 International Journal of Clinical Medicine, 2013, 4, 472-478 http://dx.doi.org/10.4236/ijcm.2013.410083 Published Online October 2013 (http://www.scirp.org/journal/ijcm) The Effect Analysis of Different Experimental Methods for the Diagnosis of Invasive Pulmonary Aspergillosis in a Rat Model* Jiancong Lin, Wenming Xu, Ming Li, Yanli Xin, Yuanyuan Niu, Changran Zhang#, Zelong Guo Department of Internal Medicine, Huang Pu Hospital of the First Affiliated Hospital, Sun Yat-sen University, Guangzhou, China. Email: #zhcr2303@sina.com.cn Received August 28th, 2013; revised September 25th, 2013; accepted October 10th, 2013 Copyright © 2013 Jiancong Lin et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. ABSTRACT Background: Consensus on the most reliable assays to detect invasive aspergillosis from minimally or noninvasive samples has not been reached. In this study, we compared the efficacy of an enzyme-linked immunosorbent assay (ELISA) for galactomannan (GM) detection and quantitative real-time PCR assay (qRT-PCR) for the diagnosis of inva- sive pulmonary aspergillosis in a rat model. Methods: Neutropenic, male Sprague-Dawley rats (specific pathogen free; 8 weeks old; weight, 200 ± 20 g) were immunosuppressed with cyclophosphamide and infected with Aspergillus fumi- gatus intratracheally. Tissue and whole blood samples were harvested on days 1, 3, 5, and 7 post-infection and exam- ined with GM ELISA and qRT-PCR. Results: On day 7, A. fumigatus DNA was amplified from 14 of 48 whole blood samples from immunosuppressed infected rats: day 1 (0/12), day 3 (0/12), day 5 (6/12), day 7 (8/12) post infection. The sensitivity and specificity of the qRT-PCR assay were 29.2% and 100%, respectively. Receiver operating characteristic curve (ROC) analysis indicated a Ct cut-off value of 15.35, and the area under the curve (AUC) was 0.627. The GM assay detected antigen in sera obtained on day 1 (5/12), day 3 (9/12), day 5 (12/12), and day 7 (12/12) post-infection, and thus had a sensitivity of 79.2% and a specificity of 100%. The ROC of the GM assay indicated that the optimal cut-off value was 1.40 (specificity, 100%; AUC, 0.919). Conclusions: The GM assay was more sensitive than qRT- PCR assay in diagnosing invasive pulmonary aspergillosis in rats. Keywords: Invasive Pulmonary Aspergillosis; Aspergillus; Galactomannan Antigen; Quantitative Real-Time PCR; Receiver Operating Characteristic Curve; Rat Model 1. Introduction The prevalence of invasive fungal infections and conse- quent mortality has increased throughout the last 2 dec- ades, and the reported mortality from an epidemiological study covering the period from 2002 to 2006 is 28.6% [1]. According to the TRANSNET database, half of extensive fungal infections are caused by Aspergillus sp. [2]. Most fungal infections occur in immunodeficient individuals, such as transplant recipients or those with leukemia re- ceiving chemotherapy, and pathological exacerbations of lung infections and the inability to diagnose fungal infec- tions are the major causes of death in these patients [3,4]. Early diagnosis of invasive fungal infections is critical for rapid treatment [3], but the lack of sensitive and spe- cific clinical symptoms and radiological patterns for in- vasive aspergillosis (IA) hinders early diagnosis. Tradi- tional histopathological examination and fungal culture relying on invasive procedures are relatively insensitive and not commonly used in clinical diagnosis due to the challenges of sampling pulmonary fluids or tissues from critically ill patients. While examination of bronchial- veolar lavage (BAL) fluid yields a higher detection rate than examination of blood or serum [5-7], obtaining BAL fluid is invasive, and many patients with IA have other severe diseases which limit BAL collection. Consensus on the most reliable assays to detect IA from minimally or noninvasive samples has not been reached, and is a major topic of current research [8,9]. Galactomannan (GM) is a cell wall component of As- pergillus, and its production increases during Aspergillus growth. Assays that detect fungal antigens such as GM *Yanli Xin and Jiancong Lin equally contributed to this manuscript. The authors declare that they have no competing interests. #Corresponding