 Open Journal of Genetics, 2013, 3, 174-182 OJGen http://dx.doi.org/10.4236/ojgen.2013.33020 Published Online September 2013 (http://www.scirp.org/journal/ojgen/) Genetic analysis for geographic isolation comparison of brown bears living in the periphery of the Western Carpathians Mountains with bears living in other areas Ján Graban1*, Jana Kisková1, Pavol Pepich2, Robin Rigg3 1Institute of the High Mountain Biology, University of Žilina Tatranská Javorina, Tatranská Javorina, Slovak Republic 2Institute of Foreign Languages, University of Žilina, Žilina, Slovak Republic 3Slovak Wildlife Society, Liptovský Hrádok, Slovak Republic Email: *graban@uniza.sk Received 24 April 2013; revised 26 May 2013; accepted 10 June 2013 Copyright © 2013 Ján Graban et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. ABSTRACT Populations of the European brown bear (Ursus arc- tos L.) differ substantially in size, degree of geo- graphic isolation and level of genetic diversity. Pre- sent patterns result from phylogeographic processes and profound human intervention. We assessed the genetic variability of a subpopulation of brown bears near the periphery of their range in the Western Carpathian Mountains and compared their genetic properties with those of bears in the core of the same population and elsewhere. Samples were collected non-invasively in 2007-2008 and 2010 in Strážovské Vrchy Protected Landscape Area (PLA) in Slovakia (included in the NATURA 2000 networking pro- gramme). Seven polymorphic microsatellite loci (UaMU26, UaMU64, G10B, G1D, G10L, UaMU50 and UaMU51) were amplified using a nested PCR in order to assess the following parameters: variability, allelic combinations, heterozygosity, number of alleles and inbreeding coefficient. Sufficient brown bear DNA for analysis was obtained from 57 out of 140 samples (41%), among which 45 different genotypes were identified. Loci had a mean of 2.71 ± 0.76 alleles. Average observed heterozygosity was 0.59. The in- breeding coefficient was negative for all but one o f th e analysed loci (2007-2008). In the year 2010 was nega- tive three of seven loci. These results imply that gene flow with other parts of the population has been maintained in the reduced level and the isolation level of bears in the study area was not so low. Neverthe- less, the genetic variability of bears in Strážovské Vrchy PLA was lower than that reported from other localities in the Carpathian Mountains. The results are discussed in the context of behavioural ecology and conservation genetics. Keywords: Carpathian Mountains; European Brown Bear; Ursu s arcto s L.; Genetic Diversity; Microsatellite Markers; Non-Invasive Sampling 1. INTRODUCTION The brown bear (Ursus arctos) re-colonised the entire European continent after the Last Glacial Maximum [1,2], and yet its current distribution shows a discon- tinuous pattern as a result of various human activities [3,4]. In contrast to the relatively large and contiguous populations with higher expected heterozygosity and allelic diversity in Eastern Europe, the Balkans and Scandinavia [5] population fragments in the western part of the continent are extremely small and isolated, with low levels of genetic variability and vulnerable to genetic drift and inbreeding [6-8]. An additional level of complexity is added by the ex- istence of three mitochondrial subclades [6,9,10]. They may be the result of different founder populations having passed through bottlenecks prior to rapid recolonisation during the Holocene [11] or recent human-induced popu- lation fragmentation due to habitat loss and killing [12,13]. It has been suggested that one of the glacial refugia from which brown bears of the eastern lineage (subclade 