Natural Science
Vol.5 No.12(2013), Article ID:41302,6 pages DOI:10.4236/ns.2013.512152
Phylogenetic relationships among small barbus, barbus (actinopterygii: Cyprinidae) from malagarasi and Pangani river basins, Tanzania
![]()
Department of Aquatic Sciences and Fisheries, University of Dar es Salaam, Dar es Salaam, Tanzania; mwitachacha@udsm.ac.tz
Copyright © 2013 Chacha J. Mwita. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In accordance of the Creative Commons Attribution License all Copyrights © 2013 are reserved for SCIRP and the owner of the intellectual property Chacha J. Mwita. All Copyright © 2013 are guarded by law and by SCIRP as a guardian.
Received 21 May 2013; revised 21 June 2013; accepted 28 June 2013
Keywords: Phylogenetic Relationship; Barbus Species; Malagarasi and Pangani Systems; Tanzania
ABSTRACT
The phylogenetic relationships among “Barbus” species (Barbus jacksonii, B. paludinosus and B. trimaculatus) from Malagarasi and Pangani River systems were examined by partial mitochondrial cytochrome b (cyt.b) gene. Sequences of the same samples from elsewhere in Africa obtained from the South African Institute of Aquatic Biodiversity (SAIAB’s) collection were also included in the analysis. Molecular technique was necessary due to phenotypic plasticity and extensive overlap of morphological features among the “Barbus species”. The findings of this study revealed that B. paludinosus from hale and Pangani populations occurred in two distinct genetic lineages though separated by a distance of approximately 10 km apart. Similar results were also noted amongst the population of B. paludinosus in Songati and Muumbara located on Lake Sagara in Malagarasi River basin. B. trimaculatus and B. jacksonii were indistinguishable such that it is envisaged that the two species are one and the same. The Barbus spp. from Africa included in the current study showed genetic affinity to neither European Barbus spp. nor South African redfins of the Pseudobarbus to conclude that the studied species of barbs are taxonomically quite different that needs further investigation to warrant renaming.
1. INTRODUCTION
The genus “Barbus” is a complex one with members distributed widely, occuring in almost every freshwater. The genus comprises of more than 800 species worldwide and about 50 of them are reported from Africa [1]. In Tanzania, the genus barbus is represented by 37 species including: Barbus jacksonii, B. paludinosus and B. trimaculatus by name, but few are found in different habitats such as rivers, streams and Lakes. The genus has high morphological plasticity [2], and contains different levels of ploidy [3] which in most cases delimits identification of the members of the genus “Barbus”. For instance, [4] clearly splits the “Barbus species” into five mitochondrial lineages corresponding to Barbus sensu stricto (tetraploid, which is further subdivided into subgenus Barbus and Luciobarbus), the hexaploid species, the Ethiopian tetraploid species, the African diploid species and the Asian diploid species. On the other hand [5] points out that the genus Barbus includes only the species from Europe, North Africa and Southwestern Asia (peri-Mediterrenean Barbus), and thus the Tropical and South African species (other than those from North Africa) belong to several other genera yet to be taxonomically named. Henceforth, [5] and Skelton (per. Com., 2012) concluded that the current systematic of the genus Barbus can no longer be maintained. However, more barbus species needs to be studied prior to introducing new nomenclature.
This study therefore, aimed at estimating the phylogenetic relationship among the “Barbus species” occurring in Malaragasi and Pangani river basins years posed uplifting of the East African topology, and related the phylogeny to the wider context of the genus elsewhere in Africa, south of the Sahara. This was achieved through the partial sequencing of mitochondrial cytochrome b gene. The cytochrome b gene has, in the past, provided complementary and informative sequence data sets for determining phylogenetic relationships between morphologically similar species. Therefore, it was expected to be a suitable phylogenetic marker to address the questions posed in this study.