author. Copyright © 2013 SciRes. IJCM  The Effect Analysis of Different Experimental Methods for the Diagnosis of Invasive Pulmonary Aspergillosis in a Rat Model 473 by enzyme-linked immunosorbent assay (GM assay) or Aspergillus DNA by polymerase chain reaction (PCR) are emerging diagnostic methods; however, their speci- ficity and sensitivity require additional characterization and refinement. The sensitivity of the GM assay ranges from 60% to 100% for infected Aspergillus samples, and the specificity ranges from 80% to 100% [10-14]. The cut-off value has a significant impact on diagnosis be- cause of cross-reactivity [8,12,15-17]. Furthermore, treat- ment with anti-fungal therapies decreases the fungal load and reduces the GM concentration, which can fall below the detectable limit of the GM assay [18]. Given the ad- vantage of high sensitivity, a PCR assay coupled with an ELISA assay (PCR-ELISA) has been used for detection of Aspergillus [19,20]. However, poor specificity has been noted due to several interference factors, including environmental contamination [19]. In addition, the PCR- ELISA assay is more cumbersome for clinical laborato- ries [5]. Recently, a quantitative real-time PCR (qRT-PCR) assay was developed that may overcome the shortcom- ings of other methods [8,21]. qRT-PCR analysis of se- rum samples of patients with hematological malignancies at risk for IA was shown to have a sensitivity of 72.7% [5]. A commercially available qRT-PCR for the detection of Aspergillus DNA (MycAssay™ Aspergillus) has shown a sensitivity of 60% - 70% and a specificity of 95% for the detection of IA [22]. However, a study by Scotter et al. [19] indicated that GM testing by ELISA and the PCR-ELISA were more capable of early detection of fungal infection than RT-PCR examination. The aim of the study is to compare the sensitivity and specificity of the GM assay to the Aspergillus-specific nucleic acid qRT-PCR assay in a rat model of pulmonary IA. 2. Methods 2.1. Aspergillus fumigatus Preparation Lyophilized Aspergillus fumigatus (A. fumigatus) was recovered in 0.5 mL sterile broth, cultured in Sabouraud agar medium at 37˚C for 48 h, and subsequently at 30˚C for 3 - 5 days. Spores were eluted from the agar surface with 10 mL PBS containing 0.05% Tween-80. The sus- pension was filtered through 8 layers of sterile gauze to remove hyphae. The spore suspension was then trans- ferred to a 15-mL tube and centrifuged at 10,000 ×g for 15 min. The supernatant was discarded, and the pellet containing spores was resuspended in normal saline. Spore count was determined on a blood count plate, and the concentration was adjusted to 8 × 105 spores/L. Spore viability was determined by culture of serial dilutions of the spore suspension. 2.2. Induction of Pulmonary Aspergillosis in Rats Neutropenic rats were infected with Aspergillus as de- scribed by Zhang et al. [23]. Briefly, male Sprague- Dawley rats (specific pathogen free; 8 weeks old; weight, 200 ± 20 g) were housed with food and water ad libatum according to animal care guidelines. Rats received one of four treatments: immunosuppression and A. fumigatus infection (n = 48), immunosuppression (no infection, n = 6), infection (no immunosuppression, n = 6) and vehicle control (no immunosuppression, no infection, n = 12). Persistent immunosuppression was induced in the indi- cated groups by intraperitoneal injection of cyclophos- phamide, as described by Leenders et al. [24]. The dos- age of cyclophosphamide was determined by a prelimi- nary experiment. Cyclophosphamide (50 mg/kg i.p.) was injected 5 days prior to the Aspergillu s spore suspension inoculation. A second cyclophosphamide injection (40 mg/kg) was performed 1 day prior to inoculation, and a third injection (30 mg/kg) was performed on day 3 after inoculation. Control groups were injected with an equivalent volume of normal saline. After treatment with cyclophosphamide, all rats were consistently kept in a clean environment, and injected with levofloxacin, 10 mg/kg/d. Immunosuppressed and normal rats of the indicated groups were infected with A. fumigatus (8 × 105 spores) after anesthesia (chloral hydrate (3.5 mL/kg i.p.) and intubation. After connecting the endotracheal tube with