American Journal of Molecular Biology
Vol. 3 No. 1 (2013) , Article ID: 27191 , 6 pages DOI:10.4236/ajmb.2013.31004
Identification and characterization drought tolerance of gene LEA-D11 soybean (glycine max L. Merr) based on PCR-sequencing
![]()
1Doctoral Programme, Post Graduate University of Brawijaya, Malang, Indonesia
2Faculty of Agriculture University of Brawijaya, Malang, Indonesia 3Faculty of Mathematic & Basic Science University of Brawijaya, Malang, Indonesia
Email: evikasandi@yahoo.com
Received 17 November 2012; revised 17 December 2012; accepted 24 December 2012
Keywords: Drought Tolerance; LEA-D11 Gene; Soybean
ABSTRACT
Drought is one of the most damaging abiotic stress. Different plants response differently to drought stress. Abiotic stresses such as drought induced diverse physicological and molecular responses in plants. These responses include changes in gene expression. One of drought tolerance gene is a gene encoding dehydrin which is belongs to the group II or D-11 LEA protein family. LEA-D11 gene produce dehydrin protein which has a role in stabilization of membrane structures and protection of macromolecules in the presence of drought. The aims of the study was to identify and to characterize the LEA-D11 gene in various soybean varieties. This research used seven varieties of soybean: Tanggamus, Nanti, Seulawah, Tidar (drought tolerant), Wilis and Burangrang (drought moderate) and Detam-1 (drought susceptible). DNA genome of those varieties was isolated using the methods from Doyle & Doyle [1]. DNA amplification was conducted using Polymerase Chain Reaction (PCR) with specific primers designed based on GmLEA-D11 gene sequence database from the NCBI. The DNA targets were sequenced using automatic sequencing machine, ABI 3130xl Genetic Analyzer, in Eijkman Institution. The result of this study showed that the sequences of GmLEA-D11 gene possessed by drought tolerance varieties (Tanggamus, Nanti, Seulawah and Tidar) and moderately tolerance (Wilis and Burangrang) were similar. However, the sequence of GmLEA-D11 gene detected in the drought susceptible variety Detam-1 was different from the two groups. Similarity between drought tolerance and moderately tolerance indicate that there is not only LEA-D11 gene responsible to drought tolerance but also others. The primer and sequences GmLEA-D11 gene can be used as molecular marker and capable of differentiating between drought susceptible and drought moderate to drought tolerant.
1. INTRODUCTION
Abiotic stress such as drought, salinity, and frozen cause greatly damage and decrease yield. Under severe condition, these adverse environmental stresses can result in death of plant. Plants must respond and adapt to these adverse environmental condition to avoid or decrease cell injury caused by water deficit. Among the diversity of reponses, plants can adapt to water deficit by the induction of specific gene [2,3], including the changing of gene expression related drought tolerance. One of the gene related drought tolerance is LEA-D11 gene encoding family dehydrin protein [4,5].
Dehydrin are part of these LEA proteins (group II) and are built up by many charged and polar amino acids without cystein and tryptophan ever occurring [6]. Dehydrin are expressed during the late stages of embryogenesis [7,8] and also accumulated in vegetative tissues in response to water deficit [9]. Dehydrin have been found to accumulate in the cytoplasm, nucleus, plasma membrane and mitochondria [8,10-12].
Protein produced by drought-inducible genes which are identified through the recent microarray analysis can be classified into two groups [13]. The first group include proteins that most probably function in abiotic stress tolerance, the second group is comprised of regulatory protein. One of the gene products may play a role in drought tolerance is late embryogenesis abundant (LEA) protein. LEA is a functional protein which plays a role in stabilization of membrane structures and protected macromolecules [8]. Transgenic plant carrying genes for drought tolerance has been developed by the introduction of LEA gene, prolin synthesis and betaine [14-16]. Dehydrin like protein may also have role similar to compatible solute (such as proline, sucrose and glycine be taine) in osmotic adjustment. Another possible role of stress proteins is to bind with the ion accumulated (ion sequestering) under drought stress and to control solute concentration in the cytoplasma [17].
