Advances in Bioscience and Biotechnology
Vol.5 No.2(2014), Article ID:42416,9 pages DOI:10.4236/abb.2014.52015
A comprehensive meta-analysis of the association between three IL1B polymorphisms and rheumatoid arthritis
1Zhejiang Provincial Key Laboratory of Pathophysiology, School of Medicine, Ningbo University, Ningbo, China
2The Affiliated Hospital, Ningbo University, Ningbo, China
3Bank of Blood Products, Ningbo No.2 Hospital, Ningbo, China
Email: #yemeng@nbu.edu.cn, #duanshiwei@nbu.edu.cn
Copyright © 2014 Dongjun Dai et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In accordance of the Creative Commons Attribution License all Copyrights © 2014 are reserved for SCIRP and the owner of the intellectual property Dongjun Dai et al. All Copyright © 2014 are guarded by law and by SCIRP as a guardian.
Received 3 November 2013; revised 18 December 2013; accepted 5 January 2014
KEYWORDS
Rheumatoid Arthritis; Meta-Analysis; Polymorphism; IL1B-511; Dominant Model
ABSTRACT
Rheumatoid arthritis (RA) is an immune-mediated chronic inflammatory disease that causes huge destruction to human body. IL1B encodes key mediator IL-1β protein, which plays an important role in the pathogenesis of inflammatory syndromes. The aim of this study was to evaluate the association between IL1B polymorphisms and RA. A meta-analysis was performed on the association between three IL1B polymorphisms (IL1B-31: rs1143627; IL1B-511: rs16944; IL1B + 3954: rs1143634) and RA. A trend of significant association was observed between IL1B + 3954 and RA (p = 0.06, odd ratio (OR) = 1.19, 95% confidential interval (CI) = 1.00 - 1.42). A significant association was found in Europeans under the dominant model between IL1B-511T and RA (p = 0.03, OR = 0.89, 95% CI = 0.81 - 0.99). Our meta-analysis indicated that IL1B − 511-T played a protective role against RA in Europeans, and that IL1B + 3954-T had the potential to increase the risk of RA. Future large-scale studies should be considered to confirm the association between IL1B polymorphisms and RA.
1. INTRODUCTION
Rheumatoid arthritis (RA) is an immune-mediated chronic inflammatory disease [1] that can lead to low bone mineral density [2], depression [3], obstructive lung disease [4] and Cardiovascular diseases [5], causing huge destruction to human body. Twin studies estimated that heritability of RA liability was up to 60% [6]. Familybased studies demonstrated that genetic factors played a more important role in the development of RA than environmental factors did [7,8].
IL1B encodes IL-1β that is one of the distinct polypeptides molecules of IL-1, a key mediator in the pathogenesis of inflammatory syndromes such as RA [9]. IL1B is 7020 bp in length and contains 826 polymorphisms according to the NCBI dbSNP database. Among them, IL1B-31 [10-14], IL1B-511 [9-11,14-26] and IL1B + 3954 [9-11,14,16-23,25-29] are the most studied in the association with RA.
Inconsistent results exist in the current association between IL1B variants and RA. For IL1B-31, there were 1 study with significant association result in European population [10] and 4 studies with non-significant association results in European [11-13] and Asian populations [14]. For IL1B-511, there were 3 significant comparisons in European population [10, 21, 24] and 13 nonsignificant comparisons in European [9,11,15,16,18,19, 22,26], Asian [14,17,23], Latin American [20] and African [25] populations. For IL1B+3954, there were 3 significant comparisons in European [21] and Asian [23,29] populations and 15 non-significant comparisons in European [9-11,16,18,19,21,22,26-28], Asian [14,17], and African populations [25].
Discrepancy among the association studies might be due to the different ethnic background, inefficient sample size [30], or the uncorrected physiological status among the association studies [31]. Meta-analysis is often used to enhance statistical power and to draw a more convincing conclusion by pooling up the research data from individual association study [32]. The goals of our metaanalyses were to find out the causes of the above inconsistent findings among various case-control association studies, and to evaluate the contribution of IL1B polymorphisms to RA.
2. MATERIALS AND METHODS
2.1. Data Collection
A systematic literature searching was performed in PubMed/MEDLINE without language restriction, using the keywords “rheumatoid arthritis IL1Bassociation” and “rheumatoid arthritis IL1B polymorphism” to identify available articles. We also checked Chinese databases (WanFang, WeiPu and CNKI) using the same keywords. The inclusion criteria of the literatures for the metaanalyses comprise the following items: (1) It was an original case-control study with an assessment of the association between IL1B polymorphisms and RA risk in humans; (2) It contains sufficient information to infer the odd ratios (ORs) and 95% confidential intervals (95% CI); (3) Genotype distribution of each polymorphism in controls met Hardy-Weinberg equilibrium (HWE). All of the association studies between IL1B polymorphisms and RA were fully considered and carefully selected in July 2013. We extracted or calculated the following information from each study: Genetic locus, first author’s name, year of publication, ethnicity, numbers of cases and controls, control source, HWE for controls, the result of individual studies about the association of IL1B − 31, IL1B − 511 and IL1B + 3954 with RA and power analysis for each of the involved studies.
2.2. Statistical Analysis
Arlequin program was used to test HWE [33]. Power and Sample Size Calculation program was applied to calculate the power of each study [34]. Review Manager 5 was used for the meta-analysis [35]. Statistical heterogeneity was tested using Cochran’s Q statistic and I² test [36] to decide the type of analysis to be used in the metaanalysis. For the studies with minimal to moderate heterogeneity (I2 < 50%), the fixed-effect model would be used for the meta-analysis. For the studies with significant heterogeneity (I2 > = 50%), the random-effect model would be used. Funnel plots are also drawn to observe the potential publication bias.
