 Advances in Bioscience and Biotechnology, 2013, 4, 21-35 ABB http://dx.doi.org/10.4236/abb.2013.410A3004 Published Online October 2013 (http://www.scirp.org/journal/abb/) RNA species whose transcription is totally silent in pre-MBT stage are not mRNA but rRNA and possible involvement of weak bases (ammonium salts and/or amines) in the transcriptional silence of rRNA genes during the pre-MBT stage in Xenopus early embryos Koichiro Shiokawa1,2 1Department of Judo Therapy, Faculty of Medical Technology, Teikyo University, Utsunomiya, Japan 2Department of Biosciences, School of Science and Engineering, Teikyo University, Utsunomiya, Japan Email: shiokawa@nasu.bio.teikyo-u.ac.jp Received 23 July 2013; revised 24 August 2013; accepted 15 September 2013 Copyright © 2013 Koichiro Shiokawa. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. ABSTRACT In Xenopus laevis embryogenesis, fertilized eggs un- dergo 12 cycles of synchronous divisions and reach the stage called midblastula transition (MBT). It has long been believed that during the first 12 cycles of cleavage (pre-MBT stage), transcriptional activity of the zygotic nuclei is totally absent. However, hetero- geneous mRNA-like RNA is synthesized in pre-MBT stage embryos, and exogenously-injected bacterial CAT genes with SV40 promoter are expressed from the cleavage stage. Nevertheless, the synthesis of rRNA as detected by rRNA-specific 2’-O-methylation does not take place in pre-MBT embryos and starts only from the latter half of the MBT stage, corroborating the fact that formation of definitive nucleoli as well as the transcription of microinjected rRNA genes starts only at and after MBT stage. Thus, while mRNA-like RNA synthesis occurs from pre-MBT stage, synthesis of rRNA is controlled in the way that transcription of rRNA genes is totally silent during pre-MBT stage and is initiated only at the latter half of MBT stage. Once initiated, the rate of the synthesis of rRNA is constant throughout later stages on a per-cell basis. We searched substances which are responsible for the transcriptional silence of rRNA genes during the pre- MBT stage. Weak bases such as ammonium ion and amines selectively inhibited rRNA synthesis at the transcriptional level in post-MBT stage embryo cells. Since we found that the level of ammonia extracted from embryos is much higher in pre-MBT embryos than in post-MBT embryos, we suggest that weak bases like ammonium ion could be responsible for the transcriptional silence of rRNA genes by slightly in- creasing intracellular pH during the pre-MBT. Keywords: Pre-MBT Transcription; Absence of rRNA Synthesis; Initiation of rRNA Synthesis; Nucleolus Formation; Weak Bases; Amines; Ammonium Ion; Xenopus Embryogenesis 1. INTRODUCTION MBT is the important time point of transition from the phase of cleavage division to the phase of morphogenetic cell interactions. After 12 rounds of cleavage cell cycles, Xenopus fertilized eggs reach midblastula stage (4096 cells/embryo), or the stage of so-called midblastula tran- sition (MBT), when G1 and G2 phases reappear in the cell cycle [1,2]. During the cleavage stage, translation of maternal mRNA takes place actively [3,4]. At MBT, cell division shifts from synchronous to asynchronous one [5, 6], cell cycles shift from a checkpoint-unregulated to checkpoint-regulated state [7,8], and cells, especially those at the dorsal marginal zone, acquire motility [9]. Also, strongly activated, at least on a per-embryo basis but not necessarily on a per-cell basis, is the transcriptio- nal activity from zygotic nuclear genes [10-17]. Also, the per-embryo activity of the transcription of exogenously- introduced bacterial CAT (chloramphenicole acetyltrans- ferase) genes with pan-expression promoter of SV40 vi- rus starts to increase from cleavage stage on [18,19]. Since the reports of Newport and Kirschner [5,6], it has long been believed that there is no transcription at all before MBT stage. However, recent results about this OPEN ACCESS 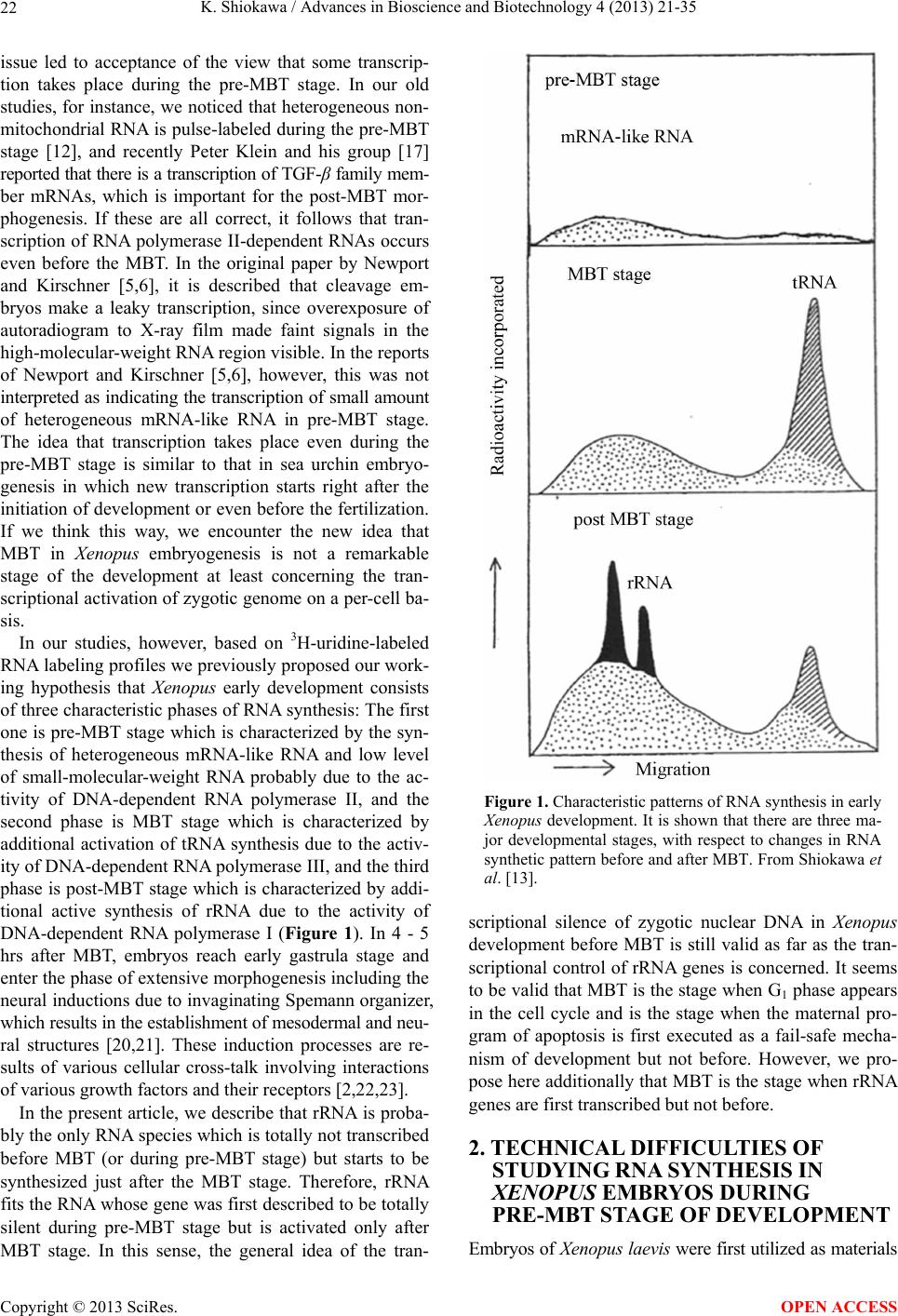 K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 22 issue led to acceptance of the view that some transcrip- tion takes place during the pre-MBT stage. In our old studies, for instance, we noticed that heterogeneous non- mitochondrial RNA is pulse-labeled during the pre-MBT stage [12], and recently Peter Klein and his group [17] reported that there is a transcription of TGF-β family mem- ber mRNAs, which is important for the post-MBT mor- phogenesis. If these are all correct, it follows that tran- scription of RNA polymerase II-dependent RNAs occurs even before the MBT. In the original paper by Newport and Kirschner [5,6], it is described that cleavage em- bryos make a leaky transcription, since overexposure of autoradiogram to X-ray film made faint signals in the high-molecular-weight RNA region visible. In the reports of Newport and Kirschner [5,6], however, this was not interpreted as indicating the transcription of small amount of heterogeneous mRNA-like RNA in pre-MBT stage. The idea that transcription takes place even during the pre-MBT stage is similar to that in sea urchin embryo- genesis in which new transcription starts right after the initiation of development or even before the fertilization. If we think this way, we encounter the new idea that MBT in Xenopus embryogenesis is not a remarkable stage of the development at least concerning the tran- scriptional activation of zygotic genome on a per-cell ba- sis. In our studies, however, based on 3H-uridine-labeled RNA labeling profiles we previously proposed our work- ing hypothesis that Xenopus early development consists of three characteristic phases of RNA synthesis: The first one is pre-MBT stage which is characterized by the syn- thesis of heterogeneous mRNA-like RNA and low level of small-molecular-weight RNA probably due to the ac- tivity of DNA-dependent RNA polymerase II, and the second phase is MBT stage which is characterized by additional activation of tRNA synthesis due to the activ- ity of DNA-dependent RNA polymerase III, and the third phase is post-MBT stage which is characterized by addi- tional active synthesis of rRNA due to the activity of DNA-dependent RNA polymerase I (Figure 1). In 4 - 5 hrs after MBT, embryos reach early gastrula stage and enter the phase of extensive morphogenesis including the neural inductions due to invaginating Spemann organizer, which results in the establishment of mesodermal and neu- ral structures [20,21]. These induction processes are re- sults of various cellular cross-talk involving interactions of various growth factors and their receptors [2,22,23]. In the present article, we describe that rRNA is proba- bly the only RNA species which is totally not transcribed before MBT (or during pre-MBT stage) but starts to be synthesized just after the MBT stage. Therefore, rRNA fits the RNA whose gene was first described to be totally silent during pre-MBT stage but is activated only after MBT stage. In this sense, the general idea of the tran- Figure 1. Characteristic patterns of RNA synthesis in early Xenopus development. It is shown that there are three ma- jor developmental stages, with respect to changes in RNA synthetic pattern before and after MBT. From Shiokawa et al. [13]. scriptional silence of zygotic nuclear DNA in Xenopus development before MBT is still valid as far as the tran- scriptional control of rRNA genes is concerned. It seems to be valid that MBT is the stage when G1 phase appears in the cell cycle and is the stage when the maternal pro- gram of apoptosis is first executed as a fail-safe mecha- nism of development but not before. However, we pro- pose here additionally that MBT is the stage when rRNA genes are first transcribed but not before. 