2. MATERIALS AND METHODS
2.1. Study Area
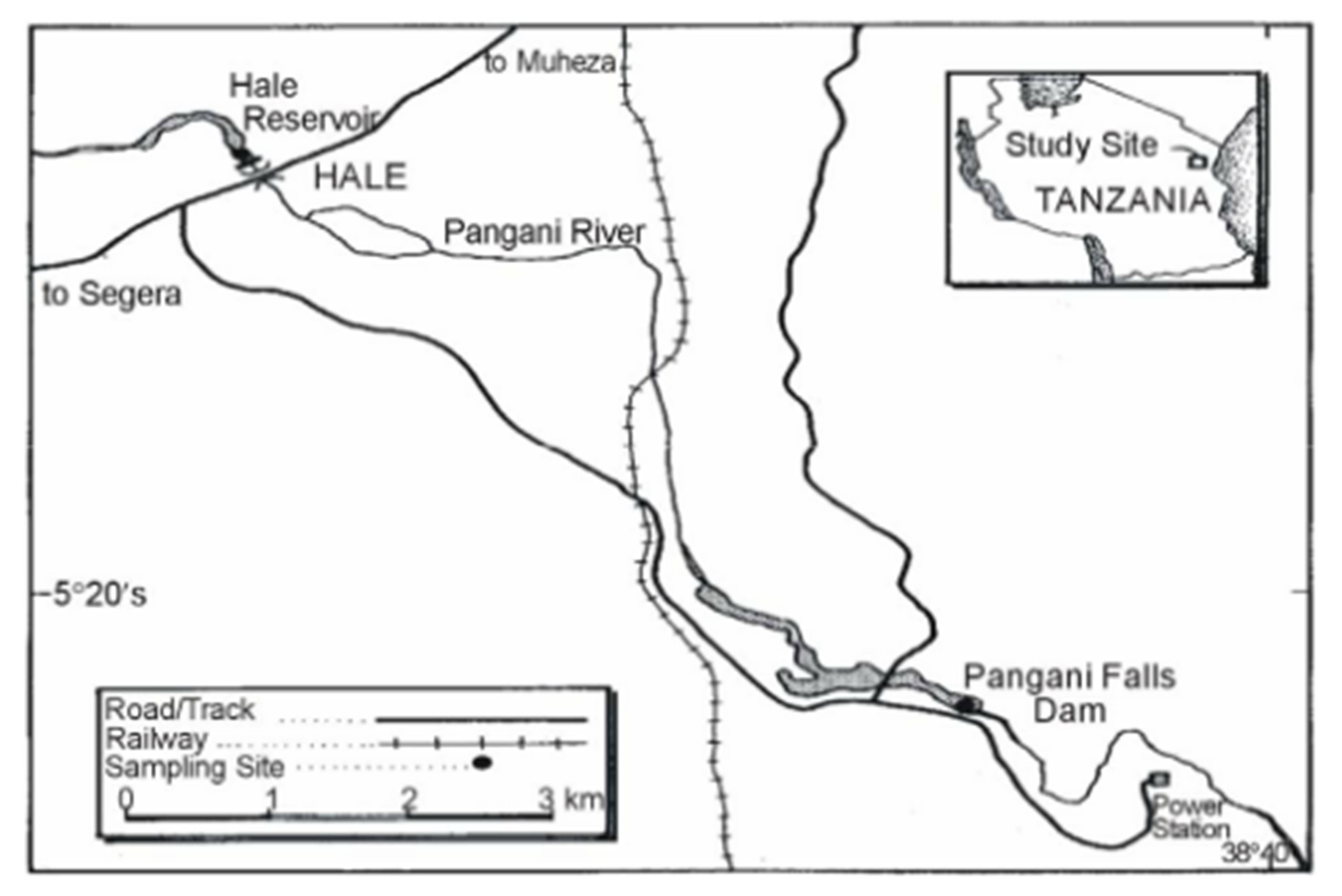
The study was based in Pangani and Malagarasi river basins. The Pangani basin is located in the North Eastern side of Tanzania between latitude 3˚03'S and 5˚59'S and longitude 36˚23'E and 39˚13'E with an area of about 53,600 km2 (Figure 1(a)). The basin is composed of five river sub-catchments, namely Pangani, Zigi, Umba, Mkulumuzi and Msangazi with main rivers draining to the Indian Ocean. Lakes in the basin are Chala, Jipe, Duluti, Manga and Karamba. Nyumba ya Mungu, Mabayani and Kalimawe Dams are manmade reservoirs. On the other hand, the Malagarasi basin is located in the North Western side of Tanzania (Figure 1(b)). Samples were collected from Lakes Sagara and Nyamagoma within the Malagarasi-Muyovozi Wetland systems. The lakes are located 200 km North East of Lake Tanganyika. The sites were chosen due to the fact that they are located in the Malagarasi-Muyovozi Ramsar site which is the first Ramsar site in Africa since 2000. Secondly, the Malagarasi River is one of the Potential fish biodiversity hot spots in Africa [6].