a syringe, 0.1 mL of the spore suspension was injected. Rats were kept erect and rotated for 30 seconds, ensuring that the inoculation entered the trachea and was distrib- uted evenly in both lungs. Rats were sacrificed after 1 - 7 days and whole blood was collected by heart puncture. Rats which received both immunosuppression and in- fection (n = 48) were sacrificed on days 1, 3, 5, and 7 after inoculation, and were referred to as group 1, 2, 3, and 4, respectively (n = 12/group). Four ml of whole blood was collected and 3 mL blood was used to deter- mine Aspergillus DNA using the qRT-PCR method, and 100 μL of serum was prepared to measure the GM con- centrations using the ELISA method. Lung tissue was collected for biopsy and tissue culture. 2.3. Isolation of A. fumigatus DNA from Blood A.fumigatus DNA was isolated as described previously [25]. We used the physical method of grinding, similar to the bead beating, to break the cell walls of the fungi to release fungal DNA. The efficiency of fungal DNA ex- traction was about 96.8%. In brief, 3 mL of whole blood were treated with 1 mL of EDTA anticoagulant, lysed in 1 mL of erythrocyte lysis buffer (0.0l mol/L Tris-HCl pH Copyright © 2013 SciRes. IJCM  The Effect Analysis of Different Experimental Methods for the Diagnosis of Invasive Pulmonary Aspergillosis in a Rat Model 474 7.6, 0.01 mol/L NaCl, 0.005 mol/L MgCl2), mixed thor- oughly, and further treated with 1 mL of lysis buffer twice, allowing extensive erythrocyte lysis. After centri- fuged at 8000 ×g for 10 min, the pellet was washed with normal saline, treated with l00 μL proteinase K lysis buffer (20 mM Tris-HC1, 2.0 mM EDTA, 1.0% Triton X-l00, 2 mg/mL proteinase K), and subsequently 50 μL lysostaphin at a concentration of 50 μg/mL. The lysate was incubated at 60˚C for 60 min, treated with an equi- valent volume of phenol, chloroform, and isoamyl alco- hol. Thirty μL of A fumigatus DNA was precipitated us- ing ethanol, centrifuged at 12000 rpm/min for 20 min, and dissolved in Tris-EDTA buffer (10 mM Tris-Cl, pH 7.5 with 1 mM EDTA) for reserve. 2.4. Polymerase Chain Reaction and Product Analysis Specific primers were based on the target mRNA se- quence in GeneBank, harboring the CDS region of the mitochondrial translation optimization gene Mto1 of A. fumigatus. Primers were designed using Primer Express 2.0 software: forward primer, 5’-tttctccacccaggaacgtt-3’; reverse primer, 5’-cgaatccggagaggtgatacc-3’; probe, 5’- FAM-cagttgtgatgacgacacgcccagt-TAMRA-3’. Primers were synthesized using the ABI 3900 high-throughput DNA synthesizer. To determine the specificity of the qRT-PCR reaction, the aforementioned primers were assessed for their ability to amplify DNA from A. niger, A. flavus, A. terreus, Candida albicans, Cryptococcus neoformans, Staphylococcus aureus, and Pseudomonas aeruginosa [26]. The qRT-PCR reaction (50 μL) included 10 μL of 5 × reaction buffer (10 mM Tris-HCl pH 8.0, 50 mM KCl, 2 mM MgCl2), 10 pmol of each primer (10 pmol/uL), 2μL dNTPs (10 mM), 3U Taq DNA polymerase, and 4 μL cDNA or positive standard. Reaction was performed as follows: 93˚C for 3 min, and 40 cycles of 93˚C 30 s, 55˚C 45 s. Real-time PCR was carried out in an auto- mated fluorescent quantitative PCR cycler (ABI 7500), and the amplification curve was analyzed based on the exponential amplification and Ct value (cycle threshold). The Ct was defined as the number of cycles required for the fluorescent signal to cross the threshold (i.e., exceed background level). The Ct value was dose-dependent on the positive standard The standard curve quantification method used the target gene synthesized in Sangon (Shanghai). Ten-fold serial dilutions of the positive standard were prepared to produce the quantitative qRT-PCR gradients. Distilled water was used as a negative control. The optical density (OD) 260/280 of the purified DNA was >1.8, and thus qualified. DNA concentration (copies/μL) was calculated with OD 260 over the fragment length, namely the posi- tive standard. After dilution of recombinant plasmid, qRT-PCR amplification was performed following the optimal procedures. The LOD was measured according to amplification curve derived from qRT-PCR. The cor- relation between Ct value and DNA copy was Ct = −3.347424 × log copy number + 35.885406. 