 Vol.5, No.9, 1047-1055 (2013) Natural Science http://dx.doi.org/10.4236/ns.2013.59129 Analytical expressions of the concentrations of substrate and product in enzyme inhibition process Sankaranarayanan Muthukumar, Lakshmanan Rajendran Department of Mathematics, The Madura College, Madurai, India; raj_sms@rediffmail.com Received 3 June 2013; revised 3 July 2013; accepted 10 July 2013 Copyright © 2013 Sankaranarayanan Muthukumar, Lakshmanan Rajendran. This is an open access article distributed under the Crea- tive Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. ABSTRACT The initial and boundary value problem in en- zyme reactions mechanism for inhabitation proc- ess is discussed. Approximate analytical exp re s- sions for the concentrations of substrate and product are presented. Approximate analytical solutions of non-linear reaction equations con- taining non-linear terms related to enzymatic re- action mechanism are solved using Homotopy perturbation method. The relevant analytical ex- pression for the substrate and product concen- tration profiles is discussed in terms of dimen- sionless reaction diffusion parameters α, β, γE, and γS. Numerical solution is also obtained us- ing Matlab program. Our analytical expression compared with numerical estimation and good agreement is noted. Keywords: Initial and Boundary V a lue Problems; Enzyme Kinetics; Non-Linear Re action Equations; Homotopy Perturbation Method; Michael is-Menten Kinetics 1. INTRODUCTION Enzymes are catalysts and increase the speed of a chemical reaction without themselves undergoing any permanent chemical change [1-7]. They are neither used up in the reaction nor do they appear as reaction products. An enzyme enhances the rate of the reaction it influences. The catalysts is not consumed as a result of the reaction, nor does it alter the equilibrium constant [8,9]. In this two-step reaction, the enzyme combines with substrate to form the complex, which can either dissociate again into unchanged substrate and enzyme or go on to the second step and form the products and unchanged enzyme. The reverse of the second step would lead to the synthesis of ES from enzyme and products, but this process can gen- erally be ignored unless the products are allowed to ac- cumulate. Also that the total substrate and product con- centration in the enzyme kinetics in biochemical systems has been modeled by system of nonlinear ordinary dif- ferential equations. An enzyme inhibitor is a molecule, which binds to enzymes and decreases their activity. Since blocking an enzyme’s activity can kill a pathogen or correct a meta- bolic imbalance, many drugs are enzyme inhibitors. Th ey are also used as herbicides and