2.2 Collection of Fish Samples
In total 46 individuals of Barbus species from Pangani and Malagarasi rivers, plus 24 DNA sequences of the same species from Mozambique, Zambia, Congo and Malawi used in previous studies obtained from SAIAB’s collection were examined. The fishes were caught by traps, gillnets, monofilaments and some of them were obtained from local fishers.
2.3. Sample Preservation
The obtained samples were soaked into 99.8% alcohol contained in zip bag. The zip bags were placed into the plastic bags and stored in the buckets for easy carrying and transport. In the laboratory, the samples were transferred into zip rock containing 60% - 70% alcohol and stored at room temperature.
2.4. DNA Sources and Extraction
Genomic DNA was extracted from the muscles (on one left side) of each specimen following the protocol Wizard Genomic DNA Purification Kit (Promega). In addition, some other analysis materials were obtained from the collection at South African Institute of Aquatic Biodiversity (SAIAB).
 (a)
(a)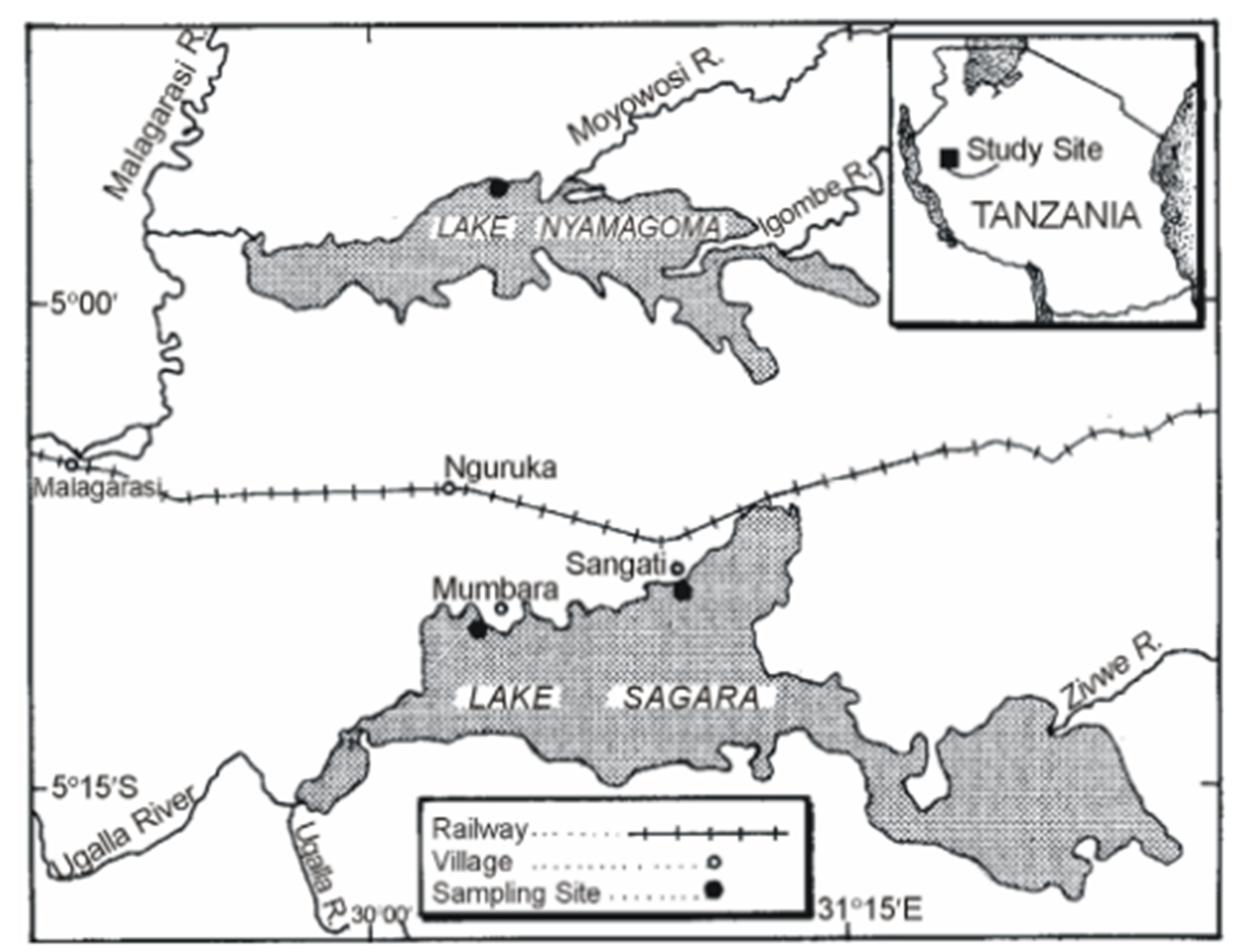 (b)
(b)
Figure 1. Map of Tanzania showing the sampling points at A: Malagarasi and B; Pangani river basins.