2.5. GM Antigen Detection (GM Assay) The GM assay was performed using the Platelia Asper- gillus kit (Bio-Rad Corporation, France), following the manufacturer’s instructions for the preparation of sam- ples. Optical densities (OD) at a primary wavelength of 450 nm and a secondary wavelength of 620 nm were determined. In each experiment, positive serum control, negative serum control, standardized control, and serum samples were run in triplicate. The mean OD derived from the standardized control was used as a standard (GM OD Index = OD of sample/OD of standard serum), and kept in the range of 0.3 to 0.8. All experiments were internally controlled: The ratio of OD from the positive control to OD from the standard was over 2, and the ratio of OD from the negative control to the OD of the stan- dard was below 0.4. 2.6. Histopathological Examination and Tissue Culture Rats that received both cyclophosphamide and infection were sacrificed on 1, 3, 5, and 7 days after A. fumigatus inoculation (n = 12 each group). Rats in the other groups were sacrificed on day 7. Blood samples and right lung tissue were used for fungal culture described above. The left lung tissue was fixed in 10% formalin and embedded in paraffin. The paraffin sections were primarily stained with hematoxylin & eosin (HE), and further with peri- odic acid-Schiff stain followed by histopathological ex- amination. 2.7. Statistical Analysis The values of the GM concentration for each group were reported as mean ± standard deviation. Wilcoxon rank sum test was used to analyze the difference between each two groups. A receiver operating characteristic (ROC) curve was created with the statistical software used for all analyses (SPSS version 16.0). A value of p < 0.05 was considered to indicate statistical significance. 3. Results 3.1. Histopathology of Pulmonary A. fumigatus Infection Immunosuppressed, infected rats showed progressive accumulation of hyphae in the alveoli (Figure 1). Many Copyright © 2013 SciRes. IJCM  The Effect Analysis of Different Experimental Methods for the Diagnosis of Invasive Pulmonary Aspergillosis in a Rat Model Copyright © 2013 SciRes. IJCM 475 (a) (b) (c) (d) (e) Figure 1. Histopathological analysis. Lung tissues were harvested from immunosuppressed, infected rats on day 1(a), 3(b), 5(c), and 7(d) (100× magnification), and from the control groups on day 7(e) (400× magnification). Lung tissue from healthy controls. Aspergillus spores and mild inflammation were found in the lung tissue of the neutropenic rats on the 1st day post infection (Figure 1(a)). On the 3rd day, the alveoli exhib- ited accumulated spores, a small presence of hyphae, hyperplasia of alveolar epithelial cells, and broadened alveolar septa. Cellulose exudate and hemorrhage in the alveolar spaces were also observed (Figure 1(b)). On the 5th day post-infection abnormal morphology of the al- veolar structures was apparent, and the spores had bloomed: hollow, colorless and acute angle-branching hyphae and granuloma were present (Figure 1(c)). Clear exudate was located in the alveolar spaces, and severe hemorrhage was evident in the capillaries. The alveolar septa had broadened, and part of the alveolar structure was damaged. On the 7th day, the aforementioned cha- racteristics had become more apparent and the alveolar structure was no longer clear (Figure 1(d)). The alveoli of the vehicle control group had normal morphology (Figure 1(e)) No apparent inflammation was noted in nonimmunosuppressed infected mice. 3.2. Detection of A. fumigatus DNA Isolated from Immunosuppressed-Infected Rats No A. fumigatus target sequence was amplified from the uninfected control group (immunosuppression and no infection, n = 6), and vehicle control group (no immuno- suppression, no infection, n = 12). The target sequence was successfully amplified in 14 out of 48 samples from immunosuppressed-infected rats. The 14 positive sam- ples were obtained on day 5 (6/12), and day 7 (8/12): no positive samples were obtained on day 1 or day 3. No DNA was identified in the no immunosuppression/infec- tion control samples (0/6) or in the no infection/immu- nosuppression control samples (0/6). Amplification curve and quantitative