Paper Menu >>
Journal Menu >>
 Journal of Geographic Information System, 2011, 3, 18-49 doi:10.436/jgis.2011.31003 Published Online January 2011 (http://www.SciRP.org/journal/jgis) Copyright © 2010 SciRes. JGIS Adjusting Second Moment Bias in Eigenspace Using Bayesian Empirical Estimators, Dirichlet Tessellations and Worldview I Data for Predicting Culex Quinquefasciatus Habitats in Trinidad Models for West Nile Vectors Benjamin G. Jacob1, Dave D. Chadee2, Robert J. Novak1 1School of Medicine, Division of Infectious Diseases, University of Alabama at Birmingham, Birmingham, Alabama, USA 2Department of Life Sciences, University of the West Indies, St. Augustine, Trinidad and Tobago, West Indies E-mail: bjacob@uab.edu, dave.chadee@sta.uwi.edu, rjnovak@uab.edu Received October 24, 2010; revised November 12, 2010; accepted December 23, 2010 Abstract Temporally weighted regression models with a spatial autoregressive component may estimate nonlinearities in spatiotemporal-sampled data of Culex quinquefasciatus, a major vector of West Nile Virus (WNV) which can help implement control strategies by determining optimal predictors associated to prolific habitats. The design of this kind of mixed model can specifically incorporate spatial autocorrelation whilst including the influence of other aspatial predictor variables. Currently, the lack of an estimation theory that allows for het- eroscedasticity and corresponding joint hypothesis testing in the presence of spatial dependence in georefer- enced Cx. quinquefasciatus habitat data is a serious shortcoming in WNV research. In this paper we used spatially lagged and simultaneous autoregressive models based on multiple predictor variables of immature Cx. quinquefasciatus and Worldview 1 (WV-1) data to help implant a remote habitat-based surveillance sys- tem in Trinidad. Initially, we used Geomatica Ortho Engine® v. 10.2 for extracting a Digital Elevation Model (DEM) from the WV-1 raw imagery. Results of the DEM analyses indicated a statistically significant inverse linear relationship between total sampled Cx. quinquefasciatus data and elevation (m) (R2 = −0.439; p < 0.0001), with a standard deviation of 10.41. Additional field-sampled information was derived using data from an orthogonal grid-matrix constructed in an ArcInfo 9.3® and overlaid onto the WV-1 data. A unique identifier was placed in the centroid of each grid cell. Univariate statistics and Poisson regression models were then generated using the georeferenced covariates in SAS/GIS®. Coefficient estimates were also used to define expectations for prior distributions in a Bayesian estimation matrix using Markov Chain Monte Carlo (MCMC) specifications. A spatial residual trend analyses was then performed using autocorrelation indices which linked tabular data in SAS PROCLMIXED® with the egg-raft count data in ArcInfo®. The estimation matrix identified prolific habitats based on the covariate distance to the nearest house. An Ordinary kriged-based interpolator was then constructed in Geostatistical Analyst Extension of ArcGIS 9.3® based on the adjusted Bayesian estimates. For total Cx. quinquefasciatus egg-raft count, first order trend was fitted to the semivariogram at a partial sill of 5.931 km, nugget of 6.374 km, lag size of 7.184 km, and a range of 31.02 km using 12 lags. We assessed the performance accuracy of the interpolation procedures based on the magnitude and distribution of errors between observed and model-predicted values using Voroni tessella- tions. These residuals divided the space between the individual georeferenced Cx. quinquefasciatus habitats by XY coordinates in 2-dimenisional space which revealed that the geophysical parameter error residuals in the interpolation model were within normal statistical limitations. Newer GIS software and WV-1 data can generate highly accurate predictive Cx. quinquefasciatus habitat distribution models which can target prolific habitats of based on field-sampled count data. Our results suggest it may be unnecessary to manage all Cx.  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 19 quinquefasciatus habitats to obtain significant reductions in incidence and prevalence of WNV in Trinidad. Keywords: Culex Quinquefasciatus, Trinidad, West Nile Virus, Worldview 1, ArcGIS® 1. Introduction West Nile virus (WNV) is among the Culex-borne en- cephalitides in the Flavivirus genus. The virus cycles between birds and mosquitoes, while horses, humans, and a number of other vertebrates are considered inci- dental hosts. Since WNV first appeared in the Western Hemisphere in New York in 1999, it has spread rapidly across the North American continent, causing large numbers of human cases with neurologic disease and death and even greater amounts of milder disease char- acterized principally by fever and rash [1]. West Nile Virus has been spreading southward into the Caribbean Basin and Latin America as well, where its public health impact remains poorly understood [2]. The virus was first detected in 2001, in Jamaica and the Cayman Is- lands. Thereafter, serological evidence of WNV trans- mission has been accumulating. Cross-reactive WNV antibodies in humans have been detected from Mexico, the Bahamas, and Cuba, in horses from Guadalupe, Mexico, Central America, Puerto Rico, Colombia, and also in resident birds from the Dominican Republic and Venezuela. In November 2006, 4 serologically confirmed human WNV encephalitis cases were reported in Argen- tina. West Nile Virus may be following the same patterns of southward dissemination as other arboviruses into the Caribbean and Central and South America via migratory birds. There have been attempts to develop and implement WNV surveillance in the Caribbean but, unfortunately present systems in the region are unprepared to track WNV spread. For example, in Guadeloupe, extensive surveillance for WNV infection demonstrated a high seroprevalance in equines in 2003, despite no human or equine case being reported. A protocol for epidemiol- ogical surveillance for WNV has been developed through a collaborative electronic working tool posted on the Caribbean Animal Health Network (CaribVET) website (www.caribvet.net). CaribVET is a collaboration of vet- erinary services, diagnostic laboratories, research insti- tutes, universities, and regional/international organiza- tion to improve animal health in the Caribbean through exchange of information and data management. Improv- ing collaborations and information dissemination for understanding WNV epidemiology is vital; however, WNV surveillance and diagnostics in the Caribbean re- quires tools for regional data analyses. Unfortunately, to date there have been virtually no significant findings in any established WNV surveillance programs in tropical America. In september 2007, WNV infection detected by polymerase chain reaction (PCR) in post-mortem brain tissue taken from an encephalitic horse and by virus iso- lation from a dead bird confirmed WNV transmission in southwest Puerto Rico but, since then there have been no isolates reported from other areas in Central and South America or the Caribbean. The explanation for the paucity in WNV detection in tropical America may be the lack of active WNV sur- veillance. One would hypothesize that the likelihood of identifying WNV cases would be maximized in the Car- ibbean Basin, given its location in the flight paths of in- fected migratory birds. However, due to cross-immunity of WNV with other tropically endemic Flaviruses such as St. Louis encephalitis, it is possible that WNV has not been able to establish persistent enzootic foci in the re- gion, so that reported serological evidence may be caused by frequent but unsuccessful virus reintroductions. Re- gardless, presently a WNV mosquito habitat-based in- tervention surveillance system using a vigorous quantita- tive assessment of field and remote-sampled data is war- ranted for countries of the Caribbean. A main challenge in devising an effective West Nile surveillance program in regions of the Caribbean; how- ever, is the lack of sufficient empirical knowledge on the spatiotemporal patterns of Culex quinquefasciatus larval habitats and their underlying contribution to the adult population. The main tropical vector for WNV in birds and people is Cx. quinquefasciatus in the Southeastern United States and the Caribbean [3]. Previous research has revealed that Cx. quinquefasciatus abundance will vary significantly in different areas due to hydrologic regime and the proximity to habitats and blood-meal hosts [4-7]. These factors are not evenly distributed spa- tiotemporally, so it is logical to expect a variation in ab- undance and distribution of virus-positive mosquitoes within different ecosystems in the Caribbean. Recently, the use of remote sensing (RS) and GIS for the quantitative prediction of geographical distributions of agent, reservoir, and vector species has been advo- cated to augment traditional WNV mapping methods [4,5,8]. For example, the United States Geological Sur- vey (USGS) and Center of Disease Control (CDC) have employed GIS and RS to prepare interpretive maps showing WNV activity in many regions of North America (http://www.cdc.gov/ncidod/dvbid/westnile/resources/wn vguidelines1999.pdf). Common GIS/RS technology for WNV data analyses include: 1) time series overlay analyses of thematic data and spatial intersection, 2)  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 20 buffer generation and neighborhood analysis, 3) vec- tor-based grid generation and network analysis; and, 4) surface modeling [9]. Once an area has been imaged, associations between risk variables and environmental- sampled covariates can be quantified using the spatial data analysis capabilities of GIS for implementing a WNV mosquito surveillance system [9-11]. Quantification of vector-host interactions incorporated on Land Use Land Cover (LULC) maps generated in GIS time series overlay analyses suggest that Culex habitat productivity vary greatly over short distances, depending on the space and time at which field measurements are made [5,12-15]. For example, Jacob et al. (2009a) gener- ated a land use land cover (LULC) classification in Erdas Imagine based on Landsat-7 ETM+ data acquired July 2003 and Landsat-5 TM data acquired July 1991 for examining two ornithophillic mosquito vector species Culex pipiens and Culex restuans in residential areas of Urbana-Champaign, Illinois [4]. A maximum likelihood unsupervised classification and LULC detection was performed using a cross-tabulation detection method and a grid-based algorithm stratified by drainage for detection of environmental predictors associated to field-sampled egg-raft abundance count data. The resulting LULC change matrix revealed that between 1991 and 2003 there was a total of 12.1% LULC change in the study site. Of 20,166 egg-rafts sampled, 76.7% was in land cover classified as maintained urban followed by 14.8% in maintained non-urban. The well-drained stratum har- bored 56.4% of the total sampled data. Additionally, a remote stratification of the urban land cover site based on QuickBird visible and near infrared (NIR) data revealed that 83.3% of the egg-raft distribution was in LULC classified as residential high canopy coverage. In another study, Jacob et al. (2009b) constructed multiple geospa- tial models based on LULC and meteorological variables to predict adult abundance of the same Culex species in Des Planes mosquito abatement district in northern Cook County, Illinois [5]. An ecological database of the study site was constructed in ArcGIS 9.2® also using multitem- poral QuickBird visible and NIR data overlaid with total adult Culex abundance and WNV infection rates from 15 georeferenced gravid traps sampled weekly from May 2002 to October 2005. A regression analyses of LULC parameters revealed that adult Culex abundance in the georeferenced gravid traps was positively associated with temperature and negatively associated with precipitation. In 2002, the overall accuracy of the model was 73.3%, 65.1% in 2003, 47.2% in 2004 and 60.0% in 2005. Temperature and precipitation explained > 80.1% of the variation in each of the models. Estimates from the mul- tivariate analysis also revealed that land cover that was classified as forest LULC along with middle range built environment (< 40.1%) predicted heightened infected vector activity. Because abiotic factors such as LULC are measurable in GIS using submeter resolution data, these pre-epidemic abiotic signatures may be quantified for predicting arboviral epidemic transmission in isolated areas of the Caribbean. These epidemic signatures may be tracked, measured, quantified, and presented visually to help forecast human arboviral epidemics. While re- mote models of the observed spread of WNV have been used to evaluate potential vector and host mechanisms of dispersal and to define the suitability of the various landscape features and meteorological variables for mosquito transmission of WNV in many portions of the US, there is no literature using LULC and high resolu- tion satellite data for evaluating ecologically sampled Cx. quinquefasciatus habitats in any region of the Caribbean. Recently, digital elevation models (DEMs) have become popular with the use of GIS for surface modeling flood and swamp water mosquito abundance in many different geographic regions [16-18]. A digital elevation model (DEM) is a digital file consisting of terrain elevations for ground positions at regularly spaced horizontal intervals. It is a digital representation of ground surface topography which is usually represented as a raster or as a triangular irregular network. DEMs include a number of production strategies for determining terrain-related parameters associated to prolific Cx. quinquefasciatus habitats such as manual profiling from photogrammetric stereo models, digitizing of contours, digitizing topographic map contour plates, converting hypsographic and hydrographic tagged vector files and also for performing autocorrelation via automated photogrammetric systems. To obtain DEMs from satellite images for determining vector amplification and host population dynamics for mapping the spatiotemporal distribution of WNV from high resolution data two methods are possible: along-track stereoscopy from the same orbit using fore and aft images and across-track stereoscopy from two adjacent orbits [16]. The four possibilities of stereo image processing for DEM generation are Fore-Aft, Fore-Nadir, Aft-Nadir and Fore-Aft-Nadir images as a pair [19]. The simultaneous acquisition of along-track stereo data has a strong advantage in terms of radiometric variation versus the multi-date acquisition of across-track stereo data which has been traditionally employed in past research for identifying terrain-related explanatory covariates of floodwater mosquito habitats [20]. The across-track approach has been applied frequently since 1980, first with Landsat from two adjacent orbits, then with SPOT using across-track steering capabilities and finally with IRS-I C/D. But these satellite data were limited due to spatial resolution. Fortunately, along-track stereoscopy  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 21 has recently gained renewed popularity using higher resolution satellite data systems including JERS-i’s optical sensor(OPS), German Modular Opto-electronic, Multi-Spectral Stereo-Scanner (MOMS), ASTER, IKON- OS, QuickBird, Orthoview, SPOT-5, FORMOSAT II, Cantosat and the latest addition of Digitalglobe’s Worldview (WV)-1. These data can define georeferenced terrain covariates associated with prolific Cx. quinque- fasciatus habitats. For example, Jacob et al. (2010) used multispectral QuickBird data and DEM-based GIS methods to evaluate stream flow direction for determining satellite-derived characteristics of drainage basin physio- graphic predictors associated with Culex erraticus, a mosquito vector of eastern equine encephalitis virus (EEEV) and Northern Cardinals, an avian host associated with WNV and EEEV in Tuskegee, Alabama [16]. Eastern equine encephalitis virus is a virulent arbovirus maintained in an enzootic cycle between local avian fauna and ornithophilic mosquitoes in coastal regions of the eastern United States [1,14,16]. Euclidian distance- to-nearest hydrological body was calculated as the distance in geo-space from a grid cell to a stream grid cell defined by a Stream Raster Grid. A Terrain Analysis using a DEM (i.e., TauDEM) was then performed in ArcGIS 9.2® to retrieve multiple geomorphological parameters. Additionally, a three-dimensional model of the study area was constructed based on the DEM using ArcScene extension of ArcGIS®. A Pearson correlation analyses computed the ratio of covariance between the sampled variables to the product of their standard deviations. Substituting estimates of the covariances and variances based on the Cx. erraticus samples provided the sample correlation coefficient which revealed significant inverse linear relationships between the total field-sampled adult count data and elevation (R2 = −.4711; p < .0001) with a standard deviation (SD) of 11.16. The analyses of the Northern Cardinal data and elevation also revealed a similar statistical relationship (R2 = −.5831; p < .0001) with a SD of 11.42. Even though the usefulness of selected static catchment predictor covariates (e.g., stream, density, and slope) in assessing Culex habitats has been verified in previous research, characteristics of drainage networks and basin physiographic parameters have not been applied in modeling WNV mosquito abundance and distribution in any country of the Caribbean. Predictive interpolation algorithms have also not been used for forecasting productive Cx. quinquefasciatus habitats based on field and remote-sampled data in any country of the Caribbean. One of the most powerful and potentially valuable GIS/RS vector mosquito data mod- elling mechanisms is that of automated prediction. Gen- erally, given relevant spatiotemporal field and re- mote-sampled explanatory predictor variables with ap- propriate annotation, a WNV mosquito model can be constructed for making accurate forecasts of one or more dependent “target” quantities