 Journal of Biosciences and Medicines, 2013, 1, 5-10 JBM http://dx.doi.org/10.4236/jbm.2013.13002 Published Online December 2013 (http://www.scirp.org/journal/jbm/) OPEN ACCESS Effective cofactor complex purification using nanobeads Makoto Umeda1, Tatsuya Uebi1, Naoya Maekawa1, Yuka Masaike2, Hiroshi Handa2, Takeshi Imai1 1Department of Aging Intervention, National Center for Geriatrics and Gerontology (NCGG), Obu, Japan 2Faculty of Bioscience and Biotechnology, Tokyo Institute of Technology, Yokohama, Japan Email: dai.ncgg@gmail.com Received September 2013 ABSTRACT Drug target factor complex identification is necessary for evidence based drug discovery. Previous study showed that using small chemical immobilized mag- netic nanobeads the chemical target factors were ef- fectively purified and identified. Here we succeeded to purify the chemical target factor complex, so called cofacto r(s). Arginine exhibits a variety of biological activities through a complex and highly regulated set of pathways that remain incompletely understood at both the whole-body and the cellular levels. The aim of this study is to develop and validate effective puri- fication system for arginine target complex. New ar- ginine target protein (arginine interacting factor 4, AIF4) was purified and identified. Using recombinant AIF4 protein and arginine-immobilized magnetic nanobeads, AIF4 cofactor, AIF4-BP1, we re purified. Interaction of AIF4 and AIF4-BP1 was detected in arginine-dependent manner, suggesting arginine re- ceptor complex formation. This nanobeads technolo- gy is more than 30-fold efficient purification efficient than general purifi ca ti on te chnology. Keywords: Ar ginine; Na nobeads 1. INTRODUCTION The drug target proteins identification is the way of evi- dence based drug discovery. Small chemical immobilized nanobeads were developed as purifying target protein [1]. Previous studies showed that prostaglandin and vitamin K2 target proteins were effectively purified in one step [2,3]. As the second chemical target protein purification, arginine is classified as a semi-essential amino acid be- cause the ability of the body to synthesize sufficient quantities to meet its needs varies according to develop- mental age and the incidence of disease or injury [4,5]. The sources of free arginine within the body are dietary protein, turnover of body proteins, and endogenous syn- thesis. Arginine is synthesized from citrulline by the se- quential action of the cytosolic enzymes arginosuccinate synthetase and arginosuccinate lyase. On the other hand, arginine can be catabolized by four sets of enzymes in mammalian cells: nitric oxide synthases (NOS), arginase, arginine-glycine amidinotransferase and arginine decar- boxylase, and be converted to a variety of biologically active compounds such as nitric oxide (NO), creatine phosphate, agmatine, polyamines, ornithine and citrulline [6]. Arginine has many effects in the body that include modulation of immune function [6], hormone secretion [7], insulin secretion [8], and endothelial function [9]. Thus, arginine is a biological active dietary compound that mediates numerous physiological activities directly or via me t a bolized pro duc ts [10]. Metabolic enzymes that use arginine as substrates are well-known arginine interacting factors (AIFs), however, much less