2. TECHNICAL DIFFICULTIES OF STUDYING RNA SYNTHESIS IN XENOPUS EMBRYOS DURING PRE-MBT STAGE OF DEVELOPMENT Embryos of Xenopus laevis were first utilized as materials Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 23 for molecular biological studies of developmental chang- es in nucleic acid metabolism, especially in the field of the control of RNA synthesis by Brown and Littna [24, 25]. These authors first isolated 32P-labeled RNA by phe- nol treatment. Gurdon and Brown [26] proved that em- bryos, which do not contain rRNA genes and therefore do not form nucleoli, do not synthesize rRNA, and thus showed that nucleoli formation is the cytological mani- festation of the functioning of ribosomal RNA genes. Shiokawa and Yamana [27,28] and Shiokawa et al. [29] introduced dissociated embryonic cell system as a new experimental system for studies of Xenopus embryonic RNA synthesis, and after comparing the pattern of RNA synthesis between 14CO2-labeled whole embryos and (3H) uridine-labeled dissociated embryonic cells, Shiokawa and Yamana [28] showed that developmental activation of rRNA synthesis is not affected by the artificial cell dis- sociation and proceeds normally in the dissociated cell system just as in the whole embryos. The utilization of the dissociated cell system is to overcome the presence of impermeable surface coat which prevents uptake of various radioactive RNA precursors into the whole embryo. The inability of the whole em- bryo to uptake radioactive precursors made detection of newly synthesized RNA very difficult [28,29]. Further- more, the previous idea of the total transcriptional silence of pre-MBT stage embryos also partly came from the situation in which cleavage stage embryos contain only a very small number of nuclei. This latter situation of the occurrence of only a small number of nuclei at the stages before MBT (pre-MBT stage) tends to lead to estimation of apparently undetectable or extremely low transcrip- tional activity per embryo even though transcription per nucleus may not be so small as compared with that in cells in post-blastula stage embryos. Also, it is a very characteristic feature of Xenopus fertilized eggs that they contain several 1000-folds amount of ribosomes per cell pre-formed by active transcription of extra-chromoso- mally amplified ribosomal DNA [30], and this tends to make it difficult to detect new formation of ribosomes in early stages. 3. DEVELOPMENTAL CONTROL OF RRNA GENE EXPRESSION AS STUDIED USING DISSOCIATED XENOPUS EMBRYONIC CELLS Ribosomes are supramolecular structures which are large cellular machinery to produce proteins. Cellular activity to produce ribosomes sensitively reflects the cellular physiological conditions for cell growth. In eukaryotic cells ribosomes consist of 60S and 40S particles, and RNA moiety of 60S particles consists of 28S rRNA and two smaller RNAs, 5.8S RNA and 5S RNA, and that of 40S particles consists only of 18S rRNA. In Xenopus oocytes rRNA producing genes, rDNA, are specifically amplified at stage III of oogenesis [30] and mature oo- cytes accumulate ca. 1012 ribosome particles which are 103 times larger than that of the adult-type cell and is enough for supporting protein synthesis for development up to the stage 42 feeding tadpoles [26]. Gurdon and Brown [26] proved that anucleolate X- enopus mutant embryos which do not have rDNA [31,32] neither synthesize rRNA nor form nucleoli, and thus showed that nucleoli formation is the cytological mani- festation of the functioning (transcription) of rDNAs. Shiokawa and Yamana [27,28] and Shiokawa et al. [29] used dissociated embryonic cells, instead of whole em- bryos, as an experimental system for studies of Xenopus embryonic RNA synthesis. Embryos were dejellied by treatment with sodium thioglycolate (pH 8.3), and then dissociated in the Ca-free medium containing 0.02 M EDTA [28]. During culture isolated cells formed aggre- gates of various sizes depending on the cell density, but the relative rate of rRNA synthesis was the same, irre- spective of the size of the aggregates finally formed [28]. Dissociated embryonic cells were labeled very extensive- ly by simply adding radioactive precursor of RNA into their culture medium, although to label whole embryos 14CO2 has to be used [24,28]. Shiokawa and Yamana [28] compared the overall RNA synthetic pattern in 3H- uridine-labled dissociated cells and 14CO2-labeled whole embryos, and concluded that as long as the overall pro- file of RNA-labeling pattern, which mainly reflects rRNA synthetic activity, is concerned, developmental change in the pattern of RNA synthesis is the same between the dissociated cell system and the whole embryos. Shioka- wa and Yamana [28] found that per embryo activity of rRNA synthesis in dissociated blastula increases in the dissociated cell system just like that in the developing whole embryos, as embryos develop from the blastula stage to the neurula stage. 4. MBT IS ALSO CHARACTERIZED BY THE FIRST STAGE WHEN MATERNALLY PRESET PROGRAM OF APOPTOSIS IS EXECUTED AS A FAIL-SAFE MECHANISM OF DEVELOPMENT MBT is characterized not only as the stage when G1 phase first appear in the cell cycle, but also the stage when maternally-preset program of apoptosis is executed. The execution of apoptosis in Xenopus embryos in the very early development could be summarized as in Fig - ure 2. As shown in Figure 2, it appears that embryonic Copyright © 2013 SciRes. OPEN ACCESS 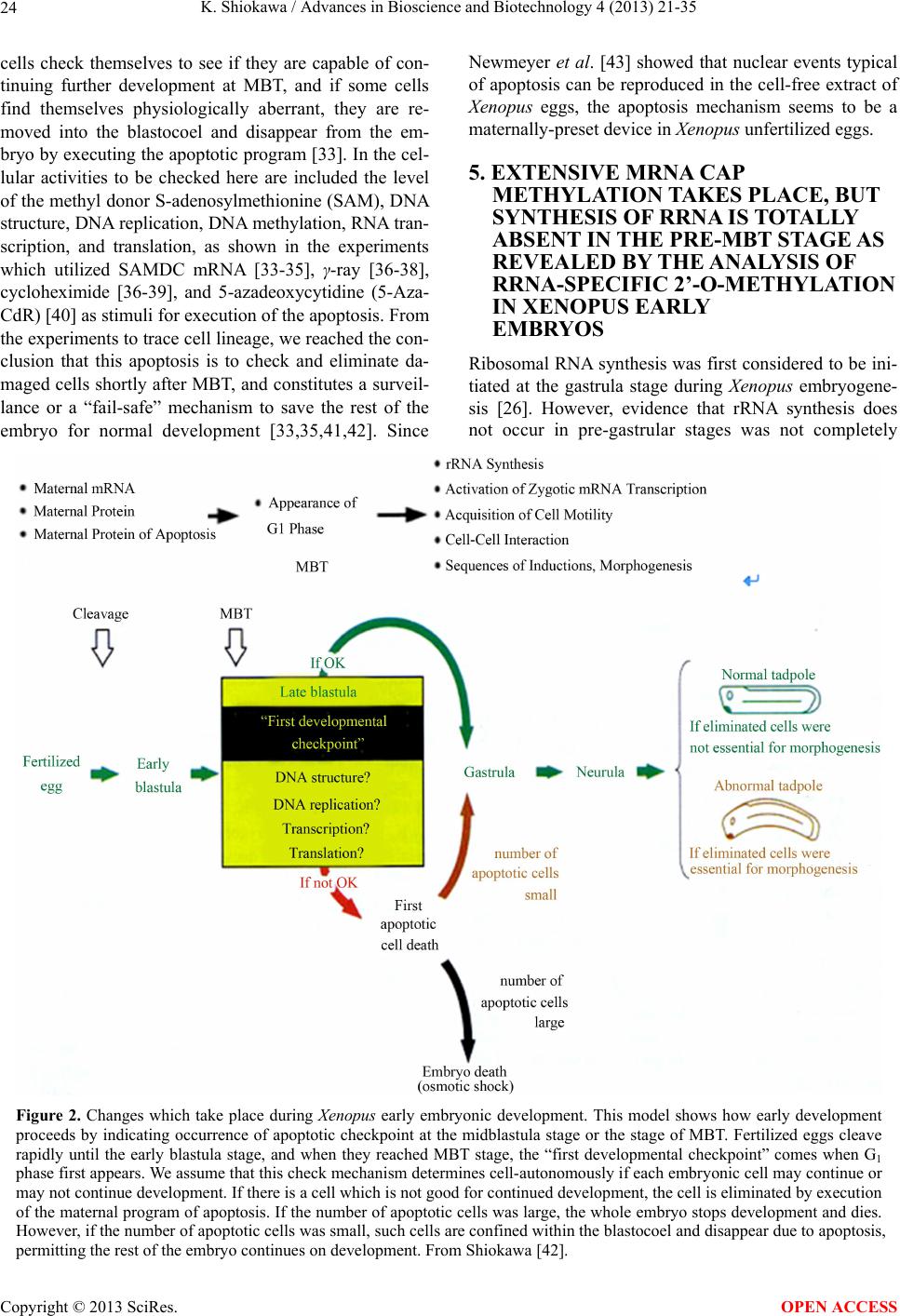 K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 Copyright © 2013 SciRes. 24 Newmeyer et al. [43] showed that nuclear events typical of apoptosis can be reproduced in the cell-free extract of Xenopus eggs, the apoptosis mechanism seems to be a maternally-preset device in Xenopus unfertilized eggs. cells check themselves to see if they are capable of con- tinuing further development at MBT, and if some cells find themselves physiologically aberrant, they are re- moved into the blastocoel and disappear from the em- bryo by executing the apoptotic program [33]. In the cel- lular activities to be checked here are included the level of the methyl donor S-adenosylmethionine (SAM), DNA structure, DNA replication, DNA methylation, RNA tran- scription, and translation, as shown in the experiments which utilized SAMDC mRNA [33-35], γ-ray [36-38], cycloheximide [36-39], and 5-azadeoxycytidine (5-Aza- CdR) [40] as stimuli for execution of the apoptosis. From the experiments to trace cell lineage, we reached the con- clusion that this apoptosis is to check and eliminate da- maged cells shortly after MBT, and constitutes a surveil- lance or a “fail-safe” mechanism to save the rest of the embryo for normal development [33,35,41,42]. Since 5. EXTENSIVE MRNA CAP METHYLATION TAKES PLACE, BUT SYNTHESIS OF RRNA IS TOTALLY ABSENT IN THE PRE-MBT STAGE AS REVEALED BY THE ANALYSIS OF RRNA-SPECIFIC 2’-O-METHYLATION IN XENOPUS EARLY EMBRYOS Ribosomal RNA synthesis was first considered to be ini- tiated at the gastrula stage during Xenopus embryogene- sis [26]. However, evidence that rRNA synthesis does ot occur in pre-gastrular stages was not completely n Figure 2. Changes which take place during Xenopus early embryonic development. This model shows how early development proceeds by indicating occurrence of apoptotic checkpoint at the midblastula stage or the stage of MBT. Fertilized eggs cleave rapidly until the early blastula stage, and when they reached MBT stage, the “first developmental checkpoint” comes when G1 phase first appears. We assume that this check mechanism determines cell-autonomously if each embryonic cell may continue or may not continue development. If there is a cell which is not good for continued development, the cell is eliminated by execution of the maternal program of apoptosis. If the number of apoptotic cells was large, the whole embryo stops development and dies. However, if the number of apoptotic cells was small, such cells are confined within the blastocoel and disappear due to apoptosis, permitting the rest of the embryo continues on development. From Shiokawa [42]. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 25 conclusive, because RNA fractionation methods routine- ly used in previous experiments were sucrose density gradient centrifugation and agarose or polyacrylamide gel electrophoresis, and these methods did not complete- ly separate rRNA from heterogeneous mRNA-like RNA, which is synthesized very actively especially in embryos during blastula stages [10,11]. 18S and 28S rRNA molecules contain a relatively large number of 2’-O-methyl groups (90 per 40S pre- rRNA), and almost all of these 2’-O-methyl groups are conserved during pre-rRNA processing and in the ma- tured rRNAs. Most eukaryotic mRNAs, on the other hand, contain no 2’-O-methyl group but the one in the cap terminus position. Therefore, rRNA and mRNA can be separately quantitated if nucleotides having 2’-O-me- thylation were determined in nuclease digests of these RNAs. Since rRNA precursor is methylated immediately after its transcription, the detection of rRNA-specific 2’-O-methylation is a sensitive method to detect rRNA gene products shortly after its transcription. Thus, the methods to detect rRNA-specific 2’-O-methylation is the methods to detect rRNA synthesis without being affected by the presence of inevitably-contaminating mRNA in the rRNA samples. We labeled Xenopus embryonic cells with (methyl-3H) methionine at the morula, blastula, gastrula and neurula stages, and purified 3H-labeled high-molecular-weight RNA from each embryos on sucrose density gradient centrifugation. We digested the RNAs pooled from the high-molecular-weight RNA regions with RNases A and T2, and analyzed the nucleotide digests on DEAE-Se- phadex columns. Figure 3 shows a set of results of such experiments. Judging from the sucrose gradient profiles (inset in each figure) rRNA synthesis indicated by the presence of distinct 18S and 28S rRNA radioactivity peaks occurred only at post-gastrula stages. In the pro- files of morula (definitely, this stage is the pre-MBT stage) and blastula RNAs, however, radioactivity was widely distributed in the high-molecular-weight RNA region, and one can not discriminate if this is the labeling of rRNA or mRNA. We found here that about 80% of the radioactivity was resistant to alkali hydrolysis, which implies that most of the label in the high-molecular- weight RNA region at these early developmental stages (morula and blastula stages) was due to methylation of DNA which was fractionated also in this region. Incor- poration of the label into 4S RNA, which is another RNA with ample 2’-O-methylation, occurred at all the stages examined (from morula to neurula stages). The DEAE- Sephadex column chromatographic profiles of morula cell RNA labeled for 3 hr showed methylation peaks at regions of charge value-2 (mononucleotide due to base methylation) and charge value-5 (mRNA cap), but there was no appreciable amount of 2’-O-methylation in the region of charge value-3 (rRNA-specific dinucleotides; NpmNp) or charge value-4 (rRNA-specific trinucleotides; NmpNmpNp) nucleotides. This shows that morula cells do not synthesize rRNA at all, although they synthesize a considerably large amount of capped mRNA. In RNA from early blastula cells which were labeled for 4 hrs until the end of late blastula stage, the largest component obtained in the DEAE column chromatography was the charge value-3 compound (rRNA-specific dinucleotides). Here, the charge value-4 compound (specific to 2’-O- methylation in 28S rRNA) was also detected though in a much smaller amount. These results show that rRNA synthesis occurs already in embryos at the blastula stage. The relative amount of the charge value-5 (mRNA cap) component showing cap methylation in the blastula stage was smaller than in morula stage. At the gastrula stage, the relative amounts of charge value-3 and charge value- 4 substances, both indicating rRNA synthesis, increased greatly and the relative amount of the cap methylation became much smaller. The methylation profile of neurula cells was essentially the same as that of gastrula cells. 6. RRNA SYNTHESIS IS ACTIVATED IN THE LATTER HALF PERIOD OF BLASTULA STAGE AS DEMONSTRATED BY ANALYSIS OF 2’-O-METHYLATION Since we found that embryos starts to synthesize rRNA during the 4 hr from early blastula to late blastula stage, we divided this period into two 2 hr-periods, and studied rRNA synthesis by detecting 2’-O-methylation in each period. The results obtained showed that embryos con- tained only a trace amount of rRNA-specific dinucleo- tides in the former 2 hrs of the blastula stage but con- tained a very large amount of rRNA-specific dinucleo- tides in the latter 2 hrs, indicating that rRNA synthesis starts in the latter half of the MBT stage (Figure 4). These results are consistent with the timing of the ex- pression of exogenously-injected ribosomal RNA genes [44]. By contrast, the extent of cap formation and base- methylation were not greatly different in the former and the latter half periods of blastula stage, indicating that only rRNA synthesis is activated during the latter 2 hr period of the blastula stage. From these results we con- cluded that rRNA synthesis starts during the midblastula to late blastula stage [10,11]. 7. ONCE STARTED IN THE LATTER HALF PERIOD OF THE MBT STAGE, THE RATE OF RRNA SYNTHESIS PER CELL IS CONSTANT THROUGHOUT LATER STAGES From the amounts of incorporation of the (methyl-3H)- roup into rRNA-specific dinucleotides (NmpNp) and g Copyright © 2013 SciRes. OPEN ACCESS 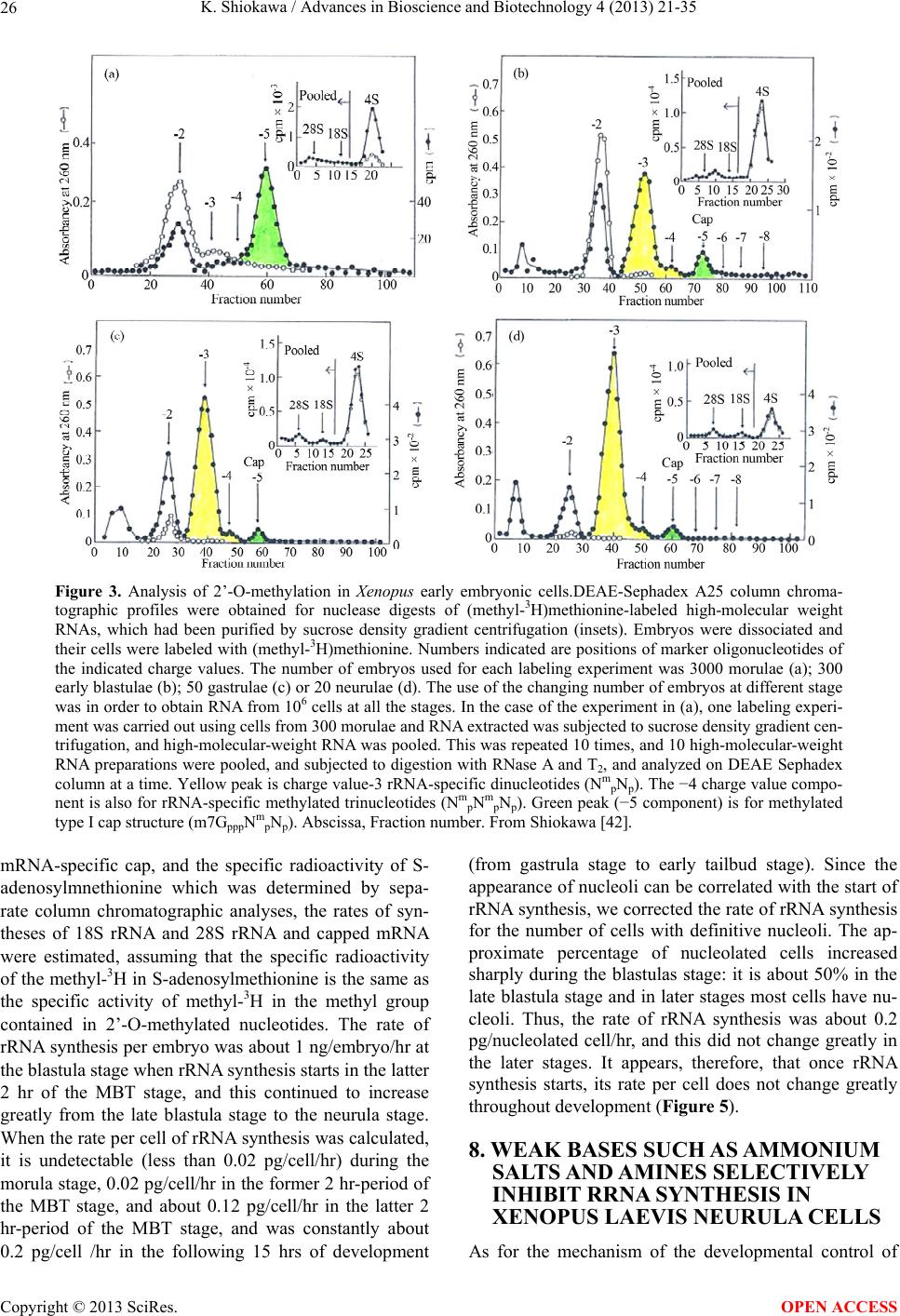 K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 26 Figure 3. Analysis of 2’-O-methylation in Xenopus early embryonic cells.DEAE-Sephadex A25 column chroma- tographic profiles were obtained for nuclease digests of (methyl-3H)methionine-labeled high-molecular weight RNAs, which had been purified by sucrose density gradient centrifugation (insets). Embryos were dissociated and their cells were labeled with (methyl-3H)methionine. Numbers indicated are positions of marker oligonucleotides of the indicated charge values. The number of embryos used for each labeling experiment was 3000 morulae (a); 300 early blastulae (b); 50 gastrulae (c) or 20 neurulae (d). The use of the changing number of embryos at different stage was in order to obtain RNA from 106 cells at all the stages. In the case of the experiment in (a), one labeling experi- ment was carried out using cells from 300 morulae and RNA extracted was subjected to sucrose density gradient cen- trifugation, and high-molecular-weight RNA was pooled. This was repeated 10 times, and 10 high-molecular-weight RNA preparations were pooled, and subjected to digestion with RNase A and T2, and analyzed on DEAE Sephadex column at a time. Yellow peak is charge value-3 rRNA-specific dinucleotides (Nm pNp). The −4 charge value compo- nent is