of interest. These quanti- ties of interest may be continuous real values (a scenario often referred to as regression” or “interpolation”), in vector mosquito data analyses or, it can be in the format of discrete labels (“aquatic habitat classification” or “habitat distribution pattern recognition”). Both these cases are referred to as “supervised learning” in the GIS/RS vector mosquito data learning vernacular. Pre- dictive interpolation algorithms can also map urban variation in vectors based on differentials in environ- mental-sampled parameters which can identify and vali- date models linking human ecological factors, while controlling for potential spatial autocorrelation in model residuals [21-25]. Spatial autocorrelation, the correlation among values of a single variable strictly attributable to their relatively close positions on a two-dimensional sur- face is frequently encountered in the analyses of spatio- temporal georeferenced vector mosquito data [4,8,21, 26,27]. Autocorrelation violates the ordinary least squares (OLS) assumption that the error terms are un- correlated. While it does not bias the OLS coefficient estimates in a predictive vector mosquito habitat distri- bution model, the standard errors tend to be underesti- mated and the t-scores overestimated when the autocor- relations of the errors at low lags are positive [16]. Kriged-based interpolation may analyze spatiotempo- ral-sampled WNV mosquito data in the presence of fine-scale autocorrelation. Kriging is mathematically related to regression analyses. Both theories derive a best linear unbiased estimator based on assumptions of co- variance and make use of the Gaussian-Markov theorem to prove independence amongst georeferenced vector mosquito data exploratory parameters [26]. Kriging vec- tor mosquito data is commonly made by interpolation of a single realization of a randomized study area based on field and remote observations in multivariate environ- mental-sampled datasets. Under the regionalized as- sumption, local habitat variation is not generally un- structured but, it is spatially dependent at some scale as georeferenced points within a given habitat distance apart depend on one another in a statistical sense (16) Any variable is stationary when its distribution is invari- ant under translation, that is, the stationary region is the location to make estimates (26). Stationary variables provide a preliminary model decision for uncertainty about the sample value at a specific sampled habitat lo- cation z(x) (16). For any space lag (h) increment in a particular direction, the distribution of z(x1), z(x2) ... z(xn) habitat based observations is the same as that of z(x1 + h), z(x2 + h)...z(xn + h), where z(x) is regarded as the sum of  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 22 three terms: (1) mean, drift or trend, (2) random depend- ent variable plus (3) error (16) This, however, requires the first two moments (i.e., mean and variance) to be constant. This spatial homogeneous region in the distri- bution of the georferenced Cx. quinquefasciatus habitats then will make random functions homogeneous and self-repeating in space. The variance of the random spa- tial variable will equal covariance at zero lag habitat dis- tance (σ2 = C (0), while the covariance between z(x) and z(x + h) becomes σx,x+h = σ2f(h) = C (h) (16). This means that the stationary covariance only depends on the sam- pled habitat distance and direction between two sampled habitats and not on their georeferenced locations, or the second order stationary assumption. Hence, the expected difference between sampled values at two georferenced Cx. quinquefasciatus habitat locations separated by dis- tance h is zero. If E (μ(x)) = m then z(x) = m + e(x) and E (z(x) - z(x + h)) = 0 (23). Covariance at h lag habitat distance then will be E{(z(x) - m)(z(x + h)-m)} = E(z(x) x z(x + h)) - m2 = Cov(h), being that z(x) the value at x sampled habitat site, z(x + h) the value at x sampled habitat site plus h lag habitat distance is based on m rep- resenting the overall mean and E() symbolizing the ex- pected value (16). Although the covariance does not exist theoretically in a predictive vector mosquito habitat dis- tribution model, the variance of the difference between two sampled habitat locations will be assumed to be pre- sent, as it does not depend on the georeferenced location and Var (z(x + h) - z(x)) = E({z(x + h) - z(x)} = 2γ(h), being E(z(x) - z(x + h) = 0. That is, differences between sampled habitat sites are merely a displacement function between them (i.e, the intrinsic hypothesis). The γ(h) (i.e., dissimilarity average function) will be the key tool for the structural interpretation of the predicted Cx. Quin- quefasciatsus habitats as well as for uncertainty estima- tion parameters. Mathematically, the variogram is de- fined as γ(h) = 0.5xVar(z(x + h) - z(x) (26). Since, the mean of z(x + h) - z(x) is zero for intrinsic variables then the variance is just the mean square difference. Conse- quently, the variogram for a predictive vector habitat distribution model is as follows: γ(h) = 0.5xE{(z(x + h) - z(x))2}(16). These predictive interpolation vector mos- quito habitat distribution models can measure unknown local mean habitat values as having a local linear or quadratic trend by combining regression of the depend- ent variable (e.g., larval/pupal count data) based on aux- iliary variables, such as terrain-related parameters (i.e., slope), RS imagery, and thematic maps, with kriged re- siduals where auxiliary predictors are used directly to solve the weights. For instance, in our previous example of research conducted in Tuskegee, Alabama for exam- ining vector-host activities of Culex erraticus, a kriged- based interpolator was also constructed in Geo-statistical Analyst Extension of ArcGIS 9.2® for predicting prolific habitats based on adult abundance count data [16]. For total adult Cx. erraticus count, a first-order trend ordi- nary kriging process was fitted to the semivariogram at a partial sill of 5.764 km, nugget of 6.114 km, lag size of 7.472 km, with a range of 32.62 km using 12 lags, sug- gesting that this species forage several kilometers beyond their habitats. The values that the semivariogram model attains at the range (i.e., the value on the y-axis) is called the sill, while the nugget or nugget variance, C0, occurs when a curve is fitted to the set of points that lie within the range and sill and the model used has a non-zero in- tercept [28]. Furthermore, to improve interpolation accuracy in a kriged-based algorithm, predictive mean standard error distributions can be quantified for attaining asymptoti- cally optimal predictors. Traditional kriging is motivated by an expected squared prediction error from a stochastic model [28]. Presently many algorthmitic paradigms of computational geometry have been applied efficiently in Geomatics and GIS using Dirichlet/Voronoi/Thiessen tessellation techniques for assessing interpolation models for quantifying the maximum error variance and the ex- tended prediction variance [28]. A Voronoi diagram is a special kind of decomposition of a metric space deter- mined by distances to a specified discrete set of objects in the space, (e.g., dataset of Culex habitat ground refer- ence coordinates). Generalization towards dynamic op- erations and kinematic motion have been investigated for about a decade using Dirichlet neighborhood analyses. Voroni residuals are amenable to assessment of good- ness-of-fit tests in a predictive vector mosquito habitat distribution models as they are approximately Gamma distributed and tend to be far less skewed than Pearson’s residuals and other residual diagnostic tests [16]. Such tessellations have been used in grid generation in vol- umes and surfaces, data compression, image segmenta- tion and edge detection, optimal allocation of resources, optimal quadrature rules, optimal representation, quanti- zation, finite difference tests and finite volume schemes. Unfortunately, there is no published literature using in- terpolation algorithms or Dirichlet tessellations for quan- tifying spatial error and predicting habitat locations using georeferenced spatiotemporal-sampled predictor vari- ables in WNV mosquito research for any country of the Caribbean. In this research, we generated multiple predictive geospatial models using field and remote-sampled ex- planatory variables of Cx. quinquefasciatus habitats in Trinidad. Elucidation of variability in mosquito produc- tivity requires spatial accounts of oviposition processes [29]. Indeed, this is important because of the recent iden- tification of sero-positive horses and domestic birds in  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 23 Trinidad [2]. Presently there is no remote habitat-based intervention for early detection of WNV activity in any regions of the Caribbean or Central and South America. A remote habitat-based intervention may provide early detection of arboviral activity by providing useful infor- mation for projection of the potential course of WNV transmission and also other arthropod-born viral diseases (e.g., St Louis encephalitis, Venezuelan equine encepha- litis) later in the season while unraveling the environ- mental factors associated with the maintenance and es- tablishment of enzootic cycles in Trinidad and in other countries of the neotropics. For mosquito larval control to be effective, however, it may be crucial to quantify individual sampled habitat variation to maximize control efforts. For example, the importance of variation in mosquito production among breeding sites in design of control programs is now broadly accepted in suppressing domestic container- breeding Aedes aegypti, a mosquito vector of Dengue in Trinidad. This research involved quantifying population densities of A. aegypti in four towns using standard house-to-house inspections of all water-holding contain- ers to determine whether persistently positive containers and premises existed over a three-month period in the wet season, from May to July 2002. From a total of 1,503 houses inspected, 15% were positive with 3% per- sistently positive over the three month period and classi- fied as ‘key premises’. From a total of 24,439 containers inspected, 1.3% were positive for A. aegypti larvae and pupae. Of 17 container types inspected, only water drums (54%), buckets (22%), tubs and basins (8%), wa- ter tanks (5%), brick holes (4%), and tires (2%) were significant larval habitats (P < 0.001) producers. The role that key premises play in the introduction and reinfesta- tion of A. aegypti-free communities was determined. The results suggested that A. aegypti control programs could be more cost effective and sustainable by targeting ef- forts on key premises and key containers to control mosquito densities and dengue transmission while re- ducing manpower needs and insecticide use. In this research, we constructed an autocovariate error matrix in SAS/GIS® for spatially targeting estimates generated from individual sampled Cx. quinquefasciatus habitat data. In our Bayesian model we did not assume that the georeferenced predictor variables were condi- tionally independent, as is commonly the case in hierar- chical generalized linear model (HGLM) construction practices for predicting WNV mosquito habitats. In our Bayesian network we assumed that the variables were conditionally independent. In practice, this assumption may not hold for WNV mosquito habitat models and may give rise to incorrect inferences. Violation of the assumption of statistical independence will give rise to incorrect model specifications [28]. Thus, we proposed the creation of a hidden node for modeling the depend- ency in the spatiotemporal-sampled parameters. In order to determine the conditional probability matrices for the hidden node, we used a gradient descent method. The objective function to be minimized was the squared-error between the measured and computed values of the in- stantiated nodes. Both forward and backward propaga- tion were used to compute the node probabilities for quantifying the dependence in the data. We also re- stricted our attention to the simultaneous autoregressive Gaussian spatial process and the autoregressive Gaussian response model (i.e., the spatial lag model), for quanti- fying latent autocorrelation error components in the Bayesian uncertainty estimates. We assumed that ob- served parameter error estimation patterns in the re- sponse variable decomposed into three statistically inde- pendent components: 1) a systematic spatial trend com- ponent that could be specified by a parsimonious set of exogenous variables; 2) a stochastic signal that reflects either an underlying spatial process and/or a set of miss- ing exogenous factors with an inherent spatial pattern; and, 3) independent white-noise disturbances. A specific subset of eigenvectors was then used to determine local and regional variation in the field and remote-sampled covariates by capturing dependencies among distur- bances generated from the Bayesian probabilistic model. Eigenvectors are a special set of vectors associated with a linear system of equations (i.e., a matrix equation) [4,5,8,10]. We constructed our remote habitat-based intervention by adoption of a landscape approach to elucidate mecha- nisms underlying Cx. quinquefasciatus productivity. Our landscape perspective was based on multiple georefer- enced explanatory variables of individual sampled habi- tats to determine their role in WNV disease transmission in Trinidad. Arguably, our framework evolved from the traditional concept of regression models generated from LULC covariates for identification of hot zones (i.e., cluster), but our remote habitat-based intervention also focused on critical spatial autoregressive modifications of the georeferenced predictors. We developed our mod- els to emphasize the role of oviposition foraging in Cx. quinquefasciatus in Trinidad. Habitat-based interventions should emphasize the link between foraging behaviors of egg-laying mosquitoes and the availability of breeding sites in evaluation of environmental management pro- grams [29]. The basic assumption in our models was that heterogeneity in immature productivity was substantially large among Cx. quinuefaciatus habitats and that the number of productive habitats was relatively fewer than that of unproductive ones in focal areas. Untargeted in- terventions are inefficient because habitats are randomly  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 24 selected without consideration of the distribution of pro- duction (29). Spatially targeting prolific breeding habitats using a remote habitat-based intervention may reduce Cx. quinuefaciatus populations in Trinidad and help imple- ment a cost-effective surveillance mechanism. Therefore, our objectives of this research were to: 1) construct a DEM to identify terrain-related explanatory covariates associated with spatiotemporal-sampled mosquito data; 2) generate regression models to determine statistical sig- nificance associated with the sampled covariates, 3) quantify latent autocorrelation components in model re- sidual variance estimates generated from an eigenfunc- tion decomposition spatial filter algorithm; and, 4) de- velop spatial interpolation models of potential prolific habitats site based on field-sampled count data. Although the discussion is centered on WNV vectors specifically in Trinidad, the framework and derived guidelines in this research may be applicable to integrated control pro- grams for other mosquito species and insect-borne dis- eases in other regions. 2. Material and Methods 2.1. Study Site The island of Trinidad is the most southerly part of the Lesser Antilles located at approximately 10° north lati- tude and 15 km from the eastern coast of Venezuela. The geology, flora and fauna are typical of the adjacent South American mainland [31,32]. Trinidad is approximately 80 km from north to south and is on average 60 km wide, roughly rectangular in shape with large promontories on the northwest and southwest of the country. The climate of Trinidad is tropical with a “rainy” season between May and November and a “dry” season from December to May. Details about climate, physical features, vegeta- tion, and population size have been previously described [33,34]. 2.2. Mosquito Egg Sampling The Cx. quinquefasciatus habitat population was sur- veyed by counting the number of egg-rafts deposited in oviposition traps. Field sampling was conducted from July 2008 to July 2009. One hundred and fifty two tem- porary, permanent, and semi-permanent mosquito larval habitats in the study site were mapped and classified, and recalibrated using a CSI-Wireless differential corrected global positioning systems (DGPS) Max receiver which yields a positional error of .179 m (+/− 392 m) [35]. Wa- ter bodies were inspected for Cx. quinquefasciatus egg- rafts. Oviposition traps were located in shady areas around residences and municipal buildings, as well as the edge of parks and woodlots. 