3. RESULT
3.1. Data Collection
As shown in Figure 1, 10 relevant studies were involved
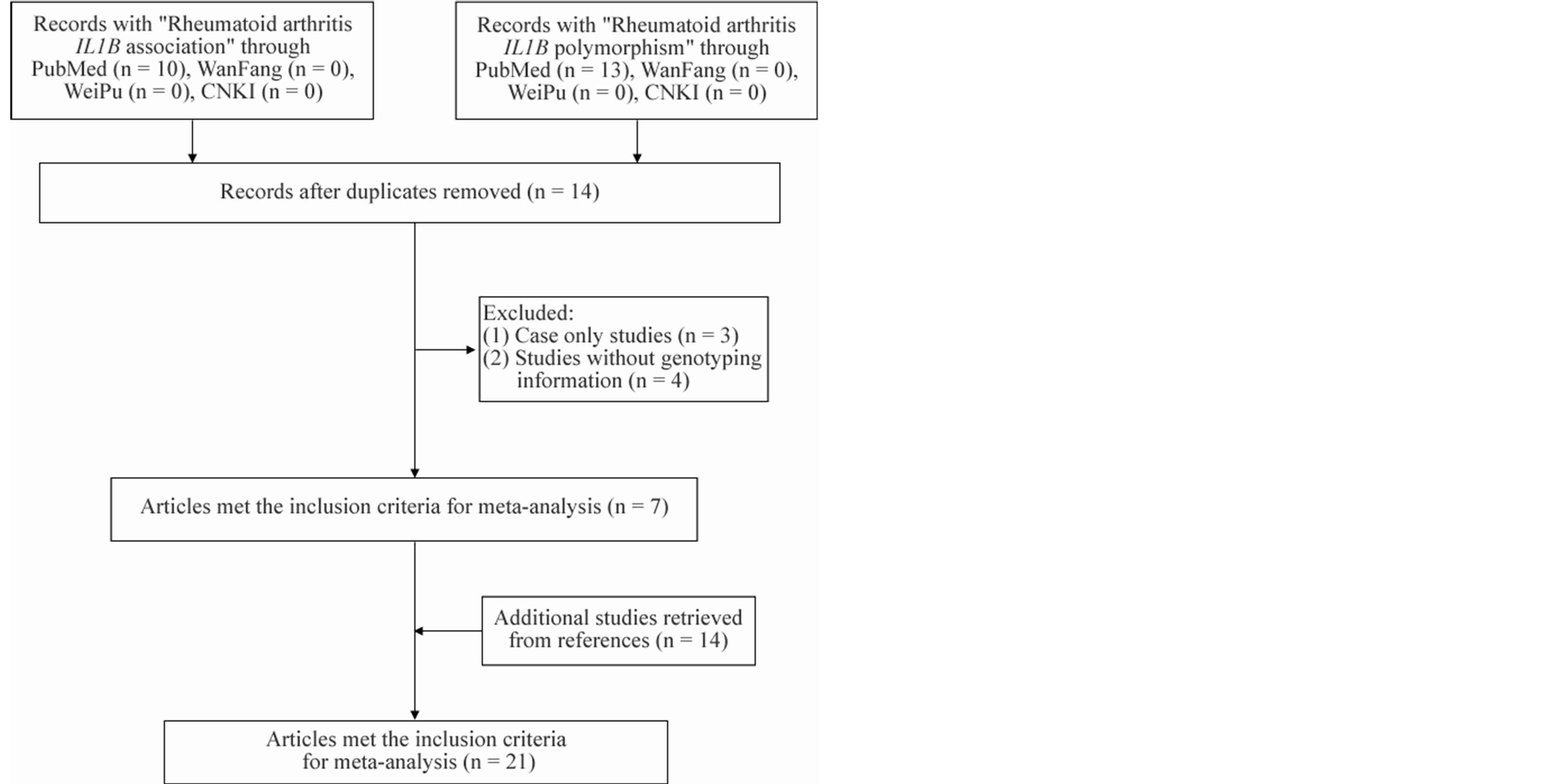
Figure 1. Flowchart of selection process for meta-analyses.
using Pubmed through the keywords “Rheumatoid arthritis IL1B association”, and 13 relevant studies were involved through the words “Rheumatoid arthritis IL1B polymorphism”. No relative study was found in Chinese databases (WanFang, WeiPu and CNKI). Among the 23 retrieved articles, we excluded 9 duplicates, 4 case-only studies [37-40], and 3 studies [41-43] for a lack of allele or genotype information. In addition, 14 additional studies [9,15-24,27-29] were retrieved from the references. Finally, a total of 21 studies [9-29] were included in our meta-analysis. The distribution of genotype in the controls met HWE (p > 0.05) in all comparisons except for one [20] with significant deviation from HWE in controls (p < 0.05) (Table 1). At last, there were 2214 RA patients and 2466 controls among 5 comparisons for the meta-analysis of IL1B-31 (rs1143627), 4491 RA patients