In addition, recently, it has been suggested that some dehydrin probably play role in antioxidative defence response directly by their radical scavenging activity [18] or indirectly by their capability of binding toxic metals and preventing production of ROS [19]. Dehydrin scavenged the hydroxyl radical and peroxyl radical, but did not superoxide anion and hydrogen peroxide [20]. Several residue such as Lys, His, Glyn d Ser, maybe related to the radical scavenging because the residue were modified when the dehydrin scavenging the hydroxyl radical. Dehydrin may protect cellular components from oxidative stress [21].
Identification and characterization of drought tolerance gene for developing molecular marker and selecting genetic variation in plants are very useful. The aims of this study is to identify and to characterize drought tolerance LEA-D11 gene in soybean varieties which tolerant, moderate and susceptible of drought.
2. MATERIAL AND METHOD
Growth Condition and Plant Material. Seven soybean varieties were utilized: Tanggamus, Nanti, Seulawah, Tidar (tolerant drought), Wilis, Burangrang (moderate drought), Detam-1 (susceptible drought). The experiment consisted of two treatments. Plants were grown in pots in a greenhouse. Control plants were well-watered throughout the experiment at about 100% field capacity; the drought stress treatment was conducted by maintaining soil water at about 25% field capacity throughout early vegetative growth until seed fulfill. After the last watering, soil water content was measured daily by weighing. The volume of water added afterward was calculated based on the weight difference between the soil before and after plant transpired in one day.
DNA Isolation. Total DNA was extracted from young soybean leaf, using the method of Doyle dan Doyle [1]. Fresh leaf with the weight of 0.1 - 0.2 g was grinded with addition of liquid nitrogen, and then 700 μL CTAB buffer was added and incubated for 30 minute in waterbath 65˚C. The DNA then was extracted using the mixture of chloroform: isoamyl alcohol (24:1). DNA was precipitated using 0.1 volume ammonium acetat and 2.5 volume ethanol absolute. The concentration and purity of extracted DNA was determined used spectrofotometric at the wavelength of 260 and 280 nm.
Primer Design. Primers were designed based on the sequence of complete CDS (coding DNA sequence) of GmLEA-D11 (ID: AM421515) from NCBI (The National Center for Biotechnology Information) database using the Oligo Analyzer 1.0.2., Oligo 1.1. software. The sequences of the primer were: forward 5’-ATGATCAGGGTCGCAAGGTC-3’, and reverse 5’ CTTGTCACTGTGTCCTCCAG-3’ with the amplification product of 700 bp.
Polymerase Chain Reaction. The total volume of PCR mixture was 20 μL per-tube, which were consist of 11.9 μL dH2O, 2 μL buffer Taq PCR; 1.6 μL MgCl2 ; 1.6 μL dNTPs 2.5 mM, (Qiagen-Taq PCR Master Mix), 0.3 μL primer forward-reverse (10 - 100 ng/µL), 0.3 μL Taq-Polymerase (5 U/μL) and 2 μL (1 μg/μL) DNA. The PCR program was set on 93˚C for 1 minute preheating, continued with 30 cycles consisting of 1 minute denaturation at a temperature of 93˚C, 1 minute annealing at a temperature of 57˚C, and 1 minute extension at a temperature of 72˚C. A final extension was conducted for 1 minute at a temperature of 72˚C. The PCR product was visualized on 1% agarose gel.
Sequences Analysis. Sequencing of the PCR products were performed with ABI automatic sequencer (ABI 3130xl Genetic Analyzer) using fluorescence-labelled nucleotides. The sequences were analyzed using multiple sequence alignment by Sequence Scanner v1.0, ClustalW, Bioedit and BLAST (Basic Local Alignment Search Tool) programme from NCBI.