also for rRNA-specific methylated trinucleotides (Nm pNm pNp). Green peak (−5 component) is for methylated type I cap structure (m7GpppNm pNp). Abscissa, Fraction number. From Shiokawa [42]. mRNA-specific cap, and the specific radioactivity of S- adenosylmnethionine which was determined by sepa- rate column chromatographic analyses, the rates of syn- theses of 18S rRNA and 28S rRNA and capped mRNA were estimated, assuming that the specific radioactivity of the methyl-3H in S-adenosylmethionine is the same as the specific activity of methyl-3H in the methyl group contained in 2’-O-methylated nucleotides. The rate of rRNA synthesis per embryo was about 1 ng/embryo/hr at the blastula stage when rRNA synthesis starts in the latter 2 hr of the MBT stage, and this continued to increase greatly from the late blastula stage to the neurula stage. When the rate per cell of rRNA synthesis was calculated, it is undetectable (less than 0.02 pg/cell/hr) during the morula stage, 0.02 pg/cell/hr in the former 2 hr-period of the MBT stage, and about 0.12 pg/cell/hr in the latter 2 hr-period of the MBT stage, and was constantly about 0.2 pg/cell /hr in the following 15 hrs of development (from gastrula stage to early tailbud stage). Since the appearance of nucleoli can be correlated with the start of rRNA synthesis, we corrected the rate of rRNA synthesis for the number of cells with definitive nucleoli. The ap- proximate percentage of nucleolated cells increased sharply during the blastulas stage: it is about 50% in the late blastula stage and in later stages most cells have nu- cleoli. Thus, the rate of rRNA synthesis was about 0.2 pg/nucleolated cell/hr, and this did not change greatly in the later stages. It appears, therefore, that once rRNA synthesis starts, its rate per cell does not change greatly throughout development (Figure 5). 8. WEAK BASES SUCH AS AMMONIUM SALTS AND AMINES SELE CTIVELY INHIBIT RRNA SYNTHESIS IN XENOPUS LAEVIS NEURULA CELLS As for the mechanism of the developmental control of Copyright © 2013 SciRes. OPEN ACCESS 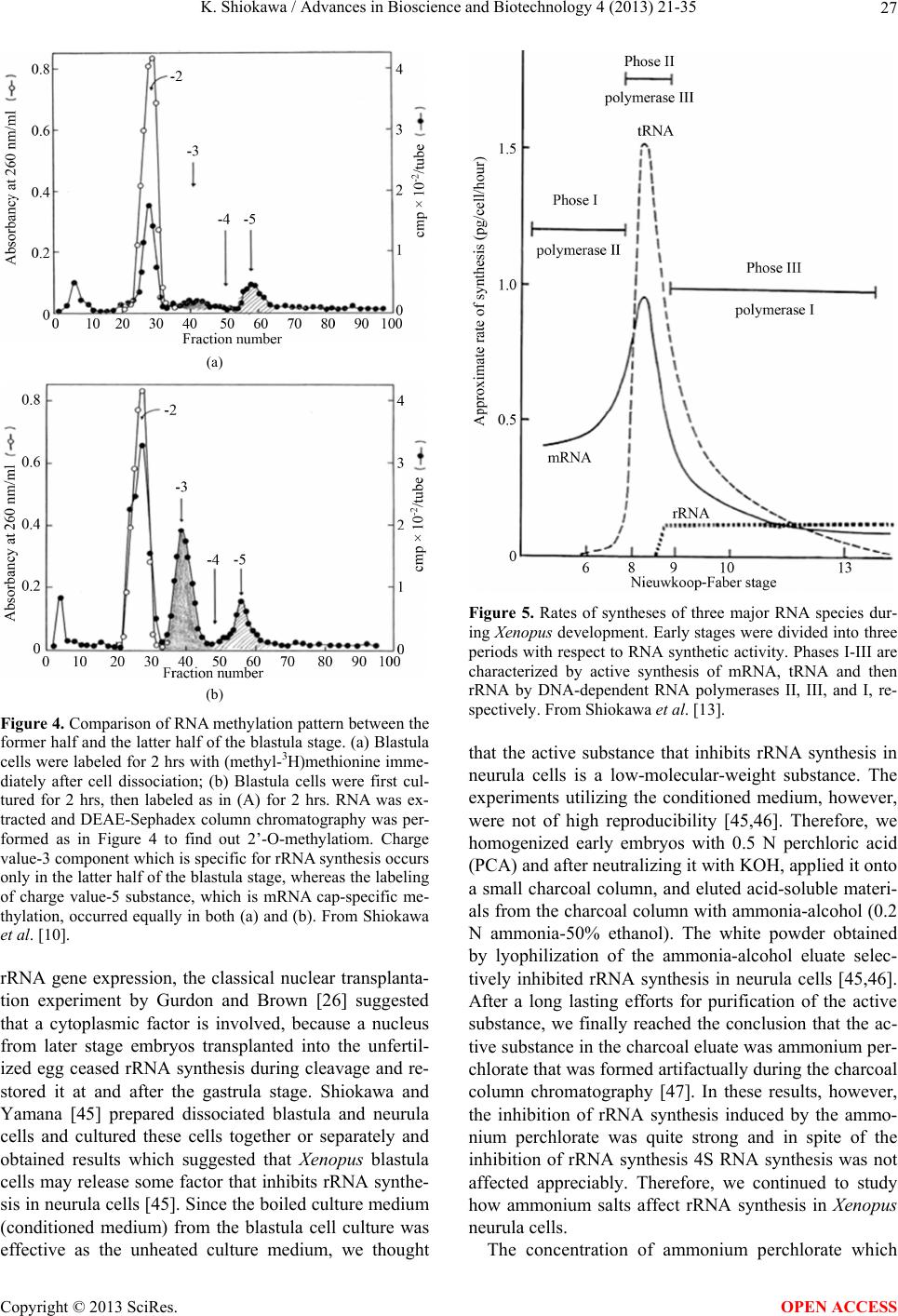 K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 27 (a) (b) Figure 4. Comparison of RNA methylation pattern between the former half and the latter half of the blastula stage. (a) Blastula cells were labeled for 2 hrs with (methyl-3H)methionine imme- diately after cell dissociation; (b) Blastula cells were first cul- tured for 2 hrs, then labeled as in (A) for 2 hrs. RNA was ex- tracted and DEAE-Sephadex column chromatography was per- formed as in Figure 4 to find out 2’-O-methylatiom. Charge value-3 component which is specific for rRNA synthesis occurs only in the latter half of the blastula stage, whereas the labeling of charge value-5 substance, which is mRNA cap-specific me- thylation, occurred equally in both (a) and (b). From Shiokawa et al. [10]. rRNA gene expression, the classical nuclear transplanta- tion experiment by Gurdon and Brown [26] suggested that a cytoplasmic factor is involved, because a nucleus from later stage embryos transplanted into the unfertil- ized egg ceased rRNA synthesis during cleavage and re- stored it at and after the gastrula stage. Shiokawa and Yamana [45] prepared dissociated blastula and neurula cells and cultured these cells together or separately and obtained results which suggested that Xenopus blastula cells may release some factor that inhibits rRNA synthe- sis in neurula cells [45]. Since the boiled culture medium (conditioned medium) from the blastula cell culture was effective as the unheated culture medium, we thought Figure 5. Rates of syntheses of three major RNA species dur- ing Xenopus development. Early stages were divided into three periods with respect to RNA synthetic activity. Phases I-III are characterized by active synthesis of mRNA, tRNA and then rRNA by DNA-dependent RNA polymerases II, III, and I, re- spectively. From Shiokawa et al. [13]. that the active substance that inhibits rRNA synthesis in neurula cells is a low-molecular-weight substance. The experiments utilizing the conditioned medium, however, were not of high reproducibility [45,46]. Therefore, we homogenized early embryos with 0.5 N perchloric acid (PCA) and after neutralizing it with KOH, applied it onto a small charcoal column, and eluted acid-soluble materi- als from the charcoal column with ammonia-alcohol (0.2 N ammonia-50% ethanol). The white powder obtained by lyophilization of the ammonia-alcohol eluate selec- tively inhibited rRNA synthesis in neurula cells [45,46]. After a long lasting efforts for purification of the active substance, we finally reached the conclusion that the ac- tive substance in the charcoal eluate was ammonium per- chlorate that was formed artifactually during the charcoal column chromatography [47]. In these results, however, the inhibition of rRNA synthesis induced by the ammo- nium perchlorate was quite strong and in spite of the inhibition of rRNA synthesis 4S RNA synthesis was not affected appreciably. Therefore, we continued to study how ammonium salts affect rRNA synthesis in Xenopus neurula cells. The concentration of ammonium perchlorate which Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 28 selectively inhibits rRNA synthesis turned out to be ca. 2.5 mM. We compared the effects of the same concen- tration (2.5 mM) of ammonium perchlorate and sodium perchlorate in Xenopus neurula cells, with the finding that only the former but not the latter inhibited rRNA synthe- sis selectively (Figure 6). This implied that the effective part of the ammonium perchlorate is not the perchlorate moiety, but the ammonium moiety. We cultured Xenopus neurula cells in the medium which contained 2.5 mM of ammonium chloride and ammonium sulfate, and found that both of these inhibit rRNA synthesis, with little in- hibition on 4S RNA synthesis [47-49]. We tested ammo- nium dihydrogenphosphate, ammonium monohydrogen- phosphate, and ammonium phosphate, using as referenc- es, potassium dihydrogenphosphate, potassium monohy- drogenphosphate and potassium phosphate all at 2.5 mM. We found that only ammonium ion-releasing salts were effective in inhibiting rRNA synthesis, and the effective- ness was roughly proportional to the number of ammo- nium ions to be released. We also found that while am- monium acetate and ammonium aspartate selectively inhibited rRNA synthesis at 2.5 mM, again, their potas- sium salt were ineffective. We also found that ammo- nium salts of all the four major ribonuleotide monopho- sphates (5’-AMP, 5’-GMP, 5’UMP, and 5’-CMP) at 5 mM were effective in selectively inhibiting rRNA syn- thesis in neurula cells. We also tested the effects of amin- es such as monomethylaminehydroperchloride, dime- thylaminehydroperchloride, and trimethylaminehydrop- erchloride, and found that all of these are active as selec- tive inhibitors of rRNA synthesis at 2.5 mM. Similar po- sitive results were obtained when three kinds of ethyla- mines were tested. From these results, we concluded Figure 6. Effects of ammonium perchlorate and sodiumn per- chlorate on RNA synthetic pattern of neurula cells.Neurula cells were labeled for 3 hrs with 20 μCi of 3H-uridine for 3 hrs in the presence of 2.5 mM of ammonium perchlorate (B) or sodium perchlorate (C); RNAs were extracted and electropho- resed on 0.5% agarose-2.4% polyacrylamide gels; (A) Control neurula cells. Dotted peaks are for 28S and 18S rRNAs, and black peaks are for 4S RNA (tRNA). From Shiokawa et al. [47]. that weak bases such as ammonium ion and amines are similarly effective as selective inhibitors of rRNA syn- thesis in Xenopus neurula cells. 