Advances in Microbiology
Vol. 2 No. 1 (2012) , Article ID: 18085 , 5 pages DOI:10.4236/aim.2012.21004
Genome Shuffling of Pseudomonas sp. Ioca11 for Improving Degradation of Polycyclic Aromatic Hydrocarbons
Biotechnology Department, Indian Oil Corporation Ltd., Research and Development Centre, Faridabad, India
Email: upretim@iocl.co.in
Received December 20, 2011; revised January 9, 2012; accepted January 18, 2012
Keywords: PAHs; Biodegradation; Genome Shuffling; Mutagenesis
ABSTRACT
A Pseudomonas sp. strain IOCa11 exhibiting broad substrate profile for polycyclic aromatic hydrocarbons (PAHs) degradation was isolated by enrichment techniques from oil-contaminated soil. We applied genome shuffling of Pseudomonas sp. strain IOCa11 to achieve improved degradation of PAHs. The initial mutant population was generated by nitrosoguanidine treatment and population exhibiting improved phenotype was subjected to multiple round of protoplast fusion in order to allow recombination between genomes. Mutant, designated as SF-IOC11-16A, obtained after recursive protoplast fusion showed substantial improvement in ability to degrade PAHs in liquid media. It could degrade 98% DBT in 72 hours in comparison to 74% by the wild parent strain. Similar improvement in degradation of the naphthalene (NAP), phenanthrene (PHE), and benzo (α) pyrene (BAP) was also observed in shuffled strain. The shuffled strain was also able to grow at higher concentration of PAHs and degrade them efficiently. The results indicate that genome shuffling can successfully be used to improve the PAHs degradation capability of bacteria.
1. Introduction
Polycyclic aromatic hydrocarbons (PAHs) are well known recalcitrant environmental pollutants and have major concern because of their mutagenic and/or carcinogenic effects [1]. The major sources of PAHs in the environment are the combustion of organic matter and the processing and use of fossil fuels. Biological degradation is widely accepted as the primary dissipation mechanism for most organic pollutants including PAHs in the environment. Numerous microorganisms have been shown to bio-transform and to mineralize PAHs [2-4]. However, bacteria with higher degradation ability are essential for fast removal of contaminant from the environment [2]. Methods for strain improvement range from random mutagenesis to highly rational methods of genetic and metabolic engineering [5]. Genomic shuffling is a novel and faster approach for strain improvement. It involves generation of the mutant strains that have improved phenotypes followed by multiple rounds of protoplast fusion to allow recombination between genome [6]. Improved progenies are selected and subjected to next round of shuffling. Genome shuffling thus accelerates directed evolution by accelerating recombination between diverse selected and improved populations [7-11]. Since, PAHs degradation is mediated by multiple genes, approach like genomic shuffling may prove more effective than other approaches in which usually alters the expression of one or few gene. In the present study, a bacterium was isolated from the oil-contaminated soil and random mutagenesis using nitroguanidine followed by recursive protoplast fusion (genome shuffling) was attempted to improve its PAHs degradation capabilities.
2. Materials and Methods
2.1. Isolation of Microorganism
The bacterial strain was isolated by the enrichment culture technique from the soil collected from oil-contaminated site at Indian Oil R&D. A 10-g sample of soil was inoculated into 100 ml of minimal salt medium (MSM) [12]. The strain was stored as frozen stock cultures at –70˚C in 25% glycerol. The bacterium was identified by partial 16S rRNA gene sequencing [12].
2.2. Growth on Hydrocarbons and Biodegradation
The ability of the isolate to PAHs as sole source of carbon and energy was assessed by inoculating it into a 24 well micro-titer plate containing 500 mL of BSM in each well and 0.02% (w·v–1) of test PAH as sole carbon source [13]. Stock solutions of PAHs were prepared in absolute ethanol. Each well was inoculated with 10-µL pre-culture (Luria-Bertani broth grown OD 600 = 0.8, approximately 108 cells ml–1) and incubated at 30˚C and 200 rev·min–1. Growth was followed for one week by measuring turbidity for the increase in cell density in comparison to controls and the cell viability counts (c.f.u. ml–1) on nutrient agar plates. No difference in turbidity and c.f.u. of the well and that of the controls were considered as no growth (−), whereas, increase in turbidity and viable cell counts (at least 10 fold) were considered as growth (+). Results of the micro-titer plate were verified by growing bacteria in 500 ml Erlenmeyer flasks containing 100 ml of MSM and PAH and monitoring turbidity and the colony forming units (c.f.u. ml–1).