are known about the AIFs that directly mediate the physiological effects of arginine. In our previous pa- per, using nanobeads technology three independent AIFs are purified and identified from HeLa cell extract [4]. Abilities of all the three arginine-AIF complexes were confirmed. To further study arginine signaling, using recombinant AIF protein, cofactor of arginine-AIF com- plex was purified and identified in this paper. This co- factor (designated AIF-BP1) was purified from cell ex- tract directly, and identified. 2. MA TERIALS AND METHODS 2.1. Materials 1-Ethyl-3-(3-dimethylaminopropyl) carbodiimide hydro- chloride (EDC), triethylamine, N,N-di methyl-4-a min o- pyridine (DMAP), N,N-dimethylformamide (DMF), acetic anhydride, dithiothreitol and iodoacetamide were purchased from Nacalai Tesque (Kyoto, Japan). Ethyle- neglycol diglycidyl ether (EGDE) was purchased from Wako Chemicals (Osaka, Japan). Trypsin was obtained from Promega (Madison, WI, USA). HEK293 cells were obtained from American Type Culture Collection (Ma- nassas, VA, USA). 2.2. Effective AIFs Purification FG beads were prepared as previously described [11]. Epoxy groups on FG beads were aminolysed by NH4OH  M. Umeda et al. / Journal of Biosciences and Medicines 1 (2013) 5-10 Copyright © 2013 SciRes. OPEN ACCESS and coupled to EGDE to produce FGNEGDE beads [12, 13]. Epoxy groups on FGNEGDE beads were amino- lysed by NH4OH (FGNEGDEN), carboxylated with suc- cinic anhydride (FGNEGDENS) and activated with NHS to produce NHS-activated FGNEGDENS beads [14]. NHS-activated FGNEGDENS beads (2.5 mg) were in- cubated with 1.5 mM arginine methyl ester (AME) in 500 L of DMSO containing 10 (V/V)% triethylamine at 37˚C for 3 hours. Released NHS during the coupling reaction was collected together with unbound AME. Coupling yield was determined by separating NHS re- leased from the beads on a Symmetry C18 cartridge, 5 m (Walters, Milford, MA, USA). Usually 40 (mol)% of AME was immobilized onto the beads. The amount of AME immobilized onto the beads can be controlled by the amount of AME added into the immobilization reac- tion. Unreacted NHS-activated carboxylate was masked with 1 M 2-ethanolamine (pH 8.0) at 4˚C for 24 hours. AME-immobilized beads were suspended in distilled water and stored at 4˚C until use [4,14]. Whole cell extracts of human umbilical vein endo- thelial cell (HUVEC) were prepared as followed; the cell pellets were washed with PBS several times, and solubi- lized with binding buffer (20 mM HEPES-NaOH pH 7.9, 10% glycerol, 200 mM KCl, 1 mM MgCl2, 0.2 mM CaCl2, 0.2 mM EDTA, 1 mM DTT and 0.2 mM PMSF) with 1% of n-octyl-β-D-glucoside (n-octylglucoside), and centrifuge 1300 g for 5 minutes and supernatant was recovered. The supernatant was dialyzed against binding buffer without n-octylglucoside for 4 hours for elimina- tion of n-octylglucoside. AME-immobilized beads or control beads (200 μg) were equilibrated with binding buffer and incubated with 200 μL of the whole cell extracts at 4˚C for 4 hours using RT-50 rotator (15 rpm, TAITEC, Saitama, Japan). After washin g with bindi n g buf fe r, they contained 10 mM argi- nine [4]. Affinity purified AIFs were separated by SDS- PAGE and gels subjected to silver staining. The specific protein bands were excised, reduced with 10 mM DTT followed by alkylation w ith 55 mM