Journal of Biophysical Chemistry
Vol.1 No.1(2010), Article ID:1880,7 pages DOI:10.4236/jbpc.2010.11004
Mode of pollen spread in clonal seed orchard of Pinus koraiensis
1Northeast Forest University, Harbin, China; ffj9018@sina.com
2Institute of Applied Ecology, Chinese Academy of Sciences, Shenyang, China; *Corresponding Author: hansj@iae.ac.cn
3Swiss Federal Research Institute WSL, Birmensdorf, Switzerland
Received 14 January 2010; revised 20 March 2010; accepted 31 March 2010.
Keywords: Pinus koraiensis; seed orchard; nuclear microsatellites markers; paternity analysis; pollen spread
ABSTRACT
The patterns of effective pollen dispersal and the relationships between pollen dispersal and genetic composition in Pinus koraiensis are still unclear. Hence, we investigated the mode of pollen dispersal of P. koraiensis in a clonal seed orchard in Lushuihe Forestry Bureau, Jilin Province, using SSR molecular markers technique and the method of maximum likelihood. A total of 13 pairs of nuclear microsatellites polymorphic primers were used in the paternity analysis. We analyzed 100 progenies and 150 paternities. A total of 56 alleles were detected in 13 loci with 3-6 alleles (4.3 alleles in average) in a single locus. The primers of SsrPt_ctg7170 and SsrPt_ctg5333 had the maximum (6) and the minimum (3) alleles, respectively. The averaged values of observed heterozygosity, expected heterozygosity and polymorphism information content of experimental groups were 0.648, 0.601 and 0.533, respectively. The mating distance of P. koraiensis followed an approximately normal distribution. The most effective pollen for the female parent came from male parent trees 15-45 m away with an average mating distance of 32.60 m and the longest of 67.88 m. The pollen source of offspring was not random but greatly affected by the wind direction during the pollination season of P. koraiensis. These results have important implications for seed orchard design to improve the genetic quality of seeds and seed production.
1. INTRODUCTION
Pinus koraiensis, a national protected tree species in China, is a dominant species in the climax vegetation, mixed needle broad-leaved forests, in eastern mountainous area in northeastern China. Currently, the distribution of this tertiary relict plant ranges from 35° to 52°N and from 126° to 143°E, concentrating mainly in Changbai Mountain, the Lesser Khingan Mountain, Wanda Mountain, and Changkuantsailing in northeastern China, northern regions in North Korea, Japan and the southern part of the Russian Far East. The slow-growing P. koraiensis with late sexual maturity is very difficult to regenerate in nature.
Ma et al. [1] showed that the area of natural P. koraiensis forest has dramatically decreased in China. Owing to the ecological and economic significance of P. koraiensis, plantations of this species have recently been widely establishing in northeastern China. Therefore, large amount of good seeds with good genetic properties are still urgently needed for successful afforestation. For this purpose, seed orchards of P. koraiensis have been established in northeastern China.
Many factors affect the physioecological quality of seeds produced in a seed orchard. For example, gene flow, to a large extent, influences the genetic structure of a forest population. Pollen movement is one of the main factors impacting the gene flow of groups. It involves the form of pollination, and number, size and shape of pollen. Pollen related to gene flow associated with the plant mating system and outcrossing rate affects the size of plant gene flow directly. The model of pollen spread determining, to a large extent, genetic compositions of open-pollinated families; can affect the levels of genetic diversity and the size of effective population [2-4]. Therefore, pollen gene flow is an important factor to impact yield and quality of seeds in seed orchards. However, the related research on P. koraiensis is scarce.
Recently, paternity analysis has become the most common method for directly estimating gene flow [5,6]. In the present study, SSR molecular markers technique was used to paternally identify the clonal half-sib progenies in P. koraiensis seed orchard, to analyze the source of offspring pollen. The present study aimed 1) to reveal the rules of the pollen spread in P. koraiensis seed orchard, and 2) to quantitatively investigate outcrossing rate and self-crossing rate of progenies. The results are helpful to guide the maximum mating of the fine genotypes among the forest population, and have some significance for the design of seed orchard and the management of genetic resources [7,8].