For solid MSM the PAHs (except naphthalene) were dissolved in 5% (w·v–1) in diethyl ether and sprayed on surface of MSM agar. Naphthalene was provided as crystals directly placed on the plate lid. Growth on PAHs in solid media was considered positive by the formation of clear zone around the growing colonies or appearance of pigments.
Quantitative degradation of representative PAHs i.e., naphthalene (NAP), phenanthrene (PHE), and benzo (α) pyrene (BAP), dibenzothiophene (DBT) was studied in 250 ml flask containing 50 ml of MSM supplemented with 200 mg·l–1 of individual test PAH. Experimental flasks were inoculated with 2% (v·v–1) inoculum (105 c.f.u. ml–1) and incubated in dark on rotatory shaker (200 rev·min–1) for one week. Un-inoculated flasks and flasks without PAH served as controls. Cell growth was measured by the viable cell counts (colony forming units; c.f.u. ml–1) on nutrient agar plates. Degradation of PAHs was measured by reversed-phase HPLC on a C18 column. Separation was achieved by isocratic elution in acetonitrile: water (70:30), with a flow rate of 0.8 mL/min and UV absorbance detector [14-16].
2.3. Genome Shuffling
Mutagenesis was carried out with nitrosoguanidine (NTG) as described by Sambrook et al. 1989 [17]. Briefly, mutagen was added into 5 ml of the log phase culture in a 15-ml centrifuge tube to a final concentration of 0.3 g·L–1 and the culture suspension was further incubated in a water bath at 37˚C for 15 min. The final concentration of treated cells was about 5 ´ 108 c.f.u. ml–1. Diluting the culture 10 times with pre-chilled, fresh mineral salts medium subsequently terminated the treatment. The mutated cells were centrifuged and transferred to the MSM containing 400 ppm of DBT (representative PAH). After 24 h incubations at 30˚C, the culture broth was serially diluted and plate on MSM-agar plates containing 400 ppm of DBT. The plates were incubated at 30˚C for 48 hours. Protoplast was prepared according to Kumar et al., 2004 [18]. Shuffling was carried out according to Patnaik et al., 2002 with required modification [19]. Briefly, protoplasts were fused by suspension in buffer (0.5 M sucrose, 10 mM Tris-HCl, 20 mM MgCl2) containing 15% dimethyl sulphoxide and 60% PEG-6000. The resulting suspension was incubated at 30˚C for 30 min. The fused protoplast preparation was diluted with regeneration media (MSM containing 0.5 M sucrose) and protoplasts were harvested by centrifugation at 3000 rpm for 25 min at 25˚C. The protoplast were re-suspended in regeneration media and shaken at 200 rpm for 12 h before plating on MSM agar plates containing DBT. The plated were scraped to generate a pooled fusion library. The formation of protoplasts, their fusion and their subsequent regeneration was repeated six times with pooled regenerated cells from one fusion being the inoculum for the subsequent protoplast culture. At the end each round of protoplast fusion, the mutants were also evaluated in liquid media containing similar concentration of PAH that used for their screening in agar plate. Non-shuffled controls were prepared by the recursive formation and regeneration of protoplasts without exposure to PEG.
3. Results and Discussion
In the present study, several microbes with ability to utilize PAHs, as carbon source was isolated from hydrocarbon contaminated soil. The strain IOCa11 was selected for further study because of its ability to grow on wide range of hydrocarbon and utilize them as carbon source. Alignment of the 16S rRNA gene sequences of strain IOCa11 with sequences obtained by doing a Blast searching revealed maximum similarity (99%) to Pseudomonas sp. Bacterial strain was tested in liquid and solid media for its ability to utilize various PAHs. These chemicals represent the most common PAH pollutants and are the main components of crude oils. Strain IOCa11 takes around 24 hours to show the visible growth in liquid and solid media. The bacterium could grow naphthalene, dibenzothiophene, phenanthrene, 4,6-dimethyl dibenzothiophene, phenyl sulfide, bezo (α) pyrene, anthracene salicylate, catechol, pyrazole, toluene, phenol, carbazole. When colonies were grown on PAH coated agar plates, zone of the clearing appeared, indicating PAH degradation.
Ability of this strain to utilize aromatics was further supported by an increase in bacterial growth concomitant with a decrease in hydrocarbon concentration. Results indicated at 200 ppm concentration of PAH the bacterium could degrade around 79% ± 6%, 74% ± 5%, 59% ± 2% and 64% ± 3% degradation of naphthalene, phenanthrene, BAP and DBT respectively in 72 hours (Figure 1). The bacterium could degrade two rings PAHs more efficiently than the 3 and 5 rings PAHs and lesser amount of degradation was achieved at higher concentration of PAH (Table 1).