3. RESULT AND DISCUSSION
3.1. Identification of GmLEA D-11 Gene on Various Soybean
Using the primer derived from the sequence of GmLEAD11 gene, PCR products with the size of about 701 bp were produced. The results showed that both of the DNA genome of soybean varieties treated with drought stress treatment and the control can be amplified by the primer (Figure 1). These indicates that the tolerant, moderate
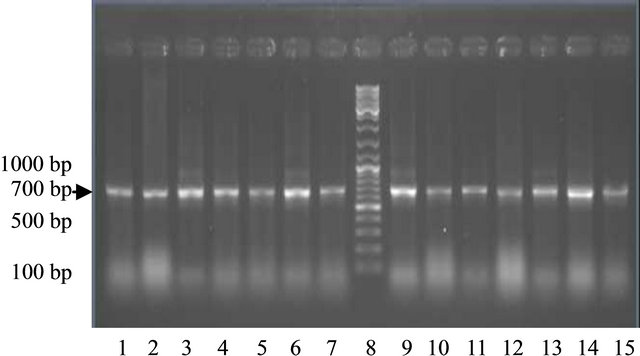
Figure 1. The PCR product in some varieties of soybean plants using primers LEA-D11 Lanes 1-7 (control); 1: Tanggamus; 2: Nanti; 3: Seulawah; 4: Tidar; 5: Wilis; 6: Burangrang l; 7: Detam; 8: Marker. Lane 9-15 (drought); 9: Tanggamus 10: Nanti; 11: Seulawah; 12 : Tidar; 13: Wilis; 14: Burangrang; 15: Detam 1.
and susceptible drought varieties both in control and drought stress treatment posses LEA-D11 gene.
Drought did not alter LEA-D11 gene, this is indicated by the appearance of bands at 700 bp in control and drought condition. Basically, a gene provides the instructions for making a protein and proteins influence the characteristics of plants. Gene is genetic material which more stable than protein. Environmental stresses do not change the gene but may change the expression of the gene such as protein alteration. However gene variation can be induced by mutagenic agents such as radiation and certain chemicals [22].
Comparing the sequence of Tanggamus varieties (drought tolerant) to the sequence of LEA-D11 of soybean in the NCBI datase resulting in the high homology of those sequences (Table 1).
The gene sequences of Tanggamus varieties had 100% similarity with Glycine max LEA-D11 gene for dehydrin. This means that the gene is amplified genes LEA-D11.
3.2. Comparison of LEA-D11 Sequence of Several Varieties of Soybeans
Sequence alignment between GmLEA-D11 Tanggamus varieties (drought tolerant) with other varieties used in this experiment (Nanti, Seulawah, Tidar, Wilis, Burangrang and Detam 1) treated with drought stress and the control without drought stress (Figure 2). The results showed that both in control and drought stress condition the sequence of LEA-D11 possessed by drought tolerant soybean varieties Tanggamus, Nanti, Seulawah and Tidar are not different from the sequence of GmLEA-D11 possessed by moderately tolerant varieties Burangrang and Wilis, however some sequence differences were detected in the drought-susceptible varieties, Detam-1.
Comparing the sequence of GmLEA-D11 gene possessed by Tanggamus with other soybean varieties, Nanti, Seulawah, Tidar, Wilis, Burangrang and Detam-1 under conditions without stress (control = K) with a variety Tanggamus, Nanti, Seulawah, Tidar, Wilis, Burangrang and Detam 1 in stress conditions (treatment = C) shows 6 mutation site. These mutation site were only found in Detam 1 but were not detected in other varieties. The changes of DNA sequence occur in Detam alter the amino acid in mutation site number 2 and 4. There is no changing the amino acids in mutation site number 1, 3, 5 and 6.
Mutation site 2 and 4 shows the nitrogen base changes. Mutation site number 2 shows the changing of amino acid from proline to serine, and mutation site 4 shows the changing of amino acid valine to Ileusine. This suggests that the difference in some nitrogen bases in DNA sequences have changed expression in response to drought stress become drought susceptible. However the sequences of GmLEA-D11 identified in this experiment were similar to the gene sequences possessed by drought tolerant varieties Tanggamus, Nanti, Seulawah, Tidar and moderately drought varieties Wilis and Burangrang. That similarity indicate that there is not only LEA-D11 gene which is responsible to drought tolerance but also other gene. There are hundreds of genes induced by drought stress has been identified [13].