and 4006 controls among 16 comparisons for the metaanalysis of IL1B-511 (rs16944) in 7 studies, and 4338 RA patients and 3742 compared controls among 16 comparisons for the meta-analysis of IL1B + 3954 (rs1143634) (Table 2).
3.2. Meta-Analyses of Three Polymorphisms and RA Risk
As showed in Table 2, among the overall analysis, a trend of significant association was observed between IL1B + 3954-T and RA (p = 0.06, OR = 1.19, 95% CI = 1.00 - 1.42, Figure 2, Table 2). And there was a significant heterogeneity for IL1B + 3954-T (p = 0.0003, I2 = 65%, Figure 2, Table 2). A further subgroup meta-analysis under the dominant model identified a significant association of IL1B-511-T and RA (p = 0.03, OR = 0.89, 95% CI = 081 - 0.99, Figure 2, Table 2). No publication
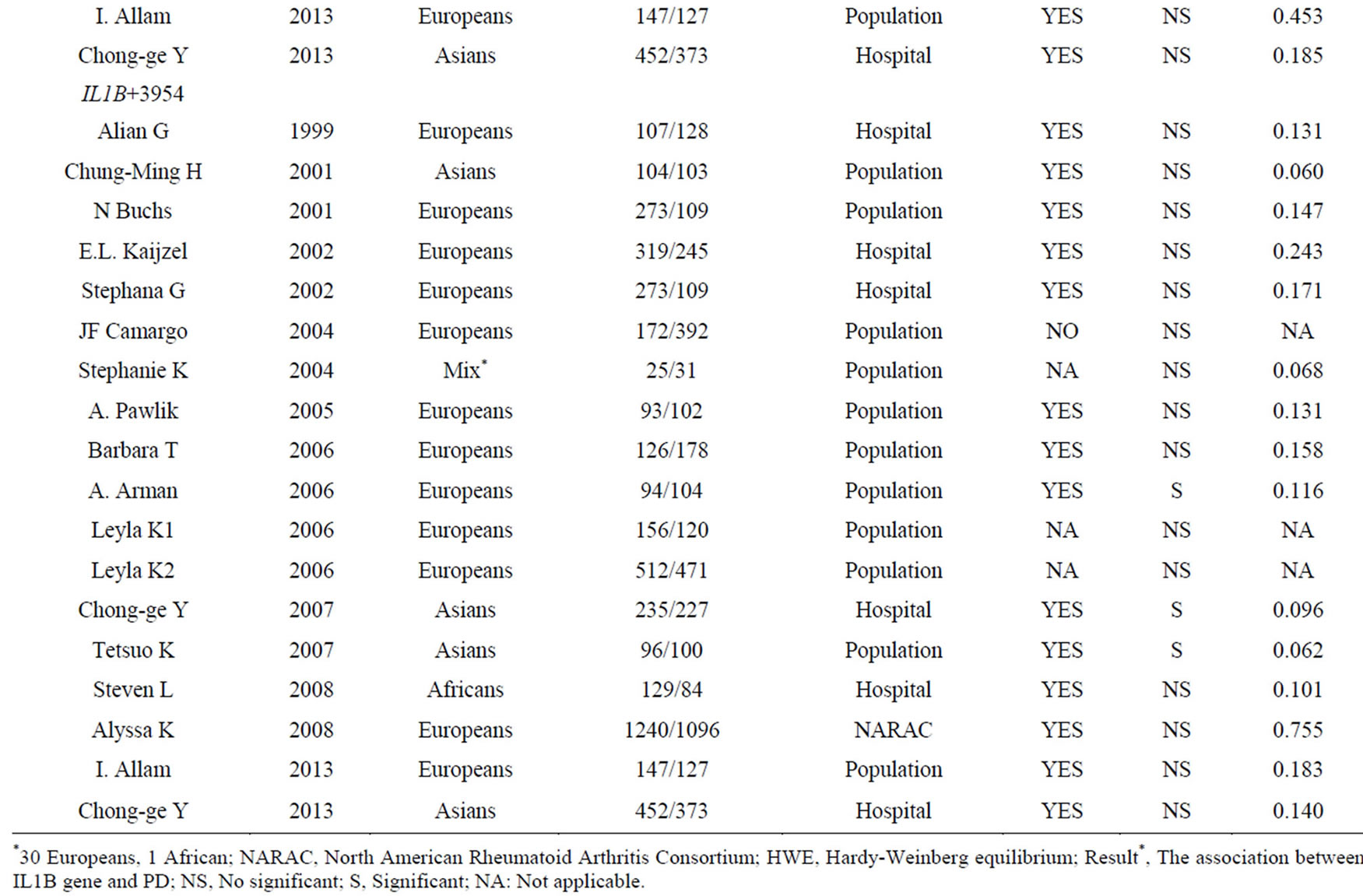

Table 1. Characteristics of studies in the meta-analyses of IL1B − 31, IL1B − 511 and IL1B + 3954 polymorphisms with RA.
.
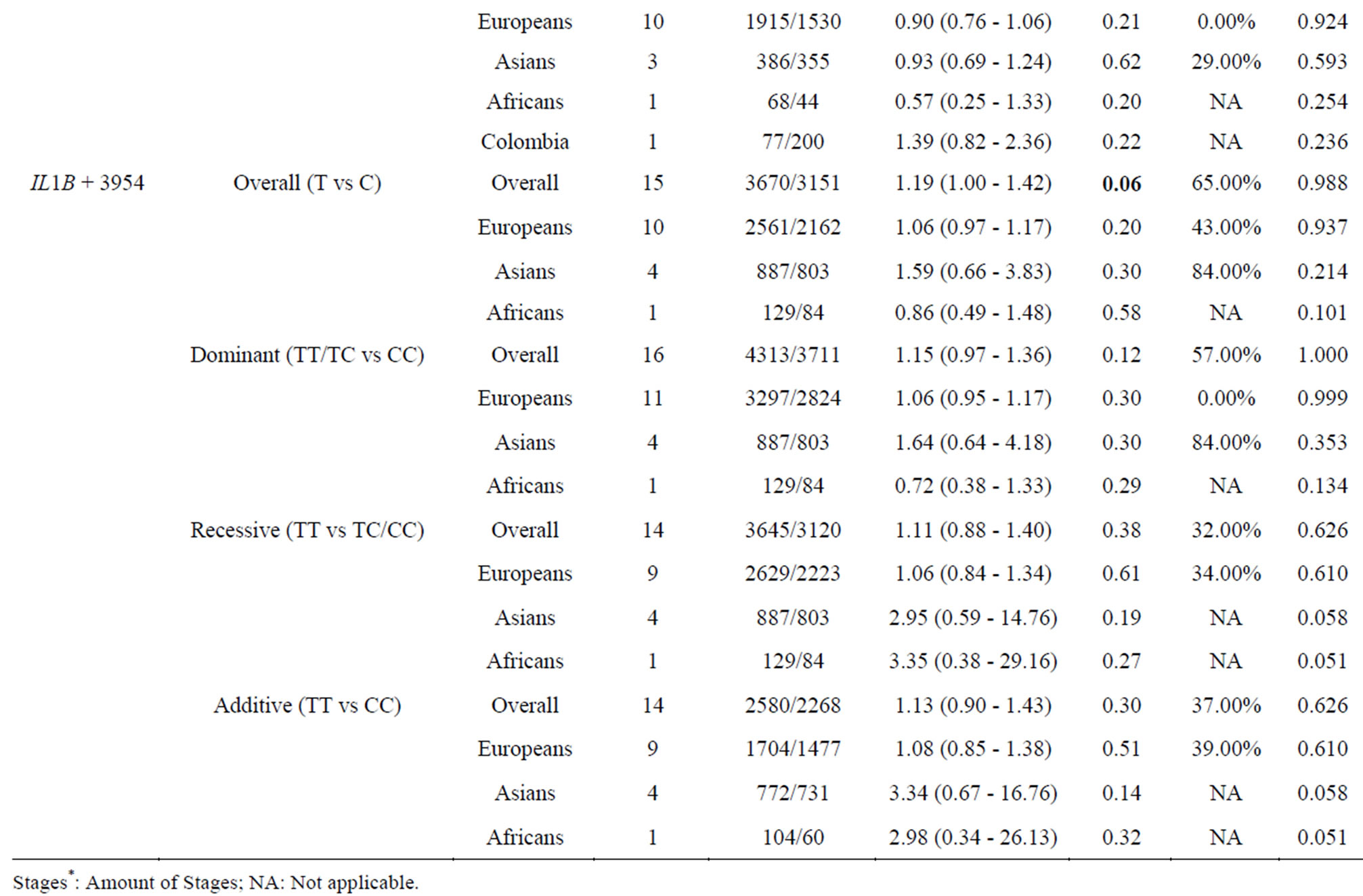

Table 2. Meta-analyses of the IL1B − 31, IL1B − 511 and IL1B + 3954 polymorphisms with RA.