2. MATERIALS AND METHODS
2.1. Study Site, Plant Materials and Sampling Methods
Point-centre quarter method, according to Lian et al. [9] and Kong [10], was used to take samples. The experimental site was set at sub-district No. 5, zone No. 2, located in the center of Hongwei Seed Orchard in Lushuihe Forestry Bureau, Jilin Province. The orchard with an area of about 10000 m2 contains 727 individuals of 61 clones planted with 3 m × 3 m spacing in 1989. At the beginning of the present study, the plants grown well with an average height of 4.31 m, and an average diameter at breast height of 7.8 cm. A well-conditioned plant (plant No. of 105) located in the center of the sub-district and at the cross-point of the line 16 and row 21 was randomly selected as the female parent. The sub-district No. 5 surrounding that selected female parent was divided into 4 quadrants with a total of 149 individuals and 61 clones as male parents recorded. Sixteen, 46, 48, and 39 trees, existed in the first, second, third, and in the fourth quadrant, respectively. The clones of fourth quadrant were 15, 33, 28, and 22. The female parent was also taken into account for a possible candidate as a male parent for self-pollination. Hence, a total of 150 candidates of male parent were analyzed. A total of 100 seeds were randomly selected from the cones collected from the female parent. Embryos (offsprings) were then obtained from these seeds.
Sampling was conducted in September 2007. Annual needles collected from each male parent were stored at –40°C. The pinecones of the same year were collected from the female parent and also stored at –40°C.
2.2. DNA Extraction
DNA was extracted from the needles using the modified CTAB method [11]. Embryo DNA was extracted using the traditional method of CTAB. In the extraction process, solution was extracted for many times until no white turbidity occurred.
2.3. Statistical Analysis
Data were scored by SSR co-dominant markers. Data analyses were performed with CERVUS (Version 2.0) based on the method of maximum likelihood. The number of alleles (NA), allele frequency (Pi), observed and expected heterozygosity (Ho, He), and polymorphic information content (PIC) were also calculated using CERVUS (Version 2.0).
2.4. Development of SSR Primer
A total of 13 pairs of primers were chosen from 96 pairs of allied species as described in the published documents (Pinus pinaster and Pinus taeda) [12-14]. The sequences of primers were shown in Table 1.
2.5. SSR-PCR Reaction System
The SSR-PCR reaction system was constructed as described by Lian et al. [9] Amplification was carried out in an ABI 9700 Thermocycler with following PCR program: preliminary denaturation at 94°C for 4 min, 30 cycles of denaturation (1 min, 94°C), annealing (1 min, the proper temperature), extending (1 min, 72°C), and extending at 72°C for 4 min, and finally keeping at 4°C. Amplification fragments were segregated on 6% nondenatured polyacrylamide gels.
3. RESULTS
3.1. SSR Polymorphism Analysis
A total of 13 pairs of SSR primers were amplified in the male parent candidates and offsprings (a total of 250 samples). As shown in Figures 1 and 2, SSR marker in P. koraiensis exhibited the characteristics of good stability and repeatability as well as clear bands. A total of 56 alleles were detected in 13 loci with 3-6 alleles in a single locus and 4.3 alleles in average (Table 2). The primers of SsrPt_ctg7170 and SsrPt_ctg5333 had the maximum (6) and the minimum (3) alleles, respectively (Table 2). The averaged values of observed heterozygosity, expected heterozygosity, and polymorphism information content of the experimental groups were 0.648, 0.601, and 0.533, respectively (Table 2).