Examined the drought resistance genes in soybean, and found that the sequence of GmDREB2 gene on different varieties of soybean are different, but the difference did not affect expression of the nature of drought tolerance [23]. It was suggested that not only GmDREB2 genes responsible for drought tolerant. There could be many genes that influence resistance to drought stress. [24] examined drought resistant gene DREB1 in several genotype of soybean, and discovered that the tolerance level of several soybean genotypes was not affected by variations in the sequences of DREB1 gene.
LEA-D11 gene is a gene that produces a functional protein dehydrin which is regulated by several genes. LEA genes work is influenced by other member of drought resistance gene family that can be expressed in certain circumstances, either simultaneously or alternately expressed depending on environmental conditions [6,25].
Some stress-responsive genes regulated by ABA [26- 29] shows two regulatory pathway of dehydrin accumulation in sunflower, which is ABA-dependent and ABAindependent. Transcription factors for LEA are DREB2 and DREB 1 which act to initiate the transcription of the gene [30].
Table 1. Homology sequence Tanggamus varieties comparison with soybean NCBI database.


Figure 2. The results of amino acids alignments GmLEA-D11 Tanggamus varieties with some varieties of soybean under conditions without stress and drought stress conditions. TK = Tanggamus control, NK = Nanti control, SK = Seulawah control, TC = Tidar control, WK = Wilis control, BK = Burangrang control, DK = Detam control, TC = Tanggamus drought, NC = Nanti drought, SC = Seulawah drought, TIC = Tidar drought, WC = Wilis drought, BC = Burangrang drought, DC = Detam drought.
The expression of certain gene is influenced by a number genes that can be active (on) or inactive (off) as depend on time and environment. DREB transcription factors and DRE element serves as a signal transduction under conditions of drought, salinity and cold stress. DREB transcription factors can control the expression of several target functional genes involved in plant tolerance to drought conditions, salinity and cold temperatures [31].
Evaluate the role of genes coding for dehydrin proteins (LEA-D11) during drought stress in arbuscular mycorrhizal Glycine max and Lactuca sativa [32]. The results show that GmLEA gene generally expressed only in drought stress treatment. This supports that the dehydrin is essential for plants to adapt in drought stress [25,29, 33,34]. Significantly, the introduction of many stressinducible genes transfer resulted in improved plant stress tolerance [35,36]. LEA-D11 gene specific primers designed can be used as molecular marker and capable of differentiating between drought susceptible and drought moderate or drought tolerant.
4. ACKNOWLEDGEMENTS
We sincerely thank Dr. Sri Widyarti, Molecular Biology Laboratory, Departement of Biology, University of Brawijaya, Dr. Arifin Noor, Biotechnology Laboratory, Departement of Agrotechnology, University of Brawijaya. We thank Rizza Pahlevi for technical assistant. Gratefully acknowledges financial support through Doctoral fellowship from University of Brawijaya.
REFERENCES
- Doyle, J.J. and Doyle, J.L. (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochemical Bulletin, 19, 11-15.
- Zhu, J.K., Hasegawa, P.M. and Bressan, R. (1997) Molecular aspect of osmotic stress in plant. Critical Reviews in Plant Sciences, 16, 253-277.
- Dure, L. (1993) Structural motif in LEA proteins. In: Close, T.J. and Bray, E.A., Eds., Reponse of Plants to Cellular Dehydration during Environmental Stress, American Society of Plant Physiologist, Rockville, 91-103.
- Ingram, J. and Barterls, D. (1996) The molecular basis of dehydration tolerance in plants. Annual Review of Plant Physiology and Plant Molecular Biology, 47, 277-403. doi:10.1146/annurev.arplant.47.1.377
- Thomashow, M.F. (1999) Plant cold acclimation: Freezing tolerance gene and regulatory mechanism. Annual Review of Plant Physiology and Plant Molecular Biology, 50, 571-599.