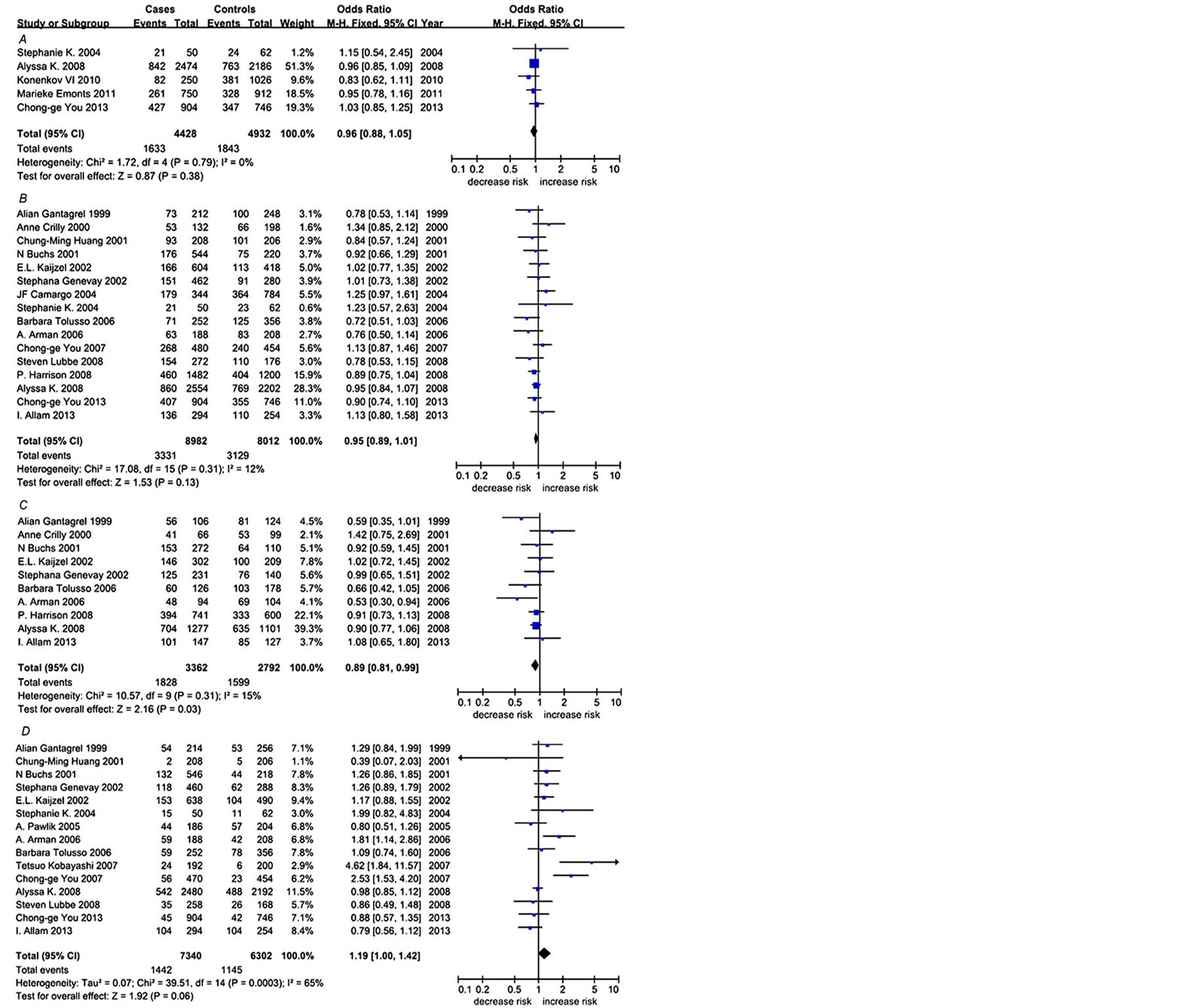
Figure 2. Forest plots of the three SNPs with RA. A) Forest plot of IL1B − 31 in overall analysis; B) Forest plot of IL1B − 511 in overall analysis; C) Forest plot of Dominant model of IL1B − 511 in Europeans; D) Forest plot of IL1B + 3954 in overall analysis.
bias was found for the meta-analyses of the three SNPs (Figure 3).
4. DISCUSSION
In the current meta-analyses, we summarized the associations of three IL1B variants with RA from 21 studies (22 stages) among 5888 cases and 5760 controls. Our results showed a trend of association between IL1B + 3954-T and RA (Table 2 and Figure 2) and a significant association under the dominant model between IL1B- 511-T and RA in Europeans (Table 2 and Figure 2).