3.2. Paternal Analysis of the Open-Pollinated Progenies
Paternity analysis of the open-pollinated progenies indicated that in a 95% credibility interval, the true male parent of 53 offsprings (accounting for 53% of the total offsprings) can be inferred from the 13 SSR primers. The cumulative exclusion probability was 94.59%, of which the primer of SsrPt_ctg 7170 had the highest exclusion probability (55%). Analysis of male parent using CERVUS software showed that three clones were the same as the female parents existed in the suspected male parents of P. koraiensis. There were no male parents from the female parents or the same clones of female parents in the 100 open-pollinated offsprings, indicating that no self-crossing occurred in the experiment. The
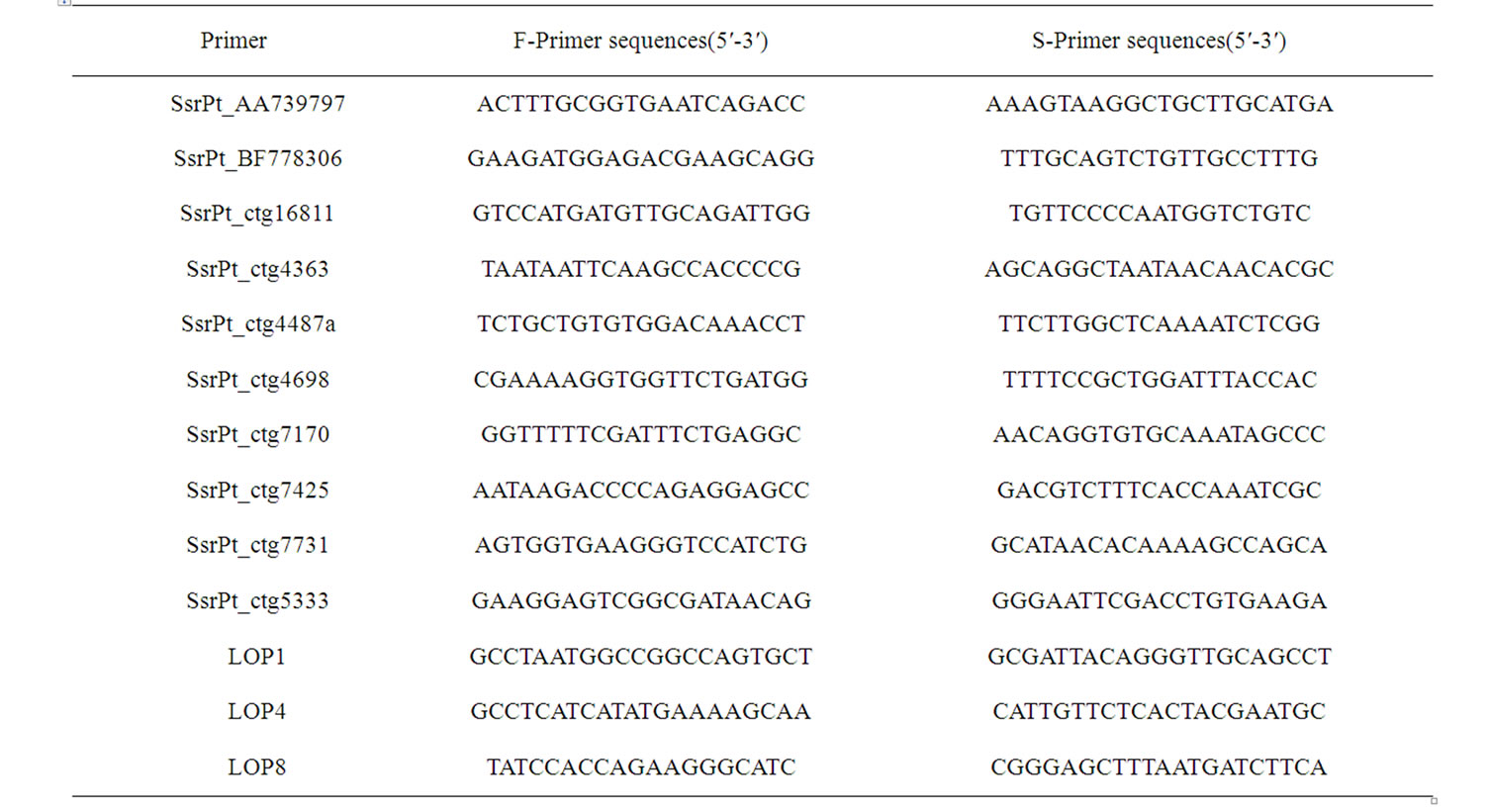
Table 1. Sequence of 13 pairs of primers from the 96 pairs of allied species.