In order to obtain mutants with an improved ability to degrade PAH, mutagenesis and recursive protoplast fuon was carried out using NTG as per the protocol described in materials and methods. The 32 mutants obtained
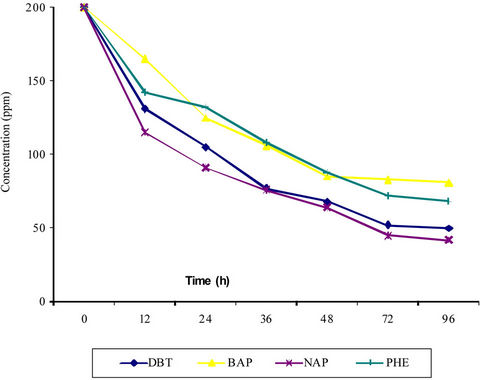
Figure 1. PAH degradation by strain IOCa11. DBT: dibenzothiophene, BAP: benzo (α) pyrene, NAP: naphthalene, PHE: Phenanthrene. Each value represents the mean of three samples with standard error < 5%.
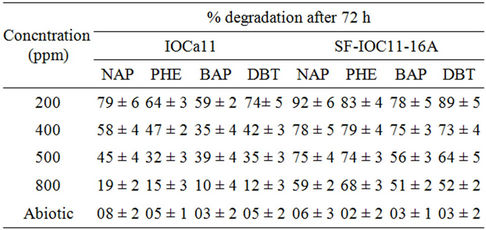
Table 1. Degradation of various PAHs at different concentration by strain IOCa11 and SF-IOC11-16A after 72 hours. DBT: dibenzothiophene, BAP: benzo (α) pyrene, NAP: naphthalene, PHE: Phenanthrene.
were used as the starting population for genome shuffling. Four successive around of protoplast fusion were carried out, and after each round the concentration of DBT on plate was increased and incubated to lesser time. On the first fusion, 12 colonies obtained on the plate containing 500 ppm of DBT after 48 hours of incubation were used for second fusion. After second fusion 10 colonies obtained on plates containing 600 ppm after 36 hours of incubation were used for third fusion. After third fusion 11 colonies obtained on plates containing 700 ppm after 30 hours of incubation were used for forth fusion (800 ppm). After forth-round growth was observed after 30 hours in three mutants. From this one fast growing mutant was evaluated further in liquid media. As a control, the population of mutant strains was also spread on various levels for specified incubation period without protoplast fusion to determine whether additional mutation caused by exposure to DBT on the plates could lead to improved growth on DBT. No colonies were observed when the mutant population without protoplast fusion was respread on plates containing 800 ppm after 48 hours of the incubations.
At the conclusion of the shuffling the ability of the strain isolated after all four rounds to grow on DBT in liquid media was assessed. Results show the growth, extent and time of degradation was progressively improved during shuffling (data not shown). Figure 2 depicts PAH degradation of one selected and most potential strain, termed as SF-IOC11-16A obtained after fourth round of shuffling. The SF-IOC11-16A grows faster in MSM containing PAH and its ability to degrade PAH was significantly higher than the wild type. It could degrade 98% DBT in 72 hours in comparison to 74% by the wild parent strain when the initial concentration of DBT in media was 200 ppm. When the concentration of the DBT in media was increased up to 800 ppm, the degradation by shuffled strain was 52% in comparison to 12% by the wild parent strain. In case of the naphthalene the wild strain was capable of degrading 79% at 200 ppm initial concentration while the shuffled strain could degrade 92% in 72 hours. When the concentration of the naphthalene in media was 800 ppm, the degradation by shuffled strain was 59% in comparison to 19% by the wild parent strain. Similar improvement in degradation of the BAP and PHE was also observed in shuffled strain in comparison to wild strain, (Table 1). In non-shuffled controls we could not observed any improvement in growth and DBT degradation ability indicating substantial improvements was associated with shuffling.