iodoaceta mide. Band slices were digested with trypsin (12 g/mL) overnight and desalted with ZipTip C18 (Millipore, MA, USA). The extracted peptides were then separated via nano-flow liquid chromatography (LC, Paradigm MS4, Tokyo, Ja- pan) using a reverse phase C18 column (Magic C18, AMR, Tokyo, Japan). The LC eluent was coupled to a micro- ionspray source attached to a LCQ Advantage MAX mass spectrometer (Thermo Electron Corporation, MA, USA). All MS/MS spectra were searched using the Tur- boSEQUEST algorithm within the BioWorks 3.2 soft- ware (Thermo Electron Corporation, MA, USA; [13]). 2.3. Production of Recombinant Protein T7 promoter tagged DNA fragments of AIF4 and AIF4- BP1 were amplified by PCR with two primers; T7 pro- moter fused to 5’ sequence of each cDNA and poly T fused to 3’ complimentary sequence of each cDNA. These DNA fragments were used to synthesize 35S-radio- labeled recombinant proteins in a coupled transcrip- tion/translation system according to the protocol of man- ufacture (Promega, WI, US A; [15]). His- or GST-tagged fusion proteins were prepared as following manufacture’s instructions (GE health care Japan). Bacteria expressing the fusion proteins were re- suspended in PBS and sonicated mildly, and debris was pelleted [15]. 2.4. Arginine Signaling Complex Purification AME-immobilized beads or control beads (200 μg) were equilibrated with binding buffer and incubated with both recombinant hvcAIF4 protein and 200 μL of the whole cell extracts at 4˚C for 4 hours using RT-50 rotator (15 rpm, TAITEC, Saitama, Japan). Affinity purified-AIF- BPs were separated by SDS-PAGE and gels subjected to silver staining, and identified with LC/MS analysis [1 5]. 2.5. Protein-Pr otein Interaction Analysis Beads binding assay was performed to evaluate arginine- protein and protein-protein interaction as follows; AME- immobilized beads or control beads, arginine-agarose or control agarose (200 μg) were equilibrated with binding buffer and incubated with 200 μL of the radiolabeled proteins (hvcAIF4, hvcAIF4-BP1) or recombinant pro- teins (His-hvcAIF4, GST-hvcAIF4-BP1) at 4˚C for 4 hours using RT-50 rotator (15 rpm, TAITEC, Japan). Af- ter washing with binding buffer, bound proteins were eluted by the elution buffer (100 mM arginine in binding buffer). Eluants were subjected to SDS-PAGE. The gels were dried, and autoradiography was then performed to visualize the radiolabeled protein [4,15]. Protein-protein interaction in cell line analysis w ith immuno-precipitated (Ip)-Western blotting was performed as described pre- viously [15]. 3. RESULTS AND DISCUSSION 3.1. Identification o f hv cAIF4 from HUVEC (Figure 1) Our previ o us study demonstr a t e d tha t AIF1, 2 an d 3 were effectively purified and identified from HeLa cell extract directly. The AIF1 (=phosphofructokinase, PFK) is a kinase enzyme that phosphorylates fructose 6-phosphate in glycolysis, and the AIF2 & 3 (=RuvBL1 & 2) were transcription factors, suggesting the variety of arginine- signali n g [ 1] . In the case o f HUVEC, using AME-immobilized ma g- netic nanobeads (Figure 1(a)), arginine-binding protein 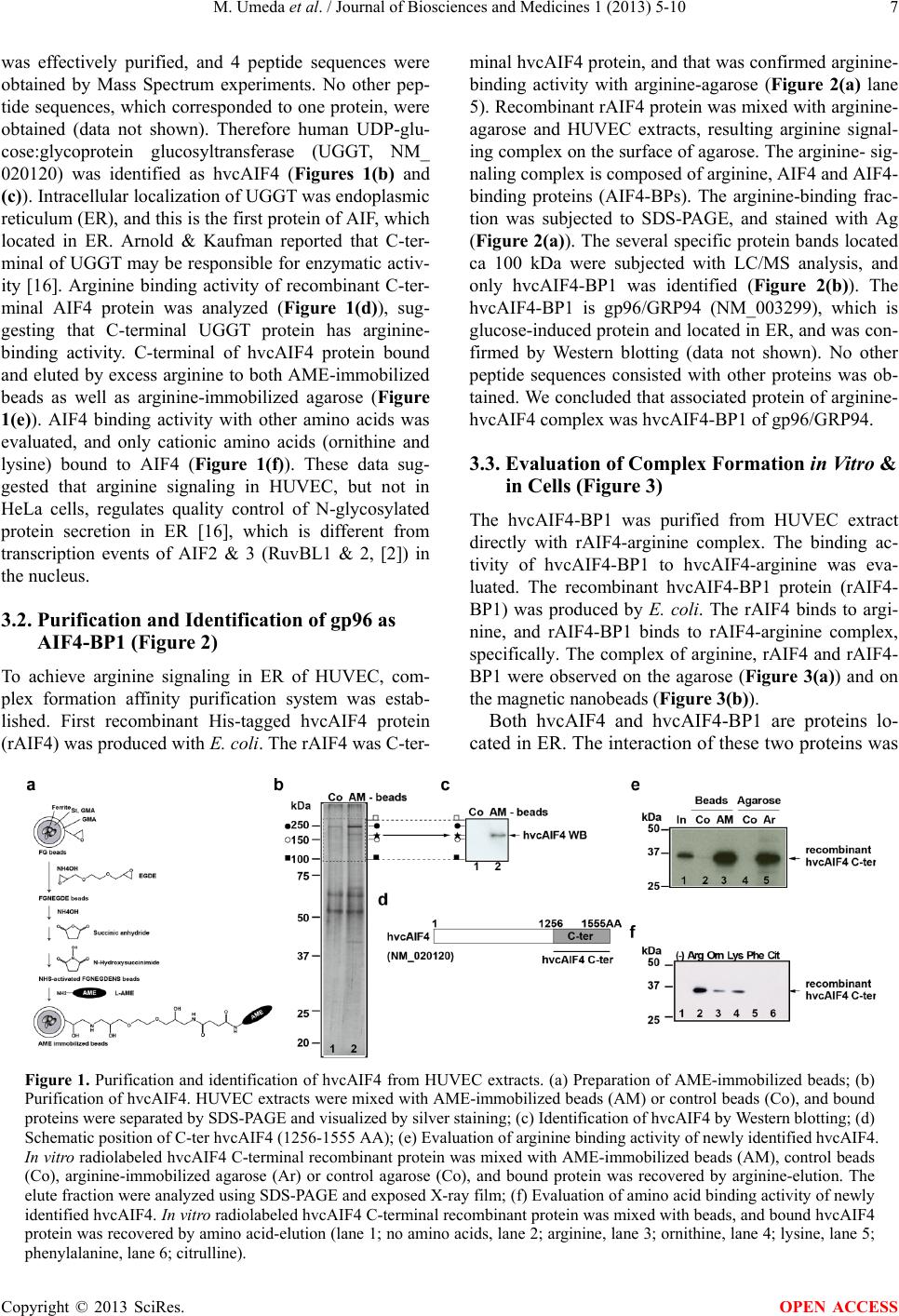 M. Umeda et al. / Journal of Biosciences and Medicines 1 (2013) 5-10 Copyright © 2013 SciRes. OPEN ACCESS was effectively purified, and 4 peptide sequences were obtained by Mass