Single nucleotide polymorphisms (SNPs) occur in a high frequency in the human genome, which may affect the function of genes [44]. IL1B + 3954 in the exon 5 and IL1B − 511 in the promoter are two key polymorphisms of IL1B that play an important role in inflammatory diseases [9]. Previous studies proved that IL1B-511-T in-
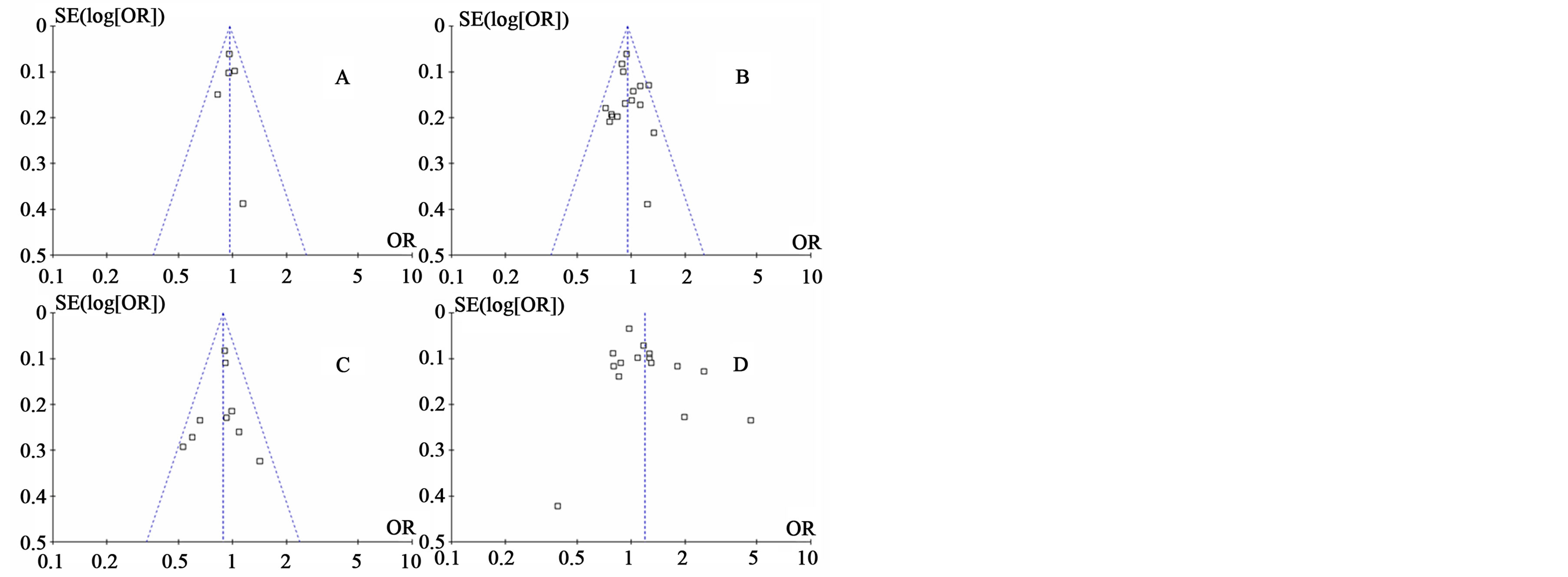
Figure 3. Funnel plots of three SNPs with RA. A) Funnel plot of IL1B − 31 in overall analysis; B) Funnel plot of IL1B − 511 in overall analysis; C) Funnel plot of Dominant model of IL1B − 511 in Europeans; D) Funnel plot of IL1B + 3954 in overall analysis.
creased LPS-induced IL-1β production by 2 - 3 folds and showed higher levels of IL-1Ra [45]. Different conclusions were shown for IL1B + 3954-T. Some researches indicated that it might increase plasma levels of IL-1β [10,46], but some others found it had no influence or reduced IL-1 levels [16,20,47]. IL1B − 511 and IL1B + 3954 showed a wide association with diseases like gastric cancer [48,49], breast cancer [50], aspirin-tolerant asthma [51], left ventricular systolic dysfunction [52], hip osteoarthritis [53] and RA [10,21-24,29].
Several other RA association studies observed dominant effect among a handful of SNPs such as −607A/C polymorphism of IL-18 gene [54], −670A/G polymorphism of FAS gene [55], rs1343151 and rs10489629 of IL-23R gene [56] and −173G/C polymorphism MIF gene [57]. The significant association of IL1B-511-T polymorphism under the dominant model may provide a new hint in the pathogenesis of RA.
Significant heterogeneity showed in the overall analysis (I2 = 65%, Table 2) and dominant model (I² = 57%, Table 2) of IL1B + 3954. A subgroup study by ethnicity (Table 2) showed that significant heterogeneity was only found in Asians (I2 = 56% in dominant model of IL1B − 511 in Europeans, I2 = 84% in overall analysis and dominant model of IL1B + 3954 in Europeans). Frequency of IL1B-511-C and in Asians (Hapmap-HCB) is 0.547 that is lower to 0.642 in Europeans (Hapmap-CEU). And the allele frequency of IL1B+3954-C and in Asians (Hapmap-HCB) is 0.988 that is much higher to 0.792 in Europeans (Hapmap-CEU). A further analysis of the two polymorphisms showed a non-significant ethnic difference between Asians and Europeans (IL1B − 511: Fst = 0.0094; IL1B + 3954: Fst = 0.0988). A further power analysis suggested there was a lack of power for the subgroup meta-analysis in Asians (power = 0.701 in IL1B- 511, power = 0.214 in IL1B + 3954, Table 2), suggesting that the non-significant association in Asians might be due to the small sample size in the existing case-control association studies in Asian population. In contrast, the power in the meta-analysis in European populations for IL1B − 511 and IL1B + 3954 polymorphisms are 0.998 and 0.937 (Table 2).
Compared with the previous two meta-analysis studies [24,58] about the polymorphisms of IL1B and RA, our meta-analyses included 13 and 8 more case-control studies than the studies by P. Harrison et al. [24] and Young LEE et al. [58], respectively. Our research showed that IL1B − 511 was significantly associated with RA in Europeans under the dominant model, and a trend association of IL1B + 3954 with RA. Moreover, our study grouped Turkish population into Caucasians instead of Asians in the subgroup meta-analysis according to the fact that the ancestors of major Turkish population were from Europe [59-61]. We performed HWE test for the controls in all the involved studies, and excluded one [20] that was included in previous meta-analysis [58]. With an enhanced power and stricter selection criteria, our meta-analyses produced a more reliable conclusion than the previous meta-analysis studies.
However, our study presented several limitations that needed to be carefully considered. Firstly, there are only a limited number of associations in non-Caucasian populations. A lack of power in the non-Caucasian studies suggested that non-significant results in Asians and other population needed to be taken with caution. Future studies with larger samples size are required to establish the association of IL1B polymorphisms with RA. Secondly, RA is a complex disease that different physiological status of RA may exist in the cases. All the existing casecontrol studies didn’t perform a stratified analysis by the RA disease stage. This may partially explain the discrepancies in the current case-control studies. Thirdly, genetic heterogeneity may exist in IL1B since there are 826 known IL1B polymorphisms. Our meta-analyses only focused on three IL1B SNPs that might not fully represent the overall contribution of IL1B variations. Other IL1B polymorphisms needed to be analyzed for their contribution to RA in the future. Fourthly, the positive findings of current study might not reach a very precise statistical significance by the certain extent multiple testing in our analyses.
In conclusion, our meta-analysis observed a trend association of IL1B + 3954-T with RA and a significant association under the dominant model between IL1B − 511-T and RA in Europeans. Further researches are required to confirm our findings and to discover the underlying mechanisms of other polymorphisms of IL1B that might contribute to the risk of RA.
ACKNOWLEDGEMENTS
The research was supported by the grants from: National Natural Science Foundation of China (31100919, 81371469), Natural Science Foundation of Zhejiang Province (LR13H020003), K. C. Wong Magna Fund in Ningbo University, and Ningbo social development research projects (2012C50032).