2.3. Habitat Base Mapping Initially, Landsat images were collected using SLC-off mode (http://edc.usgs.gov/#Find_Data/Products_and_ Data_Available/ETM). We used the USGS Global Visualization Viewer (GloVis) system (http://glovis.usgs. gov/) to search for images for performing a land cover classification. Path 233/row 53 covered the Trinidad study area. Unfortunately, the images acquired from GloVis, were all cloudy, due to the climate conditions of the island. Cloudy image is an issue for remote classification and change detection for identification of mosquito habitats [24]. We did find three Landsat 7 passes of path 233/row 53 which were less than 10% cloud free. Theses collection periods were April 30, May 16, and June 1, 2005. These data; however, were not able to discriminate the land cover due to poor resolution (i.e., 30m resolution) associated with the overlaid geo- referenced Cx. quinquefasciatus habitats coordinates. We acquired WV-1 data (www.digitalglobe .com) for the study site on July 2010. Operating at an altitude of 496 kilometers, WV-1 has an average revisit time of 1.7 days and is capable of collecting up to 750,000 km2 per day of half-meter resolution imagery. The satellite is equipped with state-of-the-art geolocation accuracy capabilities and exhibits stunning agility with rapid targeting and efficient in-track stereo collection. Various collection sizes of surveying are available: frame mode, route survey (along coastline, roads and linear objects), areal survey (60 × 60 km), and stereoscopic areas on a single pass as well (www. digitalglobe.com). In this research, the WV-1 imagery was radiometrically and sensor corrected but was not projected to a plane using a map projection or datum. The sensor correction blended the pixels from all the detectors into synthetic arrays to form the image of the study site. The geolocation accu- racy specification of the data was 4.0 to 5.5 m CE 90 at nadir excluding terrain and off-nadir effects. The im- agery was delivered at full-swath cut into 14 km lengths. The WV-1 imagery was classified using the Iterative Self-Organizing Data Analysis Technique (ISODATA) unsupervised routine in ERDAS Imagine v.8.7™ (At- lanta, USA) which is commonly used to classify land covers associated with disease foci [18,26,29]. The im- ages were co-registered manually using ground control points (GCPs) and by applying a first order polynomial algorithm with a nearest neighbor re-sampling method. The Universal Transverse Mercator (UTM) Zone 20P datum WGS-84 projection was used for all of the spatial datasets. Base maps were prepared from the field-sampled Cx.  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 25 quinquefasciatus habitat data and WV-1 imagery for the study site using the DGPS ground coordinates. We gen- erated GIS data layers in the vector shapefile format, ArcInfo® Coverage format and raster GRID format for use in the GIS software package ArcGIS v 9.3®. We constructed a discrete tessellation of the region using a digitized grid-based algorithm (Figure 1). For remote identification of vector mosquito habitats the first step is often to construct a discrete tessellation of the region [36]. A unique identifier was placed in each grid cell. Each Cx. quinquefasciatus habitat/polygon was assigned a unique identifier. Field attribute tables were then asso- ciated to the polygons. 2.4. Environmental Data Analyses Base maps were also generated using the spatiotemporal- sampled Cx. quinquefasciatus habitat data which was stored in a Vector Control Management System® (VCMS) (Clarke Mosquito Control Products, Inc. 159 N. Garden Avenue, Roselle, IL 60172) database. The VCMS sup- ported the mobile field data acquisition in the study site through a PocketPC™. All two-way, remote synchroni- zation of data, geocoding and spatial display were proc- essed using the embedded GIS Interface Kit™ that was built using ESRI’s MapObjects™ 2 technology. The VCMS database plotted and updated the DGPS ground coordinates of the Cx. quinquefasciatus habitats and supported exporting data in spatial format in which any combination of the sampled habitats and explanatory covariates were described in ESRI shapefile format for use in GIS. The database displayed this information onto a user-defined field base map. 2.4.1. Spatial Hydrological Model The latest version of PCI Geomatics Orthoengine® soft- ware was used to generate a DEM using the WV-1 data and the georeferenced Cx. quinquefasciatus habitat co- variates (Figure 2). OrthoEngine offers an industry- leading variety of control sources including, manual en- try, geocoded imagery, geocoded vectors, chip database, digitizing tablet, or a text file (www. pcigeomatics.com). We used the DGPS coordinates to orient the images to our map coordinate system. We used PCI Geomatics OrthoEngine to extract the DEM. PCI software sup- ports automatic GCP/tie point (TP) collection, using Toutins rigorous models, Rational Polynomial Coeffi- cient (RPC) model construction, automatic DEM genera- tion, orthorectification and automatic mosciaking (www. pcigeomatics.com). WV-1 stereo pairs were supplied by the full scene of the Trinidad study site using Basic 1B level data de- signed for the creation of the DEM. A set of WV-1 Basic 1B stereo images were provided by Digitalglobe. The set of stereo data was in Basic 1B OR2A product format. Due to the 2 gigabyte limit of the TIFF data, WV-1 data was distributed in tile format. Each tile came with its own RPC file. Because our OR2A format was map pro- jected, the effects of any high frequency movements had already been removed from the scene resulting in a good fit for the RPCs. In order to leverage the WV-1 image for attaining to- pographic indices from the Cx. quinquefasciatus data, a geometric model was required. We generated an RPC model. The RPC model has proven to be the most popu- lar geometric model for high resolution images (www. digitalglobe.com). Since bias or errors may still have been present in the RPCs, our results were preprocessed with a polynomial adjustment using several accurate GCPs (Table 1). A zero order RPC adjustment is ade- quate to obtain an RPC model accuracy within 1m (www.digitalglobe.com). To generate a DEM for determining the terrain-related Cx. quinquefasciatus habitat parameters in the Trinidad study site, a pair of quasi-epipolar images were generated from the fore and aft stereo images to retain elevation parallax in the X-direction only. An automated image match procedure was then used to produce the DEM through a comparison of the respective gray values of these images. To find the corresponding pixels in the left and right quasi-epipolar images, we used a hierarchical sub-pixel mean normalized cross-correction matching method. The actual matching method employed gener- ated correlation coefficients between 0 and 1 for each matched pixel, where 0 represented a total mismatch and 1 represented a perfect match. A second order surface was then fitted around the maximum correlation coeffi- cients. The difference in the georeferenced Cx. quinque- fasciatus habitats between the images gave the disparity Table 1. Rational polynomial coefficient results for the WV- 1 digital elevation model. CP RMS error CP RMS error Product Number of GCPs Number of CPs X Y X Y Fore 0 19 0.4 .1 0.3 2.5 1 17 0.4 0.5 0.6 0.7 Aft 0 16 0.4 1.3 1.0 2.2 1 17 0.5 1.1 1.3 2.7 GCP: Ground Control Points, CP: Control Point, RMS: Root Mean Square  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 26 Figure 1. Gridded area of urban land cover in the Trinidad study site using WV-1 with georeferenced Cx. quinquefasciatus habitats. Figure 2. A digital elevation model (DEM) based on georeferenced Cx. quinquefasciatus habitats for the Trinidad study site. 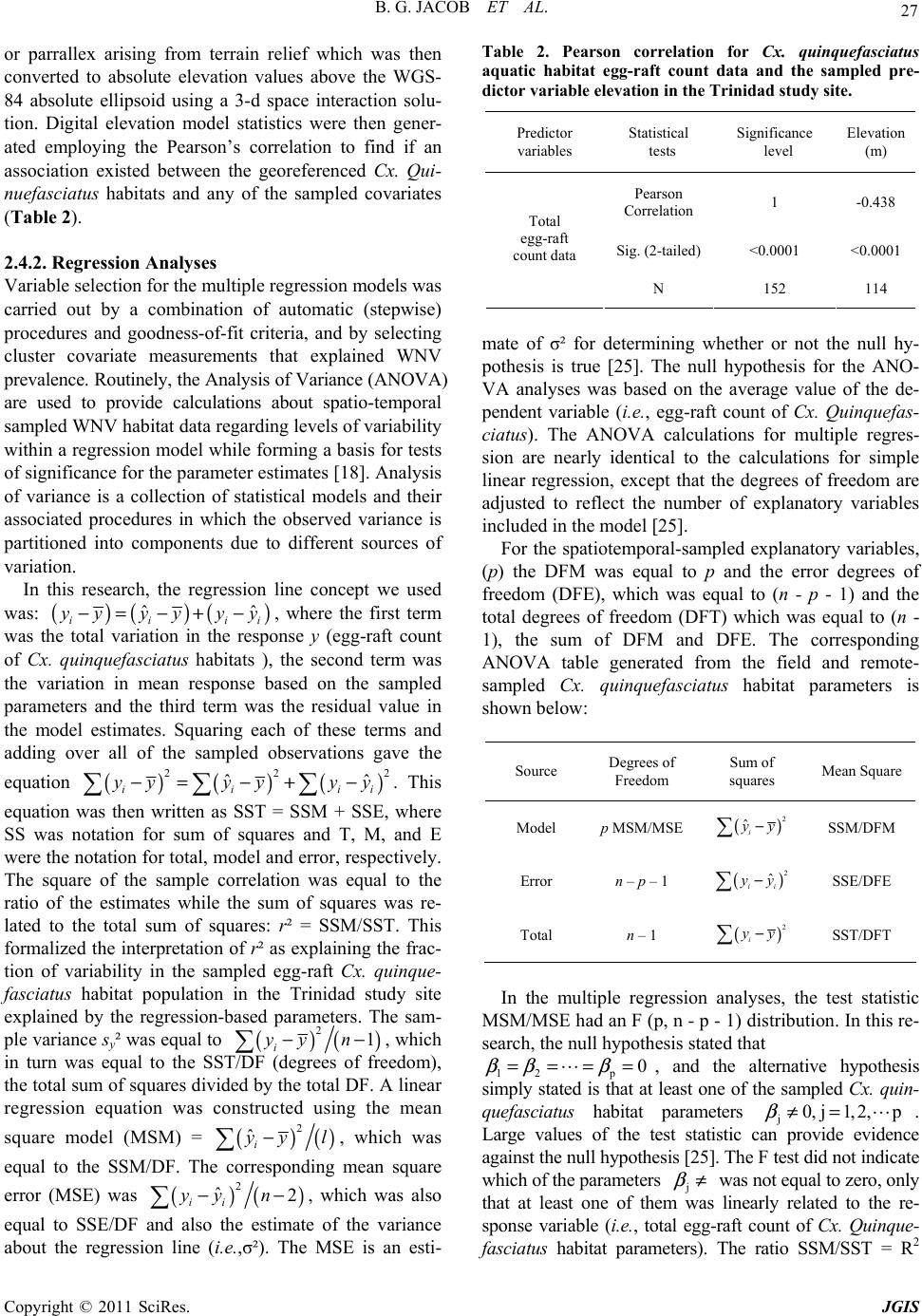 B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 27 or parrallex arising from terrain relief which was then converted to absolute elevation values above the WGS- 84 absolute ellipsoid using a 3-d space interaction solu- tion. Digital elevation model statistics were then gener- ated employing the Pearson’s correlation to find if an association existed between the georeferenced Cx. Qui- nue fa s ci a tus habitats and any of the sampled covariates (Table 2). 2.4.2. Regression Analyses Variable selection for the multiple regression models was carried out by a combination of automatic (stepwise) procedures and goodness-of-fit criteria, and by selecting cluster covariate measurements that explained WNV prevalence. Routinely, the Analysis of Variance (ANOVA) are used to provide calculations about spatio-temporal sampled WNV habitat data regarding levels of variability within a regression model while forming a basis for tests of significance for the parameter estimates [18]. Analysis of variance is a collection of statistical models and their associated procedures in which the observed variance is partitioned into components due to different sources of variation. In this research, the regression line concept we used was: ()()( ) ˆˆ iiii y yyyyy−=−+ − , where the first term was the total variation in the response y (egg-raft count of Cx. quinquefasciatus habitats ), the second term was the variation in mean response based on the sampled parameters and the third term was the residual value in the model estimates. Squaring each of these terms and adding over all of the sampled observations gave the equation ()()( ) 22 2 ˆˆ iiii yyyy yy−=−+ − . This equation was then written as SST = SSM + SSE, where SS was notation for sum of squares and T, M, and E were the notation for total, model and error, respectively. The square of the sample correlation was equal to the ratio of the estimates while the sum of squares was re- lated to the total sum of squares: r² = SSM/SST. This formalized the interpretation of r² as explaining the frac- tion of variability in the sampled egg-raft Cx. quinque- fasciatus habitat population in the Trinidad study site explained by the regression-based parameters. The sam- ple variance sy² was equal to () () 21 i yy n−− , which in turn was equal to the SST/DF (degrees of freedom), the total sum of squares divided by the total DF. A linear regression equation was constructed using the mean square model (MSM) = () () 2 ˆi yy l− , which was equal to the SSM/DF. The corresponding mean square error (MSE) was () () 2 ˆ2 ii yy n−− , which was also equal to SSE/DF and also the estimate of the variance about the regression line (i.e.,σ²). The MSE is an esti- Table 2. Pearson correlation for Cx. quinquefasciatus aquatic habitat egg-raft count data and the sampled pre- dictor variable elevation in the Trinidad study site. Predictor variables Statistical tests Significance level Elevation (m) Pearson Correlation 1 -0.438 Sig. (2-tailed) <0.0001 <0.0001 Total egg-raft count data N 152 114 mate of σ² for determining whether or not the null hy- pothesis is true [25]. The null hypothesis for the ANO- VA analyses was based on the average value of the de- pendent variable (i.e., egg-raft count of Cx. Quinquefas- ciatus). The ANOVA calculations for multiple regres- sion are nearly identical to the calculations for simple linear regression, except that the degrees of freedom are adjusted to reflect the number of explanatory variables included in the model [25]. For the spatiotemporal-sampled explanatory variables, (p) the DFM was equal to p and the error degrees of freedom (DFE), which was equal to (n - p - 1) and the total degrees of freedom (DFT) which was equal to (n - 1), the sum of DFM and DFE. The corresponding ANOVA table generated from the field and remote- sampled Cx. quinquefasciatus habitat parameters is shown below: Source Degrees of Freedom Sum of squares Mean Square Model p MSM/MSE () 2 ˆi yy− SSM/DFM Error n – p – 1 () 2 ˆ ii yy− SSE/DFE Total n – 1 () 2 i yy− SST/DFT In the multiple regression analyses, the test statistic MSM/MSE had an F (p, n - p - 1) distribution. In this re- search, the null hypothesis stated that 12p 0 ββ β ====, and the alternative hypothesis simply stated is that at least one of the sampled Cx. quin- quefasciatus habitat parameters j0,j1, 2,p β ≠=. Large values of the test statistic can provide evidence against the null hypothesis [25]. The F test did not indicate which of the parameters j β ≠ was not equal to zero, only that at least one of them was linearly related to the re- sponse variable (i.e., total egg-raft count of Cx. Quinque- fasciatus habitat parameters). The ratio SSM/SST = R2  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 28 (i.e., squared multiple correlation coefficient) was the pro- portion of the variation in the response variable that was explained by the field and remote-sampled Cx. quinque- fasciatus habitat data. The square root of R² (i.e., the multi- ple correlation coefficient) was the correlation between the sampled observations yi and the fitted values ˆi y. The relationship between WNV prevalence and each individual potential predictor variable sampled in the Trinidad study site was also investigated by single vari- able regression analysis using PROC MIXED. Since parasite prevalence data are binomial fractions, a regres- sion model was used, as is standard practice for the anal- ysis of the spatiotemporal-sampled WNV mosquito data [4]. Poisson regression analyses were also constructed in PROC MIXED to determine the relationship between Cx. quinquefasciatus habitat egg-raft count data and the sampled habitat characteristics. These regression models were generalized linear models with the logarithm as the (canonical) link function, and a Poisson distribution function. The regression analyses assumed independent counts (i.e., ni), taken at sampled habitats I = 1, 2… n. The field- sampled count data were described by a set of explana- tory variables denoted by matrix Xi, a 1 × p vector of co- variate measurements for a sampled habitat i. The ex- pected value of these data was given by μi (Xi) = ni (Xi) exp (Xi β), where β was the vector of non-redundant pa- rameters and the Poisson rates parameter was given by λi (Xi) = μi (Xi)/ni (Xi) [37]. The rates parameter λi (Xi) was both the mean and the variance of the Poisson distribu- tion for each sampled Cx. quinquefasciatus habitat loca- tion i. The dependent variable was total field-sampled egg-raft count data. The Poisson regression model as- sumed that the data was equally dispersed, meaning that the conditional variance equaled the condition mean. The procedure used maximum likelihood estimation to find the regression coefficients. The data was log-transformed before analyses to normalize the distribution and mini- mize standard error. The predictor variables used in the regression analyses are listed in Table 3. 2.4.3. Bayesian Analyses In the Bayes formulation, the specification of the Cx. quinquefasciatus habitat model was completed by as- signing priors to all unknown parameters. We used our dataset of spatiotemporal-sampled Cx. quinquefasciatus observations X = [x1,, xn] where each xi for i = 1 n was assumed to be distributed according to some dis- tribution p(xi|θ). In this research, θ was a parameter that was unknown and had to be inferred from the data. Our Bayesian procedure began by assuming that θ was dis- tributed according to some prior distribution p (θ|α), where the parameter α was a hyperparameter. Table 3. Ecological variables collected in the Trinidad study site. Variable Description Units EGG-RAFT COUNT count data (dependent variable) Number collected STUDY SITETrinidad 10 × 10 m grid cells DISHOUSE Distance to house from sampled aquatic habi- tats Meters DHABITAT Distance to house from sampled aquatic habitats Meters SHADE Amount of shade Percent of grid cell that is shaded (nearest 10%) DRAINAGE Drainage status of soils 0 = poorly drained, 1 = well drained CANOPY Canopy cover above larval habitat Percent of grid cell with canopy cover (nearest 10%) Elevation The geographic height above a habitat reference point, Meters The joint of the Cx. quinuefasciatus data was then generated using: ()()() 11 1 ,, n ni ppxx px θθ = θ= =∏X; whereby, ()() ,papθ= θXX and ()() , ii px a pxθ= θ were conditionally independent of the hyperparameter. Bayes- ian inference then determined the posterior distribution of the parameter () ,paθX using : () () () () () ()() ()() ()() ()() ()() ()() 1 1 ,, ,, ,,,, , , n i i n i i pp pppd pp pp ppdppd px p px pd θ θθ θ θα θα θα αθαθ θα θαθ θα θ αθαθθθαθ θθα θθαθ = = == == = ∏ ∏ XX XXX XX XX For the fixed regression parameters, a suitable choice was the diffuse prior, i.e., p (γ) ∝const, but a weakly informative Gaussian prior was also possible. A second- order Gaussian random walk prior was used to allow enough flexibility while penalizing abrupt changes in the function [17]. The prior was expressed as: () 22 2 tt1t2 3 ()exp f2ff 2 t f ft pf τ τα −− = −−+ where  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 29 () 1, p f ff= and 2 f τ was the variance, with diffuse priors 1 f ∝ const, 2 f ∝ const. For additional spatial effects in the sampled Cx. quin- que fa sc ia tu s data, smoothness priors were assigned whose joint distribution, s, was given by: () 22 2 t11 t 12 ()exps s 2 f ft ps τ τ τα − = −+…+ . Again, assuming diffuses priors, the variance that con- trolled the degree of smoothness in the model was 1 s, 11 s, and 2 f τ . The unstructured spatial heterogeneity term, ui, was assumed to follow an exchangeable Gaussian prior with zero mean and variance, () 22 ,.., ~0, ui u ie uN ττ . A similar prior was then assigned to the heterogeneity term for the sampled predictor variables, i.e. , () 2 ~0, iu hN τ . Finally, for the spatial components, vi, a Markov random field (MRF) prior was assigned. The conditional distribution of vi, given an adjacent covariate, vj, was a univariate normal distribution with mean equal the average vj coefficient values of vi’s neighboring sampled site and variance equal to 2 v τ divided by the number of total adjacent sampled Cx. quinquefasciatus predictors. This lead to a joint density of the form: () () 22 2 i~j exp 2 v vij pvv v τ τα −− where i ~ j denoted a sampled site i adjacent to j, and where the parameter estimates vi and vj in the adjacent sampled sites were similar. The degree of uncertainity in the spatiotemporal-sampled data was determined by the unknown precision parameter 2 v τ . By writing j jj f z β =, kk hz β =, ll uz β =, and mm vz β = for a well defined design matrix Z, a vector of regression parameters β, with all different priors, was expressed in a general Gaussian form using the expression: () 2' jj 2 1 exp β 2 j jj j pK β τα β τ with an appropriate penalty matrix Kj. The model structure was dependent on the sampled explanatory variables and smoothness of the function. In most cases, Kj is rank deficient and, hence, the prior for βj is improper [18]. For the variances 2 j τ inverse Gamma priors IG (aj, bj) was assumed, with hyper- parameters aj, bj chosen such that this prior was weakly informative. The Bayesian framework in this research was defined by conditional probabilities constructed from environ- mental-sampled Cx. quinquefasciatus habitat data. The observation nodes in the Bayesian estimation model were denoted by a vector () 12 ,,, N x xx x=, and the set of states of the observation node x j generated from the sampled data that was represented by { } 1, 2,, j j x Y∈. In this research the