2.5. PCR Amplification, Cloning and Sequencing
Two primers (Glu-F, 5’-GAA GAA CCA CCG TTG TTA TTC AA-3’) and (M93R,5’-GAA GAA CCA CCG TTG TTA TCA A-3’) were used to amplify cyt-b gene via polymerase chain reaction (PCR) [7]. PCR reactions were carried out in 50 µl reaction containing 5 µl dNTP (1 mM), 5 µl (10×) reaction buffer, 5 µl (25 mM) MgCl2, 2 µl (10 mM) each primer, 5 µl total DNA, and 0.3 µl Taq DNA polymerase. PCR cycles for amplications were performed using the following conditions: 35 cycles of denaturation at 94˚C for 2 minutes, annealing at 56˚C for 45 minutes, and extension at 72˚C for 2 minutes and 30 seconds. A final extension at 72˚C for 5 minutes was performed to completely extend the amplified product and the reactions were held at 4˚C for an average of 5 minutes. The PCR products were purified with the QIAquick (QIAGEN) kit following manufacturer’s protocol. Each PCR product was sequenced using the BigDye Terminator cycle sequencing ready reaction kit (Applied Biosystems Inc.) on an automated DNA sequencer (Applied Biosystems 377), following manufacturer’s instructions.
2.6. Phylogenetic Analysis
DNA sequences were aligned based on the inferred cytochrome b amino acid sequences. No ambiguous alignments were found and no gaps. All codon positions were included in the phylogenetic analyses. There were a total of 870 positions in the final dataset, out of which 439 were parsimonious informative. Phylogenetic analyses were conducted in MEGA4 [8]. The cytochrome b data set was subjected to the maximum parsimony (MP) method (PAUP* version 4.0 b2a [9] using heuristic searches 1000 replications (TBR branch swapping; MULPARS option in effect) with hundred random stepwise additions of taxa to find the most parsimonious trees. The robustness of the inferred trees was tested by bootstrapping [10,11] using 10,000 replications.
3. RESULTS
Three trees were derived with MP method separating the barbs of Malagarasi and Pangani basins into lineages corresponding to their respective localities. However, there were some striking findings, one individual Barbus paludinosus from the Hale separated from the rest of the group. Similarly the B. paludinosus from Hale completely separated from those of the Pangani falls though sharing the same catchment. The same findings were noted for the Muumbara and Songati populations. Two lineages were also observed for the B. trimaculatus population of Lake Nyamagoma, with one of the group coming closer to B. jacksonii of the Pangani falls (Figure 2).
When sequences of Barbus specimen (SAIAB collection) from southern part of Africa were included in the analysis, the MP method yielded a tree that separated the Barbus paludinosus of Malagarasi and Pangani basins from those of Malawi and Zambia. Similarly Barbus trimaculatus and Barbus jacksonii from the two basins of Tanzania came out independently from those of Mozambique (Figure 3).