pesticides. Not all mole- cules that bind to enzymes are inhibitors; enzyme acti- vators bind to enzymes and increase their enzymatic ac- tivity, while enzyme substrates bind and are conv erted to products in the normal catalytic cycle of the enzyme. The binding of an inhibitor can stop a substrate from entering the enzyme’s active site. Inhibitor binding is ei- ther reversible or irreversible. Irreversible inhibitors usu- ally react with the enzyme and change it chemically. In contrast, reversible inhibitors bind non-covalently and dif- ferent types of inhibition are produced depending on whether these inhibitors bind to the enzyme, the enzyme- substrate complex, or both. Many drug molecules are en- zyme inhibitors, so their discovery and improvement is an active area of research in biochemistry and pharma- cology. Mathematical modeling of enzyme kinetics is given in the books by Rubinow [10], Murray [11], Segel [12] and Roberts [13]. Recently Rajendran and his team also solved some non linear problems in enzyme reaction kinetics [14-19]. The purpose of this communication is to derive asymptotic approximate expressions for the concentra- tion of substrate and product in enzyme inhibition proc- ess. 2. MATHEMATICAL FORMULATION AND ANALYSIS OF THE SUBSTRATE INHIBITION MODELS The sub strate act as an inh ibitor for enh ancing the ra te Copyright © 2013 SciRes. OPEN ACCESS  S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 1048 of reaction. The scheme of the enzyme (E) catalysed con- version of the substrate (S) to the product (P) SP (1) is a simplified version of ESES EP (2) According to scheme (2), the substrate (S) combines reversibly with the enzyme (E) to form the enzyme-sub- strate complex (ES). The complex then dissociates into product (P) and the enzyme is regenerated. The simplest scheme of non-Michaelis-Menten kinetics may be, for example, described by adding the relationship of the in- teraction of the enzyme substrate complex (ES) with an- other substrate molecule (S) followed by the generation of the non-active complex (ESS) to the Michaelis-Men- ten scheme ES SESS (3) The above substrate inhibition process can be repre- sented by the f oll owi ng sy st em of nonli near m a ss bala nce equations: 2max 21 S S VS SS D tKSS x K (4) 2max 2, 0,, 0 1 P MS VS PP DS tKSSK x dt (5) where and stand for space and time, respectively, t ,Sxt is the concentration of the substrate, , xt is the concentration of the reaction product, d is the thick- ness of the enzyme layer, PS are the diffusion co- efficients, max is the maximal enzyme rate and DD , V and S are the Michaelis and the substrate inhibition constants, respectively. The governing Eq.4 together with the following