analysis showed that the DNA content ranged from 5 × 102 to 5.77 × 103 copies/μL of blood. The lower limit for qRT-PCR detec- tion was 100 copies/μL of blood, and was determined according to amplification curve derived from qRT-PCR. Only samples from infected animals yielded positive RT-PCR curves, and the PCR reactions spiked with DNA from A. niger, A. fla vus, A. terreus, Ca n d ida albicans, Cryptococcus neoformans, Staphylococcus aureus, and Pseudomonas aeruginosa yielded negative results. Thus, the primer set for A. fumigatus showed high specificity. Receiver operating characteristic curve (ROC) analysis indicated a Ct cut-off value of 15.35, and the area under the curve (AUC) was 0.627. 3.3. Determination of GM by ELISA A commercial kit was utilized for the determination of GM, and the mean concentrations for each group are shown in Table 1. As expected, the concentration of GM ncreased from day 1 to day 7, in agreement with the i 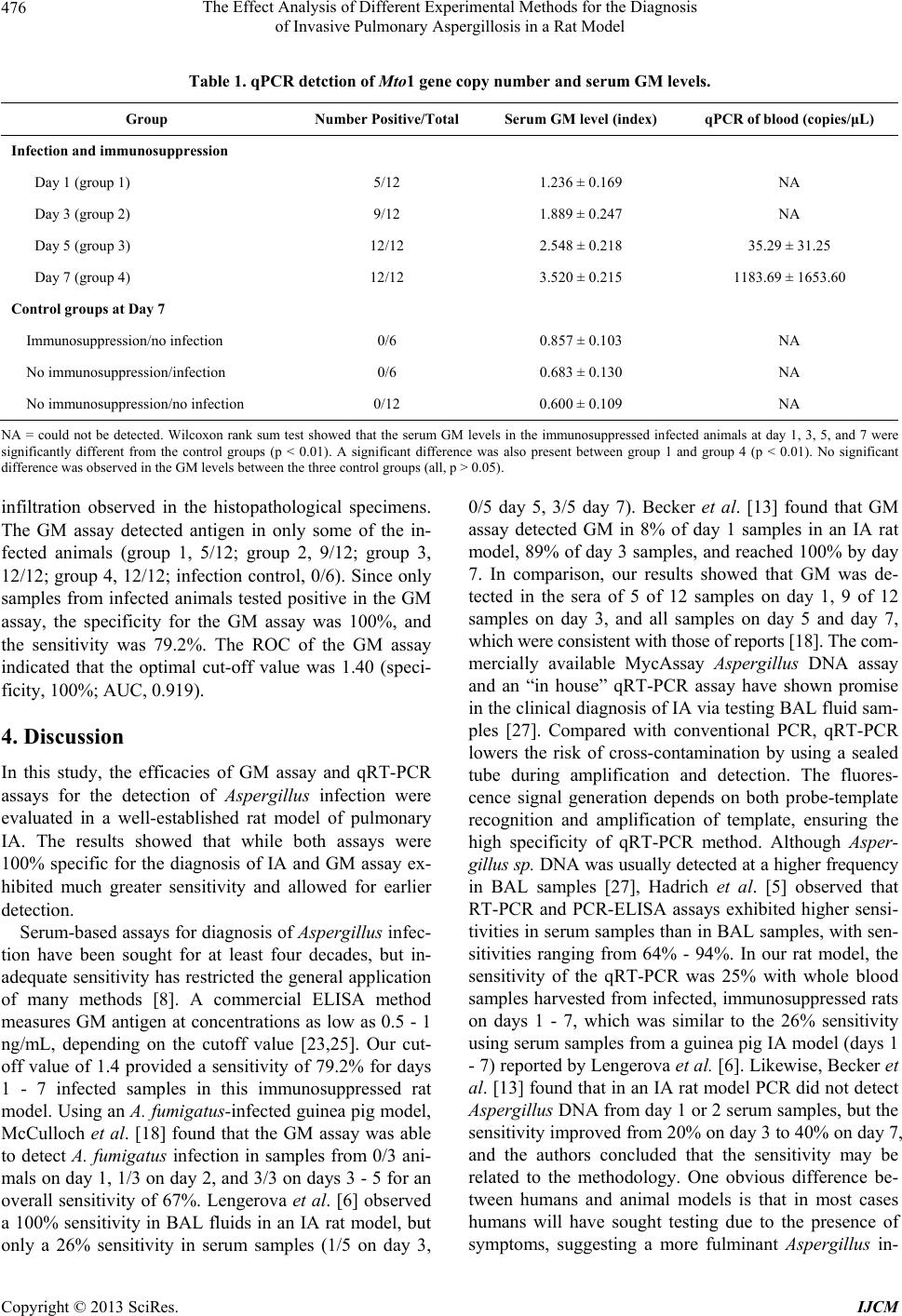 The Effect Analysis of Different Experimental Methods for the Diagnosis of Invasive Pulmonary Aspergillosis in a Rat Model 476 Table 1. qPCR detction of Mto1 gene copy number and serum GM levels. Group Number Positive/Total Serum GM level (index) qPCR of blood (copies/μL) Infection and immunosuppression Day 1 (group 1) 5/12 1.236 ± 0.169 NA Day 3 (group 2) 9/12 1.889 ± 0.247 NA Day 5 (group 3) 12/12 2.548 ± 0.218 35.29 ± 31.25 Day 7 (group 4) 12/12 3.520 ± 0.215 1183.69 ± 1653.60 Control groups at Day 7 Immunosuppression/no infection 0/6 0.857 ± 0.103 NA No immunosuppression/infection 0/6 0.683 ± 0.130 NA No immunosuppression/no infection 0/12 0.600 ± 0.109 NA NA = could not be detected. Wilcoxon rank sum test showed that the serum GM levels in the immunosuppressed infected animals at day 1, 3, 5, and 7 were significantly different from the control groups (p < 0.01). A significant difference was also present between group 1 and group 4 (p < 0.01). No significant difference was observed in the GM levels between the three control groups (all, p > 0.05). infiltration observed in the histopathological specimens. The GM assay detected antigen in only some of the in- fected animals (group 1, 5/12; group 2, 9/12; group 3, 12/12; group 4, 12/12; infection control, 0/6). Since only samples from infected animals tested positive in the GM assay, the specificity for the GM assay was 100%, and the sensitivity was 79.2%. The ROC of the GM assay indicated that the optimal cut-off value was 1.40 (speci- ficity, 100%; AUC, 0.919). 4. Discussion In this study, the efficacies of GM assay and qRT-PCR assays for the detection of Aspergillus infection were evaluated in a well-established rat model of pulmonary IA. The results showed that while both assays were 100% specific for the diagnosis of IA and GM assay ex- hibited much greater sensitivity and allowed for earlier detection. Serum-based assays for diagnosis of Aspergillus infec- tion have been sought for at least four decades, but in- adequate sensitivity has restricted the general application of many methods [8]. A commercial ELISA method measures GM antigen at concentrations as low as 0.5 - 1 ng/mL, depending on the cutoff value [23,25]. Our cut- off value of 1.4 provided a sensitivity of 79.2% for days 1 - 7 infected samples in this immunosuppressed rat model. Using an A. fumigatus-infected guinea pig model, McCulloch et al. [18] found that the GM assay was able to detect A. fumigatus infection in samples from 0/3 ani- mals on day 1, 1/3 on day 2, and 3/3 on days 3 - 5 for an overall sensitivity of 67%. Lengerova et al. [6] observed a 100% sensitivity in BAL fluids in an IA rat model, but only a 26% sensitivity in serum samples (1/5 on day 3, 0/5 day 5, 3/5 day 7). Becker et al. [13] found that GM assay detected GM in 8% of day 1 samples in an IA rat model, 89% of day 3 samples, and reached 100% by day 7. In comparison, our results showed that GM was de- tected in the sera of 5 of 12 samples on day 1, 9 of 12 samples on day 3, and all samples on day 5 and day 7, which were consistent with those of reports [18]. The com- mercially available MycAssay Aspergillus DNA assay and an “in house” qRT-PCR assay have shown promise in the clinical diagnosis of IA via testing BAL