Figure 4 combined published sequences of different Barbus species from various African countries and Central Europe obtained from the geneBank for comparison. Cyt.b sequences separated Barbus species into the clade of African Barbus species and the European Barbus species. In this relationship, Barbus paludinosus is highly related to Barbus pleurogramma from L. Tana Ethiopia (93% bootstrap values). The South African redfins such as P. asper, P. quanthlambae, P. burchelii and P. burgi believed to be closely related to small African Barbus species clustered together to form their own clade (78% bootstrap values). The Europe species of the genus Barbus and other related species formed their own clade
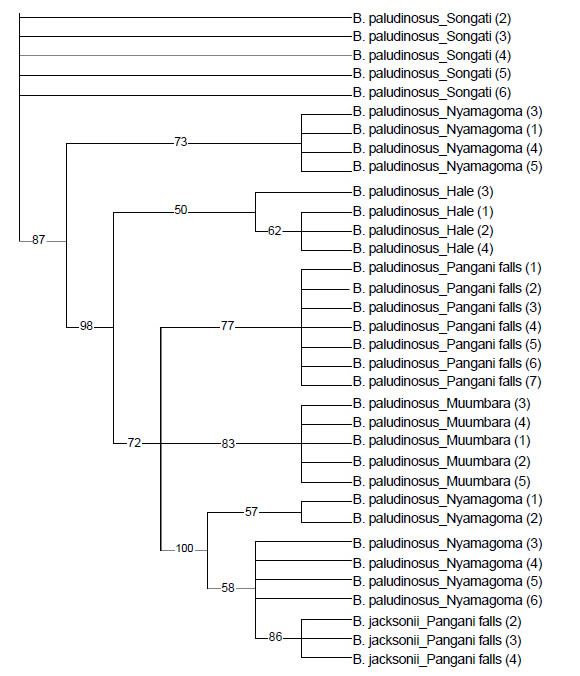
Figure 2. Phylogenetic relationships among species of the genus “Barbus” from Malagarasi and Pangani river systems as estimated by the MP. Numbers in bracket refers to respective sample catalogue from that particular locality.
(Figure 4).
4. DISCUSSION
The phylogenetic relationships of Barbus species from different localities in Malagarasi and Pangani river basins were inferred with cytochrome b nucleotide sequences. Malagarasi and Pangani rivers are believed to belong to the same catchment prior to the uplifting of the Tanzanian central plateau [12,13], and that fish species in the two river systems have had similar intraspecific genetic composition due to interbreeding and natural mixing of their populations [14]. The results obtained in the present study however, showed that the current populations of Barbus species from the two rivers are of two different genetic lineages (Figure 2) due to the consequences of geographical separation of the central plateau in the Miocene era [12].
The population of Barbus species from Hale and Pangani falls located on Pangani River is genetically distinct though the stations are located at approximately twenty kilometers apart. Hale and Pangani falls were built for hydro-electric power generation. The river water from Hale reservoir has been diverted and flows through the turbines of an electric plant and joins the main river at some point before Pangani falls where another intake of
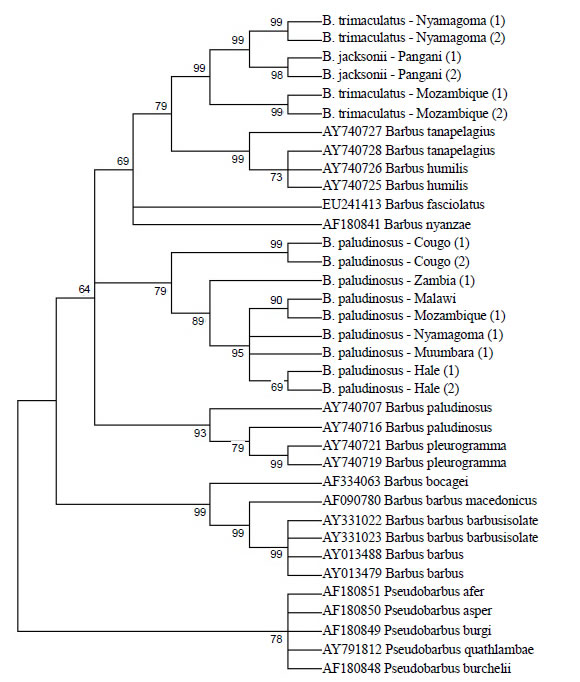

Figure 3. Phylogenetic relationships among the species of the genus “Barbus” from Mozambique, Congo DRC, Tanzania and Zambia estimated Maximum Parsimony. Numbers in bracket refers to respective sample catalogue from that particular locality.
the plant is situated. The interconnection and free movement of Barbus species across the two structures has therefore been broken. The genetic differences observed in this wetland are probably the result of this blockage. This is supported by [15], who stipulate that a habitat discontinuity of only 35m is sufficient to isolate two populations.