initial and the boundary conditions form the initial boundary value problem 0t ,00 and ,00Sx Px (6) The boundary conditions are: 00 0 ,0, ,, 0,,0, 0, x SdS SdtS S Pt pdtx (7) At steady state the Eqs.4 and 5 becomes 2max 2 d0 1 d S MS VS S DKSSK x (8) 2max 2 d0 1 d P MS VS P DKS SK x (9) with the boundar y conditions d 00, 0 d S Px x (10) 0 0, xdSxd S (11) The density of the biosensor current is given by: 0 eP x P inFD x (12) The following sets of dimensionless variables are in- troduced: 2 2max 00 22 2 00 max d , , , , d , , E SM S MMS PM V xS P XS P dS SDK SS V KKK DK (13) Thus dimensi onl ess Eqs.8 and 9 becomes 2 2 22 d0 d1 ES S XSS (14) 2 2 22 d0 d1 S P XSS (15) The dimensionless boundary conditions becomes d 0, 0, 0 d S XP x (16) 1, 0, 1XPS (17) The dim e ns i onless current is given by 00 d d eP X id P nFDs X (18) 3. ANALYTICAL EXPRESSION OF CONCENTRATIONS OF SUBSTRATE AND PRODUCT USING HPM After recently, many authors have applied the HPM to solve the non-linear problem in physics and engineering sciences [20-23]. Recently this method is also used to solve some of the non-linear problem in biological and physical sciences [24-27]. This method is a combination of homotopy in topology and classic perturbation tech- niques. Ji-Huan He used the HPM to solve the Lighthill equation [28], the Duffing equation [29] and the Blasius equation [30]. The HPM is unique in its applicability, accuracy and efficiency. The HPM uses the imbedding parameter p as a small parameter, and only a few itera- tions are needed to search for an asymptotic solution. Using this method (see Appendix A), we can obtain the concentrations of substrate and product which are the solution to Eqs.14 an d 15 (see Appendix B) as follows: Copyright © 2013 SciRes. OPEN ACCESS  S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 104 9 22 1 22 2 273 4 222 sech sech sech cosh sinh cosh 2 cosh3 SS SX kNMNM 2 MX N kX MXkk MX kMX NNNN (19) 275 2 65 12 223 22 64 22 2 2 57 6 12 2232 53 222 sech1 24 321cosh 48 1cosh2 1cosh3 36 9 3 2 sech 42 4 sinh cosh 84 NM kk PX M kk kk M MMM MM kk M MM kk k kk N XM MMMM kk kX MX MX MM 3 4 k MX 2 6 424 22 322 2 636 22 22 cosh 2 2 cosh3 936 98 3sech 4436 MX M k kkk MX 5 1 2 k k MMMM kkk NM MM MM M (20) where 2 2, 1 E M and 23 12 22 3 2 22 22 32 22 224 4 3 222 2222 2 222 22 4 2 45 22 sech tanh 2 sechcosh 2 6 3 sechcosh3sech tanh 32 8 sechsech sech 2 3 sechsech , 28 sech, s 32 E E EE ESS EE ES NM kMM NM MM NM NM Mm NMN N MMM NM NM kM NM N kMk M M 2 422 22 67 22 22 32 ech, sech ,sech , 2 sech, 6 SE E M NNM kMk M NM kM (21) The analytical expression of dimensionless current us- ing the Eq.17 is given by 26 12 222 53 22 67 4 22 32 sech 14 1cosh 1cosh2 84 1cosh3 24 36 9 k kk NM MMM