- Close, T.J. (1997) Dehydrin: A commonly in the response of plants to dehydration protein. Physiologia Plantarum, 100, 291-296. doi:10.1111/j.1399-3054.1997.tb04785.x
- Dure, L., Crouch, M., Harada, J., Ho, T.H.D., Mundy, J. and Quatrano, R. (1989) Common amino acid sequence domains among the lea protein of higher plants. Plant Molecular Biology, 12, 475-486. doi:10.1007/BF00036962
- Close, T.J. (1996) Dehydrin: Emergence of a biochemical role of a family of plant dehydration protein. Physiologia Plantarum, 97, 795-803. doi:10.1111/j.1399-3054.1996.tb00546.x
- Bray, E.A. (1997) Plant responses to water deficit. Trends in Plant Science, 2, 48-54. doi:10.1016/S1360-1385(97)82562-9
- Danyluk, J., Perron, A., Houde, M., Limin, A., Fowler, B., Benhamou, N. and Sarhan, F. (1998) Accumulation of an acidic dehydrin in the vicinity of the plasma membran during cold acclimation of wheat. The Plant Cell, 10, 623-638.
- Heyen, B.J., Alseikh, M.K., Smith, E.A., Torvik, C.F., Selas, D.F. and Randall, S.K. (2002) The calcium-binding activityn of vacuole associated, dehydin-like protein is regulated by phosporylation. Plant Physiology, 130, 675- 687. doi:10.1104/pp.002550
- Hara, M., Terashima, S., Fukaya, T. and Kuboi, T. (2003) Enhancement of cold tolerance and inhibition of lipid peroxidation by citrus dehydrin in transgenic tobacco. Planta, 217, 290-298.
- Shinozaki, K. and Yamaguchi-Shinozaki, K. (1996) Molecular responses to drought and cold stress. Current Opinion in Biotechnology, 7, 161-167. doi:10.1016/S0958-1669(96)80007-3
- Imai, R., Chang, L., Ohta, A., Bray, E.A. and Takagi, M. (1996) A lea-class gene of tomato confers salt and freezing tolerance when overexpressing in Saccharomyces cerevisae. Gene, 170, 243-248. doi:10.1016/0378-1119(95)00868-3
- Xu, D., Duan, X., Wang, B., Hong, B., Ho, T.H.D. and Wu, R. (1996) Expression of a late embryogenesis abundant protein gene, HVA 1 from barley confers tolerance to water deficits and salt stress in trangenic rice. Plant Physiology, 110, 249-257.
- Sivamani, E., Bahieldin, A., Wraith, J.M., Al-Niemo., T. Dyer, W.E., Ho, T.H.D. and Qu, R. (2000) Improved biomass productivity and water use efficiency under water deficit condition in transgenic wheat contituvely expressing the barley HVA 1 gene. Plant Science, 155, 1-9. doi:10.1016/S0168-9452(99)00247-2
- Dure, L. (1993) A repeating 11-mer amino acid sequence domains among the LEA protein of higher plant. Plant Journal, 3, 363-369. doi:10.1046/j.1365-313X.1993.t01-19-00999.x
- Hara, M., Fujinaga, M. and Kuboi, T. (2004) Radical scavenging activity and oxidative modification of citrus dehydrin. Plant Physiology and Biochemistry, 42, 657-662. doi:10.1016/j.plaphy.2004.06.004
- Hara, M., Fujinaga, M. and Kuboi, T. (2005) Metal binding by citrus dehydrin with histidine-rich domains. Journal of Experimental Botany, 56, 2695-2703. doi:10.1093/jxb/eri262
- Hara, M. (2009) The multifunctionality of dehydrins: An overview. Plant Signalling Behaviour, 5, 503-508.
- Gosal, S.S., Wani, S.H. and Manjit, S. (2009) Biotechnology and drought tolerance. Journal of Crop Improvement, 23, 19-54. doi:10.1080/15427520802418251
- Novak, F.J. and Brunner, H. (1992) Plant breeding induced mutation technology for crop improvement. IAEA Bulleting, 4, 25-33.