fluid sam- ples [27]. Compared with conventional PCR, qRT-PCR lowers the risk of cross-contamination by using a sealed tube during amplification and detection. The fluores- cence signal generation depends on both probe-template recognition and amplification of template, ensuring the high specificity of qRT-PCR method. Although Asper- gillus sp. DNA was usually detected at a higher frequency in BAL samples [27], Hadrich et al. [5] observed that RT-PCR and PCR-ELISA assays exhibited higher sensi- tivities in serum samples than in BAL samples, with sen- sitivities ranging from 64% - 94%. In our rat model, the sensitivity of the qRT-PCR was 25% with whole blood samples harvested from infected, immunosuppressed rats on days 1 - 7, which was similar to the 26% sensitivity using serum samples from a guinea pig IA model (days 1 - 7) reported by Lengerova et al. [6]. Likewise, Becker et al. [13] found that in an IA rat model PCR did not detect Aspergillus DNA from day 1 or 2 serum samples, but the sensitivity improved from 20% on day 3 to 40% on day 7, and the authors concluded that the sensitivity may be related to the methodology. One obvious difference be- tween humans and animal models is that in most cases humans will have sought testing due to the presence of symptoms, suggesting a more fulminant Aspergillus in- Copyright © 2013 SciRes. IJCM  The Effect Analysis of Different Experimental Methods for the Diagnosis of Invasive Pulmonary Aspergillosis in a Rat Model 477 fection at time of sampling than the early time points in animal models. Presumably, the longer incubation period would favor Asp ergillus replication, and may promote alveolar damage and seepage of more Aspergillus spores into the blood. In this study, analysis of qRT-PCR results showed that an AUC of 0.627 produced the maximum specificity, and although highly consistent with the pathological changes in lung tissue the qRT-PCR method was not sufficiently sensitive (25.9%) to solely rely on for early diagnosis. In contrast, the AUC of the GM assay was significantly higher (0.919), and was able to detect Aspergillus in 79% of day 1 - 7 samples. Interestingly, Torelli et al. [27] de- scribed a cohort of patients with suspected IA, and >90% of patients with BAL samples that were positive for As- pergillus DNA by MycAssay and RT-PCR also had GM-positive BAL samples. Together, it seems that GM assay and qRT-PCR assays both are capable of early de- tection of Aspergillus infection but further study for es- tablishing a gold standard is warranted [28]. 5. Acknowledgements The research was supported by Science and Technology Planning Project of Guangdong Province, China, No. 2011B090400118 , No. 2009B03081137 and Guangzhou Public Health Bureau (Grant No. 2009-YB-181, 2009- YB-183). REFERENCES [1] C. Hahn-Ast, A. Glasmacher, S. Muckter, A. Schmitz, A. Kraemer, G. Marklein, P. Brossart and M. von Lilienfeld- Toal, “Overall Survival and Fungal Infection-Related Mortality in Patients with Invasive Fungal Infection and Neutropenia after Myelosuppressive Chemotherapy in a Tertiary Care Centre from 1995 to 2006,” Journal of An- timicrobial Chemotherapy, Vol. 65, No. 4, 2010, pp. 761- 768. http://dx.doi.org/10.1093/jac/dkp507 [2] A. H. Groll and L. McNeil Grist, “Current Challenges in the Diagnosis and Management of Invasive Fungal Infec- tions: Report from the 15th International Symposium on Infections in the Immunocompromised Host: Thessalo- niki, Greece, 22-25 June 2008,” International Journal of Antimicrobial Agents, Vol. 33, No. 2, 2009, pp. 101-104. http://dx.doi.org/10.1016/j.ijantimicag.2008.08.014 [3] M. von Eiff, N. Roos, R. Schulten, M. Hesse, M. Zuhls- dorf and J. van de Loo, “Pulmonary aspergillosis: Early Diagnosis Improves Survival,” Respiration, Vol. 62, No. 6, 1995, pp. 341-347. http://dx.doi.org/10.1159/000196477 [4] G. Chamilos, M. Luna, R. E. Lewis, G. P. Bodey, R. Che- maly, J. J. Tarrand, A. Safdar, I. I. Raad and D. P. Kon- toyiannis, “Invasive Fungal Infections in Patients with Hematologic Malignancies in a Tertiary Care Cancer Cen- ter: An Autopsy Study over a 15-Year Period (1989- 2003),” Haematologica, Vol. 91, No. 7, 2006, pp. 986-989. [5] I. Hadrich, C. Mary, F. Makni, M. Elloumi, H. Dumon, A. Ayadi and