kk 3 5 M M MM kk kM MM k (22) 4. NUMERICAL SIMULATION The nonlinear differential Eqs.19 and 20 are also solved by using numerical methods. The function ode45 in Mat- lab software which is a function of solving two-point boundary value problems (BVPs) for ordinary different- tial equations is used to solve those equations. Its nu- merical solution is compared with the solution obtained by using Homotopy perturbation method and it gives a satisfactory result. The Matlab program is also given in Appendix C. 5. DISCUSSION Eqs.19 and 20 are the new and simple analytical ex- pressions of concentrations of the substrate and the prod- uct calculated using Homotopy perturbation method. The dimensionless analytical expressions of concentration substrate SX andproduct X versus the di- mensionless distance for various values of dimen- sionless reaction parameters are compared with numeri- cal solution in Figures 1 and 2. X From these figures, it is inferred that the value of the concentration of substrate increases gradually when increases. Also the concentration of substrate increases when X decreases and decreases. The concentra- tion of substrate is uniform when 0.1 E . From the Figure 2, it is inferred that the product have minimum value at 0X and 1X . Also it is maximum at X = 0.5. Concentration of product increases when the pa- rameter increases. Dimensionless current ver- sus dimensionless parameters ,, and S is plot- ted in Figure 3. From these figure it is inferred that the current increases when and decreases or γE and γS increases. 6. CONCLUSION The analytical expressions for the concentration of substrate and product in substrate inhibition process are derived using new Homotopy perturbation method. We have also presented an analytical expression for the s te a d y- state current. The extension of procedure to various inhi- bition models for steady-state and non-steady state con- ditions seems possible. These analytical results will be useful for the optimization and the design of enzyme inhibition process. Copyright © 2013 SciRes. OPEN ACCESS 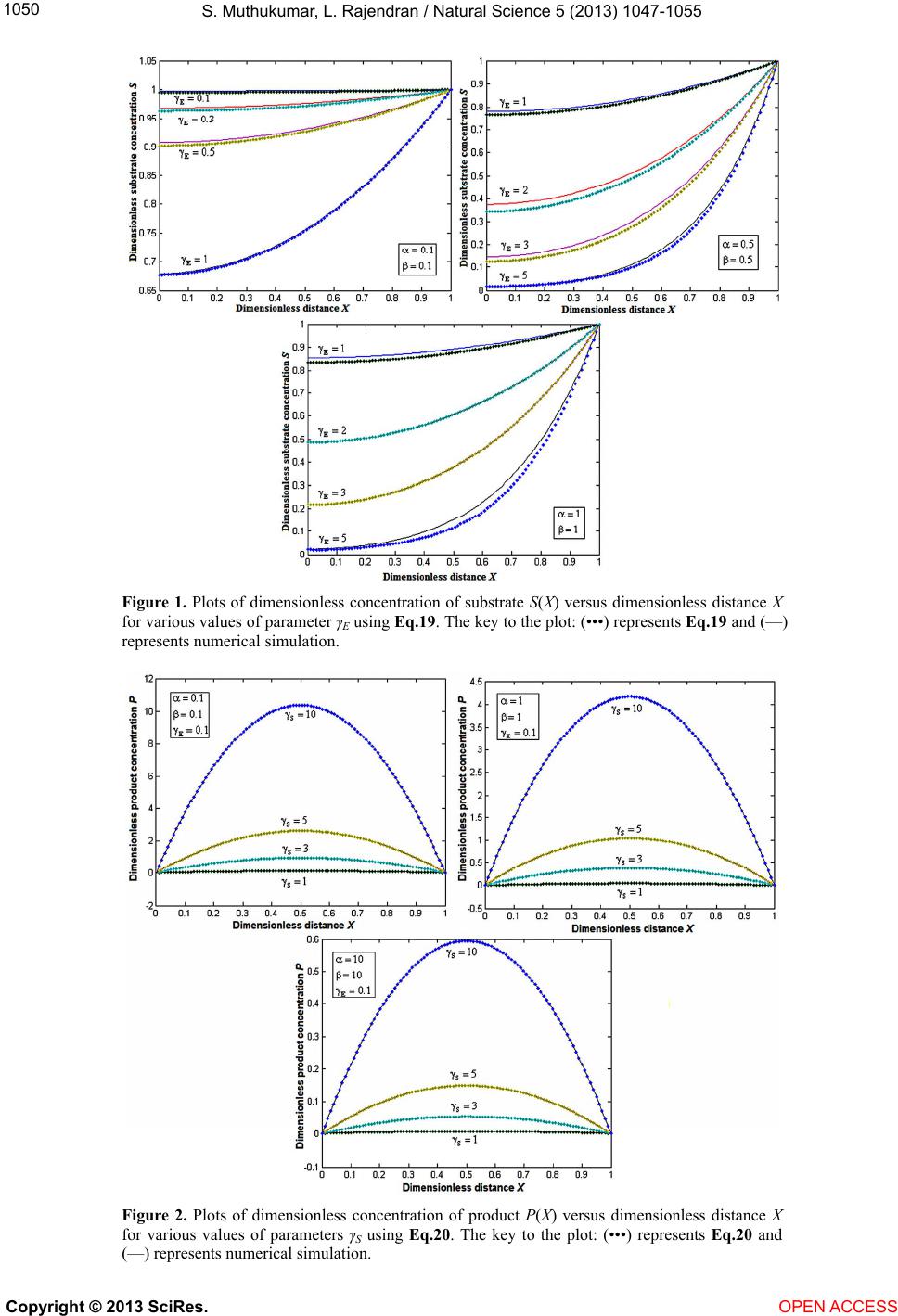 S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 Copyright © 2013 SciRes. 1050 Figure 1. Plots of dimensionless concentration of substrate S(X) versus dimensionless distance X for various values of parameter γE using Eq.19. The key to the plot: (•••) represents Eq.19 and (—) represents numerical simulation. Figure 2. Plots of dimensionless concentration of product P(X) versus dimensionless distance X for various values of parameters γS using Eq.20. The key to the plot: (•••) represents Eq.20 and (—) represents numerical simulation. OPEN ACCESS 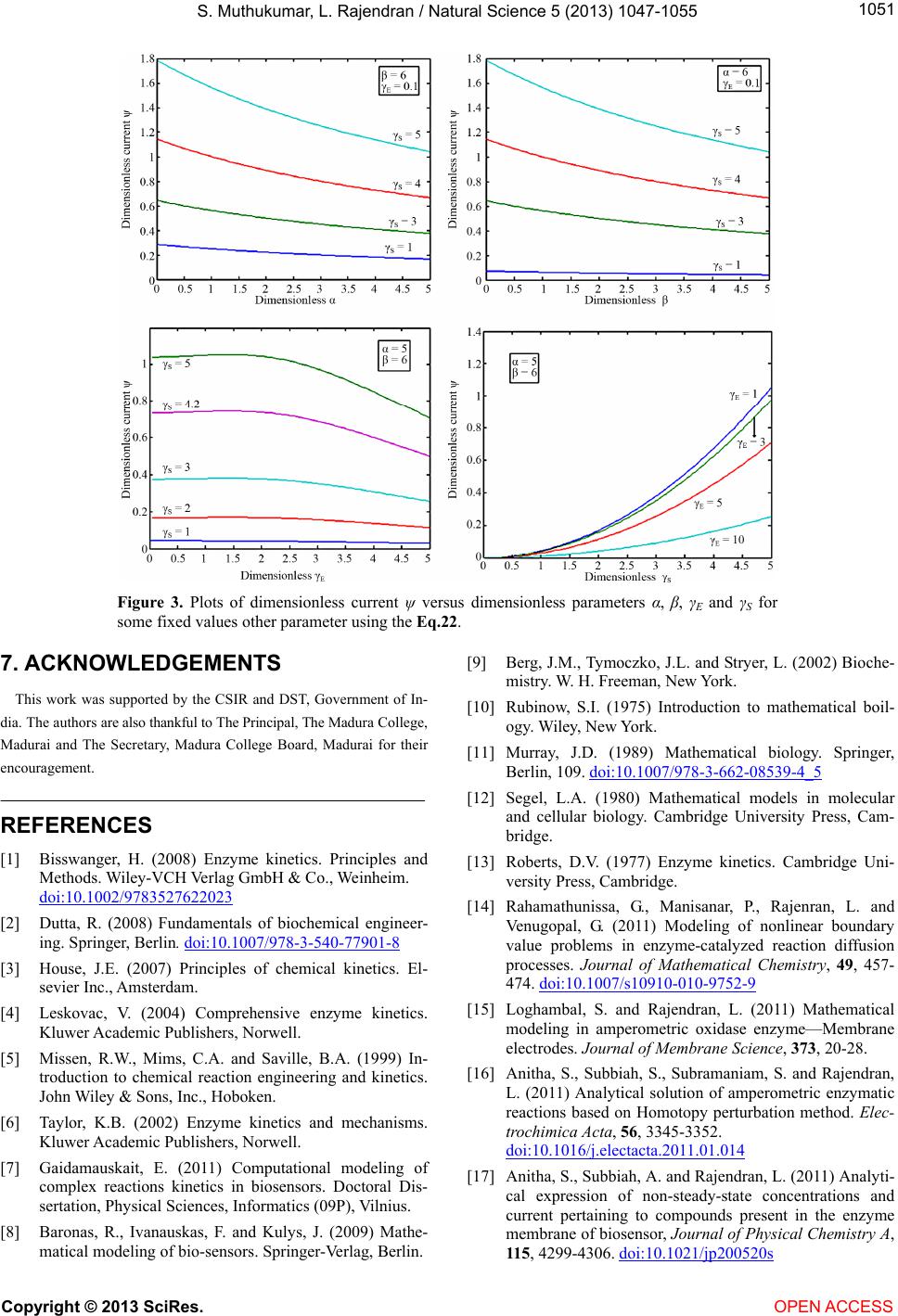 S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 1051 Figure 3. Plots of dimensionless current ψ versus dimensionless parameters α, β, γE and γS for some fixed values other parameter using the Eq.22. 