hidden nodes were denoted by { } 1, 2,, kk zT∈. The probability that the state of the hidden node k z was I, 1k iT≤≤ , was expressed as () () ,:k ki P zi α == . Because () { } ,,1,2,, k ki iT α = is a probability distribution, () , 11 k T ki i α == , holds for 1, 2,,kK= [10]. The conditional probability in this research was that the jth observation node xj was l, 1 j iY≤≤ , which was based on the condition that the states of hidden nodes were () 12 ,, , N zzz z=, gener- ated by () (,|) | jlz j bPxl==. We defined () , :{ } ki αα −, () { } ,| :jlz ββ =, and let { } , ωα β = be the set of all the spatiotemporal-sampled Cx. quinquefasciatus habitat parameters in the study site. Then the joint probability that the states of observation nodes were () 12 ,,, N x xx x= and the states of hidden nodes were () 12 ,, , N zzz z= based on () () () 11 ,,, kN kj kj Pxzkz jxz ωα β == =∏∏ . The marginal probability that the states of observation nodes were x was generated using () { } () () 1 11 1 ,,,| k KK N T kj k kk j Pxzkzjx z ωαβ = == = = ∏∏ ∏, and we used the notation { } { } 12 122 1111 1: kK K KTTTT zkz zz k z ==== = == ∏ for the summation over all states of hidden nodes. We assumed the sampled Cx. quinquefasciatus habitat pre- dictor variables { } 12 ,,, nn X XX X= were independ- ently and identically taken from the true distribution () o px . In Bayesian learning, the prior distribution ϕ ω on the parameter ω is set [18]. As such, the posterior distribution () n pX ω was computed from the spatio- temporal-sampled Cx. quinquefasciatus habitat dataset and the prior by () () () () () n 1expnH ωφω n n pX ZX ω =− which was generated using the expression () () 11 H11log({) |) o nninpXi ωω ↑ ↓ ==≡ , and () n Z X (i.e., the normalization constant). The Bayesian predictive distribution () n pxX was pro- vided by averaging the model over the posterior distribu- tion as follows, () { ()() { } nn pxXpx p xXd ωω ω =. The Bayesian stochastic complexity () n F X was de- fined by () n F X = –logZ () n X which was used as a criterion by which the model was selected and the hy- perparameters in the prior were optimized. We let [] n x E be the expectation over all the sampled Cx. quinquefas- ciatus parameters. The Bayesian stochastic complexity had the following asymptotic form using  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 30 ()() () () [log1log log log1EXnFXnn mnO λ ↑↑ ↓≈−− + where λ and m were the rational number and the natural number respectively, and the two were determined by the singularities of sampled predictor variables. In regular models, 2λ is equal to the number of parameters and m = 1, while in non-identifiable models, 2λ is not larger than the number of parameters and m ≥ 1 [38]. However, Bayesian frameworks using field and remote-sampled vector mosquito data requires integration over the poste- rior distribution, which typically cannot be performed analytically [4]. In this research, we let { } , nn X Z be the sampled Cx. quinuqefasciatus parameter estimates corresponding to the hidden error variables in the equation n Z = { } 12 ,,, n Z ZZ. The variational Bayesian framework approximated the Bayesian posterior () , nn pZ X ω of the hidden variables and the Cx. quinuqefasciatus pa- rameters using the variational posterior () , nn qZ X ω , which was factorized using () () () , nn nn n qZXQZ XrX ωω =, where () nn QZ X and () n rX ω were posteriors based on the inconspicuous error variables and the sam- pled data respectively. The variational posterior () , nn qZ X ω was chosen to minimize the functional F [q] defined by [] ()()() () () ,log, ,, n nnnn n nn Z qZ XqZ XpoX qd pX FZ ωω ω ω =, which was then further defined by the equation ()( ) () (,|)||,| s FXnKqZnXnpZn Xn ωω ↑↑↑↑↑ =+ using the sampled parameters, where ()() () ,| ||,| K qZnXnpZn Xn ωω ↑↑ ↑↑ and the true Bay- esian posterior was () ,| nn pZ X ω and the variational posterior was () , nn qZ X ω . This led to the functional [ ] F q being minimized under the constraint and then the variation posteriors, () n rX ω , and () nn QZ X which was computed using the equations () () () 1exp nnn r rXlogpXZ Q C ωϕωω =< > and () () 1exp , nn nn Q Q ZXlogpXZr C ω =< > where r C and Q C were the normalization constants. It is important to note that these equations gave only nec- essary conditions for the functional [ ] F q to be mini- mized in the Cx. quinuqefasciatus habitat model. The variational posteriors were computed by an iterative al- gorithm. We defined the variational stochastic complex- ity () n F X by the minimum value of the functional [ ] F q which was () [] min r,Q n F X Fq=, based on the dif- ference between () n F X and the Bayesian stochastic complexity () n F X. We then generated variational posterior for the esti- mation matrix. We assumed that the prior distribution () ϕ ω of the Cx. quinquefasciatus habitat parameters { } ,ab ω = was the conjugate prior distribution. Then () ϕ ω was generated by the equation () () () () 01 0 1, 0 Γ,1,2,, Γ k kkk T k kTzkz T aakK φ φ ϕφ − = == ∏, and covariates estimates were provided by () () () () 0 0ξ1 (|) 1 0 Γξ ,1,2,, Γξ j nnjj jY jz jzYx Y bbjN ϕ − = == ∏, which were Dirichlet distributions with hyperparame- ters generated using 0 φ > 0 and 0 ξ > 0. The Dirichlet distribution [i.e., Dir(α)] is a family of continuous multi- variate probability distributions parameterized by the vector α of positive reals which can generate the multi- variate generalization of the beta distribution and conju- gate prior of the categorical distribution and multinomial distribution in Bayesian statistics for predictive vector mosquito habitat models [4]. The Dirichlet distribution is the multinomial extension to the beta distribution for a binomial process which can also be used in quantifying probabilities in predictive vector mosquito habitat prob- ability models [39]. We then let () n δ be 1 when n = 0 and 1 otherwise, and then defined the sampled parameter uncertainty es- timates using () () () () ,1 : k nk z ik kz iQ nZz δ = =− . In this re- search we also used () () () () () () () 11 ,: j K njk x ij ik ik jx zQ nXxZz δδ == −=− ∏ . In these equations, () j i X was the state of the jth ob- servation node and () k i Z was the state of the kth hidden node. The variational posterior distribution of parameters { } ,ab ω = was given by using the equation () () () () () 0 k ,1 0 , 1 z 0 k,z 1 Γ(n ) | Γn x kk k k k k Tnkz nk kkz z T z T ra Xa φ φ φ +− = = + =+∏ ∏ where each of the spatiotemporal-sampled Cx. Quinqe- fasciatus habitat covariates were generated using the equation () () () () () () () :1 1 nzxi n kKZikZkQ ↑ ↑ ↓↓ ↑ ↓↓↓ ↓ ==≡ =≡ − ∏  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 31 It then followed that 1(, |) j j j x y x z zj xx nn = = for 1,,jN= and () ,kk z x z kz z nn − = denoted the sum over () i zi k≠. In this research we evaluated the statistical efficiency of the MCMC sequence in WinBUGS® using the fol- lowing steps: Step1: Sample S(m) from f(S|X,Z(m-1)) Step2: Sample P(m) from f(P|X,S(m),Z(m-1)) Step3: Sample Z(m) from f(Z|X,P(m),S(m),Φ(m-1)) where X was the field and remote-sampled Cx. Quinque- fasciatus habitat data and Z was randomly assigned a starting value Z(1) using a uniform prior distribution. Step 1 was performed by drawing from an inverse Wishart distribution. In Bayesian statistics inverse Wi- shart distribution is used as the conjugate prior for the covariance matrix of a multivariate normal distribution [9]. The probability density function of the inverse Wi- shart was: () () 1 ψB/2 12 /2 2 ψ||| 22 trace mp m mp p Be m τ − − −++ where B and ψ were positive definite matrices, and p τ (·) was the multivariate gamma function. In this re- search the distribution of the inverse of a Wishart-distrib- uted matrix was generated from () ~, A Wm and , which was of size of the matrix constructed from the field and remote-sampled Cx. quinquefasciatus habi- tat parameters. Under these circumstances 1 BA − = had an inverse Wishart distribution, () 1 1 ~,BWm − −. We calculated the probability density function: () () 1 /2 1 2 2 |||exptrB 2 |, 22 mp m mp p B pB mm ψψ ψ τ ++ −− − = where 1 ψ − =and p τ (·) was the multivariate gamma function of the sampled data. A probability density func- tion or density of a continuous random variable is a func- tion that describes the relative likelihood for this random variable to occur at a given point [25]. The marginal and conditional distributions from the inverse Wishart-distributed matrix was quantified using () 1 ~, A Wm ψ −. We then partitioned the matrices for determining if ψ was conformable with each other using: 11 12 21 22 AA AAA = , 11 12 21 22 ψψ ψψψ = where Aij and ij ψ were Pi x Pj matrices. We then determined if: 1) 11 A was independent of 1 11 12 AA − and 221 A⋅, when 1 22 122211112 AAAAA − ⋅=− which was the Schur com- plement (i.e., a submatrix within a larger matrix) of 11 A in A; 2) 11 A~ () 1 11 2 ,Wmp ψ −−; 3) 12 111 111222 1x111222 111 AA |A~(,A) pp MN ψψψ −−− ⋅⋅ ⊗ , where (,) pxq MN ⋅⋅ was a matrix normal distribution gen- erated from the spatiotemporal-sampled Cx. quinque- fasciatus habitat parameters; 4) () 1 22 111 A~ ,Wm ψ − ⋅. We used the Conjugate distribution to make inference about a covariance matrix whose prior () p had a () 1 11,Wm ψ − distribution. If the observations 1,, n X xx= are independent p-variate Gaussian vari- ables drawn from a () 0,N distribution, then the con- ditional distribution p X has a () 1,Wnm ψ −++A distribution, where A =XXT is n times the sample co- variance matrix [3]. Because the prior and posterior dis- tributions are the same family, the inverse Wishart dis- tribution was the conjugate to the multivariate Gaussian generated from the georeferenced Cx. quinquefasciatus habitat parameters. Due to its conjugacy to the multivariate Gaussian it was possible to “integrate out” the Gaussian’s parameter using: () ()() 2 22 X, X, 2 A2 m p np mn p PmPPmd mn m ψψ ψτ πψτ + = + = + This task was simplified by assuming that the spatio- temporal-sampled Cx. quinquefasciatus habitat data analyses had covariance matrices with common eigen- vectors. If covariance’s differ among sources, the inverse Wishart draws often produce invalid, especially for sources that are small components of the mixture [9]. In this research we developed a different approach, noting that a covariance matrix S can be decomposed into a vector of standard deviations, V, and a correlation matrix, R using: S = diag(V)Rdiag(V) where diag(V) was a ma- trix with diagonal elements V and zeros elsewhere. This decomposition permitted the independent sampling of V and R. We simulated the standard deviations, V, from an inverse gamma distribution: () () 2 1, 2 ,,~ , 22 mklk k kl s n n pV XZIG αβ − ++ , where nk was the number of individual habitats assigned to cluster k by Z(m-1) (i.e., an estimate of the Cx. quinquefasciatus habitat sample size), s2 k,l was the sample variance of  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 32 element l in cluster k as assigned by Z(m-1), and α and β were constants, both set to the non-informative prior value of 1. Quasi-likelihood techniques in a logistic re- gression equation and Bayesian prior distributions can enumerate intra-cluster correlations in urban vector mosquito aquatic habitats [4]. We simulated the elements of the correlation matrices, R, from a hyperbolic-tangent transformed distribution using : () () () () () 1 ,, ,, 1 tanh|,~ˆ tanh , m ki jki j k pRXRZN n − where ,, ˆki j R was the current estimate of the correlation between the ith and jth elements (i ≠ j) in a cluster k given Z(m-1), and nk was the same. After sampling values for both V and R, we reassembled the covariance matrix, Sk, for each Cx. quinquefasciatus habitat cluster, thus, com- pleting Step 1. Step 2 required drawing values for the elemental means, P. The field and remote-sampled data X had an approximate multivariate normal distribution as such Step 2 was performed using the vector of mean concen- trations for cluster k. The multivariate normal distribu- tion using the Cx. quinquefasciatus habitat parameters was given by the sample means calculated from X, gen- erated from the cluster assignments, Z(m-1)) and the co- variance Sk from Step 1. If cluster k was empty at m-1 (that was, no individual sampled habitat parameters were assigned to k by Z(m-1)), then the grand mean and covari- ance matrix of X were used as the sample mean and co- variance matrix of k. Step 3 involved drawing new cluster assignments us- ing each individual sampled Cx. quinquefasciatus habitat in the Trinidad study site. To do so, it was necessary to calculate Pr(zi = k) for every i, k combination (zi was the ith element of Z). Again we assumed multivariate normal distributions where Z(m) was simulated from: ()() () () () () () ()() () , 1 Pr| ,,, |,, |,, kk immm mm ii kkk Kmm ii k K zkXPS fX PSzk f XPSz k φ φ φ ′′ ′ − = = =′ = and where the () f⋅ terms on the right-hand side were multivariate normal likelihoods. Thus, the likelihood that the ith element of X was present in the Cx. Quinquefas- ciatus habitat population k, sampled in the study site normalized by the sum of likelihoods was quantified for all potential sources influencing presence of immatures. Finally, the sample Φ(m) from f(Z|X, P(m), S(m), Z(m)) was used to generate asymptotically efficient estimates from the spatiotemporal-sampled data. 2.6. Eigenvector Mapping We conducted a residual analyses using a misspecification perspective for the error estimation models generated in SAS/GIS® assuming that the basic regression model * yX β ε =+ had autocorrelated disturbances * ε , which was decomposed into a white-noise component, ε , and a set of unspecified and/or misspecified models that had the structure * yXBE ε γ ε = =++ . In this research, the misspecification term was E γ . Quantification of topographic patterns generated from the distribution of the georeferenced Cx. quinquefasciatus habitat covari- ates was required to describe independent key dimen- sions and underlying spatial processes for defining a pat- tern in the misspecification term. This was accomplished by expanding the matrix term: () 1 0 kk k I VV ρρ −∞ = −= , as an infinite power series, which was feasible under the assumption that the under- lying spatial process in the data was stationary; note that in this research 00 V ρ gave the identity matrix I . The simultaneous autoregressive error model was then re- written asyVyX VX ρβρβ ε −=− + . Substituting these transformation gave: () () 1 yIV XVX ρβρβ ε − =−− + () 0 kk k yVXVX ρβρβ ε ∞ = =−+ () 11 000 kkkkkk kk k yVXVX V ρβρβ ρ ε ∞∞ ∞ ++ == = =−+ () 110 0 1 kk kkkk kkk kk k misspecification term yX VXVXV yX V βρ βρβρ ε βρεε ∞∞∞ == = = ∞ = ⋅ =+ −+ =+ + The misspecification term () 1, , kk Vk ρε =∞ re- mained uncorrelated with the exogenous variable, X , as the standard OLS assumption of the disturbances, ε , were uncorrelated with the predictor variables generated from the model. The spatial lag model was expressed as: () IVyX ρβ ε −=+. Substituting the transformation gave: () () 0 1 kk k kk k misspecification term yVX yX VX ρβε βρ β εε ∞ = ∞ = ⋅ =+ =+ ++ The misspecification term ()( ) 1, , kk VX k ρβε +=∞ included the exogenous  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 33 variables X . Consequently, the exogenous variables were correlated with the misspecification term. Under this condition, standard OLS results for the basic regres- sion model* yX β ε =+, generated from the spatiotem- poral-sampled covariates provided biased estimates ˆ β of the underlying regression parameters β . The correlation, or lack thereof, between the exoge- nous variables and the misspecification terms of both habitat models were used to design spatial proxy vari- ables, so that the properties of either model could be sat- isfied. We considered two different projection matrices. Different sets of eigenvectors were established using a spatial regression model. This model feature enabled us to also use the eigenvector spatial filtering approach for predictions of the endogenous variable y. We selected an evaluation criterion using the stan- dardized Moran’s statistic. The standardized Moran’s coefficient (MC) for regression residuals * ˆ ε was de- fined as () () () 2 iiji j j iij ji i wXX XX N IwXX −− =− , where N was the number of sampled habitats and their respective parameter estimates indexed by i and j respectively; X was the spatiotemporal-sampled egg-raft count data; X was the mean of X; and wij was a matrix of spatial weights. The expected value of Moran’s I under hy- pothesis of no spatial autocorrelation was: () 1 1 EI N − =−. Its variance equaled: () ()( )() () 435 2 12 3 iij j NSS S Var I NN Nw − = −− − where () () () () () () () () 2 1 2 2 4 1 32 2 1 2 2 12 4 2 51 1 1 2 1 33 3 1 6 21 ij ji ij ij ji ij j i i i i iij j iij j Sww ww S Nxx S Nxx NNSNS w S w SS NS − − =+ + = − = − −+− + = =− + In this research, negative (positive) values indicated negative (positive) spatial autocorrelation. Values ranged from -1 (indicating perfect dispersion) to +1 (perfect correlation). A zero values indicated a random spatial pattern. For statistical hypothesis testing predictive WNV mosquito habitat modeling, MC values can be trans- formed to z-scores in which values greater than 1.96 or smaller than −1.96 indicate spatial autocorrelation that is significant at the 5% level [8,29]. Geary’s Contiguity Ratio (GC) another measure of spatial autocorrelation was also generated. GC is in- versely related to the Moran’s statistic but it is not iden- tical. Moran's indices are measure of global spatial auto- correlation, while Geary’s C is more sensitive to local spatial autocorrelation in vector mosquito data analyses [16]. In this research, the sampled Cx. quinquefasciatus habitat data was quantified by GC which was defined as () () () 2 2 1 2 iijij j i i NwXX C WXX −− =− where N was the number of the sampled habitats and explanatory parame- ters indexed by i and j respectively; X was the egg-raft count data; X was the mean of X; wij was a matrix of spatial weights; and W was the sum of all wij. The value of GC lies between 0 and 2. 