3a) re-colonised most of continental Eurasia was in the Western Carpathian Mountains of present day Slovakia [2,11,14]. Alternatively, bears may have survived the glacial period in the cold tundra-steppe of central Europe [12]. Regardless of when and how they arrived, brown bears persisted in the Western Carpathians in a large, continuous population until the Middle Ages [15] and in most forested upland areas of present-day Slovakia *Corresponding author. OPEN ACCESS  J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 175 through to the 19th century [16]. Deforestation, over- hunting and eradication programmes then precipitated a catastrophic decline in range and numbers culminating in the 1930s, when it was estimated that only c.20 - 60 indi- viduals remained in an isolated, relict population [16]. A 30-year moratorium on hunting facilitated popula- tion recovery [17]. The bears continued to increase in number and reoccupy parts of their former range despite the resumption of limited hunting from the 1960s [18]. The Western Carpathian bear population now numbers several hundred individuals and extends across all moun- tain ranges of central and northern Slovakia [19]. How- ever, although it was thought that connection with the much larger Eastern Carpathian population was re-es- tablished in the 1980s [20], the two populations show a high degree of differentiation most likely resulting from genetic divergence during the c.100 years of their divi- sion and evidence of renewed gene flow is sparse [21]. Populations that have passed through a recent demo- graphic bottleneck would generally be expected to have lost genetic diversity through stochastic drift [22]. How- ever, medium-high levels of allelic variation have been found in the nuclear DNA of Western Carpathian bears and their level of genetic diversity seems to be within the range commonly observed in different populations of brown bears and other mammals [15,21,23], which sug- gests the population bottleneck may not have been quite so severe as feared by contemporary observers. The main aim of our study was to examine the degree of genetic diversity and geographic isolation of bears in a subpopu- lation near the periphery of their current distribution in the Western Carpathians and compare it with that of bears in core areas of the population and in select Euro- pean subpopulations. Fieldwork was conducted in the Strážovské vrchy mountain range (49˚23'702"N, 18˚73'46"E), which ex- tends over an area of c. 900 km2 in northwest Slovakia (Figure 1) and forms part of the Inner Western Carpa- thian Mountains. Brown bears were apparently absent until the mid 1960s [17,19] but re-colonised during the period 1967-1984, when the recovering Western Carpa- thian bear population expanded its range 40 km north- westwards [24]. Intense development in molecular technology, have moved bear ecology into a significant development in which genetic analyses can be performed with ease and with great informative value. Environmentalists can now routinely utilize genetic information obtained from DNA to formulate questions about the behavioural ecology and conservation genetics of bear populations. The highly variable microsatellites markers that have been analysed in this study offer an effective tool for individual identi- fication. These DNA fingerprints can be used in an eco- logical context for the dendrogram construction based on the degree of microsatellite profiles similarity to link- ing individuals. Then we can get a view on the degree of genetic relatedness of individuals grouped into clusters. Implementation of population genetic features of the PLA Strážovské vrchy provides an overview of the mi- gration rate between neighboring populations of the brown bears. 