7. ACKNOWLEDGEMENTS This work was supported by the CSIR and DST, Government of In- dia. The authors are also thankful to The Principal, The Madura College, Madurai and The Secretary, Madura College Board, Madurai for their encouragement. REFERENCES [1] Bisswanger, H. (2008) Enzyme kinetics. Principles and Methods. Wiley-VCH Ve rlag GmbH & Co., Weinheim. doi:10.1002/9783527622023 [2] Dutta, R. (2008) Fundamentals of biochemical engineer- ing. Springer, Berlin. doi:10.1007/978-3-540-77901-8 [3] House, J.E. (2007) Principles of chemical kinetics. El- sevier Inc., Amsterdam. [4] Leskovac, V. (2004) Comprehensive enzyme kinetics. Kluwer Academic Publishers, Norwell. [5] Missen, R.W., Mims, C.A. and Saville, B.A. (1999) In- troduction to chemical reaction engineering and kinetics. John Wiley & Sons, Inc., Hoboken. [6] Taylor, K.B. (2002) Enzyme kinetics and mechanisms. Kluwer Academic Publishers, Norwell. [7] Gaidamauskait, E. (2011) Computational modeling of complex reactions kinetics in biosensors. Doctoral Dis- sertation, Physical Sciences, Informatics (09P), Vilnius. [8] Baronas, R., Ivanauskas, F. and Kulys, J. (2009) Mathe- matical modeling of bio-sensors. Springer-Verlag, Berlin. [9] Berg, J.M., Tymoczko, J.L. and Stryer, L. (2002) Bioche- mistry. W. H. Freeman, New York. [10] Rubinow, S.I. (1975) Introduction to mathematical boil- ogy. W iley, New York. [11] Murray, J.D. (1989) Mathematical biology. Springer, Berlin, 109. doi:10.1007/978-3-662-08539-4_5 [12] Segel, L.A. (1980) Mathematical models in molecular and cellular biology. Cambridge University Press, Cam- bridge. [13] Roberts, D.V. (1977) Enzyme kinetics. Cambridge Uni- versity Press, Cambridge. [14] Rahamathunissa, G., Manisanar, P., Rajenran, L. and Venugopal, G. (2011) Modeling of nonlinear boundary value problems in enzyme-catalyzed reaction diffusion processes. Journal of Mathematical Chemistry, 49, 457- 474. doi:10.1007/s10910-010-9752-9 [15] Loghambal, S. and Rajendran, L. (2011) Mathematical modeling in amperometric oxidase enzyme—Membrane electrodes. Journal of Membrane Science, 373, 20-28. [16] Anitha, S., Subbiah, S., Subramaniam, S. and Rajendran, L. (2011) Analytical solution of amperometric enzymatic reactions based on Homotopy perturbation method. Elec- trochimica Acta, 56, 3345-3352. doi:10.1016/j.electacta.2011.01.014 [17] Anitha, S., Subbiah, A. and Rajendran, L. (2011) Analyti- cal expression of non-steady-state concentrations and current pertaining to compounds present in the enzyme