1 means no spatial autocor- relation while 0 refers to a chaotic random distribution. Smaller (larger) than 1 means positive (negative) spatial autocorrelation [8]. In this research, distance between sampled habitats was defined in terms of an n-by-n geographic weights matrix, C, whose cij values were; 1 if the sampled Cx. quinquefasciatus egg-raft count locations i and j in the Trinidad study site were deemed nearby, and 0 other- wise. Adjusting this matrix by dividing each row entry by its row sum gave C1, where 1 was an n-by-1 vector of one’s which converted this matrix to matrix W. The resulting autoregressive model specification with no sampled covariates present (i.e., the pure spatial auto- regression specification) took on the following form: () μ1-ρρ =++Y1WYε (2.1), where μ was the scalar conditional mean of Y and ε was an n-by-1 error vec- tor residuals of the sampled data which in this research was represented as statistically independent and identi- cally distributed (iid) normally random variates. The spatial covariance matrix for equation (2.1), using the sampled covariates, was ()()()() 12 E-μ'-μ- ρ'- ρσ − == Y1 Y1ΣIWIW , where E (⋅) indicated the calculus of expectations, I was the n-by-n identity matrix denoting the matrix transpose operation, and 2 σ was the error variance. Two different autoregressive parameters appeared in the spatial covariance matrix. When autocorrelation is present in residual data, a more explicit representation of both effects leads to a more accurate interpretation of empirical results [38]. Consequently, we used an SAR model specification  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 34 ()() 12 diag diag - '- σ, − = ΣIρWIρW (2.2) where the diagonal matrix of autoregressive parameters, diag ρ contained two sampled habitat parameters: ρ+ displaying positive spatial dependency, and - ρ for those sampled habitat pairs displaying negative spatial dependency based on the aggregation of the Cx. Quin- quefasciatus egg-raft count data. For example, by letting 2 σ = 1 and employing a 2-by-2 regular square tessella- tion, 2 11 22 11 22 11 22 11 22 00ρ000 1000 000ρ00 0100 0000ρ0 0010 00000ρ 0001 + + − − =− Σ for the vector 1 2 3 4 y y y y enabled positing a positive rela- tionship between the sampled egg-raft count data and the explanatory covariates, y1 and y2, a negative relationship between covariates, y3 and y4, and, no relationship be- tween covariates y1 and y3 and between y2 and y4. This covariance yielded: () diag diag diag diag μ(ρ ρ) ρ ρ ++ −− ++ −− =− − ++ + YI II1 IIWYε where I+ was a binary 0-1 indicator variable which de- noted those parameter estimates which displayed positive spatial dependency, and I- was a binary 0-1 indicator variable denoting those sampled covariates displaying negative spatial dependency, where I+ + I- = 1. Express- ing the preceding 2-by-2 example in terms of an equation yielded: 1 2 3 4 y1000 1000 y0100 0100 μρ y0010 0000 y00010000 000011000 000010100 ρρ 001010000 000110000 0000 0000 ρ0010 0001 + −+ − =− −+ + 11 11 22 11 22 22 11 33 22 11 44 22 00yε 00y ε 00y ε 00yε + if either ρ0 += then I+ was = 0 and I- = I or if ρ0 −= then I- = 0 and I+ = I. In this research, this spatial structuring was achieved by constructing a linear combination of a subset of the eigenvectors of a modified geographic weights matrix, using (I - 11 '/n)C(I - 11 '/n) that appeared in the nu- merator of the MC. Spatial filtering furnishes an ap- proach that incorporates spatial structuring into an inter- cept term that accounts for latent autocorrelation [38]. A subset of eigenvectors was selected with a stepwise re- gression procedure. Because (I - 11 '/n)C(I - 11 '/n) = 'EΛE, where E was an n-by-n matrix of eigenvectors and Λ was an n-by-n diagonal matrix of the corre- sponding eigenvalues, the resulting egg-raft count model specification was given by: k μ =+ +Y1Eβε, where μ was the scalar mean of Y, Ek was an n-by-k matrix containing the subset of k < < n eigenvectors selected with a stepwise regression technique and β was a k-by- 1 vector of regression coefficients. A number of the eigenvectors were extracted from the matrix (I - 11'/n)C(I - 11 '/n) which were affiliated with geographic patterns of the sampled Cx. quinquefas- ciatus habitat covariates portraying a negligible degree of spatial autocorrelation. Consequently, only k of the n eigenvectors were of interest for generating a candidate set for a stepwise regression procedure. One way of con- structing a candidate set is to include an eigenvector in the set only when the vector’s corresponding MC repre- sents a minimum degree of spatial autocorrelation [38]. In this research, we used a spatial statistical threshold of |MC/MCmax| > 0.25 where MCmax was the maximum MC for a given surface partitioning, as then each candidate eigenvector represented a level of autocorrelation tend- ing to account for the redundant information in the data- set. As redundant information spatial autocorrelation represents pseudo-replicated data linking it to missing values estimation and interpolation, as well as to notions of effective sample size and degrees of freedom [38]. 2.6.1. Spatial Analyses In this research, spatial linear prediction of the prolific Cx. quinquefasciatus habitat and its spatiotemporal sampled covariates was performed using an Ordinary kriged-based interpolator in ArcGIS 9.3® using Geostatistical Analyst. The Semivariogram Properties dialog box has several models to choose from. We set the Kriging method to Ordinary. A default value for lag size was initially set to the default output cell size. For Major range, Partial sill, and Nugget, a default value was calculated internally. The length of the longer axis to reach the sill is called the major range, and the length of the shorter axis to reach the sill is called the minor range; the angle of rotation of the line formed the major range [28]. The optional output variance of prediction raster contained the kriging variance  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 35 at each output raster cell. Assuming the kriging errors are normally distributed in a predictive vector mosquito habitat distribution model there is a 95.5 percent prob- ability that the actual z-value at the cell is the predicted raster value, plus or minus two times the square root of the value in the prediction raster [16]. Low values within the output variance of prediction raster indicated a high degree of confidence in the predicted estimates. High values in interpolation based algorithms may indicate a need for more data points (26). Geostatistical techniques were used to interpolate the value Z(x0) of a prolific Cx. quinquefacistus habitat site, Z(x) at an unobserved sampling site, x0, where zi was Z(xi), i = 1 n based on the dimensions of the sam- pled habitat locations [i.e., (x1, , xn)]. The Ordinary interpolator also computed the best linear unbiased esti- mator, Ž(xo) of Z(x0), for the Cx. quinquefacistus data based on a stochastic model quantified by the variogram γ(x, y), by expectation μ(x) = E[Z(x)], and by the covari- ance function c(x,y) of the random field. In this research, the kriging estimator was given by a linear combination of the sampled field and remote parameters using () ()() 1 ˆn oioi i Z xwxZx = =; whereby, zi = Z(xi) was the weights wi (xo), and i = 1 n was the variance used to minimize any biased condition. The dependent variable was the spatiotemporal-sampled Cx. quinquefasciatus egg-raft count data, which were transformed to fulfill the diagnostic normality test prior to performing the kriging. The kriging weights of the algorithm were used to fulfill the unbiasedness condition in the spatial interpolation of the ecological-dependent variables using: 1 n i i λ == which was given by the equation system: () () () () () () * 11 1111 * 1 , ,,1 ,,1 , 1110 1 n nnnn n x x xx xx xx xx x x γ λγγ λγγ γ − = The Ordinary kriging was given by: () () () () () () () () ** ** 1 11 11 1 ** 1 ˆ var(() ()) ,, ,,1 ,,1 ,, 110 11 n nnn nn Zx Zx x xxx xx xx xxxx x xxx γγ γγ γγ γγ − −= The semivariogram generated described the depend- ence, between the Cx. quinquefasciatus parameters as a function of the distance between the sample sites. This allowed for abundance estimations at any point in the Trinidad study site. The value of prevalence, Z, at the coordinate (x0, y 0) was estimated from the n nearest habitat sampling values using Zobs(x1, y1), Zobs(x2, y 2). Zobs(xn, yn) and the linear formula which generated: () () 00 1 ˆ, , n i obsii i Z xyaZ xy = = The ai was found by the Lagrange multiplier λ and solving the system using: () () ,,0 1, 1 n iij i iahh jn γλγ =+= = under the con- straint. 1 n i a= , where hi,j denoted the distance between any two sampled Cx. quinquefasciatus habitat locations in the Trinidad study site at (xi, yi) and (xj, yj), and hj,0 was the distance between the two habitats (x0, y0). In the field of calculus of variations in mathematics, the method of Lagrange multipliers can be used to solve certain infi- nite-dimensional constrained optimization problems [26]. In this research the semivariance was defined as γ (h). The magnitude of the semivariance was dependent on the distance between sampling sites. Semivariance of the deviance residuals of the Cx. quinquefasciatus data was calculated and a variogram was constructed to determine if there was evidence of residual spatial correlation in the data. The plot of the semivariances as a function of dis- tance from a point is referred to as a semivariogram [26]. In this research, parameters of a fitted mathematical function (i.e., the semiovariogram model) included gen- erating a range, a nugget and a sill. Since our data was spatiotemporal, the square of the difference between ex- pected values at habitat points had to be added: 2γ(x,y) = C(x,x) + C(y,y) - 2C(x,y) - (E(Z(x)) - E(Z(y))2. If the co- variance function of a stationary process exists, it is re- lated to variogram by 2γ(x,y) = C(x,x) + C(y,y) - 2C(x,y) [26]. We checked for spatial dependence also in the sampled Cx. quinquefasciatus habitat data. We then de- termined the best fit variogram equation based on the number of lags (i.e., bins) and the lag distance (the mini- mum value of h) used in the calculation of experimental variogram equation. Lag size is the width (distance) of the bins into which these vectors are grouped (www. esri.com). In this research the lag size was dependent upon the minimum and maximum distances between sam- pled Cx. quinquefasciatus habitats in the study site. In an effort to uncover our variogram’s structure, similar lags were grouped together (i.e., pairs of habitat that points aligned in roughly the same direction and roughly the same distance from each other) into the bins. The semivariogram nugget coefficient allowed an in- terpretation of the required scale for defining spatial variability of the Cx. quinquefasciatus habitat data for providing minimized unbiased prediction error for re- siduals on the logit scale. Standard error in the models was calculated. Variations of kriging can produce noise- -free predictions (www.esri.com). The kriging interpola- tor produced a map that was smooth and free of “jumps” at the sampled Cx. quinquefasciatus habitat locations. The algorithms incorporated in Geostatistical Analyst  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 36 provided exact filtered value predictions at the sampled habitat locations. This prevented discontinuities in the habitat predictions and their associated standard errors. A simple quantitative measure of the interpolation was de- termined by using a root mean square error (RMSE) for the models. The kriging procedure returned the observed values at the sampled Cx. quinquefasciatus habitat locations and their interpolated values using the intensity and shape of the variograms. Using a neighborhood and/or distance search radius provided the standard errors of the interpo- lated values. Prediction errors have often been used to optimize sampling design by identifying areas where sampling effort should be increased or decreased [26]. In this research, prediction errors were generated as a func- tion of the variogram models. A simple quantitative measure of the uncertainty in the interpolation model was determined by generating the RMSE values for the models. Because we were working in two-dimensional space, the semivariogram and covariance functions changed not only with distance but with direction (i.e. , anisotropy). We quantified sampled habitats points and the vector that separated them. This vector had a distance on the x-coordinate as well as the y-coordinate. Alternatively, the vector had a distance and an angle in polar coordinates. Anisotropy was then described for the semivariogram. The isotropic model was the same in all directions; whereas, the anisotropic model reached the sill more rapidly in some directions than others. In Geostatistical Analyst, an outline of the range was given over the empirical semivariogram surface. Having created a covariance function with which to form the kriging weights matrix, the next task was to tessellate the region into patches. Voronoi maps were used to explore the distribution of the sampled Cx. quinquefasciatus habitat data for global and local outliers and to quantify the covariation among the spatiotemporal-sampled data. Voronoi diagram is a special kind of decomposition of a metric space determined by distances to a specified discrete set of objects in the space [28]. Voronoi polygons were also used for interpolating neighborhoods and patches. A Voronoi diagram was constructed with the sample points as the centers of the polygons using the Weighted Voronoi Diagram Extension in ArcGIS 9.3®. Using the Generate tab, we generated a weighted Voronoi diagram from the geo-sampled point features. The Graphical User Interface (GUI) has two tabs: Generate and Update (www.esri.com).The Voronoi Diagram was generated whereby a sampled Cx. quinquefasciatus habitat was associated with pi (spatiotemporal-sampled covariate); whereby, P = {p1, , pn}, where 2 ≤ n ≤ ∞ and xi ≠ xj for i ≠ j, i, j ∈ In. The region was given by V(pi) = {x: || x - xi || ≤ || x - xj || for j ≠ i, i ∈In} was the ordinary Vo- ronoi polygon associated with pi or the Voronoi polygon of pi and the set given by V = {V(pi),.., V(pn)} was the planar ordinary Voronoi diagram generated by P or sim- ply Voronoi diagram of P. We also defined a planar or- dinary Voronoi diagram with half planes as follows where we let P = {pi, , pn} ⊆ R2, where 2 ≤ n ≤ ∞ and xi ≠ xj for i ≠ j, i, j ∈In. We called the region () () () ln , iij ji Vp Hpp=∩ which was the ordinary Vo- ronoi polygon associated with pi and set V(P) = {V(p1),, V(pn)} the planar ordinary Voronoi diagram generated by P. A raster image showing normal Euclidean distance and adjusted Euclidean distance was generated, as well as a Voronoi polygon shapefile. The Cx. quinquefasciatus ha- bitat point attributes were then transferred to Voronoi polygons automatically by appending the spatial attrib- utes of one layer to another. Once a distance raster is generated, the user can use new points to update the dis- tance raster and create updated Voronoi polygons [28]. The Voronoi tessellation produced a set of polygons Vi with area f i (i=1,..., n). The (unknown) global mean zD was estimated by a weighted mean of the spatiotempo- ral-sampled Cx. quinquefasciatus habitat values using: 11 n gii ii n i mz f f == = . The extension error of each po- lygon was calculated by using a discrete version of ()()() 22,, , EVVVv σ γ ν γγ ν =−−. If the elementary error terms are uncorrelated, errors can be combined to an estimate of the global error: 22 21 1 i n E i i gn i i f f σ σ = = = where the summation is over all polygons Vi [28]. By conven- tion we assumed a normal probability distribution for the global error D g zm− and achieved a 90% confidence limits for D Z , 1.65* 1.65* g gD gg mzm σσ −≤≤+ . The global estimation error decreased with an increasing number of samples although some local deviation from this tendency did occur due to large polygons. As the number of samples increased the global estimation error converged to zero. 3. Results The ANOVA used in data analyses tested the null hy- pothesis that the sampled parameters of the immature Cx. quinquefasciatus population in the Trinidad study site means were equal (i.e., H0: µ1 = µ2 = = µa), by com- paring two estimates of variance. If the null hypothesis is false, then Mean Square Between (MSB) estimates gen- erated from a predictive vector mosquito habitat dis- tribution model is something larger than σ² [25]. In this research the MSE was an estimate of variance for deter- mining whether or not the null hypothesis was true,  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 37 while the second estimate MSB was based on the vari- ance of the sample means. We tested the null hypothesis as follows: if the null hypothesis was true, then MSE and MSB would be about the same since they are both esti- mates of the same quantity (i.e., σ2); however, if the null hypothesis was false then MSB would have been ex- pected to be larger than MSE, since MSB is estimating a quantity larger than σ². The ANOVA model encom- passed all possible sources of variation in sampled Cx. quinquefasciatus habitat data which allowed further test- ing of our research hypotheses. In this research the sig- nificance test of spatiotemporal-sampled data involved the ratio of MSB to MSE: F = MSB/MSE. The F-statistic was used to calculate the p-value to determine deviations from normality. If the kurtosis is greater than 0, then the F tends to be too small and we cannot reject the null hy- pothesis even though it is incorrect; the opposite is the case when the kurtosis is less than 3 [25]. The skewness of the distribution in the Cx. quinquefasciatus data did not have a sizable effect on the F statistic. If the n per cell is fairly large, then deviations from normality do not matter much at all because of the central limit theorem, which implicitly states the sampling distribution of the mean approximates the normal distribution, regardless of the distribution of the variable in the population [25]. The P value was the estimated probability of rejecting the null hypothesis (H0). Traditionally for WNV mos- quito models, if p < .05 the null hypothesis is rejected [18]. If