2. MATERIAL AND METHODS 2.1. Sample Collection and DNA Isolation A total of samples 57 (41.8%) out 140 from different sites of Strazovske vrchy Mts were collected. A 37 sam- ples collected during the year 2010 (faeces and hair sam- ples) and 20 samples from Strazovske vrchy territory (during the period 2007-2008) were examined. Suffi- cient brown bear DNA for analysis was obtained from 20 (2007-2008) out of 46 samples and 37 (2010) out of 94 samples collected in the field. A total of 45 different genotypes were identified among 28 samples from faeces and 29 from hair. DNA extraction from hairs was performed using 10% Chelex according to Kruckenhauser et al. [25], Depend- ing on availability and quality hairs with visible roots were used, DNA extractions from non-invasive samples were performed with the QIAamp DNA Stool Kit (QIAGEN) with a final elution volume of 100 µl. Samples were collected in the Strážovské Vrchy Pro- tected Landscape Area (Figure 1). The PLA covers c.300 km2 of which 78% is forest and 19% agricultural land. Altitude ranges from 315 to 1213 m a.s.l. On the basis of annual track surveys, opportunistic direct observations and camera trapping we estimated there to be approxi- mately 20 different bears using the area, although some of them may have had home ranges extending beyond the study area. 2.2. Microsatellites Analysis and Gender Identification Seven microsatellite loci Mu26, Mu64, G10B, G1D, G10L, Mu50 and Mu51 were amplified using poly- Figure 1. Location of the study area in relation to the dis- tribution of the Western Carpathian population of brown bears. Copyright © 2013 SciRes. OPEN ACCESS  J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 176 merase chain reaction [7] and fragment length (allele) analyses were carried out on eight-capillary sequencer (Genome Lab GeXP, BeckmanCoulter). Analyses were repeated in order to verify the reliability of individual allele length determination. Molecular sexing of the bears identified by DNA fin- gerprinting was assessed by amplification of the Sry fragment on theY chromosome [26]. In each cell, the autosomal microsatellite locus is twice as concentrated as the Sry gene. PCR products were checked on 2.0% aga- rose gels. Males show two bands, the microsatellite and Sry fragment, while females show only the ZFY/ZFX band. The primer pair used for gender determination is not bear-specific, but amplifies the Sry fragment in a wide variety of mammals, including humans [26]. To avoid contamination PCR reactions were set up by a fe- male investigator. DNA was extracted from hair using 10% Chelex solu- tion [25] and from faeces with the Qiamp DNA Stool® kit (Qiagen). To test for individuals, seven microsatellite loci (UaMU26, UaMU64, G10B, G1D, G10L, UaMU50 and UaMU51) were amplified in a nested polymerase chain reaction (PCR) [7]: a longer fragment of each locus was amplified prior to amplifying a more specific area. Two-step PCR procedures improve genotyping success rate and limit genotyping errors [27]. Observed (HO) and expected (HE) heterozygosity were calculated using Cervus 3.0 software (Field Genet- ics). Results were compared with genetic data from brown bears in core ranges of the Carpathian Mountains in Slovakia [21,28] and Romania [21] as well as in cen- tral Austria [25]. 