9. WEAK BASES ALSO SUPPRESS DEVELOPMENT AL ACTIVA TION OF RRNA SYNTHESIS IN XENOPUS BLASTULA CELLS Weak bases inhibit rRNA gene expression in Xenopus neurula cells in which rRNA synthesis is fully activated. We cultured blastula cells for 5, 10, 15 hrs in the medium which contained 2.5 mM of ammonium salts. When con- trol blastula cells were labeled with 3H-guanosine for 5 hrs a small amount of incorporation was detected in 28S and 18S rRNA with a large amount of incorporation into 4S RNA and DNA (Figures 7(A)-(C)). Ammonium chloride, but not potassium chloride, inhibited rRNA synthesis but not 4S RNA and DNA syntheses. When we first cultured blastula cells in the ammonium chloride- containing medium for 5 hrs, and then labeled them for 5 hrs in the continued presence of the ammonium chloride, much clearer inhibition of rRNA synthesis was obtained. This is an indication of the inhibition of post-MBT acti- vation of rRNA synthesis (Figures 7(D)-(F)). Essentially the same results were obtained when we cultured blastula cells for 10 hrs in the presence of ammonium chloride and then labeled them for 5 hrs (Figures 7(G)-(I)). Im- portantly, syntheses of 4S RNA (tRNA) and heteroge- neous nuclear RNA, and in addition, DNA were not in- terfered greatly under these conditions. Using 10% poly- acrylamide gel, we separated small-molecular-weight RNAs in the RNA preparations obtained and confirmed that ammonium chloride did not inhibit the labeling of 4S RNA, 5S RNA, and U1, U2, and U5 small nuclear RNAs. When we cultured blastula cells for 5 hrs in the am- monium ion-containing medium and after this transferred them into the normal medium which contained 3H-gua- nosine. We found here that rRNA synthesis can be re- stored to a large extent, indicating that the effects of ammninium salts are reversible. By HPLC analysis of the acid-soluble faction of the embryonic cells, we confirm- ed here that ammonia was incorporated into neurula cells within 1 hr. We also confirmed that ammonium salts at ca. 3 mM did not inhibit protein synthesis, cell division, and cellular reaggregating activity at least for the first 10 hrs of incubation, although after treatment for 10 hrs, cell reaggregation was found to be significantly inhibited. These data indicates that ammonium salt actually selec- tively inhibits not only the initiation of rRNA synthesis at MBT but also subsequent activation of rRNA synthe- sis in post-MBT stages. Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 29 Figure 7. Inhibition of the initiation and activation of rRNA gene expression by ammonium chloride but not by potassium chloride in Xenopus blastula cells. Blastula cells were treated with 2.5 mM of either ammonium chloride ((B), (E), (H)) or potassium chloride ((C), (F), (I)) and labeled with 3H-guano- sine. ((A)-(C)) Blastula cells were labeled for 5 hrs. ((D)-(F)) Cells were cultured for 5 hrs in the presence of ammonium chloride (E) or potassium chloride (F), and then labeled for 5 hrs with 3H-guanosine in the continued presence of the salts. ((G)-(I)) Cells were cultured for 10 hrs in the presence of the salts and then labeled for 5 hrs in the continued presence of the salts. RNAs were extracted and electrophoresed on 0.5% aga- rose-2.4% polyacrylamide gels. In this profile the radioactivity appeared also in DNA. ((A), (D), (G)) Control cells. Black peaks are for 28S and 18S rRNA, and white peaks are for 4S RNA, and shaded peaks are for DNA. From Shiokawa et al. [48]. 10. WEAK BASES INHIBIT RRNA TRANSCRIPTION, BUT NOT POST-TRANSCRIPTIONAL PROCESSING Xenopus neurula cells were first inhibited for 2.5 hrs by 5 mM of ammonium chloride or monomethylamine hydro- perchloride and then pulse-labeled for 2.5 hrs in the con- tinued presence of the weak bases [47]. We obtained active labeling of heterogeneous nuclear RNA (hnRNA), and 40 S rRNA primary transcript, in addition to the la- beling of 28S and 18S mature rRNAs and 4S RNA. Un- der these conditions, both of the weak bases inhibited la- beling of 40S pre-rRNA, 28S rRNA, and 18S rRNA al- most completely (Figure 8), suggesting that the inhibi- tion of rRNA synthesis by these weak bases is at the transcription level. We then selected conditions of partial inhibition of rRNA synthesis by these weak bases. When we labeled the control neurula cells for 1 hr (Figure 9(A)), we ob- tained 40S pre-rRNA (black peak), 30 S rRNA interme- diate (shaded peak) and 18S rRNA (dotted peak), in ad- dition to a large amount of heterogeneous mRNA-like RNA [50]. The appearance of 18S mature rRNA before 28S mature rRNA is due to the fact that the processing of the former is finished before that of the latter [51]. The identification of 30S component is based on slight re- Figure 8. Effects of ammonium salts and amine on pulse-la- beled RNA synthesis. Xenopus neurula cells were treated with 5 mM each of potassium chloride (B); ammonium chloride (C); or monomethylaminehydroperchloride (D) for 2.5 hrs and then pulse-labeled with 3H-uridine for 2.5 hrs in the continued pre- sence of the potassium and ammonium salts. RNA was extract- ed and gel electrophoresed, and radioactivity determined using the sliced gels. Densitometric scanning of gels before slicing are omitted from the radioactivity profiles. Peaks of 40S rRNA precursor are marked by black colour. A, Control untreated cells. From Shiokawa et al. [47]. Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 30 Figure 9. Effects of ammonium chloride on the labeling of 40S pre-rRNA, 30 S pre-rRNAs and 18S and 28S mature rRNAs. Neurula cells were labeled for 1 hr ((A), (B)) or for 3 hrs ((C), (D)) with 3H-uridine in the presence of 2.6 mM ammonium chloride. RNAs were gel electrophoresed for 2 hrs ((A), (B)) or 1 hr ((C), (D)). Longer electrophoresis was for obtaining better resolution in the high-molecular-weight RNA region. Densi- tometric scanning of gels before slicing are omitted from the radioactivity profiles. Black peaks are for 40S pre-rRNA, shad- ed peaks are for 30S pre-rRNA, and dotted peaks are for 28S and 18S mature rRNAs. From Shiokawa et al. [50]. tardation of this peak from 28S optical density peak on the agarose-polyacrylamide gel. When neurula cells pre- treated for 2.5 hrs with 2.6 mM ammonium chloride (which was the conditions for partial inhibition of rRNA synthesis) were labeled with 3H-uridine for 1 hr, the ex- tent of the inhibition of labeling was 60%, 59%, and 70% for 40s pre-rRNA, 30s rRNA intermediates and 18S rRNA, respectively (Figures 9(A) and (B)). When the same cul- tures were labeled for 3 hrs, labeling of 18 S and 28 S mature rRNAs but not 40S and 30S pre-rRNA became predominant, and inhibition here was 72%, 69%, and 7% for 28S and 18S rRNAs and 4S RNA, respectively (Fig- ures 9(C) and (D)). When we performed a parallel expe- riment using 1 mM trimethylammonium perchloride, the inhibition was 67%, 70%, 75%, 73% and 8%, for 40S pre-rRNA, 30S rRNA intermediate, 28S rRNA, 18S rRNA and 4S RNA, respectively. Thus, the extents of the inhibition of the labeling of 40S pre-rRNA, 30S interme- diate rRNA, 18S and 28S mature rRNA were much the same (67% - 75%), and in spite of the large inhibition of the labelinbg of rRNA species, inhibition of the labeling of 4S RNA (tRNA) was only 8%. Therefore, we con- cluded that inhibition was at the level of the formation of 40S primary transcript and [50] probably not at the proc- essing of 40S pre-rRNA. Approximately 30 - 60 min is needed for the pulse- labeled 40S pre-rRNA to be processed completely into 18S