There is no direct explanation for the striking genetic differences between the barbs populations from Songati and Muumbara which occur adjacently at Lake Sagara along the Malagarasi River. The observed genetic variation might be due to the massive vegetation cover in Lake Sagara that acts as a barrier for free movement of the small barbs and hence inbreeding segregation. Furthermore, Lake Sagara is said to dry up in 2000 and refilled in 2002 and one of the major changes that it re-

Figure 4. Evolutionary relationships among the 36 taxa from Africa and elsewhere. Numbers in bracket refers to respective sample catalogue from that particular locality.
ceived is the massive vegetation cover. Probably, the species assemblage of Lake Sagara turned out to be oligophyletic, i.e., several (riverine) lineages are likely to have entered the lake habitat when it refilled and radiated in parallel [16]. It also might have brought in certain new species that need further description. Tweddle, (pers.com) believes that there are many new species in Malagarasi River especially of the genus Barbus that are yet to be described.
[6] found major variations in the distribution of oxygen and nutrients between Lakes Nyamagoma and Sagara. These variations can influence the distribution of fauna and flora and which in long term basis can effect genetic differentiation among species. Genetic differences observed in the population of barbs from the two lakes could be the results of these variations. Also differences in rivers that flow into each of the two lakes and the size of surface water mighty influence the biology and nutrients dynamics of the two lakes to further impact the genetic variability among the biotic component of the lakes.
The barbs of the Malagarasi and Pangani wetlands in Tanzania are more related intraspecifically than they do to other barbs from outside Tanzania. For instance, Barbus paludinosus though being widely spread among the barbs [17], genetically they show great variation from one country to another (Figure 3). The reason for the variation observed is not well understood, though it might be that these barbs are different species and not one and the same as currently speculated.
The present study did not support [18] findings who concluded that the African small barbs are closely related to the South African redfins of the genus Pseudobarbus and that the small barbs be placed in the genus pseudobarbus. However, the South African redfins namely; P. asper, P. quanthlambae, P. burchelii and P. burgi clustered distinctly from the other African small barbs (Figure 4) signifying that they belong to a different clade. Barbus tanapelagious and Barbus humilis obtained from the geneBank belong to the same clade, of whether these species are indistinguishable needs further investigation.
[19] considered Barbus trimaculatus the African representation of the western Barbus jacksonii. The findings of the present study noted that Barbus trimaculatus and Barbus jacksonii are more or less morphologically similar and the two species are genetically closely related and fall onto the same subclade in the current analysis (Figures 2 and 3). Whether the two are indistinguishable needs another study. The difference in body colour observed may be the result of environmental pressures [20].
The European species formed their own clade leaving the African species in another clade (Figure 4), this proves that the African Barbus species are genetically distinct and phylogenetically unrelated to the type species of the genus Barbus (Barbus barbus). Therefore, the African Barbus species are not related to central Europen species as per results in Figure 4. Henceforth, taxonomical redescription of the African small barbs is still wanting.
5. CONCLUSION AND RECOMMENDATIONS
Barbus species even though morphologically similar, cyt.b sequence results in the current study revealed two distinct clades of Barbus paludinosus and that of Barbus trimaculatus and Barbus jacksonii. The conclusion on the relationship of Barbus jacksonii and Barbus trimaculatus needs further studies on their cytotaxonomy and biochemistry. The Barbus species are genetically related, i.e. specimens obtained from different localities in the same river, for instance, Malagarasi river, were more closely related to each other than they were related to those from Pangani river wetlands. On the other hand, when the Tanzanian Barbus species were compared to specimens obtained from elsewhere in Africa, they showed genetic affinity and seemed to be very distinct from those of other countries under study. The study also concludes that African Barbus species were distinct from South African redfins and were not related to European species.
6. ACKNOWLEDGEMENTS
I thank NUFUTZ-2007/10227 Project of the University of Dar es Salaam for funding this study. G. B. Kamugisha for collecting the samples and SAIAB for technical support.