REFERENCES
- Bossaller, L. and Rothe, A. (2013) Monoclonal antibody treatments for rheumatoid arthritis. Expert Opinion on Biological Therapy, 2013.
- Chen, J., Liu, W., Lin, Q., et al. (2013) Vitamin D deficiency and low bone mineral density in native Chinese rheumatoid arthritis patients. International Journal of Rheumatic Diseases, 2013.
- Tillmann, T., Krishnadas, R., Cavanagh, J. and Petrides, K. (2013) Possible rheumatoid arthritis subtypes in terms of rheumatoid factor, depression, diagnostic delay and emotional expression: An exploratory case-control study. Arthritis Research & Therapy, 15, R45. http://dx.doi.org/10.1186/ar4204
- Nannini, C., Medina-Velasquez, Y.F., Achenbach, S.J., et al. (2013) Incidence and mortality of obstructive lung disease in rheumatoid arthritis: A population-based study. Arthritis Care & Research (Hoboken), 65, 1243-1250. http://dx.doi.org/10.1002/acr.21986
- Koivuniemi, R., Paimela, L., Suomalainen, R. and Leirisalo-Repo, M. (2013) Cardiovascular diseases in patients with rheumatoid arthritis. Scandinavian Journal of Rheumatology, 42, 131-135. http://dx.doi.org/10.3109/03009742.2012.723747
- MacGregor, A.J., Snieder, H., Rigby, A.S., et al. (2000) Characterizing the quantitative genetic contribution to rheumatoid arthritis using data from twins. Arthritis & Rheumatism, 43, 30-37. http://dx.doi.org/10.1002/1529-0131(200001)43:1<30::AID-ANR5>3.0.CO;2-B
- Wordsworth, P. and Bell, J. (1991) Polygenic susceptibility in rheumatoid arthritis. Annals of the Rheumatic Diseases, 50, 343-346. http://dx.doi.org/10.1136/ard.50.6.343
- Choi, S.J., Rho, Y.H., Ji, J.D., Song, G.G. and Lee, Y.H. (2006) Genome scan meta-analysis of rheumatoid arthritis. Rheumatology (Oxford), 45, 166-170. http://dx.doi.org/10.1093/rheumatology/kei128
- Cantagrel, A., Navaux, F., Loubet-Lescoulie, P., et al. (1999) Interleukin-1beta, interleukin-1 receptor antagonist, interleukin-4, and interleukin-10 gene polymorphisms: Relationship to occurrence and severity of rheumatoid arthritis. Arthritis & Rheumatism, 42, 1093-1100. http://dx.doi.org/10.1002/1529-0131(199906)42:6<1093::AID-ANR5>3.0.CO;2-P
- Hall, S.K., Perregaux, D.G., Gabel, C.A., et al. (2004) Correlation of polymorphic variation in the promoter region of the interleukin-1 beta gene with secretion of interleukin-1 beta protein. Arthritis & Rheumatism, 50, 1976- 1983. http://dx.doi.org/10.1002/art.20310
- Johnsen, A.K., Plenge, R.M., Butty, V., et al. (2008) A broad analysis of IL1 polymorphism and rheumatoid arthritis. Arthritis & Rheumatism, 58, 1947-1957. http://dx.doi.org/10.1002/art.23592
- Konenkov, V.I., Golovanova, O.V., Prokof'ev, V.F., et al. (2010) Distribution of allelic variants of promotor sites of cytokine genes and endothelial growth factor gene among healthy subjects and patients with rheumatoid arthritis in a Russian Europeoid population. Vestnik Rossiiskoi Akademii Meditsinskikh, 2010, 9-14.
- Emonts, M., Hazes, M.J., Houwing-Duistermaat, J.J., et al. (2011) Polymorphisms in genes controlling inflammation and tissue repair in rheumatoid arthritis: A case control study. BMC Medical Genetics, 12, 36. http://dx.doi.org/10.1186/1471-2350-12-36
- You, C.G., Li, X.J., Li, Y.M., et al. (2013) Association analysis of single nucleotide polymorphisms of proinflammatory cytokine and their receptors genes with rheumatoid arthritis in northwest Chinese Han population. Cytokine, 61, 133-138. http://dx.doi.org/10.1016/j.cyto.2012.09.007
- Crilly, A., Maiden, N., Capell, H.A. and Madhok, R. (2000) Predictive value of interleukin 1 gene polymorphisms for surgery. Annals of the Rheumatic Diseases, 59, 695-699. http://dx.doi.org/10.1136/ard.59.9.695
- Buchs, N., di Giovine, F.S., Silvestri, T., et al. (2001) IL-1B and IL-1Ra gene polymorphisms and disease severity in rheumatoid arthritis: Interaction with their plasma levels. Genes & Immunity, 2, 222-228. http://dx.doi.org/10.1038/sj.gene.6363766
- Huang, C.M., Tsai, F.J., Wu, J.Y. and Wu, M.C. (2001) Interleukin-1beta and interleukin-1 receptor antagonist gene polymorphisms in rheumatoid arthritis. Scandinavian Journal of Rheumatology, 30, 225-228. http://dx.doi.org/10.1080/030097401316909576
- Kaijzel, E.L., van Dongen, H., Bakker, A.M., et al. (2002) Relationship of polymorphisms of the Interleukin-1 gene cluster to occurrence and severity of rheumatoid arthritis. Tissue Antigens, 59, 122-126. http://dx.doi.org/10.1034/j.1399-0039.2002.590208.x
- Genevay, S., Di Giovine, F.S., Perneger, T.V., et al. (2002) Association of interleukin-4 and interleukin-1B gene variants with Larsen score progression in rheumatoid arthritis. Arthritis & Rheumatism, 47, 303-309. http://dx.doi.org/10.1002/art.10394
- Camargo, J.F., Correa, P.A., Castiblanco, J. and Anaya, J.M. (2004) Interleukin-1beta polymorphisms in Colombian patients with autoimmune rheumatic diseases. Genes & Immunity, 5, 609-614. http://dx.doi.org/10.1038/sj.gene.6364133
- Arman, A., Yilmaz, B., Coker, A., Inanc, N. and Direskeneli, H. (2006) Interleukin-1 receptor antagonist (IL- 1RN) and interleukin-1B gene polymorphisms in Turkish patients with rheumatoid arthritis. Clinical and Experimental Rheumatology, 24, 643-648.