and 28S rRNAs [50,51]. Therefore, we first pulse- labeled neurula cells for 35 min and chased the label in rRNA species for 2 hrs. Labeled RNAs in neurula cells under these conditions were 40S pre-rRNA and hetero- geneous mRNA-like RNA (Figure 10(A)). These neu- rula cells were administered with actinomycin D (10 μg/ml) as a transcription inhibitor, and treated either with ammonium chloride (5 mM) or with trimethylammonium perchloride (5 mM) for 2 hrs in the continued presence of actinomycin D. During the chase period of 2 hrs, the label in the 40S pre-rRNA in the control culture com- pletely disappeared and 18S and 28S mature rRNAs ap- peared (Figure 10(B)). Such changes in the labeling pro- file were observed also in the cultures treated with 10 mM ammonium chloride (Figure 10(C)) and trimethyl- ammonium perchloride (Figure 10(D)). These results show that weak bases used here inhibited rRNA tran- scription but did not inhibit the processing of 40S pre- rRNA into two mature rRNAs [50]. Also, these results exclude the possibility of aberrant processing or rapid wastage of the mature rRNAs. Figure 10. Effects of ammonium chloride on the processing of 40S pre-rRNA which was labeled before administration of am- monium chloride. Neurula cells were pulse-labeled with 3H- uridine for 35 min. One culture was then immediately frozen as a zero-time control (A). Three cultures were then administrered with 10 μg/ml of actinomycin D and further cultured for 2 hrs in the presence of either 10 mM ammonium chloride (C) or 10 mM of trimethylammonium perchloride (D). Control cells were incubated for 2 hrs without being exposed to these weak bases (B). RNA were extracted and electrophoresed on agarose-poly- acrylamide gels. Densitometric scanning of gels before slicing are omitted from the radioactivity profiles. A black peak is 40S pre-rRNA and dotted peaks are 28S and 18S rRNAs. From Shi- okawa et al. [50]. Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 Copyright © 2013 SciRes. 31 11. POSSIBLE INVOLVEMENT OF PH CHANGE IN THE WEAK BASE-TREATED NEURULA CELLS From these results, the mechanism by which weak bases inhibit rRNA synthesis in Xenopus embryonic cells can be summarized as in Figure 12. In Xenop us embryos, a slight increase of intracellular pH has various important effects [52]. Such pH changes have been implicated as a necessary step in oocyte ma- turation [53], fertilization [54], and commencement of cleavages [55]. We suspected that a slight elevation of intracellular pH might be involved in the inhibition of rRNA synthesis. Since it is known that pH-mediated changes are abolished when Na+ was eliminated from the surrounding medium [53,56], we tested the effect of 3 mM ammonium chloride and 1.5 mM trimethylammo- nium perchloride on neurula cells in the medium in which all the Na+ was replaced by choline ions. Results of the labeling experiment showed that approximately 80% of the inhibitory activity of both ammonium chlo- ride (Figure 11(B)) and trimethylammonium perchloride (Figure 11(C)) completely disappeared when their ef- fects were tested in the medium whose Na+ had been re- placed by choline ions (respectively, Figures 11(E) and (F)). Therefore, these weak bases seem to exert their rRNA synthesis-inhibiting effects via slight pH elevation within the neurula cells. Figure 11. Disappearance of the inhibitory effect on rRNA synthesis of ammonium chloride and trimethylammonium- perchloride in the medium in which Na+ was replaced by choline ion. Neurula cells were treated for 2.5 hrs either in the normal medium (0.1 × Steinberg’s solution) ((A)-(C)) or in the choline+-containing Na+-free medium ((D)-(F)) with 3 mM of ammonium chloride ((B), (E)) or 1.5 mM of trime- thylammonium perchloride ((C), (F)). ((A), (D)) are untreated control neurula cells. All the cultures were administered with 3H-udirine and cells were labeled for 3 hrs. Dotted peaks are 18S and 28S rRNAs. From Shiokawa et al. [50]. We prepared acid-soluble fractions of these weak base-treated neurula cells, and found that the level of ATP as well as other ribonucleotidetriphosphates re- mained unchanged. Therefore, neither ammonium chlo- ride nor trimethylammonium perchloride disturbed the energy generating system in Xenopus neurula cells [50]. Figure 12. A working hypothesis that weak bases, most probably ammonium ion within the embryonic cells, may be involved in the regulation of rRNA transcription in Xenopus early embryos. Left: Weak bases do not inhibit TCA cycle and do not induce wastage of 40S pre-rRNA and 18S and 28S mature rRNAs. Also, processing of pre-rRNAs is not inhibited. Instead, weak bases inhibit transcrip- tion of rRNA genes (rDNA). This inhibition is probably mediated by intracellular slight pH elevation. From Shiokawa et al. [50]. Right: Developmental changes in the amount of ammonium ion within the embryo determined by amino acid analyzer using the PCA-soluble fraction of the embryo homogenates. The changing amount of the ammonium ion is expressed as ammonia (ng/egg or embryo). From Shiokawa et al. [48]. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 32 12. UPTAKE OF AMMONIUM ION AND AMINES IN XENOPUS NEURULA CELLS Using amino acid analyzer, we analyzed ammonia and amines in acid-soluble fractions of Xenopus neurula cells which had been treated with 2.5 mM of either ammo- nium chloride, ammonium phosphates, or methylamines. In the amino acid profiles, it has been shown that the peak representing ammonia became quite large in the cells treated with ammonium salts. In cells treated with monomethylamine hydroperchloride, a large amount of monomethylamine was detected, with no increase in the amount of ammonia. However, amounts of other com- ponents such as ornithine and other amino acids in the neurula cells did not change appreciably in spite of the treatments with ammonium salts and amines. These re- sults indicate that both ammonium ion and amines selec- tively inhibit rRNA synthesis after being uptaken into Xenopus neurula cells. 13. AMMONIA IS A CANDIDATE OF THE REGULATOR OF RRNA SYNTHESIS WHICH CONTROLS THE INITIATION AND ACTIVA TION O F RRNA SYNTHESIS AT AND AFTER MBT When we examined ninhydrin-positive materials in the acid-soluble fraction of Xenopus early embryos, there was a sizable amount of ammonia but not amines within the embryo. When we performed similar analyses at va- rious stages of development, we found that the level of ammonia in an egg was ca. 55 ng/egg before fertilization and this value did not change greatly during cleavage, but decreased promptly to the level as low as ca. 20 ng/ embryo. The occurrence of ca. 55 ng/egg of ammonia corresponds ca. 3 mM of ammonia as an intra-egg con- centration, and 20 ng/embryo of ammonia corresponds ca. 1 mM as an intra-embryo concentration, since the vo- lume of an egg is ca. 1 μl. The lowered level of ammonia was maintained throughout later stages until the muscu- lar response stage (Figure 12). We attempted to isolate ammonia (and amines if any) from cleavage stage embryos. Starting from 25,000 clea- vage embryos, we isolated about 10 mg of residual mate- rials after evaporating the hydrochloric acid solution that was used to capture the cellular volatile ammonia com- ponents. Mass spectrometric analysis revealed the pres- ence of ammonium chloride but not of amines. Ten mil- ligrams of ammonium chloride obtained here corre- sponds to ca. 4 mg of ammonia. We tested effects of the ammonium chloride isolated from cleavage stage em- bryos for its activity to inhibit rRNA synthesis in Xeno- pus neurula cells at 1.0 and 5.0 mM. The results showed that the synthesis of rRNA was inhibited by 60% (at 1 mM) and 90% (at 5 mM) with only a slight inhibition (less than 10%) in 4S RNA (tRNA) synthesis. These ob- servations also suggest that ammonium ion is a candidate for the factors that regulate initiation and activation of rRNA synthesis in Xenopus embryogenesis. 