the null hypothesis is true in the estimation of a model then the F ratio should be approximately one, since MSB and MSE should be about the same [25]. In order to conduct a significance test using the prolific spatiotemporal-sampled Cx. quinquefasciatus habitats predictor variables, it was thus necessary to know the sampling distribution of F. Additionally, from the sampling distribution, gener- ated from the sampled t parameters, the probability of obtaining an F as large or larger than the one was also calculated. When there are only two means to compare, the t-test and the F-test are equivalent; the relation be- tween ANOVA and t is given by F = t2 [25]. If the prob- ability values were lower than the significance level, then the null hypothesis was rejected. Significant differences by ANOVA were noted for mean numbers of Cx. Quin- quefasciatus captured throughout the sampling frame (F = 42.2, degrees of freedom [df] = 1). The Poisson regression modeled the spatiotemporal- sampled Cx. quinquefasciatus habitat count data and contingency tables. Poisson regression assumed the response variable Y had a Poisson distribution and assumed the logarithm of its expected value can be modeled by a linear combination of the spatiotemporal- sampled Cx. quinquesciatus parameters. A Poisson regression model is sometimes known as a log-linear model, especially when used to model contingency tables [25]. Our model with a single independent variable x, took on the form: () () log E|Yx bx α =+ . If Yi are independent observations with corresponding values xi of the sampled predictor variable, then a and b can be estimated by maximum likelihood if the number of distinct x values is at least 2 [25]. In this research the maximum-likelihood estimates lacked a closed-form expression and had to be quantified using numerical methods. The probability surface for maximum-likelihood Poisson regression was convex, making Newton-Raphson or other gradient-based methods appropriate estimation techniques for data analyses of the spatiotemporal-sampled Cx. quinquefas- ciatus habitat data. Linearized iteration scheme converges quickly and accurately using inversion equations solved through the Newton-Raphson method as each iteration, requires solving the finite difference equations and the linear adjoint equations only once, respectively [25]. OrthoEngine, PCI Geomatics’ Geocorrection and Ex- traction tools generated a DEM of the Trinidad study site. OrthoEngine included a number of automation tools to improve the efficiency of georeferencing the sampled Cx. quinquefasciatus data including Automatic GCP collec- tion from Chip Databases, and Automatic TP measure- ment. We stitched the image files. Stitching operation merged all the raster tiles and recalculated the image RPC from individual RPCs. Stitching automatically re- vealed the final RPCs as a binary segment in an output PIX file. Coefficients were reviewed using ‘Project Re- port Option’ in ‘Reports’ procedure step. We then se- lected geometric Model under the image information panel. The final step was the ‘Schedule Ortho Genera- tion.’ We proceeded to the ‘Orthogeneration’ processing step and selected the files to be processed. We selected an appropriate DEM file and set other processing options before generating the final orthorectified image. This allowed for the precise orthorectification of the WV-I imagery. The WV-1 over the Trinidad study site had look angles of approximately south at 30 degrees-off nadir and vertical. For simple corrections, polynomial, thin plate spline and rational functions were used. This allowed using image correlation from the stereo images. Once the images were geometrically corrected, they were mosaicked into a seamless image database, to form a conventional image map product and a GIS base layer for derivation of vector hypsographic and hydrographic covariates associated to the sampled Cx. quinquefascia- tus habitat data. The range of the elevation in the DEM had a minimum value of 0 m with a maximum value of 425 m. The spatial autocorrelation analysis results are re-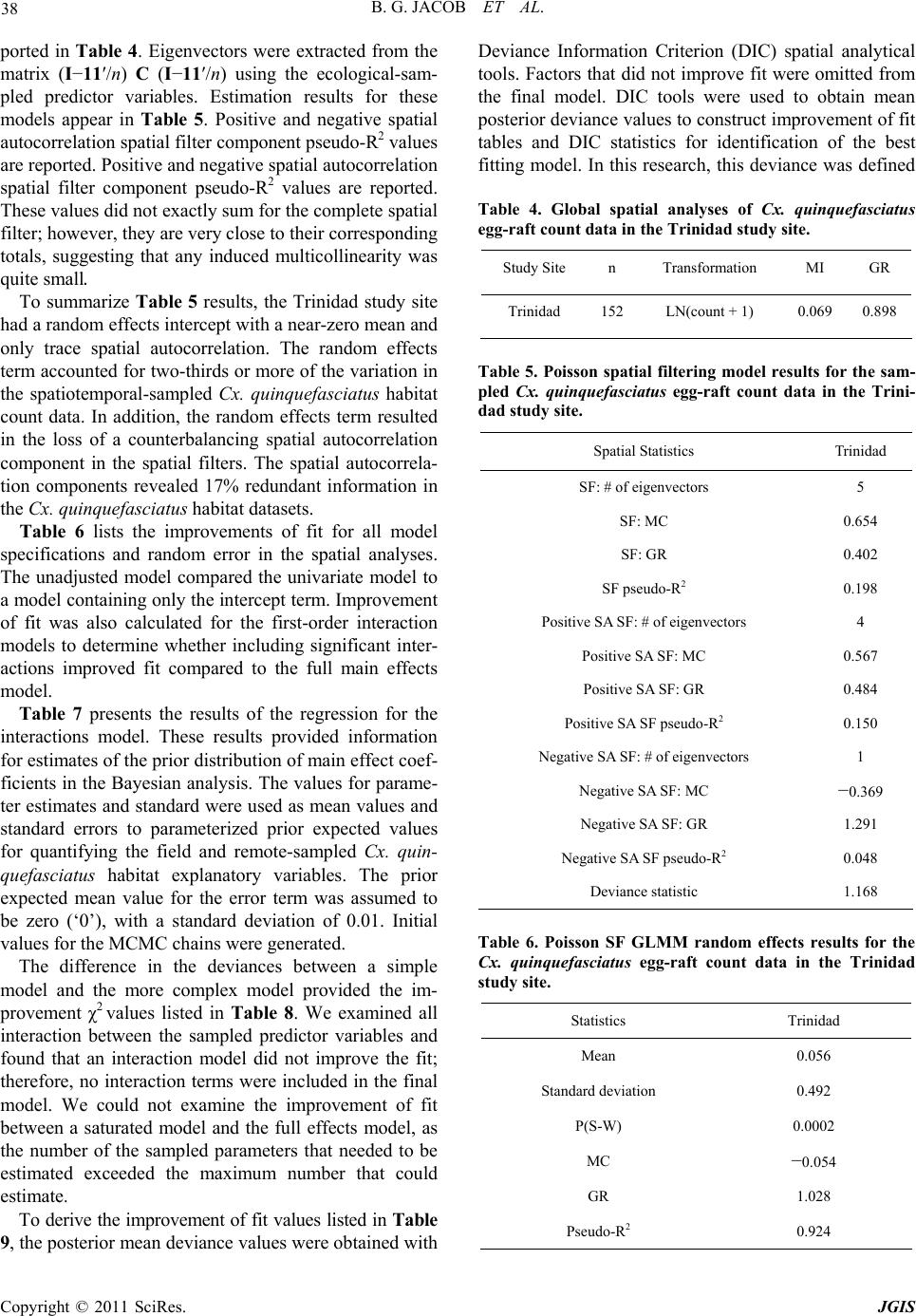  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 38 ported in Table 4. Eigenvectors were extracted from the matrix (I−11′/n) C (I−11′/n) using the ecological-sam- pled predictor variables. Estimation results for these models appear in Table 5. Positive and negative spatial autocorrelation spatial filter component pseudo-R2 values are reported. Positive and negative spatial autocorrelation spatial filter component pseudo-R2 values are reported. These values did not exactly sum for the complete spatial filter; however, they are very close to their corresponding totals, suggesting that any induced multicollinearity was quite small. To summarize Table 5 results, the Trinidad study site had a random effects intercept with a near-zero mean and only trace spatial autocorrelation. The random effects term accounted for two-thirds or more of the variation in the spatiotemporal-sampled Cx. quinquefasciatus habitat count data. In addition, the random effects term resulted in the loss of a counterbalancing spatial autocorrelation component in the spatial filters. The spatial autocorrela- tion components revealed 17% redundant information in the Cx. quinquefasciatus habitat datasets. Table 6 lists the improvements of fit for all model specifications and random error in the spatial analyses. The unadjusted model compared the univariate model to a model containing only the intercept term. Improvement of fit was also calculated for the first-order interaction models to determine whether including significant inter- actions improved fit compared to the full main effects model. Table 7 presents the results of the regression for the interactions model. These results provided information for estimates of the prior distribution of main effect coef- ficients in the Bayesian analysis. The values for parame- ter estimates and standard were used as mean values and standard errors to parameterized prior expected values for quantifying the field and remote-sampled Cx. quin- que fa sc i atu s habitat explanatory variables. The prior expected mean value for the error term was assumed to be zero (‘0’), with a standard deviation of 0.01. Initial values for the MCMC chains were generated. The difference in the deviances between a simple model and the more complex model provided the im- provement χ2 values listed in Table 8. We examined all interaction between the sampled predictor variables and found that an interaction model did not improve the fit; therefore, no interaction terms were included in the final model. We could not examine the improvement of fit between a saturated model and the full effects model, as the number of the sampled parameters that needed to be estimated exceeded the maximum number that could estimate. To derive the improvement of fit values listed in Table 9, the posterior mean deviance values were obtained with Deviance Information Criterion (DIC) spatial analytical tools. Factors that did not improve fit were omitted from the final model. DIC tools were used to obtain mean posterior deviance values to construct improvement of fit tables and DIC statistics for identification of the best fitting model. In this research, this deviance was defined Table 4. Global spatial analyses of Cx. quinquefasciatus egg-raft count data in the Trinidad study site. Study Site n Transformation MI GR Trinidad 152LN(count + 1) 0.069 0.898 Table 5. Poisson spatial filtering model results for the sam- pled Cx. quinquefasciatus egg-raft count data in the Trini- dad study site. Spatial Statistics Trinidad SF: # of eigenvectors 5 SF: MC 0.654 SF: GR 0.402 SF pseudo-R2 0.198 Positive SA SF: # of eigenvectors 4 Positive SA SF: MC 0.567 Positive SA SF: GR 0.484 Positive SA SF pseudo-R2 0.150 Negative SA SF: # of eigenvectors 1 Negative SA SF: MC −0.369 Negative SA SF: GR 1.291 Negative SA SF pseudo-R2 0.048 Deviance statistic 1.168 Table 6. Poisson SF GLMM random effects results for the Cx. quinquefasciatus egg-raft count data in the Trinidad study site. Statistics Trinidad Mean 0.056 Standard deviation 0.492 P(S-W) 0.0002 MC −0.054 GR 1.028 Pseudo-R2 0.924 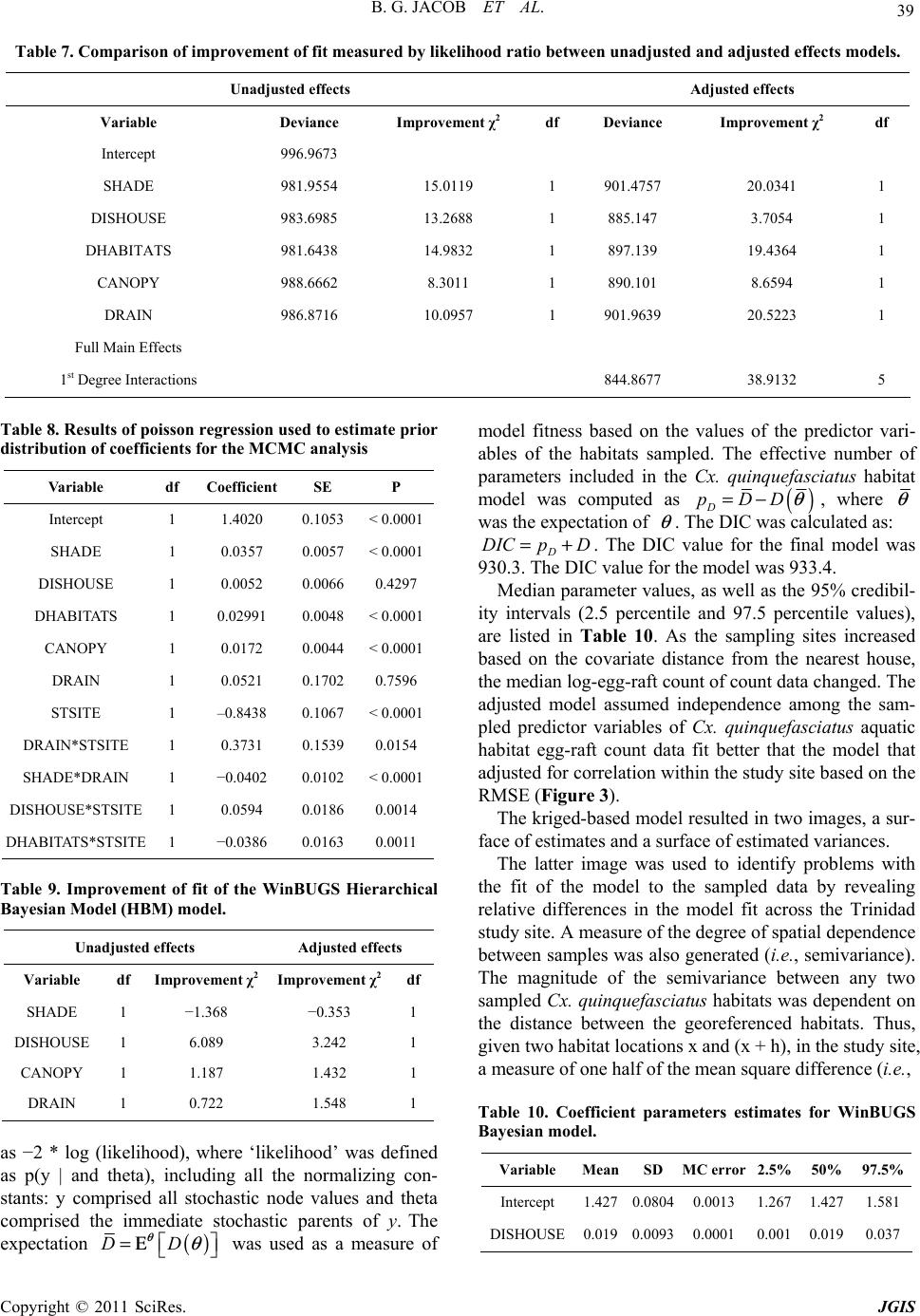 B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 39 Table 7. Comparison of improvement of fit measured by likelihood ratio between unadjusted and adjusted effects models. Unadjusted effects Adjusted effects Variable Deviance Improvement χ2 df Deviance Improvement χ2 df Intercept 996.9673 SHADE 981.9554 15.0119 1 901.4757 20.0341 1 DISHOUSE 983.6985 13.2688 1 885.147 3.7054 1 DHABITATS 981.6438 14.9832 1 897.139 19.4364 1 CANOPY 988.6662 8.3011 1 890.101 8.6594 1 DRAIN 986.8716 10.0957 1 901.9639 20.5223 1 Full Main Effects 1st Degree Interactions 844.8677 38.9132 5 Table 8. Results of poisson regression used to estimate prior distribution of coefficients for the MCMC analysis Variable df Coefficient SE P Intercept 1 1.4020 0.1053 < 0.0001 SHADE 1 0.0357 0.0057 < 0.0001 DISHOUSE 1 0.0052 0.0066 0.4297 DHABITATS 1 0.02991 0.0048 < 0.0001 CANOPY 1 0.0172 0.0044 < 0.0001 DRAIN 1 0.0521 0.1702 0.7596 STSITE 1 –0.8438 0.1067 < 0.0001 DRAIN*STSITE 1 0.3731 0.1539 0.0154 SHADE*DRAIN 1 −0.0402 0.0102 < 0.0001 DISHOUSE*STSITE 1 0.0594 0.0186 0.0014 DHABITATS*STSITE 1 −0.0386 0.0163 0.0011 Table 9. Improvement of fit of the WinBUGS Hierarchical Bayesian Model (HBM) model. Unadjusted effects Adjusted effects Variable df Improvement χ2 Improvement χ2df SHADE 1 −1.368 −0.353 1 DISHOUSE 1 6.089 3.242 1 CANOPY 1 1.187 1.432 1 DRAIN 1 0.722 1.548 1 as −2 * log (likelihood), where ‘likelihood’ was defined as p(y | and theta), including all the normalizing con- stants: y comprised all stochastic node values and theta comprised the immediate stochastic parents of y. The expectation () DD θ θ =Ε was used as a measure of model fitness based on the values of the predictor vari- ables of the habitats sampled. The effective number of parameters included in the Cx. quinquefasciatus habitat model was computed as () D pDD θ =− , where θ was the expectation of θ . The DIC was calculated as: D DICpD=+. The DIC value for the final model was 930.3. The DIC value for the model was 933.4. Median parameter values, as well as the 95% credibil- ity intervals (2.5 percentile and 97.5 percentile values), are listed in Table 10. As the sampling sites increased based on the covariate distance from the nearest house, the median log-egg-raft count of count data changed. The adjusted model assumed independence among the sam- pled predictor variables of Cx. quinquefasciatus aquatic habitat egg-raft count data fit better that the model that adjusted for correlation within the study site based on the RMSE (Figure 3). The kriged-based model resulted in two images, a sur- face of estimates and a surface of estimated variances. The latter image was used to identify problems with the fit of the model to the sampled data by revealing relative differences in the model fit across the Trinidad study site. A measure of the degree of spatial dependence between samples was also generated (i.e., semivariance). The magnitude of the semivariance between any two sampled Cx. quinquefasciatus habitats was dependent on the distance between the georeferenced habitats. Thus, given two habitat locations x and (x + h), in the study site, a measure of one half of the mean square difference (i.e., Table 10. Coefficient parameters estimates for WinBUGS Bayesian model. Variable MeanSD MC error 2.5% 50%97.5% Intercept 1.4270.08040.0013 1.267 1.427 1.581 DISHOUSE0.019 0.00930.0001 0.001 0.019 0.037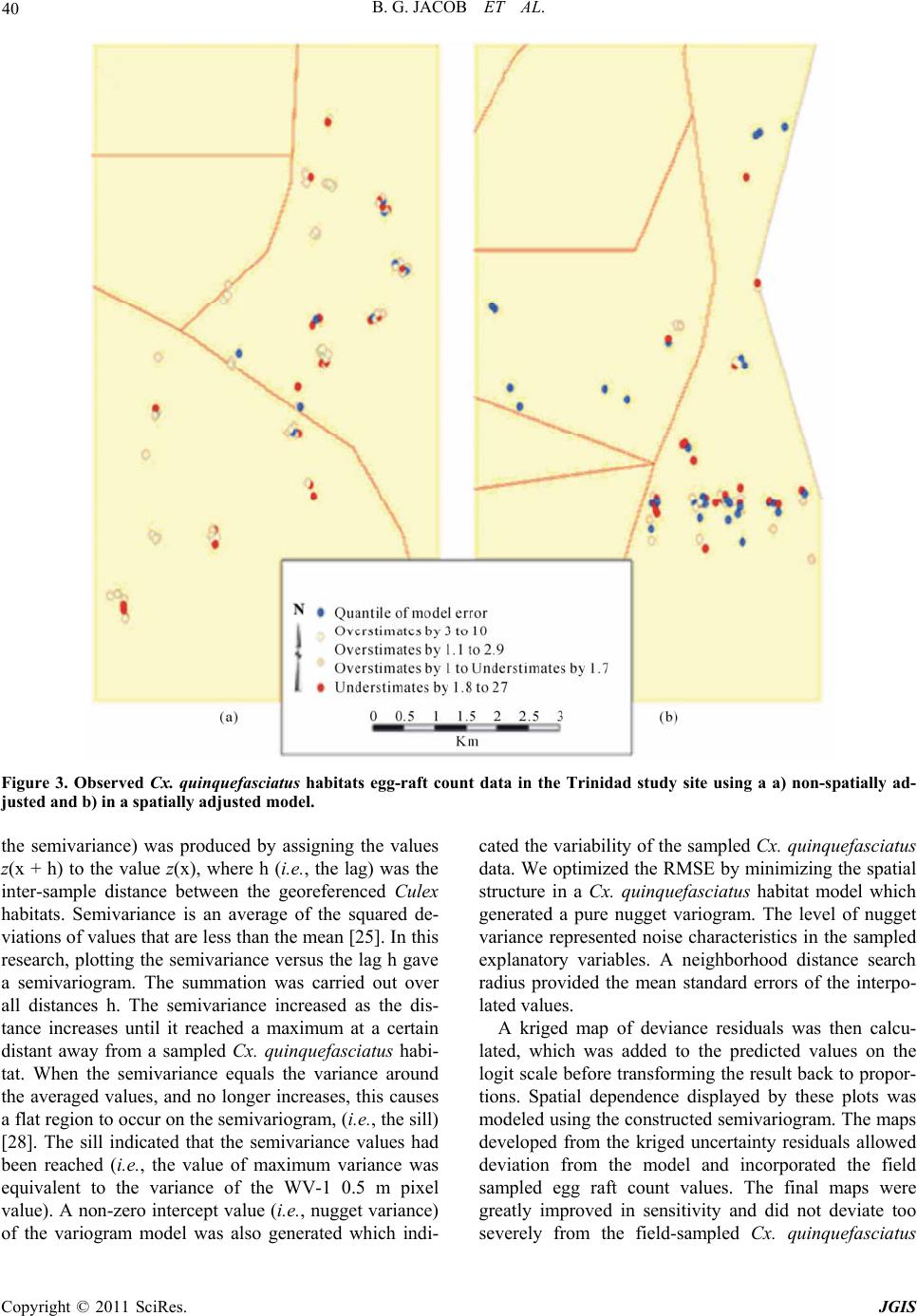  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 40 Figure 3. Observed Cx. quinquefasciatus habitats egg-raft count data in the Trinidad study site using a a) non-spatially ad- justed and b) in a spatially adjusted model. the semivariance) was produced by assigning the values z(x + h) to the value z(x), where h (i.e., the lag) was the inter-sample distance between the georeferenced Culex habitats. Semivariance is an average of the squared de- viations of values that are less than the mean [25]. In this research, plotting the semivariance versus the lag h gave a semivariogram. The summation was carried out over all distances h. The semivariance increased as the dis- tance increases until it reached a maximum at a certain distant away from a sampled Cx. quinquefasciatus habi- tat. When the semivariance equals the variance around the averaged values, and no longer increases, this causes a flat region to occur on the semivariogram, (i.e., the sill) [28]. The sill indicated that the semivariance values had been reached (i.e., the value of maximum variance was equivalent to the variance of the WV-1 0.5 m pixel value). A non-zero intercept value (i.e., nugget variance) of the variogram model was also generated which indi- cated the variability of the sampled Cx. quinquefasciatus data. We optimized the RMSE by minimizing the spatial structure in a Cx. quinquefasciatus habitat model which generated a pure nugget variogram. The level of nugget variance represented noise characteristics in the sampled explanatory variables. A neighborhood distance search radius provided the mean standard errors of the interpo- lated values. A kriged map of deviance residuals was then calcu- lated, which was added to the predicted values on the logit scale before transforming the result back to propor- tions. Spatial dependence displayed by these plots was modeled using the constructed semivariogram. The maps developed from the kriged uncertainty residuals allowed deviation from the model and incorporated the field sampled egg raft count values. The final maps were greatly improved in sensitivity and did not deviate too severely from the field-sampled Cx. quinquefasciatus  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 41 habitat data. The exponential model fitted to the semi- variogram at a partial sill 0 and nugget of 5.8 km, lag size 6.5 km number of lags 12 with a range of 73.6 km which formed the basis of the model to map predictions of residuals in an 73.6 km radius around each sampled Cx. quinquefasciatus habitat (Figure 4). Voronoi polygons were then modeled. An approxi- mation, based on the midpoints of the legs and center of the triangle were used to evaluate the area covered by each Voronoi polygon. A new sample point was calcu- lated using the importance metric (Gamma) of each pixel in the Voronoi polygons generated according to () , , , , , uv uv p uv uv uv C Γ =Γ where (u, v) were the coordinates of a WV-I pixel in the polygon, Gamma(u, v) was the importance metric of that pixel, and C(p) was the coor- dinates of the new polygon center (i.e. the sampled Cx. quinuqefasciatus habitat). To do this calculation on each polygon we used a scan conversion. We ran a scan con- version and examined each pixel in the frame buffer for each polygon. The Voronoi tessellation provided a spa- tial trend analyses of the error in the model which re- vealed that all coefficients were within normal statistic- cal limitations (Figure 5). 4. Discussion The ANOVA tested the significant differences between means in the Cx. quinquefasciatus habitat data. In re- peated measures ANOVA; however, containing factors such as spatiotemporal-sampled Cx. quinquefasciatus with more than two levels, additional special assump- tions may be required for quantification of sampled pre- dictor variables; specifically, the compound symmetry assumption and the assumption of sphericity. The com- pound symmetry assumption requires that the variances derived from the data (i.e., pooled within-group) and Figure 4. An ordinary kriged-based interpolation of egg-raft abundance of Cx. quinquefasciatus habitats in the trinidad study site.