membrane of biosensor, Journal of Physical Chemistry A, 115, 4299-4306. doi:10.1021/jp200520s Copyright © 2013 SciRes. OPEN ACCESS  S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 1052 [18] Uma Maheswari, M. and Rajendran, L. (2011) Analytical solution of non-linear enzyme reaction equations arising in mathematical chemistry. Journal of Mathematical Che- mistry, 49, 1713-1726. doi:10.1007/s10910-011-9853-0 [19] Meena, A., Eswari, A. and Rajendran, L. (2011) Mathe- matical modeling of biosensors: Enzyme-substrate Inter- action and bio-molecular interaction. In: Serra, P.A., Ed., New Perspectives in Biosensors Technology and Applica- tions, InTech, Rijeka. doi:10.5772/19513 [20] Ghori, Q.K., Ahmed, M. and Siddiqui, A.M. (2007) Ap- plication of homotopy perturbation method to squeezing flow of a Newtonian fluid. International Journal of Non- linear Sciences and Numerical Simulation, 8, 179-184. [21] Ozis, T. and Yildirim, A. (2007) A comparative study of He’s homotopy perturbation method for determining fre- quency-amplitude relation of a nonlinear oscillator with discontinuities. International Journal of Nonlinear Sci- ences and Numerical Simulation, 8, 243-248. [22] Li, S.J. and Liu, Y.X. (2006) An improved approach to nonlinear dynamical system identification using PID neural networks. International Journal of Nonlinear Sci- ences and Numerical Simulation, 7, 177-182. doi:10.1515/IJNSNS.2006.7.2.177 [23] Mousa, M.M., Ragab, S.F. and Nturforsch, Z. (2008) Ap- plication of the Homotopy perturbation method to linear and nonlinear Schrödinger equations. Zeitschrift für Na- turforschung, 63, 140-144. [24] Margret PonRani, V. and Rajendran, L. (2010) Analytical expression of non steady-state concentration profiles at planar electrode for the CE mechanism. Natural Science, 2, 1318-1325. doi:10.4236/ns.2010.211160 [25] Varadharajan, G. and Rajendran, L. (2011) Analytical solution of coupled non-linear second order reaction dif- ferential equations in enzyme kinetics. Natural Science, 3, 459-465. doi:10.4236/ns.2011.36063 [26] Venugopal, K., Eswari, A. and Rajendran, L. (2011) Ma- thematical model for steady state current at ppo-modified micro-cylinder biosensors. Journal of Biomedical Science and Engineering, 4, 631-641. doi:10.4236/jbise.2011.49079 [27] Muthukumar, S. and Rajendran, L. (2012) Concentration of species in two species oscillator using Homotophy peturbation method. Global Journal of Theoretical and Applied Mathematics Sciences, 2, 99-108. [28] He, J.H. (1999) Homotopy perturbation technique. Com- puter Methods in Applied Mechanics and Engineering, 178, 257-262. [29] He, J.H. (2003) Homotopy perturbation method: A new non-linear analytical technique. Applied