Spectrum experiments. No other pep- tide sequences, which corresponded to one protein, were obtained (data not shown). Therefore human UDP-glu- cose:glycoprotein glucosyltransferase (UGGT, NM_ 020120) was identified as hvcAIF4 (Fi g u re s 1(b) and (c)). Intracellular localization of UGGT was endoplasmic reticulum (ER), and this is the first protein of AIF, wh ich located in ER. Arnold & Kaufman reported that C-ter- minal of UGG T may be responsible for enzymatic activ- ity [16]. Arginine binding activity of recombinant C-ter- minal AIF4 protein was analyzed (Figure 1(d)), sug- gesting that C-terminal UGGT protein has arginine- binding activity. C-terminal of hvcAIF4 protein bound and eluted by excess arginine to both AME-immobilized beads as well as arginine-immobilized agarose (Figure 1(e)). AIF4 binding activity with other amino acids was evaluated, and only cationic amino acids (ornithine and lysine) bound to AIF4 (Figure 1(f)). These data sug- gested that arginine signaling in HUVEC, but not in HeLa cells, regulates quality control of N-glycosylated protein secretion in ER [16], which is different from transcription events of AIF2 & 3 (RuvBL1 & 2, [2]) in the nucleus. 3.2. Purification and Identification of gp96 as AIF4-BP1 (Figure 2) To achieve arginine signaling in ER of HUVEC, com- plex formation affinity purification system was estab- lished. First recombinant His-tagged hvcAIF4 protein (rAIF4) was produced with E. coli. The rAIF4 was C-ter- minal hvcAIF4 protein, and that was confirmed arginine- binding activity with arginine-agarose (Figure 2(a) lane 5). Recombinant rAIF4 protein was mixed with arginine- agarose and HUVEC extracts, resulting arginine signal- ing complex on the surface of agarose. The arginine- sig- naling complex is composed of arginine, AIF4 and AIF4 - binding proteins (AIF4-BPs). The arginine-binding frac- tion was subjected to SDS-PAGE, and stained with Ag (Fi gu re 2 (a)). The several specific protein bands located ca 100 kDa were subjected with LC/MS analysis, and only hvcAIF4-BP1 was identified (Figure 2(b)). The hvcAIF4-BP1 is gp96/GRP94 (NM_003299), which is glucose-induced protein and located in ER, and was con- firmed by Western blotting (data not shown). No other peptide sequences consisted with other proteins was ob- tained. We concluded that associated protein of arginine- hvcAIF4 complex was hvcAIF4-BP1 of gp96/GRP94. 3.3. Evaluation of Complex Formation in Vitro & in Cells (Figure 3) The hvcAIF4-BP1 was purified from HUVEC extract directly with rAIF4-arginine complex. The binding ac- tivity of hvcAIF4-BP1 to hvcAIF4-arginine was eva- luated. The recombinant hvcAIF4-BP1 protein (rAIF4- BP1) was produced by E. coli. The rAIF4 binds to argi- nine, and rAIF4-BP1 binds to rAIF4-arginine complex, specifically. The complex of arginine, rAIF4 and rAIF4- BP1 were observed on the agarose (Figure 3(a)) and on the magnetic nanobeads (Figure 3(b)). Both hvcAIF4 and hvcAIF4-BP1 are proteins lo- cated in ER. The interaction of these two proteins was Figure 1. Purification and identification of hvcAIF4 from HUVEC extracts. (a) Preparation of AME-immobilized beads; (b) Purification of hvcAIF4. HUVEC extracts were mixed with AME-immobilized beads (AM) or control beads (Co), and bound proteins were separated by SDS-PAGE and visualized by silver staining; (c) Identification