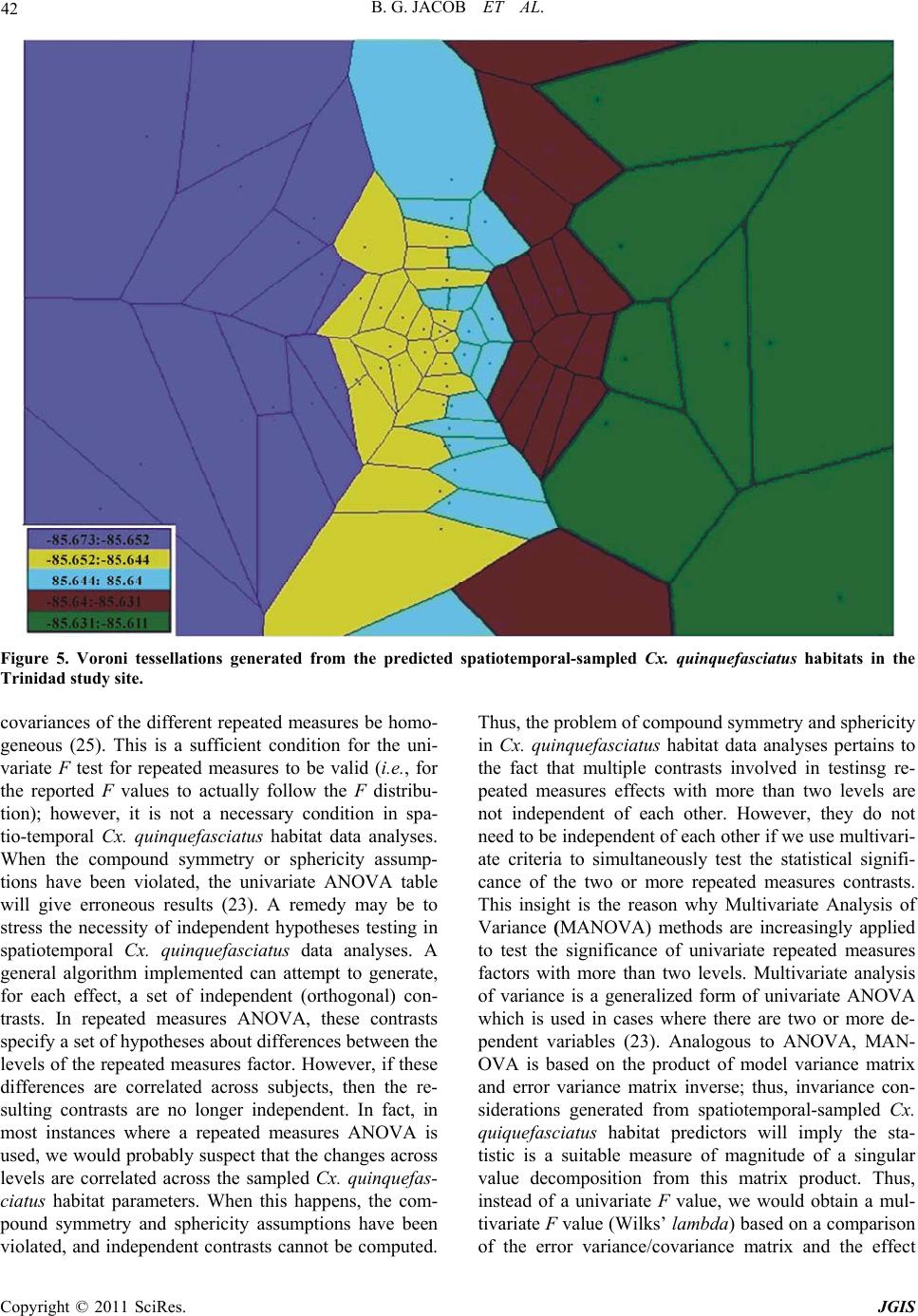  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 42 Figure 5. Voroni tessellations generated from the predicted spatiotemporal-sampled Cx. quinquefasciatus habitats in the Trinidad study site. covariances of the different repeated measures be homo- geneous (25). This is a sufficient condition for the uni- variate F test for repeated measures to be valid (i.e., for the reported F values to actually follow the F distribu- tion); however, it is not a necessary condition in spa- tio-temporal Cx. quinquefasciatus habitat data analyses. When the compound symmetry or sphericity assump- tions have been violated, the univariate ANOVA table will give erroneous results (23). A remedy may be to stress the necessity of independent hypotheses testing in spatiotemporal Cx. quinquefasciatus data analyses. A general algorithm implemented can attempt to generate, for each effect, a set of independent (orthogonal) con- trasts. In repeated measures ANOVA, these contrasts specify a set of hypotheses about differences between the levels of the repeated measures factor. However, if these differences are correlated across subjects, then the re- sulting contrasts are no longer independent. In fact, in most instances where a repeated measures ANOVA is used, we would probably suspect that the changes across levels are correlated across the sampled Cx. quinquefas- ciatus habitat parameters. When this happens, the com- pound symmetry and sphericity assumptions have been violated, and independent contrasts cannot be computed. Thus, the problem of compound symmetry and sphericity in Cx. quinquefasciatus habitat data analyses pertains to the fact that multiple contrasts involved in testinsg re- peated measures effects with more than two levels are not independent of each other. However, they do not need to be independent of each other if we use multivari- ate criteria to simultaneously test the statistical signifi- cance of the two or more repeated measures contrasts. This insight is the reason why Multivariate Analysis of Variance (MANOVA) methods are increasingly applied to test the significance of univariate repeated measures factors with more than two levels. Multivariate analysis of variance is a generalized form of univariate ANOVA which is used in cases where there are two or more de- pendent variables (23). Analogous to ANOVA, MAN- OVA is based on the product of model variance matrix and error variance matrix inverse; thus, invariance con- siderations generated from spatiotemporal-sampled Cx. quiquefasciatus habitat predictors will imply the sta- tistic is a suitable measure of magnitude of a singular value decomposition from this matrix product. Thus, instead of a univariate F value, we would obtain a mul- tivariate F value (Wilks’ lambda) based on a comparison of the error variance/covariance matrix and the effect  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 43 variance/covariance matrix. The problem of compound symmetry and sphericity pertains to the fact that multiple contrasts involved in testing repeated measures of Cx. quinquefasciatus habitat effects with more than two lev- els are not independent of each other but, fortunately a multivariate criteria can simultaneously test the statistical significance of the two or more repeated measures con- trasts. We thus endorse a MANOVA approach for spati- otemporal data analyses of Cx. quinquefasciatus habitat parameters as it simply bypasses the assumption of compound symmetry and sphericity altogether. The spatial hydrological model generated from the en- vironmental-sampled Cx. quinquefasciatus covariates revealed that elevation was an important variable in the DEM model. Anderson et al. (2004) captured signify- cantly more Culex quinquefasciatus in a variety of traps (bird-baited, MMX mosquito magnet, CO2-baited CDC trap) in the canopy (7.6 m) than at near ground level (1.5 m) [40]. In our research local differences among WV-1 DEM grid cells were also analyzed to calculate slope and other land surface parameters using relative and absolute accuracy estimates for determining the statistical signifi- cance of terrain-related parameters associated to the Cx. quinquefasciatus habitats. In this research the slope gra- dient was defined as the maximum rate of change in alti- tude (i.e., tan (Θ), aspect (ψ) as the compass direction of this maximum rate of change). Slope is defined by a plane tangent to a topographic surface, as modeled by the DEM at a point, whereby, it is classified as a vector; as such it has a quantity (gradient) and a direction (aspect). More analytically, the slope gradient at a sampled Cx. quinquefasciatus habitat in the Trinidad study site was the first derivative of elevation (Z) with respect of the slope (S), where S was in the aspect direction (ψ). We calculated the first derivative of a function at a sampled habitat as the angular coefficient of the tangent to the function of a sampled habitat. The resolution of our WV-1 DEM quantified the georeferenced explanatory covariates at each 0.5m pixel (i.e., absolute accuracy). At the same time the relative accuracy of the sampled data generated a geometrically correct reference frame for validating the covariate elevation associated with the sampled Cx. quinquefasciatus habitats. WV-1 DEMs can allow for hydrologic modeling, view-shed determination, slope/aspect analyses, and 3-d surface visualization of georeferenced terrain-related parameters associated to spatiotemporal-sampled Cx. quinquefasciatus habitat data. In the regression model, PROC MIXED quantified correlations among error measurements using random effects estimates and random regression coefficients. PROC MIXED used three options for the method of es- imation including: 1) Maximum Likelihood (ML); 2) Restricted or Residual maximum likelihood (REML), which was the default method; and, 3) Minimum Vari- ance Quadratic Unbiased Estimation (MIVQUE). In this research, the ML was the regular maximum likelihood method as the Cx. quinquefasciatus parameter estimates maximized the likelihood function, while the REML was a variant of maximum likelihood. The sample mean was the maximum likelihood estimator of the sampled habitat population mean, based on the egg-raft count data. The sample variance was also a close approximation to the maximum likelihood estimator of the immature sampled variance. REML estimated the variance parameters using a marginal likelihood which produced unbiased estimates. REML estimators generated estimates from the spatio- temporal-sampled parameters by maximizing only that part of the likelihood function that was invariant to the fixed effects part of the linear model. The variance of a quadratic function of the random variables in the linear model was then minimized to obtain locally best unbi- ased estimators of the variance components. The matri- ces of these quadratics were computed from relationship matrices and from prior estimates of variances used in the model equations. In the Bayesian matrix we were able to instantiate the nodes directly which was equivalent to supplying, a measurement for a radius based on the distribution of the georeferenced Cx. quinquefasciatus habitats. We were able to quantify an uncertainty estimate in our likelihood information by supplying a probability distribution over a set of radii generated from the sampled data. We used a backward propagation to compute the node probabilities. The error gradients were then propagated in any direc- tion throughout the Bayesian framework. This type of probability propagation allowed choosing between the sampled predictor variables using the computed prob- ability of each sampled covariate value based on one or more of the nodes. Our Bayesian approach allowed flexible model fitting and estimation and geomapping of all “high risk” Cx. quinquefasciatus habitats based on sampled egg-raft count data in the study site. The MCMC algorithm for the Cx. quinquefasciatus egg-raft count model produced a sequence of parameter vectors that represented random draws from the posterior distribution. The DIC com- prised two goodness-of-fit measures and the posterior distribution of the deviance, which was the number of effective sampled habitat covariates for measuring com- plexity in our Cx. quinquefasciatus model. Habitats with high egg-raft counts were compared with the results of a Monte Carlo simulation, which established the probabili- ties and occurrences of these habitats. Our results indi- cated that likelihood weights influenced the resulting posterior distributions of the field and remote-sampled  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 44 Cx. quinquefasciatus egg-raft count parameters, which, in turn influenced the spatial trends in the variance un- certainty estimates for model prediction. Distance from the nearest house was a significant predictor associated with prolific habitats in the Trinidad study site. Environmental factors commonly associated with Cx. quinquefasciatus habitats include the distance from the nearest human habitation. The location of WNV cases at the place of residence can be based on the fact that Culex mosquitoes are crepuscular feeders, and are most likely to come in contact with people while they are at home during the evening hours. Because egg-laying is a spatial process dependent upon the location of the focal habitat relative to sources of gravid mosquitoes, habitats closer to human inhabitations tend to receive more eggs and thus are more productive if conspecific competition is negligible [29]. Furthermore, housing age can also play a big role in the explanation of the focal areas. This is ex- pected since older houses have poorer drainage systems and open foundations which may provide larval and adult habitats. For example, houses built in the post World War II era in the study site may suffer from a general inattention. These houses may contain catch ba- sins, rich with organic material which are breeding grounds for Culex mosquitoes. Most of the Culex species require organic-rich water for larval development and are commonly found in high numbers in artificial containers, unattended pools, retention ponds, storm drains, catch basins, sewage systems and treatment plants [41,42]. Furthermore, combined sewage and street runoff dis- charges into natural waterways are considered the main sources of urban stream pollution, which are associated to WNV as Culex mosquitoes proliferate in eutrophic hydrological networks. The identification of social fac- tors that characterize these focal areas in the Trinidad study site may also provide insight into human risk and can help to target control and prevention strategies. Fu- ture analyses should explicitly test whether enzootic and bridge transmission are related to human population fac- tors includeing household income, population age, age of housing, housing density, and population density. Since it can be assumed from this analysis that the WNV is transmitted near a person’s residence in the Trinidad study site, extensive engineering to improve the runoff of water in residential areas should be considered. A framework for assessing autocorrelation in residual error from the spatial analyses was constructed in SAS/ GIS®. We generated a semiparametric filtering model using proxy variables constructed from the spatiotempo- ral-sampled georeferenced explanatory variables. In our Cx. quinquefasciatus habitat model, the misspecification terms was replaced by the proxy vari- ables for imple- menting a conditional covariance matrix employing the autoregressive specification. An unknown misspecifica- tion term can be approximated by a set of spatial proxy variables [38]. After modeling the mis- specification terms, the remaining residuals ˆ ε become white noise. This result implied that the estimated re- gression pa- rameters ˆ β were unbiased for the basic regression model, * yX β ε =+ , where * ε incorpo- rated the misspecification term and the white-noise dis- turbances. This allowed us to calibrate the autoregressive models. All of these eigenvectors exhibited weak posi- tively autocorrelated spatial patterns in the habitat pa- rameter estimates. Local weather patterns, mosquito con- trol programs and movement of hosts can have spatial ex- pressions, which cause immature Culex mosquetoes to aggregate in geographic space rendering positive auto- correlation in sampled predictor variables [8,26]. In this research, we used an Ordinary interpolation in ArcGIS® for identifying initial spatial structures in the sampled Cx. quinquefasciatus habitats and for quantify- ing the variance and autocorrelation in the data. The es- timation method determined the best linear unbiased es- timator. It was linear since the estimated values were weighted linear combinations of the spatiotemporal- sampled data and it was unbiased because the mean of the error was zero. For determining the optimal predict- tors, a semivariogram was constructed which expressed the spatial variation in the sampled data. In this research the variogram [i.e., 2γ(x,y)] was a function describing the degree of spatial dependence between the sampled Cx. quinquefasciatus habitats [i.e., Z(x)]. This was defined as the expected squared increment of the values between the sampled habitat locations x and y. Our semivariogram was nonnegative since it was the expectation of a square. The covariance function was related to variogram by 2γ(x,y) = C(x,x) + C(y,y) - 2C(x,y). In this research, the γ(x,y) = E(|Z(x) − Z(y)|2) = γ(y,x) was a symmetric func- tion, consequently, γs(h) = γs(- h) was an even function. A function is a semivariogram if, and only if, it is a con- ditionally negative definite function, i.e. for all weights 1,, N ww subject to 10 N i iw == and locations 1,, N x x, thus, it holds: () 11 ,0 NN iijj ij wxxw γ == ≤ [28]. The function corre- sponded to the fact that the variance [i.e., va r(X) of () 1 N ii j X wZ x = = generated from the sampled Cx. quinquefasciatus data was given by the negative of this double sum, which in this research was nonnegative. All spatial dependence was quantified in the model. If a sta- tionary random field has no spatial dependence (i.e. C(h) = 0 if 0h≠ the semivariogram is the constant var(Z(x)) everywhere except at the origin, where it is zero [28]. Theoretically, at zero separation distance (lag = 0), the semivariogram value is zero in a predictive vector mos-  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 45 quito habitat distribution model; however, at an infini- tesimally small sampled habitat distances, the semi- variogram often exhibits a nugget effect (i.e., measure- ment errors or spatial sources of variation at distances smaller than the sampling interval), which is some value greater than zero [16]. Additionally, since the random field was stationary and ergodic, the ()() () var lim s h hZx γ →∞ = corresponded to the variance generated by the Cx. quin- que fa sc i atu s habitat data. We defined a practical range and defined the distance at which 95% of the sill was reached for the asymptotic variogram model. The semi- variogram calculated estimates of the surface at the dif- ferent habitat locations. The increase in the semivario- gram values with increasing lags diminished with dis- tance and leveled off, at around 70 km (i.e., the range or active lag distance). This was the approximate distance at which spatial autocorrelation between the Cx. Quinque- fasciatus habitat data point pairs ceased or became much more variable. In the future sampling variograms should be constructed using spatiotemporal-sampled Cx. Quin- que fa sc i atu s habitat data. The sampling variogram, unlike the semivariogram, shows where a significant degree of spatial dependence in the sample space or sampling unit dissipates into randomness when the vari- ance terms of a temporally or in-situ ordered set are plotted against the variance of the set and the lower lim- its of its 99% and 95% confidence ranges [28]. Sam- pling variograms may quantify extraneous measurement variances before spatial dependence is verified in a pre- dictive Cx. quinquefasciatus habitat distribution model. In this research interpolation accuracy was also measured by the natural logarithm of the mean squared interpolation error and Voroni polygons which revealed all uncertainty effects while quantifying several covariate interaction terms in Euclidean space. The Voroni cells generated were a d-dimensional polytope (i.e., a convex hull of a finite set of georeferenced points). A set of hyperplanes were generated using the polygonal shapes around each sampled Cx. quinuqefasciatus habitat in the Trinidad study site. If S contains only two points, a and b, then the set of all points equidistant from a and b is a hyperplane-an affine subspace of codimension 1 [28]. The set of the polytopes covered the whole space and was the Voronoi tessellation corresponding to the georeferenced Cx. quinquefasciatus habitats. The convex Voronoi polygons were visualized as space-filling polygons constructed around the dataset of the sampled covariates (i.e.,Voronoi centers), such that each polygon contained all points closer to its Voronoi center than to the center of any other polygon. The relationship of the Voronoi centers to edges of polygons were used to test whether any convex tessellation existed in the polygons. This test amounted to finding Voronoi centers that best fit the given tessellation. In this research, Voronoi centers were found by solving two systems of linear equations. These equations represented 1) conditions on the slope of polygon edges relative to the slope of lines through Voronoi centers, and 2) conditions on the distance from edges to Voronoi centers. Least squares and constrained least-squares solutions were used to solve the two systems. In order to quantitatively characterize the error in the interpolation model, we examined normalized cell size distribution related to the homogenous and inhomogeneous fixation densities (i.e., Cx. quinquefasciatus habitat egg-raft count), which provided additional insight as to how the convex tessellations varied between the Voronoi polygons. A goodness-of-fit statistic was derived using the Voronoi tessellations which determined the error propa- gation in the multivariate model. The Voroni residuals reflected sampling errors and analytical errors based on the sampled Cx. quinquefasciatus habitat egg-raft count density. In the future new network nodes using Dirichlet tessellations in spatiotemporal-sampled Cx. quinque- fasciatus habitats should be analyzed. For example, Okabe and Satoh, (2008) have developed a software package, SANET, which can be installed as an ArcGIS add-in for “Spatial Analysis on a Network” for determining new network nodes [43]. The SANET software includes a network-based Voronoi tool, very similar to some of the network analysis tools provided in ArcGIS itself and in network-oriented packages such as TransCAD and Cube. The point set (i.e., off-network sites) can be linked to the nearest network edge, and then non-directed network Voronoi regions can be generated from georferenced Cx. quinquefasciatus habitat locations using shortest path distances. Separate inward and outward-directed Voronoi regions can then be computed and each habitat can be derived using unweighted calculations. For future research, the risk of WNV transmission in the Trinidad study site should also be investigated using wild bird roosting behavior and seasonal migratory pat- tern data. Quantifying WNV amplification at a local scale, with explicit attention to distances measurements from mosquito and hatch-year bird sites and human habitation areas may affect timing of infection in birds, mosquitoes, and humans. Seropositive resident and mi- gratory birds have been documented along the Atlantic and Mississippi flyways into the Caribbean [2]. While transient birds may suggest a means by which WNV has rapidly spread across North and Central America, resi- dent breeding birds may be more critical for zoonotic transmission in Trinidad [44-47]. In North America, high seroprevalence is associated with avian species that breed year-round and live close to humans (e.g. Ameri- can robin, House sparrow, Mourning dove, Northern  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 46 cardinal, Northern mockingbird), though other wild birds may be important [47-49]. In contrast to the urban avian species commonly associated as WNV hosts in North America, avian WNV hosts that occur in Trinidad are often non-urban, wild bird species [2]. Caribbean birds that have tested seropositive for WNV and are resident to Trinidad