ferricytochrome c is not reduced by these proteins.
Prior to our analysis of the expressed recombinant arNOX, the pattern of oscillations of human samples was poorly understood. Patterns observed consisted of 4 to 7 maxima with the average being about five. The origins of the maxima, recurring every 26 min, was eventually correctly attributed to individual arNOX forms based on 2- D gel separations (Figure 1), one maximum per form, such that superoxide appeared to be generated by different arNOX species only once every 26 min when present in dilute solution (Figure 3). This was confirmed from initial analyses of recombinant TM9SF4 where one burst of superoxide generated was observed every 26 min also when analyzed in a dilute solution (Figure 4). The origin of this single burst was subsequently traced to a burst of superoxide associated with the maximum labeled ƒ of the recurring pattern of five maxima (Figure 5). Specific activity is µmoles/min/mg protein for the recombinant arNOX purified by isoelectric focusing (Figure 4) whereas in the crude fractions only nanomolar activities were recorded (Figure 3).
The human arNOX cDNA encodes a polypeptide having a highly hydrophobic C-terminal portion organized into nine transmembrane domains with a very similar structure and sequence to members of a novel family of multispanning domain proteins designated “TM9SF” (transmembrane protein 9 superfamily) by the Human Gene Nomenclature Committee. The leader member of the TM9SF family is the Saccharomyces cerevisiae EMP70 gene product, a 70 kDa precursor that is processed into a 24 kDa protein (p24a) located in the endosomes [5]. To date, five subtypes of human TM9SF proteins have been identified, i.e., TM9SF1a [hMP70; 11], TM9SF1b, TM9SF2 [p 76; 5], TM9SF3 and TM9SF4 (D87444), which exhibit 30% - 40% amino acid sequence identity to each other and with the yeast p24a precursor [15]. This was a surprising result in that arNOX activity of body fluids of aged individuals was the result of at least five distinct proteins.
Hydropathy analysis [16] of p76 and its close relatives [17] revealed that these proteins share a unique membrane binding domain [5]. They also contain a short N-terminal hydrophobic extension characteristic of a signal sequence, followed by a mostly hydrophilic, amino terminal portion that extends up to amino acid residue 300 in certain family members. The remaining portions of these proteins are extremely hydrophobic and contain nine transmembrane domains to make them integral membrane proteins that adopt a type 1 topology. Polypeptide translocation would be initiated via their N-terminal hydrophobic signal sequence and they would ultimately be anchored in the membrane via stop-transfer sequences.
Full length members of the TM9 protein superfamily are all characterized as cell surface proteins (as are arNOX proteins) having a characteristic series of 9 membrane spanning hydrophobic helices that criss-cross the plasma membrane and also are present on endosomes [5]. The transmembrane regions are highly conserved and similar or identical in each of the five forms. There are 5 such family members known (1 with two transcript variants, 2, 3 and 4). The two transcript variants of family member 1 are very similar with the exception that member 1a transcript variant, contains additional C-terminal residues absent from transcription variant 1b. The different family members (1 through 4) are encoded by different genes and are therefore not splice variants from a single gene as for the different transcript variants of ENOX2 [18].
5. ACKNOWLEDGEMENTS
We thank Connie Phung for conduct of isoelectric focusing experiments, Dr. Christiaan Meadows, for ELISA results of Table 1, Jeff Morré and Samanthi Wickramasekra of Oregon State University for sequencing of trypsin and pepsin digests, Aya Ryuzoji and Patrick Finnegan for preparation of figures, Amanda Lecklitner for technical assistance and Peggy Runck for manuscript preparation.
REFERENCES
- Morré, D.M., Guo, F. and Morré, D.J. (2003) An agingrelated cell surface NADH oxidase (arNOX) generates superoxide and is inhibited by coenzyme Q. Molecular and Cellular Biochemistry, 264, 101-109. doi:10.1023/A:1027301405614
- Kern, D.G., Draelos, Z.D., Meadows, C., Morré, D.M. and Morré, D.J. (2010) Controlling reactive oxygen species in skin at their source to reduce skin aging. Rejuvination Research, 13, 165-167. doi:10.1089/rej.2009.0914
- Morré, D.M., Lenaz, G. and Morré, D.J. (2000) Surface oxidase and oxidative stress propagation in aging. Journal of Experimental Biology, 203, 1513-1521.
- Morré, D.M., Meadows, C. and Morré, D.J. (2010) arNOX: Generator of reactive oxygen species in the skin and sera of aging individuals subject to external modulation. Rejuvination Research, 13, 162-164. doi:10.1089/rej.2009.0919
- Schimmöller, F., Diaz, E. Mühlbauer, B. and Pfeffer, S.R. (1998) Characterization of a 76 kDa endosomal, multispanning membrane protein that is highly conserved throughout evolution. Gene, 216, 311-318. doi:10.1016/S0378-1119(98)00349-7
- Butler, J., Koppenol, W.H. and Margoliash, E. (1982) Kinetics and mechanism of the reduction of ferricytochrome c by the superoxide anion. Journal of Biological Chemistry, 257, 10747-10750.