of hvcAIF4 by Western blotting; (d) Schematic position of C-ter hvcAIF4 (1256-1555 AA); (e) Evaluation of arginine binding activity of newly identified hvcAIF4. In vitro radiolabeled hvcAIF4 C-terminal recombinant protein was mixed with AME-immobilized beads (AM), control beads (Co), arginine-immobilized agarose (Ar) or control agarose (Co), and bound protein was recovered by arginine-elution. The elute fraction were analyzed using SDS-PAGE and exposed X-ray film; (f) Evaluation of amino acid binding activity of newly identified hvcAIF4. In vitro radiolabeled hvcAIF4 C-terminal recombinant protein was mixed with beads, and bound hvcAIF4 protein was recovered by amino acid-elution (lane 1; no amino acids, lane 2; arginine, la ne 3; ornithi ne, lane 4; ly sine, la ne 5; phenylalanine, lane 6; citrulline).  M. Umeda et al. / Journal of Biosciences and Medicines 1 (2013) 5-10 Copyright © 2013 SciRes. OPEN ACCESS Figure 2. Purification and identification of AIF4-BP1. (a) Purification of hvcAIF4-BP1. Recom- binant AIF4 protein produced by E. coli. (rAIF4) and HUVEC extracts (HUVEC ext) were mixed with arginine-immobilized agarose (Ar) or control agarose (Co), and bound proteins were sepa- rated by SDS-PAGE and visualized by silver staining. Filled star pointed the protein band for hvcAIF4-BP1. Lanes 2 and 3 are loaded directly only recombinant proteins of rAIF4 and HUVEC ext, respectively, as size markers and input before purification; (b) Identification of hvcAIF4-BP1. Eight polypeptides (a-h) were identified by ion-spray mass spectr ometry. Figure 3. Evaluation of complex formation in vitro. (a) Recombinant proteins of hvcAIF4 (rAIF4) and hvcAIF4-BP1 (rAIF4-BP1) were mixed with arginine-immobilized agarose (Ar) or control agarose (Co), and bound proteins were separated by S DS-PAGE and visualized by CCB staining. Lanes 1 and 2 are loaded directly only recombinant proteins of rAIF4 and rAIF4-BP1, respectively. Arginine-agarose-binding rAIF4 or rAIF4-BP1 band is indicated as empty or filled circle, respectively; (b) Recombinant proteins of hvcAIF4 (rAIF4) and hvcAIF4-BP1 (rAIF4-BP1) were mixed with AME-immobilized beads (AM) or control beads (co), and bound proteins were separated by SDS-PAGE and visualized by silver staining. AME-immobi- lized-beads-binding rAIF4 or rAIF4-BP1 band is indicated as empty or filled circle, respectively; (c) To evaluate complex formation in cells, expression vectors of hvcAIF4 (hvcAIF4 vec), hvcAIF4-BP1 (hvc- AIF4-BP1 vec) or same amount of empty vector (−) were co-transfected to the HEK 293 cell line. The cell lysate wa s Ip with antibodies against control IgG (Lanes 1 to 4) or hvcAIF4-BP1 (Lanes 5 to 8), Ip fraction was separated by SDS-PAGE and transferred to the membrane, and visualized by Western blot with anti- hvcAIF4. The hvcAIF4 protein, which binds to hvcAIF4-BP1 in cells, is i ndicated as empty circle. *p < 0.05 significant difference, and NS; p > 0.05 not significant. analyzed in HEK293 cells. The expression vectors of hvc-AIF4 and hvcAIF4-BP1 were synthesized and trans- fected to HEK293 cells. Their protein expression was confirmed (data not shown). In anti-AIF4 antibody Ip fraction of double DNA transfected HEK293 cells (Fig- ure 3(c), lane 8), much higher amount of hvcAIF4 pro- teins were detected than control vector transfected (Fig- ure 