Genome shuffling is useful for engineering of mutitrait phenotypes that would be difficult to engineer directly because it may be improve a complex trait while still maintaining other characteristics of the strains. Since, PAH
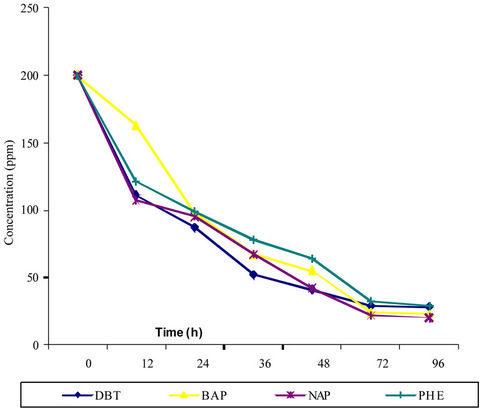
Figure 2. PAHs degradation by strain SF-IOC11-16A. DBT: dibenzothiophene, BAP: benzo (α) pyrene, NAP: naphthalene, PHE: Phenanthrene. Each value represents the mean of three samples with standard error < 5%.
degradation is mediated by multiple genes, approach like genomic shuffling may prove more effective than other approaches [20]. Genome shuffling accelerates directed evolution by facilitating recombination between members of diverse selected populations. It amplifies the genetic diversity within selected populations through extensive recombination between individual members and produce “multiparent complex progeny”. The resulting recombination represents larger combinatorial library of the original genetic diversity. Dai and Copley 2004 used genome shuffling to improve the degradation of pesticide pentachlorophenol (PCP) by Sphingobium chlorophenolicum ATCC 39723 [20] and could obtained strains that degrade PCP faster and tolerate 13 fold higher levels of PCP than the wild type strains. This approach has also used to improve the production of antibiotics in Strptomyces fradiae [6]. Patnaik et al., 2002 used genome shuffling to improve acid tolerance in Lactobacillus [19] and found three fold more lactic acid production than wild type at pH 4.0. Hida et al., 2007 had successfully applied genome shuffling of Streptomyces sp. U121 to obtain mutant with higher production (more than five fold than wild strain) of hydroxycitric acid [21].
The mechanism underlying the improved performance of the mutant strain SF-IOC11-16A may be due to mutations leading to removal of metabolite inhibition, constitutive expression of enzymes responsible for PAHs metabolism, amelioration of adverse effect of higher concentration of PAH etc. [22]. At this stage it is difficult to conclude any thing about the mechanisms but it is suspected that may be the multiple mutations are responsible for the improved phenotypes exhibited by the mutant stains after shuffling rounds. Results show that genome shuffling can be effectively used to improve the PAHs degradation ability of the bacteria.
REFERENCES
- K. C. Jones, J. A. Stratford, K. S. Waterhouse, E. T. Furlong, W. Giger, R. A. Hites, C. Schaffner and A. E. Johnston, “Increases in the polynuclear aromatic hydrocarbon content of an agricultural soil over the last century,” Environment Science Technnology, Vol. 23, No. 1, 1989, pp. 95-101. doi:10.1021/es00178a012
- C. E. Cerniglia, “Biodegradation of polycyclic aromatic hydrocarbons,” Current Opinion in Biotechnology, Vol. 4, No. 3, 1993, pp. 331-338. doi:10.1016/0958-1669(93)90104-5
- G. Fuchs, M. Boll and J. Heider, “Microbial degradation of aromatic compounds—from one strategy to four,” Nature Review Microbiology, Vol. 9, No. 11, 2011, pp. 803-816. doi:10.1038/nrmicro2652
- R. A. Kanaly and S. Harayama, “Biodegradation of HighMolecular-Weight Polycyclic Aromatic Hydrocarbons by Bacteria,” Journal of Bacteriology, vol. 182, no. 8, 2000, pp. 2059-2067. doi:10.1128/JB.182.8.2059-2067.2000
- W. P. Stemmer, “DNA shuffling by random fragmentation and reassembly: in vitro recombination for molecular evolution,” Proceedings of Notational Academy of Sciences USA, Vol. 91, No. 2, 1994, pp. 10747-10751. doi:10.1073/pnas.91.22.10747
- Y. Z. Zhan, K. Perry, V. A. Vinci, K. Powell, W. P. C. Stemmer and S. del Cardayre, “Genome Shuffling Leads to Rapid Phenotypic Improvement in bacteria,” Nature, Vol. 415, 2002, pp. 644-646. doi:10.1038/415644a