- Tolusso, B., Pietrapertosa, D., Morelli, A., et al. (2006) IL-1B and IL-1RN gene polymorphisms in rheumatoid arthritis: Relationship with protein plasma levels and response to therapy. Pharmacogenomics, 7, 683-695. http://dx.doi.org/10.2217/14622416.7.5.683
- You, C.G., Li, J.F., Xie, X.D., et al. (2007) Association of interleukin-1 genetic polymorphisms with the risk of rheumatoid arthritis in Chinese population. Clinical Chemistry and Laboratory Medicine, 45, 968-971. http://dx.doi.org/10.1515/CCLM.2007.156
- Harrison, P., Pointon, J.J., Chapman, K., Roddam, A. and Wordsworth, B.P. (2008) Interleukin-1 promoter region polymorphism role in rheumatoid arthritis: A meta-analysis of IL-1B-511A/G variant reveals association with rheumatoid arthritis. Rheumatology (Oxford), 47, 1768- 1770. http://dx.doi.org/10.1093/rheumatology/ken374
- Lubbe, S., Tikly, M., van der Merwe, L., Hodkinson, B. and Ramsay, M. (2008) Interleukin-1 receptor antagonist gene polymorphisms are associated with disease severity in Black South Africans with rheumatoid arthritis. Joint Bone Spine, 75, 422-425. http://dx.doi.org/10.1016/j.jbspin.2007.09.015
- Allam, I., Djidjik, R., Ouikhlef, N., et al. (2013) Interleukin-1 and the interleukin-1 receptor antagonist gene polymorphisms study in patients with rheumatoid arthritis. Pathologie Biologie (Paris), 2013.
- Pawlik, A., Kurzawski, M., Florczak, M., Gawronska Szklarz, B. and Herczynska, M. (2005) IL1beta+3953 exon 5 and IL-2 -330 promoter polymorphisms in patients with rheumatoid arthritis. Clinical and Experimental Rheumatology, 23, 159-164.
- Kazkaz, L., Marotte, H., Hamwi, M., et al. (2007) Rheumatoid arthritis and genetic markers in Syrian and French populations: different effect of the shared epitope. Annals of the Rheumatic Diseases, 66, 195-201. http://dx.doi.org/10.1136/ard.2004.033829
- Kobayashi, T., Ito, S., Kuroda, T., et al. (2007) The interleukin-1 and Fcgamma receptor gene polymorphisms in Japanese patients with rheumatoid arthritis and periodontitis. Journal of Periodontology, 78, 2311-2318. http://dx.doi.org/10.1902/jop.2007.070136
- Button, K.S., Ioannidis, J.P., Mokrysz, C., et al. (2013) Power failure: Why small sample size undermines the reliability of neuroscience. Nature Reviews Neuroscience, 14, 365-376. http://dx.doi.org/10.1038/nrn3475
- Xu, W.D., Zhou, M., Peng, H., Pan, H.F. and Ye, D.Q. (2013) Lack of association of IL-6 polymorphism with rheumatoid arthritis/type 1 diabetes: A meta-analysis. Joint Bone Spine, 2013.
- Tao, J.H., Zou, Y.F., Feng, X.L., et al. (2011) Meta-analysis of TYK2 gene polymorphisms association with susceptibility to autoimmune and inflammatory diseases. Molecular Biology Reports, 38, 4663-4672. http://dx.doi.org/10.1007/s11033-010-0601-5
- Excoffier, L., Laval, G. and Schneider, S. (2005) Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evolutionary Bioinformatics Online, 1, 47-50.
- Gibson, E., Fenster, A. and Ward, A.D. (2013) The impact of registration accuracy on imaging validation study design: A novel statistical power calculation. Medical Image Analysis, 17, 805-815. http://dx.doi.org/10.1016/j.media.2013.04.008
- Kawalec, P., Mikrut, A., Wisniewska, N. and Pilc, A. (2013) The effectiveness of tofacitinib, a novel Janus kinase inhibitor, in the treatment of rheumatoid arthritis: A systematic review and meta-analysis. Clinical Rheumatology, 2013.
- Coory, M.D. (2010) Comment on: Heterogeneity in metaanalysis should be expected and appropriately quantified. International Journal of Epidemiology, 39, 932. http://dx.doi.org/10.1093/ije/dyp157
- Marotte, H., Farge, P., Gaudin, P., Alexandre, C., Mougin, B. and Miossec, P. (2006) The association between periodontal disease and joint destruction in rheumatoid arthritis extends the link between the HLA-DR shared epitope and severity of bone destruction. Annals of the Rheumatic Diseases, 65, 905-909. http://dx.doi.org/10.1136/ard.2005.036913
- Pawlik, A., Herczynska, M., Kurzawski, M., Safranow, K., Dziedziejko, V., Juzyszyn, Z. and Droździk, M. (2009) IL-1beta, IL-6, and TNF gene polymorphisms do not affect the treatment outcome of rheumatoid arthritis patients with leflunomide. Pharmacological Reports, 61, 281-287.
- Ursum, J., van der Weijden, M.A., van Schaardenburg, D., Prins, A.P.A., Dijkmans, B.A.C., Twisk, J.W.R., Crusius, J.B.A. and van der Horst-Bruinsma, I.E. (2010) IL10 GGC haplotype is positively and HLA-DQA1*05- DQB1*02 is negatively associated with radiographic progression in undifferentiated arthritis. Journal of Rheumatology, 37, 1431-1438. http://dx.doi.org/10.3899/jrheum.090913
- Konenkov, V.I., Zonova, E.V., Korolev, M.A., Leonova, I., Shevchenko, A.V., Golovanova, O.V. and Prokof'ev, V.F. (2010) Pharmacogenetic criteria for the efficacy of basic anti-inflammatory therapy for rheumatoid arthritis. Terapevticheskiĭ arkhiv, 82, 56-61.