S. Ranque, “Comparison of PCR-ELISA and Real-Time PCR for Invasive Aspergillosis Diagnosis in Patients with Hematological Malignancies,” Medica l My- cology, Vol. 49, No. 5, 2011, pp. 489-494. [6] M. Lengerova, I. Kocmanova, Z. Racil, K. Hrncirova, S. Pospisilova, J. Mayer, L. K. Najvar, N. P. Wiederhold, W. R. Kirkpatrick ad T. F. Patterson, “Detection and Meas- urement of Fungal Burden in a Guinea Pig Model of In- vasive Pulmonary Aspergillosis by Novel Quantitative Nested Real-Time PCR Compared with Galactomannan and (1,3)-Beta-D-glucan Detection,” Journal of Clinical Microbiology, Vol. 50, No. 3, 2012, pp. 602-608. http://dx.doi.org/10.1128/JCM.05356-11 [7] V. R. Aquino, F. Nagel, H. F. Andreolla, F. de-Paris, M. O. Xavier, L. Z. Goldani, D. W. Denning and A. C. Pas- qualotto, “The Performance of Real-Time PCR, Galacto- mannan, and Fungal Culture in the Diagnosis of Invasive Aspergillosis in Ventilated Patients with Chronic Obstruc- tive Pulmonary Disease (COPD),” Mycopathologia, Vol. 174, No. 2, 2012, pp. 163-169. http://dx.doi.org/10.1007/s11046-012-9531-1 [8] M. J. Ruping, J. J. Vehreschild, A. Groll, C. Lass-Florl, H. Ostermann, M. Ruhnke and O. A. Cornely, “Current Is- sues in the Clinical Management of Invasive Aspergillo- sis—The AGIHO, DMykG, OGMM and PEG Web-Based Survey and Expert Consensus Conference 2009,” Myco- ses, Vol. 54, No. 5, 2011, pp. e557-e568. http://dx.doi.org/10.1111/j.1439-0507.2010.01989.x [9] Y. Zhao, S. Park, P. Warn, R. Shrief, E. Harrison and D. S. Perlin, “Detection of Aspergillus fumigatus in a Rat Model of Invasive Pulmonary Aspergillosis by Real-Time Nucleic Acid Sequence-Based Amplification,” Journal of Clinical Microbiology, Vol. 48, No. 4, 2010, pp. 1378- 1383. http://dx.doi.org/10.1128/JCM.02214-09 [10] M. Hoenigl, H. J. Salzer, R. B. Raggam, T. Valentin, A. Rohn, A. Woelfler, et al., “Impact of Galactomannan Test- ing on the Prevalence of Invasive Aspergillosis in Pa- tients with Hematological Malignancies,” Medical Myco- logy, Vol. 50, No. 3, 2012, pp. 266-269. http://dx.doi.org/10.3109/13693786.2011.603102 [11] I. Hadrich, F. Makni, F. Cheikhrouhou, S. Neji, I. Amouri, H. Sellami, H. Trabelsi, H. Bellaaj, M. Elloumi and A. Ayadi, “Clinical Utility and Prognostic Value of Galacto- mannan in Neutropenic Patients with Invasive Aspergil- losis,” Pathologie Biologie (Paris), Vol. 60, No. 6, 2012, pp. 357-361. http://dx.doi.org/10.1016/j.patbio.2011.10.011 [12] T. J. Walsh, M. C. Wissel, K. J. Grantham, R. Petraitiene, V. Petraitis, M. Kasai, A. Francesconi, M. P. Cotton, J. E. Hughes, L. Greene, et al., “Molecular Detection and Spe- cies-Specific Identification of Medically Important As- pergillus Species by Real-Time PCR in Experimental In- vasive Pulmonary Aspergillosis,” Journal of Clinical Mi- crobiology, Vol. 49, No. 12, 2011, pp. 4150-4157. http://dx.doi.org/10.1128/JCM.00570-11 [13] M. J. Becker, S. de Marie, D. Willemse, H. A. Verbrugh and I. A. Bakker-Woudenberg, “Quantitative Galactoman- nan Detection Is Superior to PCR in Diagnosing and Copyright © 2013 SciRes. IJCM  The Effect Analysis of Different Experimental Methods for the Diagnosis of Invasive Pulmonary Aspergillosis in a Rat Model Copyright © 2013 SciRes. IJCM 478 Monitoring Invasive Pulmonary Aspergillosis in an Ex- perimental Rat Model,” Journal of Clinical Microbiology, Vol. 38, No. 4, 2000, pp. 1434-1438. [14] M. M. Leeflang, Y. J. Debets-Ossenkopp, C. E. Visser, R. J. Scholten, L. Hooft, H. A. Bijlmer, J. B. Reitsma, P. M. Bossuyt and C. M. Vandenbroucke-Grauls, “Galactoman- nan Detection for Invasive Aspergillosis in Immunocom- promized Patients,” Cochrane Database of Systematic Reviews, Vol. 8, No. 4, 2008, Article ID: 007394. [15] S. F. Yeo and B. Wong, “Current Status of Nonculture Methods for Diagnosis of Invasive Fungal Infection,” Clinical Microbiology Review, Vol. 15, No. 3, 2002, pp. 465-484. http://dx.doi.org/10.1128/CMR.15.3.465-484.2002 [16] L. Millon, F. Grenouillet, J. Crouzet, F. Larosa, S. Loewert, A. P. Bellanger, E. Deconinck and F. Legrand, “False- Positive Aspergillus Real-Time PCR Assay Due to a Nu- tritional Supplement in a Bone Marrow Transplant Reci- pient with GVH Disease,” Medical Mycology, Vol. 48, No. 4, 2010, pp. 661-664. http://dx.doi.org/10.3109/13693780903451836 [17] T. J. Walsh, S. Shoham, R. Petraitiene, T. Sein, R. Schau- fele, A. Kelaher, H. Murray, C. Mya-San, J. Bacher and V. Petraitis, “Detection of Galactomannan Antigenemia in Patients Receiving Piperacillin-Tazobactam and Correla- tions between in Vitro, in Vivo, and Clinical Properties of the Drug-Antigen Interaction,” Journal of Clinical Mi- crobiology, Vol. 42, No. 10, 2004, pp. 4744-4748. http://dx.doi.org/10.1128/JCM.42.10.4744-4748.2004 [18] E. McCulloch, G. Ramage, R. Rajendran, D. F. Lappin, B. Jones, P. Warn, R. Shrief, W. R. Kirkpatrick, T. F. Patter- son and C. Williams, “Antifungal Treatment Affects the Laboratory Diagnosis of Invasive Aspergillosis,” Journal of Clinical Pathology, Vol. 65, No. 1, 2012, pp. 83-86. http://dx.doi.org/10.1136/jcp.2011.090464 [19] J. M. Scotter and S. T. Chambers, “Comparison of Galac- tomannan Detection, PCR-Enzyme-Linked Immunosor- bent Assay, and Real-Time PCR for Diagnosis of Inva- sive Aspergillosis in a Neutropenic Rat Model and Effect of Caspofungin Acetate,” Clinical and Diagnostic Labo- ratory Immunology, Vol. 12, No. 11, 2005, pp. 1322- 1327. [20] F. Suarez, O. Lortholary, S. Buland, M. T. Rubio, D. Ghez, V. Mahe, G. Quesne, S. Poiree, A. Buzyn, B. Varet, et al., “Detection of Circulating Aspergillus fumigatus DNA by Real-Time PCR Assay of Large Serum Volumes Im- proves Early Diagnosis of Invasive Aspergillosis in High- Risk Adult Patients under Hematologic Surveillance,” Journal of Clinical Microbiology, Vol. 46, No. 11, 2008, pp. 3772-3777. http://dx.doi.org/10.1128/JCM.01086-08 [21] M. L. Luong, C. J. Clancy, A. Vadnerkar, E. J. Kwak, F. P. Silveira, M. C. Wissel, K. J. Grantham, R. K. Shields, M. Crespo, J. Pilewski, et al., “Comparison of an Asper- gillus Real-Time Polymerase Chain Reaction Assay with Galactomannan Testing of Bronchoalvelolar Lavage Fluid for the Diagnosis of Invasive Pulmonary Aspergillosis in Lung Transplant Recipients,” Clinical Infectious Diseases, Vol. 52, No. 10, 2011, pp. 1218-1226. http://dx.doi.org/10.1093/cid/cir185 [22] P. L. White, M. D. Perry, A. Moody, S. A. Follett, G. Morgan and R. A. Barnes, “Evaluation of Analytical and Preliminary Clinical Performance of Myconostica Myc- Assay Aspergillus When Testing Serum Specimens for Diagnosis of Invasive Aspergillosis,” Journal of Clinical Microbiology, Vol. 49, No. 6, 2011, pp. 2169-2174. http://dx.doi.org/10.1128/JCM.00101-11 [23] C. R. Zhang, Y. C. Tang, K. Kawakami, T. T. Zhang, K. X. Zhang and J. X. Zhu, “An Experimental Study of the Therapeutic Effect of Interleukin-2 and Interleukin-12 with and without Amphotericin B on Pulmonary Fungal Infec- tion,” Zhonghua Jie He He Hu Xi Za Zhi, Vol. 27, No. 4, 2004, pp. 234-236. [24] A. C. Leenders, S. de Marie, M. T. ten Kate, I. A. Bakker- Woudenberg and H. A. Verbrugh, “Liposomal Amphote- ricin B (AmBisome) Reduces Dissemination of Infection as Compared with Amphotericin B Deoxycholate (Fun- gizone) in a Rate Model of Pulmonary Aspergillosis,” Journal of Antimicrobial Chemotherapy, Vol. 38, No. 2, 1996, pp. 215-225. http://dx.doi.org/10.1093/jac/38.2.215 [25] J. Loffler, H. Hebart, U. Schumacher, H. Reitze and H. Einsele, “Comparison of Different Methods for Extrac- tion of DNA of Fungal Pathogens from Cultures and Blood,” Journal of Clinical Microbiology, Vol. 35, No. 12, 1997, pp. 3311-3312. [26] J. Liu, Y. Shi, D. He, P. Liu, Y. Zhang, L. Xu and L. Wang, “Development of Real-Time PCR Method to De- tect Aspergillus fumigates,” Biotechnology, Vol. 19, No. 1, 2009, pp. 34-36. [27] R. Torelli, M. Sanguinetti, A. Moody, L. Pagano, M. Cai- ra, E. De Carolis, L. Fuso, G. De Pascale, G. Bello, M. Antonelli, et al., “Diagnosis of Invasive Aspergillosis by a Commercial Real-Time PCR Assay for Aspergillus DNA in Bronchoalveolar Lavage Fluid Samples from High-Risk Patients Compared to a Galactomannan En- zyme Immunoassay,” Journal of Clinical Microbiology, Vol. 49, No. 12, 2011, pp. 4273-4278. http://dx.doi.org/10.1128/JCM.05026-11 [28] M. A. Mennink-Kersten, J. P. Donnelly and P. E. Verweij, “Detection of Circulating Galactomannan for the Diagno- sis and Management of Invasive Aspergillosis,” Lancet Infectious Disease, Vol. 4, No. 6, 2004, pp. 349-357. http://dx.doi.org/10.1016/S1473-3099(04)01045-X
|