- M. H. Dai, S. Ziesman, T. Ratcliffe, R. T. Gill and S. D. Copley, “Visualization of Protoplast Fusion and quantitation of recombination in Fused Protoplasts of Auxotrophic Strains of Escherichia coli,” Metabolic Engineering, Vol. 7, No. 1, 2005, pp. 45-52. doi:10.1016/j.ymben.2004.09.002
- D. R. Singleton, L. G. Ramirez and M. D. Aitken, “Characterization of a Polycyclic Aromatic Hydrocarbon Degradation Gene Cluster in a Phenanthrene-Degrading Acidovorax Strain,” Applied Environmental Microbiology, Vol. 75, No. 9, 2009, pp. 2613-2620. doi:10.1128/AEM.01955-08
- M. H. Dai and S. Copley, “Genome Shuffling Improves Degradation of the Anthropogenic Pesticide Pentachlorophenol by Sphingobium chlorophenolicum ATCC 39723,” Applied Environmental Microbiolology, Vol. 70, No. 4, 2004, pp. 2391-2397. doi:10.1128/AEM.70.4.2391-2397.2004
- J. Gong, H. Zheng, Z. Wu, T. Chen and X. Zhao, “Genome Shuffling: Progress and applications for Phenotype Improvement,” Biotechnology Advances, Vol. 27, No. 6, 2009, pp. 996-1005. doi:10.1016/j.biotechadv.2009.05.016
- D. Pinel, F. D’Aoust, S. B. del Cardayre, P. K. Bajwa, H. Lee and V. J. Martin, “Saccharomyces cerevisiae Genome Shuffling through Recursive Population Mating Leads to Improved Tolerance to Spent Sulfite Liquor,” Applied Environmental Microbiology, Vol. 77, No. 14, 2011, pp. 4736-4743. doi:10.1128/AEM.02769-10
- M. Kumar, V. Leon, A. De Sisto, O. A. Ilzins, I. GalindoCastro and S. L. Fuenmayor, “Polycyclic Aromatic Hydrocarbon Degradation by Biosurfactant-Producing Pseudomonas sp. IR1,” Zeitschrift für Naturforschung, Vol. 61, No. 3-4, 2006, pp. 203-212.
- M. Kumar, V. Leon, A. De Sisto, O. A. Ilzins and L. Luis, “Biosurfactant production and Hydrocarbon-Degradation by halotolerant and thermotolerant Pseudomonas sp.,” World Journal of Microbiology and Biotechnology, Vol. 24, No. 7, 2008, pp. 1047-1057. doi:10.1007/s11274-007-9574-5
- A. R. Clemente, A. T. Anazawa and L. R. Durrant, “Biodegradation of polycyclic aromatic hydrocarbons by soil fungi,” Brazilian Journal of Microbiology, Vol. 32, no. 4, 2001, pp. 255-261. doi:10.1590/S1517-83822001000400001
- M. Arbabi, N. Simin and C. Anyakora, “Biodegradation of Polycyclic Aromatic Hydrocarbons (PAHs) in Petroleum Contaminated Soils,” Iran Journal Chemistry Chemical Engineering, Vol. 28, No. 3, 2009, pp. 53-59.
- S. D. Haemmerli, M. S. A. Leisola, D. Sanglard and A. Fiechter, “Oxidation of Benzo (α) pyrene by Extracellular Ligninases of Phanerochaete chrysosporium,” Journal Biological Chemistry, Vol. 261, No. 15, 1986, pp. 6900- 6903.
- J. Sambrook, E. F. Fritsch and T. Maniatis, “Molecular Cloning: A Laboratory Manual,” 2nd edition, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1989.
- M. Kumar, S. C. Agrawal and R. K. Upreti, “An improved Method for preparation of spheroplast from Ecscherichia coli,” In: P. C. Jain, ed., Microbiology and Biotechnology for sustainable Development, CBS Publishers and Distributors, New Delhi, 2004, pp. 314-325.
- R. Patnaik, S. Louie, V. Gavrilovic, K. Perry, W. P. C. Stemmer, C. M. Ryan and S. del Cardayre, “Genome shuffling of Lactobacillus for Improved Acid Tolerance,” Nature Biotechnology, Vol. 20, 2002, pp. 707-712. doi:10.1038/nbt0702-707
- M. H. Dai and S. Copley, “Genome Shuffling Improves Degradation of Anthropogenic Pesticide Pentachlorophenol by Sphingobium chlorophenolicum ATCC 39723,” Applied Environmental Microbiolology, Vol. 70, No. 4, 2004, pp. 2391-2397. doi:10.1128/AEM.70.4.2391-2397.2004
- H. Hida, T. Yamada and Y. Yamada, “Genome shuffling of Streptomyces sp. U121 for Improved Production of Hydroxycitric Acid,” Applied Microbiology and Biotechnology, Vol. 73, No. 6, 2007, pp. 1387-1399. doi:10.1007/s00253-006-0613-1
- M. Kumar, “Improving PAH degradation by Genome Shuffling,” Asian Journal of Microbiology Biotechnology and Environmental Sciences, Vol. 9, No. 1, 2007, pp. 145- 149.