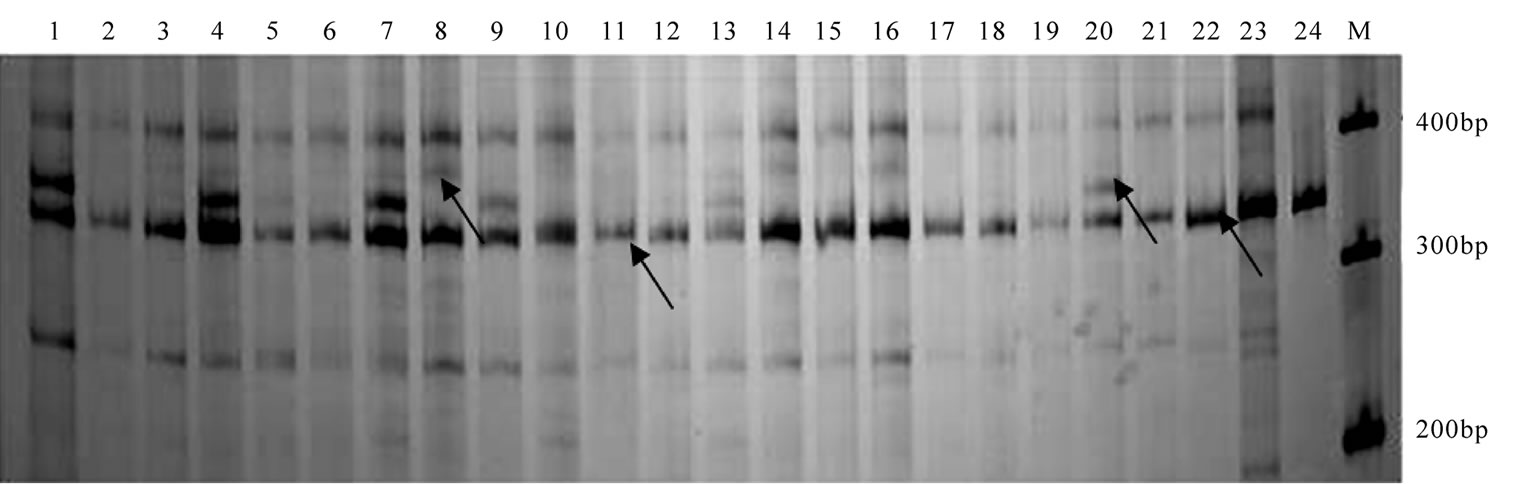
Figure 1. The bands of Pinus koraiensis amplified from the primer pairs of LOP4. Lanes 1-24 were male parents; M: marker. The arrows show the polymorphic bands.

Figure 2. The bands of Pinus koraiensis amplified from the primer pairs of Ssr_ctg4363. Lanes 1-24 were male parents; M: marker. The arrows show the polymorphic bands.
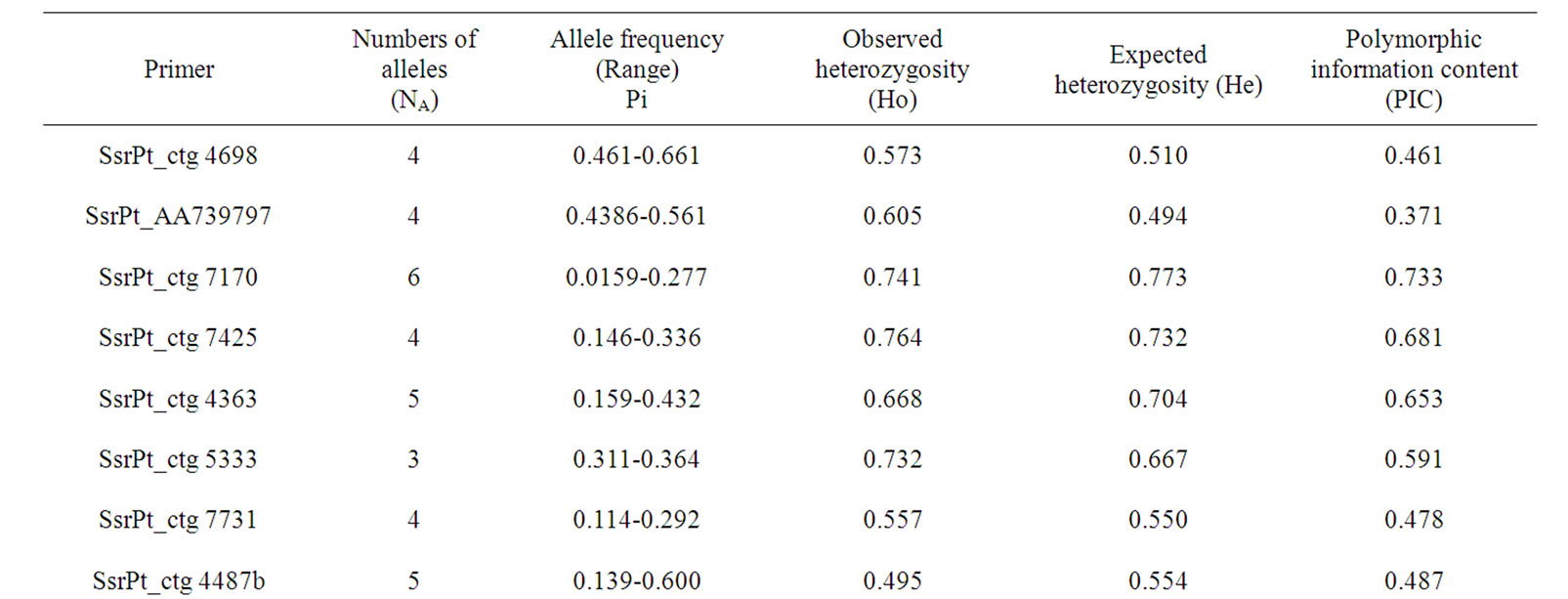
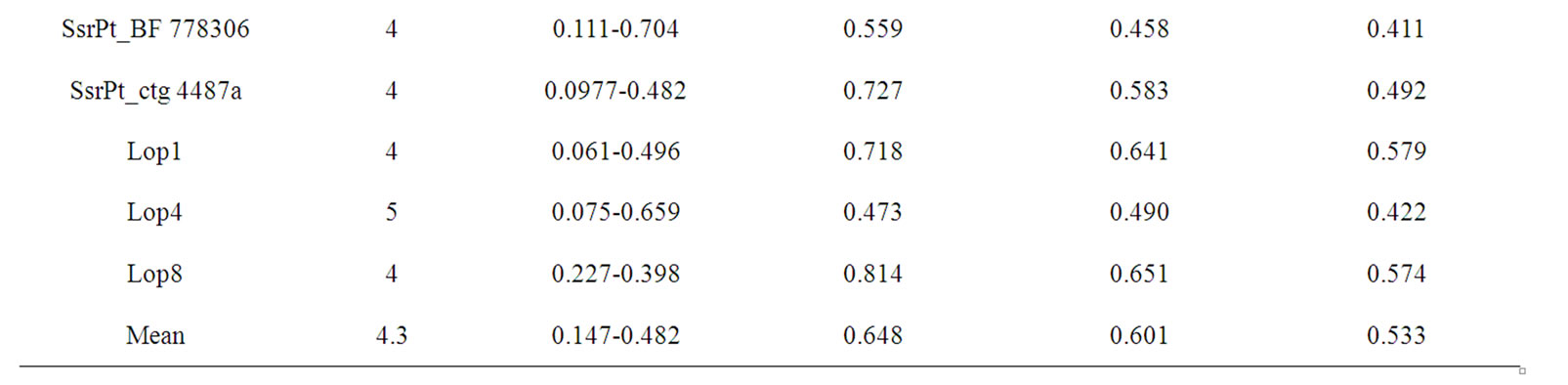
Table 2. Polymorphism of SSR in Pinus koraiensis.
results reflected that outcrossing was the main mating approach of P. koraiensis in the open-pollinated situation.
In a 95% credibility interval, 48 male parents out of the 150 candidates (32%) produced offsprings with the female parents. Among them, the male parents of clones Nos. 149 and 126 identified three sub-generations, with a male reproductive fitness of 3%. The reproductive fitness of the rest of the male parents was 1%-2%, with an average value of 1.34%. Another 102 (68%) male candidates did not provide effective pollens for progeny groups, and the male reproductive fitness was zero.
3.3. Dispersal Patterns of Effective Pollen
The spatial distribution and mating pattern of all the candidates showed that the pollen source of offspring was not random (Figure 3). The pollen from the male candidates located in the southern quadrants 3 and 4 were significantly higher than those from trees in the northern quadrants 1 and 2 (Figure 3). This may be closely related to the dominant wind direction affecting the pollination process of P. koraiensis. According to the climatological data collected in 2007, the dominant wind in the pollination season blew northeasterly (i.e. from the Southwest toward Northeast). Hence, pollen from the male parents in quadrants 3 and 4, which were located in the upwind direction of the female parent, can easily reach the female parent. On the other hand, however, quadrants 1 and 2 were located in the downwind position, pollen of male parents in these quadrants were hard to reach the female parent (Figure 3).