- Camp, N.J., Cox, A., di Giovine, F.S., McCabe, D., Rich, W. and Duff, G.W. (2005) Evidence of a pharmacogenomic response to interleukin-l receptor antagonist in rheumatoid arthritis. Genes and Immunity, 6, 467-471. http://dx.doi.org/10.1038/sj.gene.6364228
- Cimaz, R., Cazalis, M.A., Reynaud, C., Gerloni, V., Zulian, F., Biggioggero, M., Martini, G., Pontikaki, I., Fantini, F., Mougin, B. and Miossec, P. (2007) IL1 and TNF gene polymorphisms in patients with juvenile idiopathic arthritis treated with TNF inhibitors. Annals of the Rheumatic Diseases, 66, 900-904. http://dx.doi.org/10.1136/ard.2006.067454
- Sorensen, L.K., Havemose-Poulsen, A., Bendtzen, K. and Holmstrup, P. (2009) Aggressive periodontitis and chronic arthritis: Blood mononuclear cell gene expression and plasma protein levels of cytokines and cytokine inhibitors. Journal of Periodontology, 80, 282-289. http://dx.doi.org/10.1902/jop.2009.080347
- Venter, J.C., Adams, M.D., Myers, E.W., et al. (2001) The sequence of the human genome. Science, 291, 1304- 1351. http://dx.doi.org/10.1126/science.1058040
- Hurme, M. and Santtila, S. (1998) IL-1 receptor antagonist (IL-1Ra) plasma levels are co-ordinately regulated by both IL-1Ra and IL-1β genes. European Journal of Immunology, 28, 2598-2602. http://dx.doi.org/10.1002/(SICI)1521-4141(199808)28:08<2598::AID-IMMU2598>3.0.CO;2-K
- Pociot, F., Molvig, J., Wogensen, L., Worsaae, H. and Nerup, J. (1992) A TaqI polymorphism in the human interleukin-1β (IL-1β) gene correlates with IL-1β secretion in Vitro. European Journal of Clinical Investigation, 22, 396-402. http://dx.doi.org/10.1111/j.1365-2362.1992.tb01480.x
- Dominici, R., Malferrari, G., Mariani, C., Grimaldi, L. and Biunno, I. (2002) The Interleukin 1-β exonic (+3953) polymorphism does not alter in Vitro protein secretion. Experimental and Molecular Pathology, 73, 139-141. http://dx.doi.org/10.1006/exmp.2002.2435
- Marcos-Pinto, R., Dinis-Ribeiro, M., Carneiro, F., et al. (2013) First-degree relatives of early-onset gastric cancer patients show a high risk for gastric cancer: Phenotype and genotype profile. Virchows Archiv, 463, 391-399.
- Persson, C., Engstrand, L., Nyren, O., Hansson, L.E., Enroth, H., Ekström, A.M. and Ye, W.M. (2009) Interleukin 1-β gene polymorphisms and risk of gastric cancer in Sweden. Scandinavian Journal of Gastroenterology, 44, 339- 345. http://dx.doi.org/10.1080/00365520802556015
- Collado-Hidalgo, A., Bower, J.E., Ganz, P.A., Irwin, M.R. and Cole, S.W. (2008) Cytokine gene polymorphisms and fatigue in breast cancer survivors: Early findings. Brain, Behavior, and Immunity, 22, 1197-1200. http://dx.doi.org/10.1016/j.bbi.2008.05.009
- Falfan-Valencia, R., Pavon-Romero, G.F., Camarena, A., de la Luz García, M., Galicia-Negrete, G., Negrete-García, M.C. and Teran, L.M. (2012) The IL1B-511 polymorphism (rs16944 AA genotype) is increased in Aspirinexacerbated respiratory disease in Mexican population. Journal of Allergy, 2012, Article ID: 741313. http://dx.doi.org/10.1155/2012/741313
- Gueant-Rodriguez, R.M., Juilliere, Y., Battaglia-Hsu, S.F., Debard, R., Gérard, P., Reyes, P., Danchin, N. and Guéant, J.L. (2011) Association of IL1B polymorphism with left ventricular systolic dysfunction: A relation with the release of interleukin-1β in stress condition. Pharmacogenetics and genomics, 21, 579-586. http://dx.doi.org/10.1097/FPC.0b013e3283493a05
- Jotanovic, Z., Etokebe, G.E., Mihelic, R., Kårvatn, M.H., Mulac-Jericevic, B., Tijanic, T., Balen, S., Sestan, B. and Dembic, Z. (2011) Hip osteoarthritis susceptibility is associated with IL1B -511(G>A) and IL1 RN (VNTR) genotypic polymorphisms in Croatian Caucasian population. Journal of Orthopaedic Research, 29, 1137-1144. http://dx.doi.org/10.1002/jor.21378
- Chen, S., Jiang, F., Ren, J., Liu, J. and Meng, W. (2012) Association of IL-18 polymorphisms with rheumatoid arthritis and systemic lupus erythematosus in Asian populations: A meta-analysis. BMC Medical Genetics, 13, 107. http://dx.doi.org/10.1186/1471-2350-13-107
- Lee, Y.H., Bae, S.C., Choi, S.J., Ji, J.D. and Song, G.G. (2012) Associations between the FAS -670 A/G and -1,377 G/A polymorphisms and susceptibility to autoimmune rheumatic diseases: A meta-analysis. Molecular Biology Reports, 39, 10671-10679. http://dx.doi.org/10.1007/s11033-012-1957-5
- Song, G.G., Bae, S.C., Choi, S.J., Ji, J.D. and Lee, Y.H. (2012) Associations between interleukin-23 receptor polymorphisms and susceptibility to rheumatoid arthritis: A meta-analysis. Molecular Biology Reports, 39, 10655- 10663. http://dx.doi.org/10.1007/s11033-012-1955-7
- Xie, Q., Wang, S.C., Bian, G., Zhan, F.L., Xie, J.K. and Li, J. (2012) Association of MIF-173G/C and MBL2 codon 54 gene polymorphisms with rheumatoid arthritis: A meta-analysis. Human Immunology, 73, 966-971. http://dx.doi.org/10.1016/j.humimm.2012.07.043
- Lee, Y.H., Ji, J.D. and Song, G.G. (2009) Association between interleukin 1 polymorphisms and rheumatoid arthritis susceptibility: A meta-analysis. Journal of rheumatology, 36, 12-15.
- Hodoglugil, U. and Mahley, R.W. (2012) Turkish population structure and genetic ancestry reveal relatedness among Eurasian populations. Annals of Human Genetics, 76, 128-141. http://dx.doi.org/10.1111/j.1469-1809.2011.00701.x
- Huang, Y., Chen, H., Wang, J., Bunjhoo, H., Xiong, W.N., Xu, Y.J. and Zhao, J.P. (2013) Relationship between CCR2-V64I polymorphism and cancer risk: A metaanalysis. Gene, 524, 54-58. http://dx.doi.org/10.1016/j.gene.2013.04.011
- Holmes, M.E., Eisenmann, J.C., Ekkekakis, P. and Gentile, D. (2008) Physical activity, stress, and metabolic risk score in 8- to 18-year-old boys. Journal of Physical Activity & Health, 5, 294-307.
NOTES
*DD and LW are co-first authors of this work.
#Corresponding authors.