- Pahlevi, R. (2010) Study of marker specifity gene drought in variety of soybean (Glycine max) used PCR-sequencing. Thsesis, Post Graduate Programe, Brawijaya University, Malang.
- Mahmudah (2009) Identification gene drought DREB1 and P5CS in varian soybean (Glycine max) from selection in vitro used method PCR-sequencing. Post Graduate Programe, Brawijaya University, Malang.
- Cellier, F., Conejero, G., Breitler, J.C. and Casse, F. (1998) Molecular and physological responses to water deficit in drought tolerant and drought sensitive lines in sunflower. Plant Physiology, 116, 319-328. doi:10.1104/pp.116.1.319
- Choi, D.W., Zhu, B. and Close, T.J. (1990) The barley (Hordeum vulgare L.) dehydrin multigene family: Sequences, allele types, chromosome assignment, and expression characteristic of 11 Dhn genes of cv Dicktoo. Theoretical and Applied Genetics, 98, 1234-1247. doi:10.1007/s001220051189
- Cohen, A. and Bray, E.A. (1992) Nocleotide sequence of an ABA-induced tomato genes that is expressed in wilted vegetative organs and developing seeds. Plant Molecular Biology, 18, 411-413. doi:10.1007/BF00034969
- Godoy, J.A., Pardo, J.M. and Pintor-Toro, J.A. (1990) A tomato cDNA inducible by salt stress and absisic acid: Nucleotide sequence and expression pattern. Plant Molecular Biology, 15, 695-705. doi:10.1007/BF00016120
- Giordani, T., Natali, L., D’Ercole, A., Pugliesi, C., Fambrini, M., Vernieri, P., Vitagliano, C. and Cavallini, A. (1999) Expression of a dehydrin gene during embryo development and drought stress in ABA-deficient mutants of sunflower (Helianthus annuus L.). Plant Molecular Biology, 39, 739-748. doi:10.1023/A:1006194720022
- Shinozaki, K. and Yamaguchi-Shinozaki, K. (1997) Gene expression and signal transduction in water stress response. Plant Physiology, 115, 327-334. doi:10.1104/pp.115.2.327
- Qiang, L., Zhao, N.M., Yamaguchi-Shinozaki, K. and Shinozaki, K. (2000) Regulatory role od DREB transcription factors in plant drought, salt and cold tolerance. Chinese Science Bulletin, 45, 970-975. doi:10.1007/BF02884972
- Porcel, R.B., dan Miguelm J. and Ruiz-Lozano, J.M. (2004) Evaluation of genes encoding for delta 1-pyroline-5-carboxylate synthase (P5CS) during drought stress in arbuscular mycorrhizal Glycine max and Lactuca sativa plants. http://www.ncbi.nlm.nih.gov
- Robertson, M. and Chandler, P.M. (1992) Pea dehydrin: Identification, characterization and expression. Plant Molecular Biology, 19, 1031-1044. doi:10.1007/BF00040534
- Colmenero-Flores, J.M., Campos, F. and Garciarrubias, A.A. (1997) Characterization of phaseolus vulgaris cDNA clones responsive to water deficit: Identification of a novel late embryogenesis abundant-like protein. Plant Molecular Biology, 35, 393-405. doi:10.1023/A:1005802505731
- Zhang, J. and Kirkham, M.B. (2005) Enzymatic responses of the ascorbate-gluthatione cycle to drought in sorghum and sunflower plant. Plant Science, 113, 139-147. doi:10.1016/0168-9452(95)04295-4
- Umezawa, T., Fujita., M., Fujita., Y., Yamaguchi-Shinozaki, K. and Shinozaki, K. (2006) Engineering drought tolerance in plants: Discovering and tailoring genes unlock the future. Current Opinion in Biotechnology, 17, 113-122. doi:10.1016/j.copbio.2006.02.002