2.3. Statistical Methods Observed (HO) and expected (HE) heterozygosity were calculated with CERVUS software. Descriptive statistics for each locus (mean number of alleles per locus, het- erozygosities and polymorphic information content (PIC) were computed from allele frequencies. The Fisher’s exact test was used to check for genotypic linkage dis- equilibrium for all pairs of loci by employing the Markov chain method, as implemented in GENEPOP [29]. De- viations from Hardy-Weinberg (HW) proportions were evaluated through the Weir and Cockerham’s [30] and Robertson and Hill’s [31] estimates of FIS to test for heterozygote deficit with Levene’s correction for small sample size, using the method described by Guo and Thompson [32]. The difference between Hardy-Weinberg heterozygo- sity (HE) and that expected from the observed number of alleles (HEQ) was tested under the assumption of muta- tion-drift equilibrium, given the sample size. Evidence for a recent reduction of population size is assumed when HE is significantly higher than HEQ. The patterns of microsatellite mutations appear to be extremely complex [33] and the evolution mode of bear microsatellite loci was not known. Therefore, calculations were made ac- cording to three models: the Infinite Alleles Mode [34], the strict Stepwise Mutation Model [34] and the Two Phase Model (TPM) with a 5% of multi-step changes: [35] Valdes, Slatkin and Freimer [36], an offshoot of the SMM, which accounts for addition or deletion of more than one repeat unit. The loci screened in our study were evaluated for their reliability and resolving power when performing parent- age tests. A simulation of parentage analysis was con- ducted using CERVUS version 2.0 [37]. That programme uses allele frequencies from the study population to run simulations of paternity inference when multiple males are non-excluded, allowing for user-defined inputs, such as the number of males that are candidates for paternity, the proportion of candidate males that are sampled and errors in genotyping. Success rates of parentage tests were derived assuming HW equilibrium in cases where one true parent was known and in cases where neither parent is known a priori, with 80 and 95% confidence levels. The simulations (10 000 repetitions) were con- ducted by changing parameters, e.g. the proportion of loci typed, the number of candidate parents, the fraction of candidates sampled and the level of potential labora- tory mistyping. 2.4. Division of Animals into Clusters Processing of acquired data and in particular in finding answers to the question of individuals distribution on the site led them to propose procedures for evaluation of microsatellite data was processed using our original software (Java script) (will be published in Oecologia Montana). Two methods were used Neighbor-joining and UPGMA (Unweighted Pair Group Method with Arith- metic Mean) for construction of clusters graphic presen- tation based on microsatellites data processing (Figure 2). Neighbor-joining method provides not only the topo- logy but also the branch lengths of the final tree. A pair of “neighbors” is a pair of animals connected through a single interior node in an unrooted, bifurcating tree [38]. UPGMA is a simple agglomerative or hierarchical clustering method often used for the creation of phenetic trees (phenograms). UPGMA assumes a constant rate of similarity between animals. UPGMA was initially de- signed for use in protein electrophoresis studies, but is currently most often used to produce guide trees for more sophisticated phylogenetic reconstruction algorithms. The mean observed heterozygosity (HO) among the seven loci examined in the year 2007-2008 was 0.70 and 0.53 in the year 2010, the mean expected heterozygosity (HE) 0.54 in the year 2007-2008 and 0.57 in the year Copyright © 2013 SciRes. OPEN ACCESS 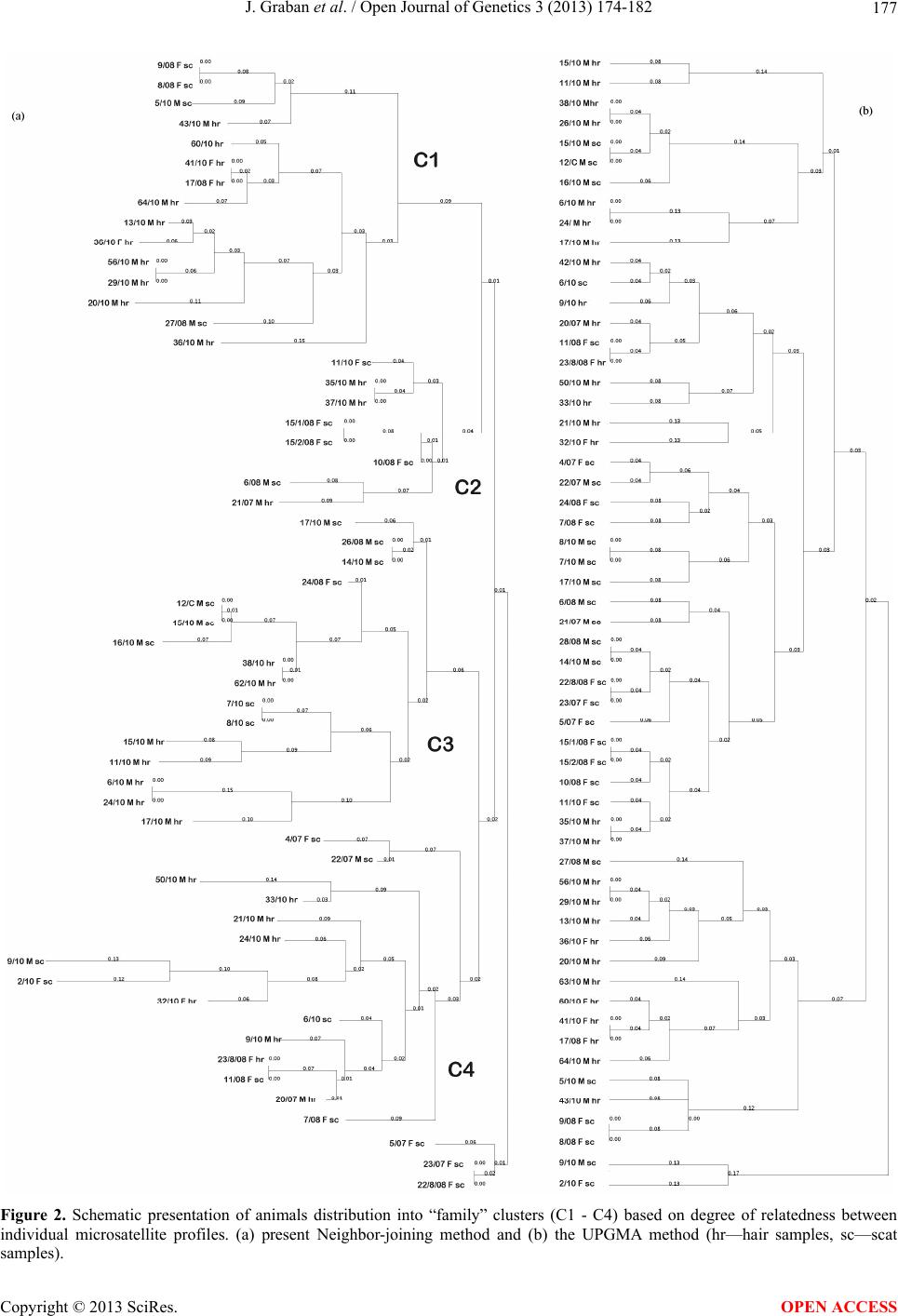 J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 Copyright © 2013 SciRes. 