14. CONCLUSION It has been long believed that RNA synthesis never takes place from zygotic nuclei during pre-MBT stage or clea- vage stage in early Xenopus embryogenesis. However, Shiokawa’s old data [12] and Klein’s new data [17] show that there is RNA synthesis in pre-MBT embryos. How- ever, not all the RNA is synthesized during the pre-MBT stage. For example, rRNA is the RNA whose genes are totally silent during the pre-MBT stage, and transcription of rRNA genes is initiated from the latter half of the MBT stage. This developmental change in the initiation of rRNA genes coincides with the developmentally con- trolled appearance of nucleoli not before but after MBT. Weak bases were found to selectively inhibit rRNA syn- thesis in neurula cells and also inhibit the developmental activation of rRNA synthesis which takes place after MBT. Based on some evidence, weak bases such as ammonia could be a factor that constitutes the control mechanism of the developmental regulation of rRNA genes at the transcriptional level, but not at the step of the post-tran- scriptional processing level. Our data suggest that ammo- nia and amines exert their effect via slight elevation of intracellular pH in the early Xenopus embryos. REFERENCES [1] Graham, C.F. and Morgan, R.W. (1966) Changes in the cell cycle during early amphibian development. Develop- mental Biology, 14, 439-460. http://dx.doi.org/10.1016/0012-1606(66)90024-8 [2] Heasman, J. (2006) Patterning the early Xenopus embryo. Development, 133, 1205-1217. http://dx.doi.org/10.1242/dev.02304 [3] Woodland, H.R. (1974) Changes in the polysome content of developing Xenopuslaevis embryos. Developmental Biology, 40, 90-101. http://dx.doi.org/10.1016/0012-1606(74)90111-0 [4] Richter, J.D., Wasserman, W.J. and Smith, L.D. (1982) The mechanism for increased protein synthesis during Xenopus oocyte maturation. Developmental Biology, 89, 159-167. http://dx.doi.org/10.1016/0012-1606(82)90304-9 [5] Newport, J. and Kirschner, M. (1982) A major develop- mental transition in early Xenopus embryos I. Characte- rization and timing of cellular changes at the midblastula stage. Cell, 30, 675-686. http://dx.doi.org/10.1016/0092-8674(82)90272-0 [6] Newport, J. and Kirschner, M. (1982) A major develop- Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 33 mental transition in early Xenopus embryos. II. Control of the onset of transcription. Cell, 30, 687-696. http://dx.doi.org/10.1016/0092-8674(82)90273-2 [7] Carter, A.D. and Sible, J.C. (2003) Loss of XChk1 func- tion triggers apoptosis after the midblastula transition in Xenopus embryos. Mechanisms of Development, 120, 315-323. http://dx.doi.org/10.1016/S0925-4773(02)00443-4 [8] Wroble, B.N. and Sible, J.C. (2005) Chk2/Cds1 protein kinase blocks apoptosis during early development of Xe- nopuslaevis. Developmental Dynamics, 233, 1359-1365. http://dx.doi.org/10.1002/dvdy.20449 [9] Minoura, I., Nakamura, H., Tashiro, K. and Shiokawa, K. (1995) Stimulation of circus movement by activin, bFGF and TGF-β2 in isolated animal cap cells of Xenopuslaevis. Mechanisms of Development, 49, 65-69. http://dx.doi.org/10.1016/0925-4773(94)00303-5 [10] Shiokawa, K., Misumi, Y. and Yamana, K. (1981) Dem- onstration of rRNA synthesis in pre-gastrular embryos of Xenopuslaevis. Development, Growth and Differentiation, 23, 579-587. http://dx.doi.org/10.1111/j.1440-169X.1981.00579.x [11] Shiokawa, K., Tashiro, K., Misumi, Y. and Yamana, K. (1981) Non-coordinated synthesis of RNA’s in pre-gas- trular embryos of Xenopuslaevis. Development, Growth and Differentiation, 23, 589-597. http://dx.doi.org/10.1111/j.1440-169X.1981.00589.x [12] Nakakura, N., Miura, T., Yamana, K., Ito, A. and Shioka- wa, K. (1987) Synthesis of heterogeneous mRNA-like RNA and low-molecular-weight RNA before the midbla- stula transition in embryos of Xenopuslaevis. Develop- mental Biology, 123, 421-429. http://dx.doi.org/10.1016/0012-1606(87)90400-3 [13] Shiokawa, K., Misumi, Y., Tashiro K., Nakakura, N., Ya- mana, K. and Oh-Uchida, M. (1989) Changes in the pat- terns of RNA synthesis in early embryogenesis of Xeno- puslaevis. Cell Differentiation and Development, 28, 17- 25. http://dx.doi.org/10.1016/0922-3371(89)90019-1 [14] Yasuda, G.K. and Schubiger, G. (1992) Temporal regula- tion in the early embryo: Is MBT too good to be true? Trends in Genetics, 8, 124-127. [15] Shiokawa, K., Kurashima, R. and Shinga, J. (1994) Tem- poral control of gene expression from endogenous and exogenously-introduced DNAs in early embryogenesis of Xenopuslaevis. The International Journal of Developmen- tal Biology, 38, 249-255. [16] Andeol, Y.A. (1994) Early transcription in different ani- mal species: Implicaitonfor transition from maternal to zygotic control in developoment. Roux’s Archives of De- velopmental Biology, 204, 3-10. http://dx.doi.org/10.1007/BF00744867 [17] Yang, J., Tan, C., Darken, R.S., Wilson, P.A. and Klein, P.S. (2002) Beta-catenin/Tcf regulated transcription prior to the midblastula transition. Development, 129, 5743- 5752. http://dx.doi.org/10.1242/dev.00150 [18] Etkin, L.D. and Balcells, S. (1985) Transformed Xenopus embryos as a transient expression system to analyze gene expression at the midblastula transition. Developmental Biology, 108, 173-178. http://dx.doi.org/10.1016/0012-1606(85)90019-3 [19] Shiokawa, K., Yamana, K., Fu, Y., Atsuchi, Y. and Hoso- kawa, K. (1990) Expression of exogenously introduced bacterial chloramphenicol acetyltransferase gene in Xeno- puslaevis embryos before the midblastula transition. Roux’s Archives of Developmental Biology, 198, 322- 329. http://dx.doi.org/10.1007/BF00383770 [20] Nagel, M., Tahinci, E., Symes, K. and Winklbauer, R. (2004) Guidance of mesoderm cell migration in the Xe- nopus gastrula requires PDGF signaling. Development, 131, 2727-2736. http://dx.doi.org/10.1242/dev.01141 [21] Ninomiya, H., Elinson, R.P. and Winklbauer, R. (2004) Antero-posterior tissue polarity links mesoderm conver- gent extension to axial patterning. Nature, 430, 364-367. http://dx.doi.org/10.1038/nature02620 [22] Shook, D., Majer, C. and Keller, R. (2004) Pattern and morphogenesis of presumptive superficial mesoderm in Two closely related species, Xenopuslaevis and Xenopus- tropicalis. Developmental Biology, 270, 163-185. http://dx.doi.org/10.1016/j.ydbio.2004.02.021 [23] Gurdon, J.B. (1988) A community effect in animal deve- lopment. Nature, 336, 772-774. http://dx.doi.org/10.1038/336772a0 [24] Brown, D.D. and Littna, E. (1964) RNA synthesis during the development of Xenopuslaevis, the African clawed toad. Journal of Molecular Biology, 8, 669-687. http://dx.doi.org/10.1016/S0022-2836(64)80116-9 [25] Brown, D.D. and Littna, E. (1966) Synthesis and accumu- lation of DNA-like RNA during embryogenesis of Xeno- puslaevis. Journal of Molecular Biology, 20, 81-94. http://dx.doi.org/10.1016/0022-2836(66)90119-7 [26] Gurdon, J.B. and Brown, D.D. (1965) Cytoplasmic regu- lation of RNA synthesis and nucleolar formation in deve- loping embryos of Xenopuslaevis. Journal of Molecular Biology, 12, 27-35. http://dx.doi.org/10.1016/S0022-2836(65)80279-0 [27] Shiokawa, K. and Yamana, K. (1965) Demonstration of “polyphosphate” and its possible role in RNA synthesis during early development of Rana japonica embryos. Ex- perimental Cell Research, 38, 180-186. http://dx.doi.org/10.1016/0014-4827(65)90439-8 [28] Shiokawa, K. and Yamana, K. (1967) Pattern of RNA synthesis in isolated cells of Xenopuslaevis embryos. De- velopmental Biology, 16, 368-388. [29] Shiokawa, K., Nada, O. and Yamana, K. (1967) Synthesis of RNA in isolated cells from Xenopuslaevis embryos. Nature, 213, 1027-1027. http://dx.doi.org/10.1038/2131027a0 [30] Brown, D.D. and Dawid, I.B. (1968). Specific gene am- plification in oocytes. Nature, 160, 272-280. [31] Tashiro, K., Shiokawa, K., Yamana, K. and Sakaki, Y. (1986) Structural analysis of ribosomal DNA homologues in nucleolus-less mutant Xenopuslaevis. Gene, 44, 299- 306. http://dx.doi.org/10.1016/0378-1119(86)90194-0 [32] Steele, R.E., Thomas, P.S. and Reeder, R.H. (1984) Anu- cleolate frog embryos contain ribosomal DNA sequences and a nucleolar antigen. Developmental Biology, 102, 409-416. Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 34 http://dx.doi.org/10.1016/0012-1606(84)90205-7 [33] Kai, M., Kaito, C., Fukamachi, H., Higo, T., Takayama, E., Hara, H., Ohya, Y., Igarashi, K. and Shiokawa, K. (2003) Overexpression of S-adenosylmethionine decarbo- xylase (SAMDC) in Xenopus embryos activates maternal program of apoptosis as a “fail-safe” mechanism of early embryogenesis. Cell Research, 13, 147-158. http://dx.doi.org/10.1038/sj.cr.7290159 [34] Shibata, M., Shing, J., Yasuhiko, Y., Kai, M., Miura, K., Shimogori, T., Kashiwagi, K., Igarashi, K. and Shiokawa, K. (1998) Overexpression of S-adenosylmethionine decar- boxylase (SAMDC) in early Xenopus embryos induces cell dissociation and inhibits transition from the blastula to gastrula stage. The International Journal of Develop- mental Biology, 42, 675-686. [35] Kai, M., Higo, T., Yokoska, J., Kaito, C., Kajita, E., Fu- kamachi, H., Takayama, E., Igarashi, K. and Shiokawa, K. (2000) Overexpression of S-adenosylmethionine decarbo- xylase (SAMDC) activates the maternal program of apop- tosis shortly after MBT in Xenopus embryos. The Inter- national Journal of Developmental Biology, 44, 507-510. [36] Hensey, C. and Gautier, J. (1997) A developmental timer that regulates apoptosis at the onset of gastrulation. Me- chanisms of Development, 69, 183-195. http://dx.doi.org/10.1016/S0925-4773(97)00191-3 [37] Sible, J.C., Anderson, J.A., Lewelly, A.L. and Maller, J.L. (1997) Zygotic transcription is required to block a mater- nal program of apoptosis in Xenopus Embryos. Develop- mental Biology, 189, 335-346. http://dx.doi.org/10.1006/dbio.1997.8683 [38] Anderson, J.A., Lewellyn, A.L. and Maller, J.L. (1997) Ionizing radiation induces apoptosis and elevates cyclin A1-Cdk2 activity before but not after the midblastula transition in Xenopus. Molecular Biology of the Cell, 8, 1195-1206. http://dx.doi.org/10.1091/mbc.8.7.1195 [39] Stack, J.H. and Newport, J.W. (1997) Developmentally regulated activation of apoptosis early in Xenopus gas- trulation results in cyclin A degradation during interphase of the cell cycle. Development, 124, 3185-3195. [40] Kaito, C., Kai, M., Higo, T., Takayama, E., Fukamachi, H., Sekimizu, K. and Shiokawa, K. (2001) Activation of the maternally preset program of apoptosis by microin- jection of 5-aza-2’-deoxycytidine and 5-methyl-2’-de- oxycytidine-5’-triphosphate in Xenopus laevis embryos. Development, Growth & Differentiation, 43, 383-390. http://dx.doi.org/10.1046/j.1440-169x.2001.00579.x [41] Shiokawa, K., Kai, M., Higo, T., Kaito, C., Fukamachi, H., Yaoita, Y. and Igarashi, K. (2000) Maternal program of apoptosis activated shortly after midblastula transition by overexpression of S-adenosylmethionine decarboxy- lase in Xenopus early embryos. Comparative Biochemis- try and Physiology Part B, 126, 149-155. [42] Shiokawa, K. (2012) Maternally-preset program of apop- tosis and caspases involved in execution of the apoptosis at midblastula transition (MBT) but not before in Xenopus laevis embryogenesis. Advances in Bioscience and Biotechnology, 3, 751-769. http://dx.doi.org/10.4236/abb.2012.326096 [43] Newmeyer, D.D., Farschon, D.M. and Reed, J.C. (1994) Cell-free apoptosis in Xenopus egg extracts: Inhibition by Bcl-2 and requirement for an organelle fraction enriched in mitochondria. Cell, 79, 353-364. http://dx.doi.org/10.1016/0092-8674(94)90203-8 [44] Busby, S.J. and Reeder, R.H. (1983) Spacer sequences regulate transcription of ribosomal gene plasmids injected into Xenopus embryos. Cell, 34, 989-996. http://dx.doi.org/10.1016/0092-8674(83)90556-1 [45] Shiokawa, K. and Yamana, K. (1967) Inhibitor of ribo- somal RNA synthesis in Xenopus laevis embryos. Deve- lopmental Biology, 16, 389-406. http://dx.doi.org/10.1016/0012-1606(67)90049-8 [46] Laskey, R.A., Gerhart, J.C. and Knowland, J.S. (1973) Inhibitor of ribosomal RNA synthesis in neurula cells by extracts from blastulae of Xenopus laevis. Developmental Biology, 33, 241-248. http://dx.doi.org/10.1016/0012-1606(73)90134-6 [47] Shiokawa, K., Kawazoe, Y. and Yamana, K. (1985) Demonstration that inhibitor of rRNA synthesis in “char- coal-extracts” of Xenopus embryos is artifactually pro- duced ammonium perchlorate. Developmental Biology, 112, 258-260. http://dx.doi.org/10.1016/0012-1606(85)90141-1 [48] Shiokawa, K., Kawazoe, Y., Nomura, H., Miura, T., Na- kakura, N., Horiuchi, T. and Yamana, K. (1986) Ammo- nium ion as a possible regulator of the commencement of rRNA synthesis in Xenopus laevis embryogenesis. De- velopmental Biology, 115, 380-391. http://dx.doi.org/10.1016/0012-1606(86)90257-5 [49] Shiokawa, K., Kawazoe, Y., Tashiro, K. and Yamana, K. (1986) Effects of various ammonium salts, amines, polyamines, and alpha-methylornithine on rRNA synthe- sis in neurula cells of Xenopus laevis and Xenopus bore- alis. Cell Differentiation, 18, 101-108. http://dx.doi.org/10.1016/0045-6039(86)90004-7 [50] Shiokawa, K., Fu, Y., Kawazoe, Y. and Yamana, K. (1987) Mode of action of ammonia and amine on rRNA synthesis in Xenopus laevis embryonic cells. Develop- ment, 100, 513-523. [51] Wellauer, P.K. and Dawid, I.B. (1974) Secondary struc- ture maps of ribosomal RNA and DNA: I. Processing of Xenopus laevis ribosomal RNA and structure of single- stranded ribosomal DNA. Journal of Molecular Biology, 89, 379-395. http://dx.doi.org/10.1016/0022-2836(74)90526-9 [52] Webb, D.J. and Charbonneau, M. (1986) Weak bases inhibit cleavage and embryogenesis in the amphibians, Xenopus (toad) and Pleurodeles (newt), asterias (starfish). Cell Differentiation, 20, 33-44. http://dx.doi.org/10.1016/0045-6039(87)90463-5 [53] Stith, B.J. and Maller, J.L. (1984) The effect of insulin on intracellular pH and ribosomal protein S6 phosphoryla- tion in oocytes of Xenopus laevis. Developmental Biology, 102, 79-89. http://dx.doi.org/10.1016/0012-1606(84)90176-3 [54] Nuccitelli, R., Webb, D.J., Lagier, S.T. and Matson, G.B. (1981) 31P NMR reveals increased intracellular pH after fertilization in Xenopus eggs. Proceedings of the National Academy of Sciences of the United States of America, 78, Copyright © 2013 SciRes. OPEN ACCESS  K. Shiokawa / Advances in Bioscience and Biotechnology 4 (2013) 21-35 Copyright © 2013 SciRes. 35 OPEN ACCESS 4421-4425. http://dx.doi.org/10.1073/pnas.78.7.4421 [55] Lee, S.C. and Steinhardt, R.A. (1981) Observation on intracellular pH during cleavage of eggs of Xenopus laevis. The Journal of Cell Biology, 91, 414-419. http://dx.doi.org/10.1083/jcb.91.2.414 [56] Wasserman, W.J. and Houle, J.G. (1984) The Xenopus oocytes: A potential role for intracellular pH. Develop- mental Biology, 101, 436-445.
|