3(c), lanes 1 & 5), single DNA transfected (Figure 3(c), lanes 2, 3, 6 & 7), or control IgG Ip (Figure 3(c), lanes 1 to 4). Taken together, arginine/AIF4/AIF4-BP1  M. Umeda et al. / Journal of Biosciences and Medicines 1 (2013) 5-10 Copyright © 2013 SciRes. OPEN ACCESS Figure 4. Arg -beads technology is the best way to purify arginine target factors complex. AIF4-BP1 purification from Arginine-bound AIF4 complex was performed with several different beads of sepharose (Se; lane 2), Ni-sepharose (Ni; lanes 3 and 4) and Arg-beads (Ar; lane 5). Purified AIF4-BP1 protein (panel a) and co-purified controlled recombinant AIF4 C-ter protein (panel b) were shown with Western blot. The summary of results of panel (a) and (b) was shown (c). Only Arg-beads was the methods to purify AIF4-BP1 from Arginine-bound AIF4 proteins complex with single step. complex was observed in vitro and in ER of cell line. 3.4. Arginine-Beads Technology Is the Best Way to Purify Arginine Target Factors (Figure 4) Using arginine immobilized agarose and magnetic nano- beads, the arginine signaling complex was purified effi- ciently. The recombinant hvcAIF4 is the His-tagged p ro- tein, which is easily purified with Ni-sepharose. Instead of arginine-agarose or nanobeads, Ni-seph aro se w as us ed for complex purification. His-tagged rAIF4 protein was similarly purified with Ni-sepharose and arginine beads (Figure 4(b), lanes 3, 4 & 5), although AIF4-BP1 was purified only in arginine beads (Figure 4(a), lane 5). It was not purified by Ni-sepharose (F i gu re 4 (a), lanes 3 & 4), indicating that nanobeads technology is much better arginine signaling complex purification than conventional ways. 4. CONCLUSION The nanobeads technology enables to efficient purify arginine target proteins and cofactors for new drug dis- covery. 5. ACKNO WL EDG EM ENTS We are grateful to our department members in NCGG for helpful dis- cussions. This work was supported by a Grant-in-Aid for the Ministry of Education, Culture, Sports, Science and Technology. REFERENCES [1] Ito, T., Ando, H. , Suzuki, T., Ogura, T., Hotta, K., Im- amura, Y., Yamaguchi, Y. and Handa, H. (2010) Identi- fication of a primary target of thalidomide teratogenicity. Science, 327, 1345-1350. http://dx.doi.org/10.1126/science.1177319 [2] Maekawa, N., Hiramoto, M., Sakamoto, S., Azuma, M., Ito, T., Ikeda, M., Naitou, M., Acharya, H.P., Kobayashi , Y., Suematsu, M., Handa, H. and Ima i, T. (2011) High- performance affinity purification for identification of 15-deoxy-12,14-PGJ2 interacting factors using magnetic nanobeads. Biomedical Chromatography, 25, 466-471. http://dx.doi.org/10.1002/bmc.1469 [3] Uebi, T., U meda, M., Maekawa, N., Karasawa, S., Handa, H. and Imai, T. (2013). Prohibitins, novel vitamin K2 target factors in osteoblast. Journal of Bioscience and Medicine, in Press . [4] Hiramoto, M., Maekawa, N., Kuge, T., Ayabe, F., Wata- nabe, A., Masaike, Y., Hatakeyama, M., Handa, H. and Imai, T. (2010) High-performance affinity chromatogra- phy method for identification of L-arginine interacting factors using magnetic nanobeads. Biomedical Chroma- tography, 24, 606-612. http://dx.doi.org/10.1002/bmc.1469 [5] Morris Jr., M.S. (2006) Arginine: beyond protein. The American Journal of Clinical Nutrition, 83, 508S-512S. [6] Angele, M.K.K., Nitsch, S.M., Hatz, R.A., Angele, P., Hernandez-Richter, T., Wichmann, M.W., Chaudry, I.H. and