177 Figure 2. Schematic presentation of animals distribution into “family” clusters (C1 - C4) based on degree of relatedness between individual microsatellite profiles. (a) present Neighbor-joining method and (b) the UPGMA method (hr—hair samples, sc—scat samples). OPEN ACCESS  J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 178 2010. Only one locus (UaMU26) had a HO lower than HE in the year 2007-2008. In the year 2010 it was locus UaMU26, UaMU64, G10B and Ua MU50 (Table 1). Locus G1D had the most alleles (four), while UaMU26, G10B and G10L showed low allelic variability (two al- leles per locus). With the exception of G10L in the year 2007-2008 and UaMU64 in the year 2010, the observed number of alleles at each locus (na) was greater than the effective number of alleles (ne). The inbreeding coeffi- cient was negative for six out of the seven loci (mean value FIS = −0.27), the exception being UaMU26 (FIS = +0.33) (2007-2008). In the year 2010 the inbreeding co- efficient was negative only for three out seven loci (mean value FIS = 0.08). Table 1. Genetic variability of brown bears in the Strážovské Vrchy Mountains, Slovakia. na—observed number of alleles, ne—effec- tive number of alleles, PIC—Polymorphic information content, HO—observed heterozygosity, HE—expected heterozygosity, FIS— inbreeding coefficient. (a) 2007-2008 Locus na n e PIC H O H E F IS UaMU26 2 1.89 0.35 0.30 0.47 0.36 UaMU64 3 2.38 0.47 0.85 0.58 −0.47 G10B 2 1.49 0.27 0.40 0.33 −0.21 G1D 4 3.03 0.59 0.85 0.67 −0.27 G10L 2 2.04 0.38 0.80 0.51 −0.57 UaMU50 3 2.56 0.51 0.80 0.61 −0.31 UaMU51 3 2.63 0.53 0.90 0.62 −0.45 Mean 2.71 2.29 0.44 0.70 0.54 −0.27 St. Dev 0.76 0.52 0.10 0.24 0.12 0.31 (b) 2010 Locus na n e PIC H O H E F IS UaMU26 2 1.89 0.35 0.07 0.47 0.85 UaMU64 3 3.13 0.59 0.54 0.68 0.21 G10B 2 1.96 0.37 0.32 0.49 0.35 G1D 4 3.33 0.64 0.79 0.70 −0.13 G10L 2 1.92 0.36 0.75 0.48 −0.56 UaMU50 3 2.50 0.51 0.50 0.60 0.17 UaMU51 3 2.27 0.48 0.75 0.56 −0.34 Mean 2.71 2.43 0.47 0.53 0.57 0.08 St. Dev 0.76 0.59 0.12 0.27 0.10 0.47 (c) 2007-2010 Locus na n e PIC H O H E F IS UaMU26 2 2 0.38 0.14 0.50 0.72 UaMU64 3 2.94 0.58 0.67 0.66 −0.02 G10B 2 1.96 0.37 0.35 0.49 0.29 G1D 4 3.33 0.64 0.81 0.70 −0.16 G10L 2 2 0.37 0.81 0.50 −0.62 UaMU50 3 2.44 0.50 0.61 0.59 −0.04 UaMU51 3 2.27 0.48 0.77 0.56 −0.38 Mean 2.71 2,42 0.47 0.59 0.57 −0.03 St. Dev 0.76 0.53 0.11 0.26 0.08 0.44 Copyright © 2013 SciRes. OPEN ACCESS  J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 179 Locus UaMU26, UaMU64, G1D, G10L, UaMU50 andUaMU51 had the equal number of different allelic combinations (three) (Table 2). The least variability was found at loci G10B (2 alleles per locus in 2 different combinations) (2007-2008). Locus UaMU64 and UaMU50 had the equal number of different allelic combinations (four), followed by UaMU26, G1D, G10B and UaMU51 with three each. The least variability was found at loci G10L (2 alleles per locus in 2 different combinations) (2010). 3. DISCUSSION AND CONCLUSIONS The brown bear is a wide-ranging species exhibiting male-biased dispersal [39]. Adult males commonly use hundreds of square kilometres in their search for food and mating opportunities and dispersing subadult male brown bears may roam over areas up to 12,000 km2 [1]. Such movements facilitate gene flow and, in the case of dispersing young males, there is evidence that it operates as a mechanism to avoid inbreeding [39]. The relatively high level of heterozygosity and low degree of inbreed- ing we found in bears in Strážovské vrchy (this study) suggests that the subpopulation is not geographically isolated and gene exchange with other segments of the population has been maintained. The most obvious potential source of migration into the study area is the Malá Fatra mountain range, which lies immediately to the east (Figure 