The mating distance between parents is the dispersal distance of effective pollens, which reflects the pattern of mating within populations. P. koraiensis is a windpollinated plant species, therefore the mating distance can be well described using the flying distance of pollen. The results showed that the mating distance of P.
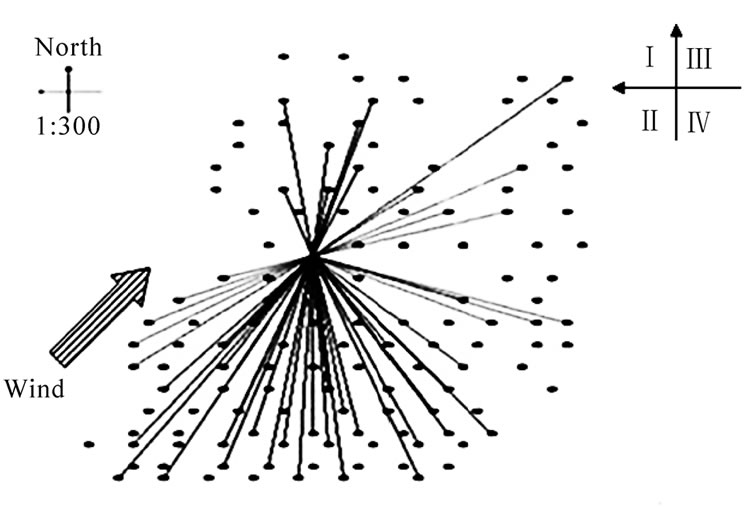
Figure 3. Spatial distribution of paternal candidates and mating pattern for the maternal tree in the experimental population of Pinus koraiensis.
koraiensis followed an approximately normal distribution (Figure 4). The proportion of successful mating of the 53 open-pollinated offsprings identified was higher in the distances of 15-30 m (36%) and 30-45 m (30%) between the female parent and the male parent, with an average mating distance of 32.60 m and the longest distance of 67.88 m (Figure 4).
3.4. Impact of Foreign Pollen
The impact of foreign pollen contamination was analyzed. In the results of electrophoresis of 13 pairs of SSR primers, six alleles in 25 offsprings (25% of the total offsprings) were found to be different to the male parents, which may be caused by foreign pollen contamination. The proportion of contamination was 25%.
4. DISCUSSION
4.1. Analysis of Paternal
The paternal analyses revealed the most likely parent of each offspring and the corresponding credibility (confidence probability) in an area of statistical inference. At a 95% credibility interval, 53 out of the 100 offsprings’ male parents have been confirmed in the present study, indicating a high inference rate resulted from the use of 13 SSR primers. Moreover, the cumulative exclusion probability was 94.59%, also indicating that SSR codominant marker is highly appropriate for paternity analysis.
The observed heterozygosity (0.648) was higher than the expected heterozygosity (0.601), indicating that the proportion of heterozygote was relatively large within the group. This finding is consistent with other studies dealing with other tree species [15,16]. The polymorphism information content (PIC) was used as an indicator of locus diversity [17]. Botstein et al. [18] pointed out that the locus is high, moderate, and low polymorphic when PIC > 0.5, 0.25 < PIC < 0.5, and PIC < 0.25, respectively. The present study showed a PIC > 0.5, indicating that SSR polymorphism of P. koraiensis is high.
The composition of male parents found in the present study was similar to the results gained from other tree species [19-22]. He et al. [23] based on their study of parents analysis using exclusion method, stated that the optimal number of expected parents was less than 100 in the population studied; otherwise the limitations of the analysis would increase. In the present study, a total of 150 male candidates were analyzed, thus the determination ability of the male parents may be influenced to some extent.
4.2. Spread of Effective Pollen
The characteristics, type, quantity, density, and distribution of parents, and also environmental factors determine the spread distance and pattern of effective pollen [22]. For instance, the intensity of pollen flow and the chance of successful mating decrease with increasing pollination distance [24]. Our results showed that the spread distance of P. koraiensis followed an approximately normal distribution, which is similar to the results reported by Sun et al. [22]. The most effective pollen for the female parents came from trees with a distance of 15-45 m (Figure 4). Moreover, wind direction significantly affects pollination [25]. In line with this view, the receivable pollens of female parent came mainly from the southern direction associated with the dominant wind in the pollination season of P. koraiensis.
Of course, it is also notable that the object of this study is a man-made seed orchard of P. koraiensis trees. Higher density of the same tree species with similar age (maturity) can provide rich pollen, which may limit the long-distance spread of pollen.