Schildberg, F.W. (2002) L-Arginine: A unique amino acid for improving depressed wound immune function following hemorrhage. European Surgical Research, 34, 53-60. http://dx.doi.org/10.1159/000048888 [7] Fisker, S., Nielsen, S., Ebdrup, L., Bech, J.N., Christian- sen, J.S., Pedersen, E.B. and Jorgensen, J.O. (1999) The role of nitric oxide in L-arginine-stimulated growth hor- mone release. Journal of Endocrinological Investigation, 22, 89-93. [8] Weinhaus, A.J., Poronnik, P., Tuch, B.E. and Cook, D.I. (1997) Mechanisms of arginine-induced increase in cy- tosolic calcium concentration in the beta-cell line NIT-1. Diabetologia, 40, 374-382. http://dx.doi.org/10.1007/s001250050690 [9] Schlaich, M.P., Parnell, M.M., Ahlers, B.A., Finch, S., Marshall, T., Zhan, W.Z. and Kaye, D.M. (2004) Im- paired L-arginine transport and endothelial function in hypersensive and genetically predisposed normotensive subjects. Circulation, 110, 3680-3686. http://dx.doi.org/10.1161/01.CIR.0000149748.79945.52 [10] Tong, B.C. and Barbul, A. (2004) Cellular and physio- logical effects of arginine. Mini-Reviews in Medicinal Chemistry, 4, 823-832. http://dx.doi.org/10.2174/1389557043403305 [11] Nishio, K., Masaike, Y., Ikeda, M., Narimatsu, H., Goken,  M. Umeda et al. / Journal of Biosciences and Medicines 1 (2013) 5-10 Copyright © 2013 SciRes. OPEN ACCESS N., Tsubouchi, S., Hatakeyama, M., Sakamoto, S., Ha nyu, N., Sandhu, A., Kawaguchi, H., Abe, M. and Handa, H. (2008) Development of novel magnetic nano-carriers for high-performance affinity purification. Colloids and Sur- faces B: Biointerfaces, 64, 162-169. http://dx.doi.org/10.1016/j.colsurfb.2008.01.013 [12] Imai, T., Sumi, Y., Hatakeyama, M., Fujimoto, K., Ka- waguchi, H., Yajima, H. and Handa, H. (1996) Selective isolation of DNA or RNA using single-stranded DNA af- finity latex particles. Journal of Colloid and Interface Science, 177, 245-249. http://dx.doi.org/10.1006/jcis.1996.0027 [13] Shimizu, N., Sugimoto, K., Tang, J., Nishi, T., Sato, I., Hiramoto, M., Aizawa, S., Hatakeyama, M., Ohba, R., Hatori, H., Yoshi kawa, T., Suzuki, F., Oomori, A., Tana- ka, H., Kawaguchi, H., Watanabe, H. and Handa, H. (2000) High-performance affinity beads for identifying drug receptors. Nature Biotechnology, 18, 877-881. http://dx.doi.org/10.1038/78496 [14] Ohtsu, Y., Ohba, R., Imamura, Y., Kobayashi, M., Hatori, H., Zenkoh, T., Hatakeyama, M., Manabe, T., Hino, M., Yamaguchi, Y., Kataoka, K., Kawaguchi, H., Watanabe, H. and Handa, H. (2005) Selective ligand purification us- ing high-performance affinity beads. Analytical Bioche- mistry, 338, 245-252. http://dx.doi.org/10.1038/78496 [15] Imai, T., Matsuda, K., Shimojima, T., Muramatsu, M., Handa, H. and Kato, S. (1997) ERC-55, a binding protein for the papilloma virus E6 oncoprotein, specifically inte- racts with vitamin D receptor among nuclear receptors. Biochemical and Biophysical Research Communications, 233, 765-769. http://dx.doi.org/10.1006/bbrc.1997.6531 [16] Arnold, S.M. and Kaufman, R.J. (2003) The noncatalytic portion of human UDP-glucose: Glycoprotein glucosyl- transferase I confers UDP-glucose binding and transfe- rase function to the catalytic domain. The Journal of Bio- logical Chemistry, 278, 43320-43328. http://dx.doi.org/10.1074/jbc.M305800200
|