or breed seasonally in Trinidad include: Ba- nanaquits (Coereba flaveola), Palm-tanagers (Phaenico- philus palmarum), and Grossquit species [2]. Since the significance of avian species in WNV transmission may relate to their host competency, natural exposure to mosquito vectors and spatiotemporal availability, an avian GIS-based model using spatial filter eigenvectors and Bayesian estimates might be able to generate glob- ally asymptotically stable parameters while quantifying the amplification cycle prior to bridge vector involve- ment. Bridge vectors are mosquito species that readily feed on humans and mammals [2]. In conclusion, correlation estimates generated from the WV-1 DEM revealed that the terrain covariate elevation was significantly associated with the Cx. quinquefasciatus habitats sampling sites. The spatial hydrological model allowed for hydrologic modeling, view-shed deter- mination, slope/aspect analyses, and 3-d surface visua- lization of the georeferenced parameters associated to Cx. quinquefasciatus habitat data. Elevation data was represented in the WV-1 DEM by including gridded data where the terrain covariate was estimated for each cell in a regular grid, a triangular irregular network, and contours. The coefficients from the decomposition of the sampled predictor variables were then entered into a Bayesian matrix. This framework allowed flexible model fitting, estimation, and mapping of all high risk habitats based on field-sampled egg-raft count data using a MCMC specification. Residual estimates were then analyzed with an eigenvector spatial filtering algorithm which trans- formed all the georferenced predictor variables containing spatial dependence into covariates free of spatial dependence by partitioning the original sampled habitat data and the sampled egg-raft abundance count data into two synthetic variates: 1) a spatial filter variate capturing latent spatial dependency and 2) a non-spatial variate that was free of spatial dependence. The geographic distribution of the sampled habitats based on the egg-raft counts exhibited positive autocorrelation in all models tested: log-egg-raft counts aggregated in geo-space. The procedure accounted for 17 % pseudo-replicated information in the datasets. All models were then adjusted based on the residual estimates form the eigenfunction decomposition algorithm and then kriged. The sill indicated that the semivariance values had been reached (i.e., the value of maximum variance was equivalent to the variance of a 0.5m pixel value), while a non-zero intercept value (i.e., nugget variance) of the varigram model was indicative of the variability of the field and remotesampled Cx. quinquefasciatus data. A simple quantitative measure of the interpolation performed was determined by generating RMSE values for the models and by constructing Voroni polygons. Optimizing the RMSE by minimizing the spatial structure in a Cx. quinquefasciatus habitat model generated a pure nugget variogram, of which the level of nugget variance represented noise characteristics in the predictor variables. Interpolation accuracy tests measured by the natural logarithm of the mean squared interpolation error and the Voroni polygons revealed all error effects of the parameter estimates in the kriged-based model and quantified the covariate interaction terms. Newer GIS software, WV-I data and spatial statistics can implement larval control strategies by targeting prolific habitats of Cx. quinquefasciatus based on field-sampled count data in Trinidad. In the future high quality spatial autoregressive and surface models should be constructed using spatiotem- poral-sampled Cx. quinquefasciatus and other mosquito habitat data and Worldview-2 (WV-2) imagery for im- proving sensitivity in LULC, elevation and kriged-based interpolation parameters in Trinidad. On October 8, 2009 Digitalglobe launched WV-2 which is the first high resolution commercial satellite to offer 8-band capability with high accuracy, agility and spectral diversity (www. digitalglobe.com). This allows the creation of accurate Cx. quinuqefasciatus habitat maps without the need of GCPs. A wide range of features can be extracted using the traditional four spectral bands but WV-2 offers the addition of the Coastal Yellow, Red edge and shorter wave NIR bands which may provide a wider range of potential derivatives for targeting prolific Cx. Quinque- fasciatus and other mosquito habitats. For example, the Coastal-blue bands can be used for bathymetry, benthic, wetland and atmospheric modeling which can assist in detecting subtle differences in land cover and coastal features associated with prolific mosquito habitats. Probably the best feature of WV-2 for geomapping Cx. quinquefasciatus and other mosquito habitats, however, is the ability to derive algorithms and functions from new band combinations for feature extraction and improved classification. For example, the Red-edge NIR band can identify plant health and age through chlorophyll produc- tion while the yellow band in conjunction with the Red Edge band can provide agricultural applications and iron oxide mineralogy which can model moisture content. Thus, WV-2 can provide detailed imagery for precise map gen- eration, LULC change detection, construction of DEMs and autoregressive predictive models. WV-2 data can further help implement a remote-based habitat surveil- lance in Trinidad and in other regions.  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 47 5. Acknowledgements We acknowledge the data collection provided by the Ministry of Health in Trinidad and Tobago and the De- partment of Life Sciences, University of the West Indies, St. Augustine, and Trinidad. This research was funded by the National Institute of Health Grant U01A154889 (Novak Robert) University of Alabama at Birmingham and from the Government of Trinidad and Tobago, GTTGF-6 (Chadee Dave). 6. References [1] D. Nash, F. Mostashari, A. Fine, J. Miller, D. O’Leary, K. Murray, A. Huang, A. Rosenberg, A. Greenberg, M. Sherman, S. Wong, G. L. Campbell, J. T. Roehrig, D. J. Gubler, W. J. Shieh, S. Zaki, P. Smith and M. Layton, “The Outbreak of West Nile Virus Infection in the New York City Area in 1999,” The New England Journal of Medicine, Vol. 344, 2001, pp. 1807-1814. doi:10.1056/NEJM200106143442401 [2] N. Komar and G. C. Clark, “West Nile Virus Activity in Latin America and the Caribbean,” Revista Panameri- cana de Salud Pública, Vol. 19, 2006, pp. 112-117. doi:10.1590/S1020-49892006000200006 [3] M. J. Turell, M. R. Sardelis, M. L. O’Guinn and D. J. Dohm, “Potential Vectors of West Nile Virus in North America. Curr,” Top Microbiol. Immunol, Vol. 267, 2002, pp. 241-252. [4] B. G. Jacob, R. L. Lampman, M. P. Ward, E. J. Muturi, J. A. Morris, E. X. Caamano and R. J. Novak, “Geospatial Variability in the Egg Raft Distribution and Abundance of Culex Pipiens and Culex Restuans in Urbana-Cham- paign, Illinois,” International Journal of Remote Sensing, Vol. 30, 2009, pp. 2005-2019. doi:10.1080/01431160802549195 [5] B. G. Jacob, W. Gu, E. J. Muturi, E. X. Caamano, J. M. Morris, R. Lampman, R. J. Novak, “Developing Opera- tional Algorithms Using Linear and Non-Linear Least Squares Estimation in Python® for Identification of Culex Pipiens and Culex Restuans Aquatic Habitats in a Mos- quito Abatement District (Cook County, Illinois),” Geo- spat Health, Vol. 3, 2009, pp. 23-31. [6] J. K. Meece, J. S. Henkel, L. Glaser, K. D. Reed, “Mos- quito Surveillance for West Nile Virus in Southeastern Wisconsin-2002,” Clinical Medicine & Research Vol. 1, 2003, pp. 37-42. doi:10.3121/cmr.1.1.37 [7] F. W. Kutz, T. G. Wade and B. B. Pagac, “A Geospatial Study of the Potential of Two Exotic Species of Mosqui- toes to Impact the Epidemiology of West Nile Virus in Maryland,” Journal of the American Mosquito Control Association, Vol. 19, 2003, pp. 190-198. [8] D. A. Griffith, “A Comparison of Six Analytical Disease Mapping Techniques as Applied to West Nile Virus in the Coterminous United States,” International Journal of Health Geographics, Vol. 4, 2005, pp. 18-26. doi:10.1186/1476-072X-4-18 [9] U. Kitron, “Risk Maps: Transmission and Burden of Vector-Borne Diseases,” Parasitol Today, Vol. 16, 2000, pp. 324-325. doi:10.1016/S0169-4758(00)01708-7 [10] D. J. Rogers and S. E. Randolph, “Studying the Global Dis- tribution of Infectious Diseases Using GIS and RS,” Na- - ture Reviews Microbiology, Vol. 1, 2003, pp. 231-237. [11] S. I. Hay, J. A. Omumbo, M. H. Craig and R. W. Snow, “Earth Observation, Geographic Information Systems and Plasmodium Falciparum Malaria in Sub-Saharan Africa,” Advances in Parasitol,Vol. 47, 2000. [12] E. P. Pegoraro, M. J. Monson, R. K. Rey, A. Barron-Gaf- ford and C. B. Osmond, “The Effect of Elevated CO2, Soil and Atmospheric Water Deficit and Seasonal Phenology on Leaf and Ecosystem Isoprene Emission,” Functional Plant Biology, Vol. 34, 2007, p. 774. doi:10.1071/FP07021 [13] J. S. Brownstein, T. R. Holfold, D. Fish, “Enhancing West Nile Virus Surveillance, United States,” Emerging Infectious Diseases, Vol. 10, 2004, pp. 1129-1133. [14] N. Komar, S. Langevin, S. Hinten, N. Nemeth, E. Ed- wards, D. Hettler, B. Davis, R. Bowen, M. Bunning, “Experimental Infection of North American Birds with the New York 1999 Strain of West Nile Virus,” Emerg- ing Infectious Diseases, Vol. 9, 2003, pp. 311-322. [15] L. R. Petersen, A. A. Marfin, D. J. Gubler, “West Nile Virus,” JAMA, the Journal of the American Medical As- sociation, Vol. 4, No. 290, 2003, pp. 524-528. [16] B. G. Jacob, N. D. Burkett-Cadena, J. C. Luvall, S. H. Parcak, C. J. W. McClure, L. K. Estep, G. E. Hill, E. W. Cupp and R. J. Novak, “Developing GIS-Based Eastern Equine Encephalitis Vector-Host Models in Tuskegee, Ala- Bama,” International Journal of Health Geographics, Vol. 9, 2010, p. 12. doi:10.1186/1476-072X-9-12 [17] E. Mushinzimana, S. Munga, N. Minakawa, L. Li, C. C. Feng, L. Bian, U. Kitron, C. Schmidt, L. Beck, G. Zhou and A. K. Githeko, “Landscape Determinants and Re- mote Sensing of Anopheline Mosquito Larval Habitats in the Western Kenya Highlands,” Malaria Journal, Vol. 5, 2006, p. 13. doi:10.1186/1475-2875-5-13 [18] J. Shaman, J. F. Day and M. Stieglitz, “Drought-Induced Amplification and Epidemic Transmission of West Nile Virus in Southern Florida,” Journal of Medical Entomol- ogy, Vol. 42, No. 2, 2005, pp. 134-141. doi:10.1603/0022-2585(2005)042[0134:DAAETO]2.0.C O;2 [19] J. R. Jensen, “Remote Sensing of Urban Suburban Infra- structure and Socio-Economic Attributes,” Photogram- metric Engineering and Remote Sensing, Vol. 5, No. 65, 2005, pp. 611-622. [20] B. G. Jacob, E. J Muturi, E. X. Caamano, J. T. Gunter, E. Mpanga, R. Ayine, J. Okelloonen, J. P. M. Nyeko, J. I. Shililu, J. I. Githure, J. L. Regens and R. J. Novak, “Hy- dro- logical Modeling of Geophysical Parameters of Ar- boviral and Protozoan Disease Vectors in Internally Dis- placed People Camps in Gulu, Uganda,” International Journal of Health Geographics, Vol. 7, 2008, pp. 1-11. doi:10.1186/1476-072X-7-11  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 48 [21] B. G. Jacob, D. A. Griffith and R. J. Novak, “Decompos- ing Malaria Mosquito Aquatic Habitat Data into Spatial Autocorrelation Eigenvectors in a SAS/GIS® Module,” Transactions in GIS Vol. 12, 2008, pp. 341-364. doi:10.1111/j.1467-9671.2008.01104.x [22] C. N. Theophilides, S. C. Ahearn, E. S. Binkowski, W. S. Paul and K. Gibbs, “First Evidence of West Nile Virus Am- plification and Relationship to Human Infections,” Inter- national Journal of Geographical Information Science, Vol. 20, 2006, pp. 103-115. doi:10.1080/13658810500286968 [23] T. G. Andreadis, J. F. Anderson and C. R. Vossbrink, “Mos- quito Surveillance for West Nile Virus in Con- necticut, 2000: Isolation from Culex Pipiens, Cx. Restu- ans, Cx. Salinarius, and Culiseta Melanura,” Emerging Infectious Diseases, Vol. 7, 2001, pp. 670-674. doi:10.3201/eid0704.010413 [24] M. Eidson, J. Miller, L. Kramer, B. Cherry and Y. Hagi- wara, “Dead Crow Densities and Human Cases of West Nile Virus New York State 2000,” Emerging Infectious Dis- eases, Vol. 7, 2001, pp. 662-664. doi:10.3201/eid0704.010411 [25] M. Kulldroff, “A Spatial Scan Statistic,” Community Statistical Theory Methods, Vol. 26, 1997, pp. 1481-1496. doi:10.1080/03610929708831995 [26] B. G. Jacob, E. Muturi, J. Mwangangi, J. Funes, J. Shililu, J. Githure and R. J. Novak, “Remote and Field Level Quantification of Vegetation Covariates for Malaria Mapping in Three Rice Agro-Village Complexes in Cen- tral Kenya,” International Journal of Health Geographics, Vol. 6, 2007, pp. 21-28. doi:10.1186/1476-072X-6-21 [27] I. Kleinschmidt, M. Bagayoko, G. P. Y. Clarke, M. Craig and D. L. Le Sueur, “A Spatial Statistical Approach to Ma- laria Mapping,” International Epidemiology Asso- ciation, Vol. 29, No. 2, 2000, pp. 355-361. [28] N. Cressie, “Aggregation in Geostatistical Problems,” Geostatistics troia, 1992. [29] W. Gu and R. J. Novak, “Habitat-Based Modeling of Impacts of Mosquito Larval Interventions on Entomo- logical Inoculation Rates, Incidence, and Prevalence of Malaria,” American Journal of Tropical Medicine and Hygiene, Vol. 73, 2005, pp. 546-552. [30] B. G. Jacob, D. A. Griffith, E. J. Muturi, E. X. Caamano, J. I. Githure, J. T. Gunter, R. J. Novak, “A Heteroskedas- tic Error Covariance Matrix Estimator Using a First-Order Conditional Autoregressive Markov Simula- tion for Deriving Asymptotical Efficient Estimates from Ecological Sampled Anopheles Arabiensis Aquatic Habi- tat Covariates,” Malaria Journal, Vol. 8, 2009, pp. 216-224. doi:10.1186/1475-2875-8-216 [31] D. Chadee, “Key Premises, a Guide to Aedes Aegypti (Diptera: Culicidae) Surveillance and Control,” Bulletin of Entomological Research, Vol. 94, 2004, pp. 201-207. doi:10.1079/BER2004297 [32] J. P. Bradbury, B. Leyden, M. Salgado-Labouriau, Lewis, W. M. Jr., C. Schubert, M. W. Binford, D. G. Frey, D. R. Whitehead and F. W. Weibezahn, “Late Quaternary Envi- ronmental History of Lake Valencia,” Ven. Science, Vol. 4, 1981, pp. 18-26. [33] J. R. Flenley, “The Equatorial Rain Forest: A Geological History,” Butterworth, London, Vol. 9, 1979, p. 162. [34] J. S. Beard, “The Mora Forests of Auburn, British West Indies,” Journal of Ecology, Vol. 33, 1946, pp. 173-192. doi:10.2307/2256464 [35] D. D. Chadee, “Indoor and Outdoor Host-Seeking Rhythms of Anopheles Albitarsis (Diptera: Culicidae) in Trinidad, West Indies,” Journal of Medical Entomology, Vol. 29, 1992, pp. 567-569. [36] B. G. Jacob, P. G. Nelson, R. Lampman, J. A. Morris, A. Raims, J. Funes, C. LaPonte and R. J. Novak, “Compar- ing Global Positioning System (GPS) Technology for Identi- fying Spatial Ecological Variation for Urban Mosquito Abatement,” Wing Beats, Vol. 16, 2005, pp. 30-33. [37] B. G. Jacob, J. Shililu, E. J. Muturi, J. M. Mwangangi, S. M. Muriu, J. Funes, J. Githure, J. L. Regens and R. J, Novak, “Spatially Targeting Culex Quinquefasciatus Aquatic Habitats on Modified Land Cover for Imple- menting an Integrated Vector Management (IVM) Pro- gram in Three Villages Within the Mwea Rice Scheme, Keny,” International Journal of Health Geographics, Vol.5, 2006, pp. 18-27. doi:10.1186/1476-072X-5-18 [38] F. A. Haight, “Handbook of the Poisson Distribution,” New York, Wiley, 1967. [39] S. Lang and A. Brezger, “Bayesian P-Splines,” Journal of Computational and Graphical Statistics Vol. 13, 2004, pp. 183-212. doi:10.1198/1061860043010 [40] D. A. Griffith, “Spatial Autocorrelation on Spatial Filter- ing,” Springer, 2003. [41] J. F. Anderson, T. G. Andreadis, A. J. Main and D. L. Kline, “Prevalence of West Nile Virus in Tree Can- opy-Inhabiting Culex Pipiens and Associated Mosqui- toes,” American Journal of Tropical Medicine and Hy- giene, Vol. 71, 2004, pp. 112-119. [42] W. K. Reisen, J. O. Lundstrom, T. W. Scott, B. F. El- dridge, R. E. Chiles, R. Cusack, V. M. Martinez, H. D. Lothrop, D. Gutierrez, S. E. Wright, K. Boyce and B. R. Hill,. “Patterns of Avian Seroprevalence to Western Equine Encephalomyelitis and St. Louis Encephalitis Vi- ruses in California, USA,” Journal of Medical Entomol- ogy, Vol. 37, 2000, pp. 507-527. doi:10.1603/0022-2585-37.4.507 [43] L. M. Calhoan, M. Avery, L. Jones, K. Gunarto, R. King, J. Roberts and T. R. Burkot, “Combined Sewage Over- flows (CSO) Are Major Urban Breeding Sites for Culex Quin- quefasciatus in Atlanta, Georgia,” American Jour- nal of Tropical Medicine and Hygiene, Vol. 77, 2007, pp. 478-484. [44] K. Sugihara, A. Okabe and T. Satoh, “Computation Method for the Point Cluster Analysis on Networks,” Geoinfor- matica, Vol. 15, No. 1, 2008, pp. 9-92. [45] J. H. Rappole, S. R. Derrickson and Z. Hubálek, “Migra- tory Birds and Spread of West Nile Virus in the Western Hemisphere,” Emerging Infectious Diseases, Vol. 6, 2000,  B. G. JACOB ET AL. Copyright © 2011 SciRes. JGIS 49 pp. 11-19. doi:10.3201/eid0604.000401 [46] A. T. Peterson, D. A. Vieglais and J. K. Andreasen, “Mi- gra- tory Birds Modeled as Critical Transport Agents for West Nile Virus in North America,” Vector-Borne and Zoono- tic Diseases, Vol. 3, 2003, pp. 27-37. doi:10.1089/153036603765627433 [47] R. B Tesh, R. Parson, M. Siirin, Y. Randle, C. Sargent, H. Guzman, T. Wuithiranyagool, S. Higgs, D. L. Vanlanding- ham, A. A. Bala, K. Haas and B. Zerinque, “Year-Round West Nile Virus Activity, Gulf Coast Region, Texas and Louisi- ana,” Emerging Infectious Diseases, Vol. 10, 2004, pp. 1649-1652. [48] T. A. Beveroth, M. P. Ward, R. L. Lampman, A. M. Rin- gia and R. J. Novak, “Changes in Seroprevalence of West Nile Virus Across Illinois in Free-Ranging Birds from 2001 through 2004,” American Journal of Tropical Medicine and Hygiene, Vol. 74, 2006, pp. 174-179. [49] R. M. Gleiser, A. J. MacKay, A. Roy, M. M. Yates, R. H. Vaeth, G. M. Faget, A. E. Folsom, W. F. Augustine, R. A. Jr. Wells and M. J. Perich, “West Nile Virus Surveillance in East Baton Rouge Parish, Louisiana,” Journal of the American Mosquito Control Association, Vol. 23, 2007, pp. 29-36. doi:10.2987/8756-971X(2007)23[29:WNVSIE]2.0.CO;2 [50] P. P. Marra, S. Griffing, C. Caffrey, A. M. Kilpatrick, R. McLean, C. Brand, E. Saito, A. P. Dupuis, L. Kramer and R. J. Novak, “West Nile Virus and Wildlife,” BioScience, Vol. 54, 2004, pp. 393-402. doi:10.1641/0006-3568(2004)054[0393:WNVAW]2.0.C O;2 |