4.3. Mating Means
The mating means are affected by many factors including pollination medium, the genetic differences between breeding individuals in groups, the spatial distribution, flowering habits, the characteristics of pollen, and climatic conditions in blooming period [22]. Our results showed that the mating means of P. koraiensis were dominated by outcrossing. Two possible reasons may contribute to outcrossing of P. koraiensis: 1) the species has dichogamy phenomena; and 2) the concentrated and homogeneous distribution of individuals in a seed orchard is beneficial to cross-pollination. Moreover, in an evolutionary view, the majority of outcrossing species would form mechanisms to avoid inbreeding depression such as self-incompatibility. Stern et al. [26] reported that self-sterility of conifers was mostly caused by the wilting of embryo after self-fertilization. The present study found that the self-crossing rate was nearly zero, implying that the self-incompatibility phenomenon may exist in P. koraiensis which needs to be test.
4.4. Pollen Contamination
The pollen contamination level (25%) in this seed or-

Figure 4. The dispersal distance of effective pollen.
chard was lower as compared with research results of other Pinus species, for example, 36%~60% for P. taeda [27], 84.5% for P. elliotii [28], and 9%~74% for P. sylvestris [29-31].
The unique binds for fragments found in the present study may come from other male parents outside the sampling plot (i.e. the sub-district No. 5). Indeed, it is most possible that the contamination pollen came from the neighboring sub-districts rather than from the surrounding natural forests since the isolation strip around the seed orchard was more than 500 m.
5. CONCLUSIONS
The present study found that the most effective pollen for the female parent came from male parent trees 15-45 m away with an average mating distance of 32.60 m and a maximum distance of 67.88 m. The pollen source of offspring was not random but greatly affected by the wind direction during the pollination season of P. koraiensis. These results have important implications for seed orchard design to improve the genetic quality of seeds and seed production.
6. ACKNOWLEDGEMENTS
This research was supported by Chinese Academy of Science Knowledge Innovation Sub-project, also supported by Heilongjiang Postdoctoral Research Start-up Funding.
REFERENCES
- Ma, J.L., Zhang, L.W., Cheng D. and Li, J.W. (1992) Geographic distribution of Pinus koraiensis in the world. Journal of Northeast Forestry University, 20, 40-47.
- Levin, D.A. and Kerster, H.W. (1974) Gene flow in seed plants. Evolutionary Biology, 7, 139-220.
- Hamrick, J.L. and Schnabel, A. (1985) Understanding the genetic structure of plant populations: Some old problems and a new approach. In: Gregorius, H.R., Ed., Population Genetics in Forestry, Springer-Verlag, New York, 5, 1-70.
- Adams, W.T., Birkes, D.S. and Erickson, V.J. (1992) Using genetic markers to measure gene flow and pollen dispersal in forest tree seed orchards. In: Wyatt, R., Ecology and Evolution of Plant Reproduction: New Approaches, Chapman and Hall, New York.
- Meagher, T.R. (1986) Analysis of paternity within a natural population of Chamaelirium leuteum. I. Identification of most-likely male parents. American Naturalist, 128(2), 199-215.
- Snow, A.A. and Lewis, P.O. (1993) Reproductive traits and male fertility in plants Empirical approaches. Annual Review of Ecology and Systematics, 24, 331-351.
- Adams, W.T. and Birkes, D.S. (1989) Matting patterns in seed orchards. Proceedings of 20th Southern Forest Tree Improvement Conference, Charleston, 26-30 June 1989, 75-86.
- Wheeler, N.C. and Jech, K.S. (1992) The use of electrophoretic markers in seed orchard research. New Forests, 6(1-4), 311-328.
- Lian, C.L., Miwa, M. and Hogetsu, T. (2001) Outcrossing and paternity analysis of Pinus densiflora (Japanese red pine) by microsatellite polymorphism. The Genetics Society of Great Britain, 87, 88-98.
- Kong, F.L. (2006) Quantitative genetics in plants. China Agricultural University Press, Beijing, 20-50.
- Wang, J., Yang C.P. and Liu, G.F. (2006) Extraction of genomic DNA from woody plants and it’s identification. Bulletin of Botanical Research, 26, 589-594.