Mathematics and Computation, 135, 73-79. doi:org/10.1016/S0096-3003(01)00312-5 [30] He, J.H. (2003) A simple perturbation approach to Blasius equation. Applied Mathematics and Computation, 140, 217-222. doi:10.1016/S0096-3003(02)00189-3 Copyright © 2013 SciRes. OPEN ACCESS  S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 1053 APPENDIX A: BASIC IDEA OF HOMOTOPY PERTURBATION METHOD In this appendix we outline the basic idea of Homo- topy perturbation method. This method has eliminated the limitations of the traditional perturbation methods. On the other hand it can take full advantage of the tradi- tional perturbation techniques, so there has been a con- siderable deal of research in applying homotopy tech- nique for solving various strongly nonlinear equations. To explain Homotopy perturbation method, let us con- sider the following function 0, Auf rr (A1) with the boundary conditions, ,0, Bu unr (A2) where A, B, r and denote a general differential operator, a boundary operator, a known analytical func- tion and the boundary of the domain , respectively. Generally speaking, the operator A can be divided into a linear part L and a nonlinear part N. Eq.A1 can there fo re, be written as 0LuNuf r (A3) By Homotopy technique, we construct a Homotopy ,: 0,1vrp R which satisfies 0 ,1 0. 0,1, Hvpp LvLu pAv fr pr (A4) or 0 , 0. 0 vpLv LupLu pNv fr (A5) where 0,1 p 0f r is an embedding parameter, and μ0 is an initial approximation of Eq.A1, which satisfies the boundary conditions. When Eqs.A4 and A5 be- come a linear equation; when it become a non- linear equation. So the changing process of p from zero to unity is just that of changing to . We can first use the embedding pa- rameter p as a small parameter. Obviously, from Eqs.A4 and A5, we will have 0p p1 00Lv Lu Av 0 ,0 0HvLvLu (A6) ,10 HvAvf r (A7) Assume that the solutions of Eqs.A4 and A5 can be written as a power series in p 2 01 2 vvpv pv (A8) Setting results in the approximate solution of Eq.A1: 1p 012 1 lim p uvvvv (A9) The combination of the perturbation method and the Homotopy method is called the HPM. APPENDIX B: SOLUTION OF EQS.14 AND 15 USING HPM To find the solution of Eq.13, the homotopy is con- structed as follows: 222 2 222 22 2 22 22 2 ddd 1ddd dd d0 dd d E E SSS BS XXX SS S BSS S XX X (B1) The approximate solution of e Eq.B1 is: 2 01 2 SS BSBS (B2) Substituting Eqs.B2 in B1 results: 22 01 01 22 201 201 2 201 01 2 22 2 01 01 01 22 201 dd 1dd d d d d dd dd 0 E E SBSSBS BXX SBSSBS X SBS BSBS X SBS SBS SBS XX SBS (B3) Comparing the coefficients of the powers of B 222 02 000 0 222 ddd :0 ddd E SSS BS XX X (B4) 222 12 111 1 222 22 22 2 00 00 00 22 22 ddd :ddd dd dd 0 dd dd E SS S BS XXX SS SS SS XX XX (B5) The initial approximations are as follows: 01 dd 0, 0,0 dd SS XxX (B6) 01 1, 1, 0XS S (B7) Solving (B4) and (B5) with boundary conditions (B6) and (B7) gives 0sech coshSMMX (B8) and Copyright © 2013 SciRes. OPEN ACCESS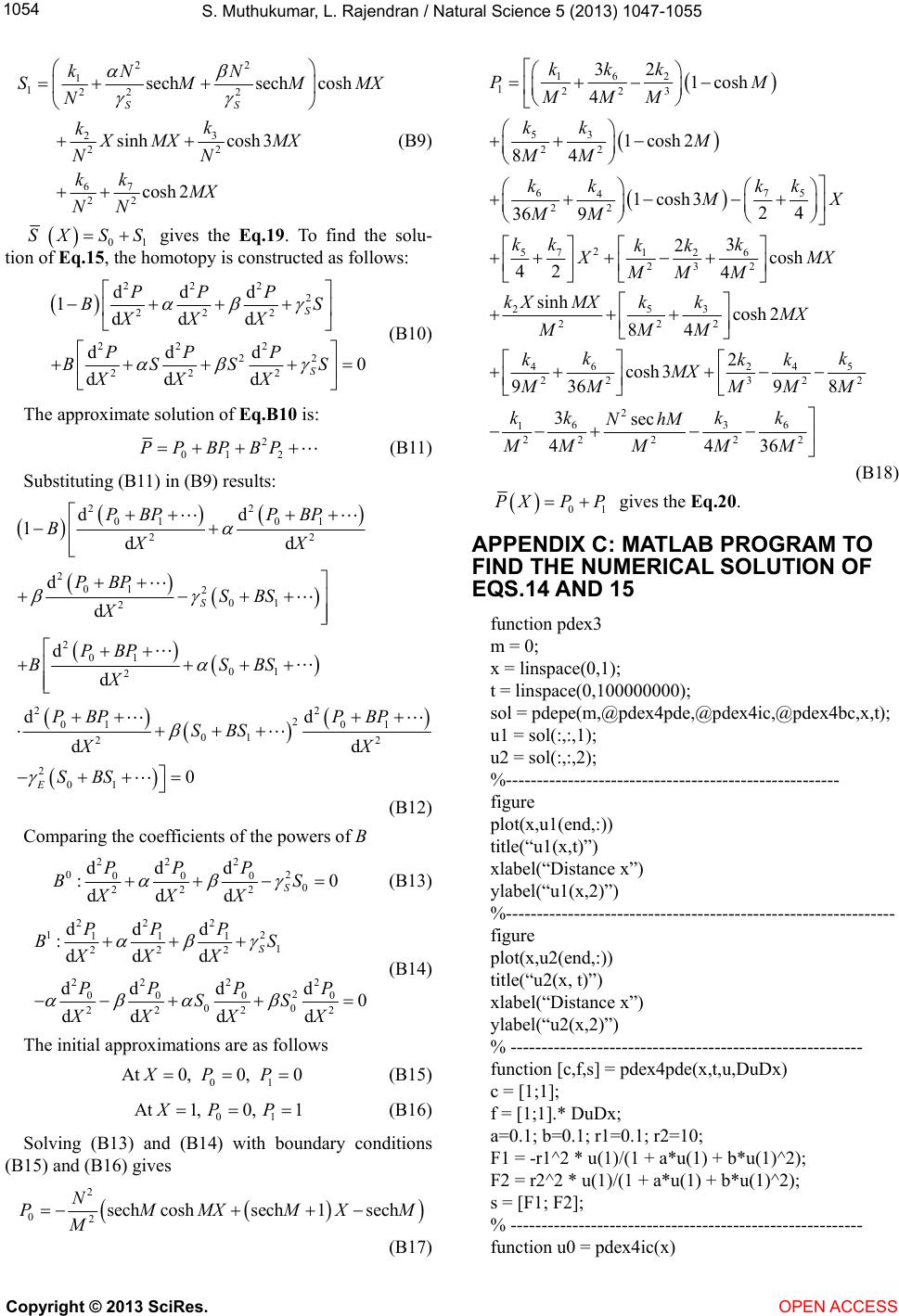  S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 1054 22 1 122 2 3 2 22 67 22 sechsech cosh sinh cosh3 cosh 2 SS kNN SMM N k kXMX MX NN kk MX NN MX (B9) 01 gives the Eq.19. To find the solu- tion of Eq.15, the homotopy is constructed as follows: SXS S 222 2 222 222 22 222 ddd 1ddd dd d0 dd d S S PPP BS XXX PPP BSSS XX X (B10) The approximate solution of Eq.B10 is: 2 01 2 PPBPBP (B11) Substituting (B11) in (B9) results: 22 01 01 22 201 201 2 201 01 2 2 2 01 01 01 22 201 dd 1dd d d d d d dd 0 S E PBP PBP BXX PBP SBS X PBP BSBS X PBP PBP SBS XX SBS 2 d (B12) Comparing the coefficients of the powers of B 222 0 000 0 222 ddd : ddd S PPP B XXX 2 0S (B13) 222 12 111 1 222 22 22 2 00 00 00 22 22 ddd :ddd dd dd 0 dd dd S PPP B XXX PP PP SS XX XX S (B14) The initial approximations are as follows 01 At 0,0,0XPP (B15) 01 At 1,0,1XP P (B16) Solving (B13) and (B14) with boundary conditions (B15) and (B16) gives 2 02sech coshsech1sech N PMMXMX M 6 12 1223 53 22 67 4 22 2 57 6 12 23 2 53 2222 6 42 321cosh 4 1cosh2 84 1cosh3 24 36 9 3 2cosh 42 4 sinh cosh 2 84 93 k kk PM MMM kk M MM kk kMX MM kk k kk XM MMM kk kX MXMX MMM k k M 5 k X 5 24 232 2 636 12222 2 2 cosh3 69 3sec 4436 k kk MX 2 8 MMM kkk kNhM MMMMM (B18) 0 PXP P 1 gives the Eq.20. APPENDIX C: MATLAB PROGRAM TO FIND THE NUMERICAL SOLUTION OF EQS.14 AND 15 functio n pdex3 m = 0 ; x = linspace(0,1); t = linspace(0,1000 00000); sol = pdepe(m,@pdex4pd e,@pdex4ic,@pdex4bc,x ,t); u1 = sol(:,:,1); u2 = sol(:,:,2); %------------------------------------------------------ figure plot(x,u1(end,:)) title(“u1(x,t)”) xlabel(“Distance x”) ylabel(“u1(x,2)”) %--------------------------------------------------------------- figure plot(x,u2(end,:)) title(“u2(x, t)”) xlabel(“Distance x”) ylabel(“u2(x,2)”) % --------------------------------------------------------- function [c,f,s] = pdex4pde(x,t,u,DuDx) c = [1;1]; f = [1;1].* DuDx; a=0.1; b=0.1; r1=0.1; r2=10; F1 = -r1^2 * u(1)/(1 + a*u(1) + b*u(1)^2); F2 = r2^2 * u(1)/(1 + a*u(1) + b*u(1)^2); M (B17) s = [F1; F2]; % --------------------------------------------------------- function u0 = pde x 4ic ( x) Copyright © 2013 SciRes. OPEN ACCESS  S. Muthukumar, L. Rajendran / Natural Science 5 (2013) 1047-1055 Copyright © 2013 SciRes. OPEN ACCESS 1055 u0 = [1;1]; % -------------------------------------------------------- function [pl,ql,pr,qr] = pdex4bc(xl,ul,xr,ur,t) pl = [0; ul(2)-0] ; ql = [1; 0]; pr = [ur(1)-1 ; ur(2)-0]; qr = [0; 0];
|