REFERENCES
- Eccles, D.H. (1992) FAO species identification sheets for fishery purposes. Field guide to the freshwater fishes of Tanzania. Rome: UNDP Project URT/87/016.
- Doadrio, I. (1990) Phylogenetic relationships and classification of west Palearctic species of the genus Barbus (Osteichthyes, Cyprinidae). Aquatic Living Resources, 3, 265-282. http://dx.doi.org/10.1051/alr:1990028
- Oellermann, L.K. and Skelton, P.H. (1990) Hexaploidy in yellowfish species (Barbus, Pisces, Cyprinidae) from southern Africa. Journal of Fish Biology, 37, 105-115. http://dx.doi.org/10.1111/j.1095-8649.1990.tb05932.x
- Doadrio, I., Bouhandad, R. and Machordom, A. (1998) Genetic differentiation and biogeography in Saharan populations of the genus Barbus (Osteichthyes, Cyprinidae). Folia Zoology, 47, 7-20.
- Turan, D., Kottelat, M. and Ekmekçi, F.G. (2009) Barbus niluferensis, a new species of barbel (Teleostei: Cyprinidae) from Nilüfer River, Turkey, with re-description of B. oligolepis. Zootaxa. Turkish Journal of Fisheries and Aquatic Sciences, 1981, 15-28.
- Nkotagu, H.H. and Ndaro, S.G.M. (2004) The Malagarasi wetland ecosystem: An integrated study. Dar es Salaam University Press, 141.
- Zardoya, R. and Doadrio, I. (1998) Phylogenetic relationships of Iberian cyprinids: Systematic and biogeographical implications. Proceedings of the Royal Society of London, 265(B), 1365-1372. http://dx.doi.org/10.1098/rspb.1998.0443
- Tamura, K., Dudley, J., Nei, M. and Kumar, S. (2007) MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution, 24, 1596-1599. http://dx.doi.org/10.1093/molbev/msm092
- Swofford, D. (2002) Phylogenetic analysis using parsimony (* and other methods) (Ver.4.0b10. ed.). Sinauer Associates, Sunderland.
- Felsenstein, J. (1981) Evolutionary trees from DNA sequences: A maximum likelihood approach. Journal of Molecular Evolution, 17, 368-376.
- Felsenstein, J. (1982) Numerical methods for inferring evolutionary trees. Review Biologie, 57, 1-26. http://dx.doi.org/10.1007/BF01734359
- Johnson, T.C., Scholz, C.A., Talbot, M.R., Kelts, K., Ricketts, R.D., N’gobi, G. and McGill, J.W. (1996) Late Pleistocene desiccation of Lake Victoria and rapid evolution of cichlid fishes. Science, 273, 1091-1093. http://dx.doi.org/10.1126/science.273.5278.1091
- Odada, E.O. and Olago, D.O. (2002) The East African Great Lakes: Limnology, palaeolimnology and biodiversity. Kluwer, London. http://dx.doi.org/10.1007/0-306-48201-0
- Stankiewiz, J. and de Wit, M.J. (2006) A proposed drainage evolution model for central Africa—Did the Congo flow East? Journal of African Earth Sciences, 44, 75-84. http://dx.doi.org/10.1016/j.jafrearsci.2005.11.008
- Rico, C. and Turner, G.F. (2002) Extreme microallopatric divergence in a cichlid species from Lake Malawi. Molecular Ecology, 11, 1585-1590. http://dx.doi.org/10.1046/j.1365-294X.2002.01537.x
- Salzburger, W. and Meyer, A. (2004) The species flocks of East African cichlid fishes: Recent adces in molecular phylogenetics and population genetics. Published online: 20 April Springer-Verlag. Naturwissenschaften Journal of Ichyology, 91, 277-290.
- Skelton, P.H. (1993) A complete guide to the freshwater fishes of Southern Africa. Southern Book Publishers, Johannesburg.
- Seegers, L. (1996) The fishes of the Lake Rukwa drainage. Tervuren, Belgium.
- Greenwood, P.H. (1962) A revision of certain Barbus (Pisces: Cyprinidae) from East, Central and South Africa. Bulletin of the British Museum of natural History, 8, 151- 208.
- Skelton, P.H. (2001) A complete guide to the freshwater fishes of southern Africa. 2nd Edition, Struik, Cape Town.