- Kishi, T., Morré, D.M. and Morré, D.J. (1999) The plasma membrane NADH oxidase of HeLa cells has hydroquinone oxidase activity. Biochimica et Biophysica Acta, 1412, 66-77. doi:10.1016/S0005-2728(99)00049-3
- Morré, D.J., Gomez-Rey, M.L., Schramke, C., Em, O., Lawler, J., Hobeck, J. and Morré, D.M. (1999) Use of dipyridyl-dithio substrates to measure directly the protein disulfide-thiol interchange activity of the auxin stimulated NADH: Protein disulfide reductases (NADH oxidase) of soybean plasma membranes. Molecular and Cellular Biochemistry, 207, 7-13. doi:10.1023/A:1006916116297
- Smith, P.K., Krohn, R.I., Hermanson, G.T., Mailia, A.K., Gartner, F.H., Provenzano, M.D., Fujimoto, E.K., Goeke, N.M., Olson, F.J. and Klenk, D.C. (1985) Measurement of protein using bicinchoninic acid. Analytical Biochemistry, 150, 76-85. doi:10.1016/0003-2697(85)90442-7
- Braman, J., Papworth, C. and Greener, A. (1996) Sitedirected mutagenesis using double-stranded plasmid DNA templates. Methods Molecular Biology, 57, 31-44.
- Singer-Krüger, B., Frank, R., Crausaz, E. and Riezman, H. (1993) Partial purification and characterization of early and late endosomes from yeast. Identification of four novel proteins. Journal of Biological Chemistry, 268, 14376-14386.
- Morré, D. J. and Morré, D.M. (2003) Cell surface NADH oxidases (ECTO-NOX proteins) with roles in cancer, cellular time-keeping, growth, aging and neurodegenerative disease. Free Radical Research, 37, 795-808. doi:10.1080/1071576031000083107
- Jiang, Z., Goldstein, N. M., Morré, D.M. and Morré, D.J. (2008) Molecular cloning and characterization of a candidate human growth-relaqted ant time-keeping constitutive cell surface hydroquinone (NADH) oxidase. Biochemistry, 47, 14028-14038. doi:10.1021/bi801073p
- Chueh, P.-J., Kim, C., Cho, N., Morré, D.M. and Morré, D.J. (2002) Molecular cloning and characterization of a tumor-associated, growth-related and time-keeping hydroquinone (NADH) oxidase (NOX) of the HeLa cell surface. Biochemistry, 41, 3732-3741. doi:10.1021/bi012041t
- Chluba-de Tapia, J., de Tapia, M., Jäggin, V. and Eberle, A.N. (1997) Cloning of a humam multispanning membrane protein cDNA: Evidence for a new protein family. Gene, 197, 195-204. doi:10.1016/S0378-1119(97)00263-1
- Kyte, J. and Doolittle, R.F. (1982) A simple method for displaying hydropathic character of a protein. Journal of Molecular Biology, 157, 105-132. doi:10.1016/0022-2836(82)90515-0
- Sugasawa, T., Lenzen, G., Simon, S., Hidaka, J., Cahen, AW., Guillaume, J.-L., Camoin, L., Strosberg, A. and Nahmias, C. (2001) The iodocyanopindolol and SM- 11044 binding protein belongs to the TM9SF multispanning membrane protein superfamily. Gene, 273, 283-289. doi:10.1016/S0378-1119(01)00587-X
- Tang, X., Tian, Z., Chueh, P.-J., Chen, S., Morré, D.M. and Morré, D.J. (2007) Alternative splicing as the basis for specific localization of tNOX, a unique hydroquinone (NADH) oxidase, to the cancer cell surface. Biochemistry, 46, 12337-12346. doi:10.1021/bi700973k
Abbreviations
arNOX: age-related NADH oxidase;
CAPS: 3 (cyclohexylamino)-1-propane sulfonic acid;
DTDP: dithiodipyridine;
ELISA: enzyme-linked immunosorbent assay;
ENOX: ECTO-NOX;
IPTG: isopropyl-beta-D-thiogalactopyranoside;
MWCO: molecular weight cutoff;
PMSF: phenylmethylsulfonyl fluoride;
SDS-PAGE: sodium dodecyl sulfate polyacrylamide gel electrophoresis;
SF: superfamily;
TFA: trifluoroacetic acid;
TM: transmembrane
NOTES
*Corresponding author.