- Zhou, Y., Bui, T., Auckland, L.D. and Williams, C.G. (2002) Undermethylated DNA as a source of microsatellites from a conifer genome. Genome, 45(1), 91-99.
- Chagné, D., Chaumeil, P., Ramboer, A., Collada, C., Guevara, A., Cervera, M.T., Vendramin, G.G., Garcia, V., Frigerio, J.M., Echt, C., Richardson, T. and Plomion, C. (2004) Cross-species transferability and mapping of genomic and cDNA SSRs in pines. Theoretical and Applied Genetics, 109(6), 1204-1214.
- Liewlaksaneeyanawin, C., Ritland, C.E., EI-Kassaby, Y.A. and Ritland, K. (2004) Single-copy, species-transferable microsatellite markers developed from loblolly pine ESTs. Theoretical and Applied Genetics, 109(2), 361-369.
- Ge, S. (1998) Isozymes and studies on population genetic variation in forest trees. Journal of Nanjing Forest University, 1, 68-77.
- El-Kassaby, Y.A. (1991) Genetic variation within and among conifer populations: review and ecaluation of methods. In: Fineschi, S., Ed., Biochemical Markers in the Population Genetics of Forest Trees, SPB Academic Publishing, Hague, 61-76.
- Nei, M. (1972) Genetic distance between populations. American Naturalist, 106, 283-292.
- Botstein, D., White, R.L., Skolnick, M. and Davis, R.W. (1980) Construction of a genetic linkage maps in man using restriction fragment length polymorphisms. American Journal of Human Genetics, 32(3), 314-331.
- Smouse, P.E. and Meagher, T.R. (1994) Genetic analysis of male reproductive contributions in Chamaelirium luteum (L.) gray (Liliaceae). Genetics, 136(1), 313-322.
- Chen, X.Y. and Song, Y.C. (2000) Analysis of parentage for naturally for naturally established seedlings of cyclobalanopsis glauca. Journal of Wuhan Botanical Research, 18, 174-180.
- Sun, Y.G. and Li, H.G. (2007) The paternity analysis for open-pollination progenies of Liriodendron L. using SSR markers. Chinese Bulletin of Botany, 24, 590-596.
- Zhang, D.M., Sun, P.G., Shen, X.H. and Ru, G.X. (2009) Paternity analysis of open-and control-pollinated seeds collected from a seed orchard of Pinus tabulaeformis. Chinese Journal of Plant Ecology, 33(2), 302-310.
- He, T.H. and Ge, S. (2001) Mating system, paternity analysis and gene flow in plant populations. Acta Phytoecologica Sinica, 25, 144-154.
- Sulaiman, Z., Collins, G., Witherspoon, J. and Sedgley, M. (2004) Identification of pollen donors for the avocado cultivar Gwen in a mixed orchard by isozyme analysis. Journal of Horticultural Science and Biotechnology, 79(4), 571-575.
- Wang, G.Y., Yi, H.B., Wang, B., Zhang, S.F., Wang, G.X., Huang, Q.C., Zhang, G.T. and Sun, Q.Y. (1995) Study on the pollen density in the seed orchard of Korean Pine. Journal of Northeast Forestry University, 23, 58-63.
- Stern, K. and Roche, L. (1974) Genetics of Forest Ecosystems. Springer-Verlag, Berlin, 65-69.
- Friedman, S.T. and Adams, W.T. (1985) Estimation of gene flow into two seed orchards of loblolly pine (Pinus taeda L.). Theoretical and Applied Genetics, 69(5-6), 609- 615.
- Squillace, A.E. and Long, E.M. (1982) Proportion of pollen from nonorchard sources. In: Franklin, E.C., Ed., Pollen Management Handbook, USDA Agriculture Handbook # 587, AVI Publishing Company Inc., Washington DC, 15-19.
- Lai, H.L. and Wang, Z.R. (1997) A study on gene flow between a Masson pine seed orchard and a plantation near the orchard. Journal of Nanjing Forestry University, 21, 37-41.
- Harju, A. and Muona, O. (1989) Background pollination in Pinus sylvestris seed orchards. Scandinavian Journal of Forest Research, 4(1-4), 513-520.
- Wang, X.R., Lindgren, D., Szmidt, A.E. and Yazdani, R. (1991) Pollen migration into a seed orchard of Pinus sylvestris and the methods of its estimation using allozyme markers. Scandinavian Journal of Forest Research, 6(1-4), 379-386.