1) and has a high density of bears [19,28]. The eastern edge of the Strá- žovské vrchy seems to present the least obstruction to wildlife movement in and out of the study area, as the unfenced primary road I/64 passes through a heavily forested landscape for the 20-km section between Ra- jecká Lesná and Kľačno, forest availability being the most important habitat constraint on bear distribution in the Western Carpathians [40]. There are, however, sev- eral other nearby ranges from which bears could rea- sonably be expected to reach Strážovské vrchy, including Vtáčnik, Kremnické vrchy and Veľká Fatra. To elucidate their movements without the need to live-trap, immobilise and fit animals with telemetry equipment, DNA profiling or ‘genetic fingerprinting’ in combination with GPS localisation of sampling sites en- ables individuals to be identified and tracked non-inva- sively [7,28]. This may help to identify important biocor- ridors in need of protection or improvement [41,42]. Ef- fective immigration can also be estimated from changes in observed and expected heterozygosity and heterozy- gote excess [13,43,44]. Although the results of the present study showed rela- tively high variability (increasing in the year 2010), mi- crosatellite analysis of brown bears in Malá Fatra Na- tional Park [28] found higher numbers of alleles per lo- cus (Table 3). Moreover, the difference between ob- served and expected heterozygosity was greater in Malá Fatra than in Strážovské vrchy and low values of FIS for each locus demonstrated a higher occurrence of het- erozygotes. Higher variability has also been found in central Slovakia, northern Slovakia and Romania [21]. The leading edge of an expanding population might be expected to become less diverse as a result of a series of genetic bottlenecks [22]. However, two loci analysed in the small central Austrian bear population [25] showed a similar level of variability to that found in Malá Fatra and central Slovakia, even though all genotyped indi- viduals were descended from just four founders. These four individuals arrived in the area 20 - 40 years ago, which is more recently than bears re-colonised the Strážovské vrchy, and there is no evidence of there hav- ing been any subsequent immigration. The founders of the central Austrian bear population Table 2. Allelic combinations found at seven microsatellite loci in genomic DNA from brown bears in the Strážovské Vrchy Moun- tains. Slovakia. The most frequently occurring combination for each locus is shown in bold. AC—allelic combinations detected. (a) 2007-2008 AC UaMU26 UaMU64 G10B G1D G10L UaMU50 UaMU51 1 182/182 177/194 114/114 171/221 143/143 118/121 110/110 2 182/198 184/194 114/126 179/179 143/171 121/125 110/136 3 198/198 184/184 179/208 171/171 125/125 110/116 (b) 2010 AC UaMU26 UaMU64 G10B G1D G10L UaMU50 UaMU51 1 182/182 177/177 114/114 171/221 143/143 118/121 110/110 2 182/198 177/194 114/126 179/179 143/171 121/121 110/136 3 198/198 184/194 126/126 179/208 121/125 110/116 4 184/184 125/125 Copyright © 2013 SciRes. OPEN ACCESS  J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 180 Table 3. The selected Carpatian Mts and central Austria Brown bear subpopulations genetic variability comparison. na—ob- served number of alleles, HO—observed heterozygosity, HE— expected heterozygosity, FIS—inbreeding coefficient of Small Fatra [28]—SF, Northern Slovakia [21]—NS, Central Slovakia [21]—CS, Central Austria [25]—CA and Romania [21]—R. Locality and locus na HO H E F IS SF-UaMU26 4 0.65 0.44 −0.48 CA-UaMU26 4 0.65 0.57 −0.13 SF-UaMU64 9 0.74 0.46 −0.61 NS-G10B 5 0.59 0.63 0.06 CS-G10B 4 0.60 0.63 0.05 CA-G10B 4 1 0.69 −0.45 R-G10B 8 0.76 0.75 −0.01 SF-G1D 4 0.63 0.43 −0.47 NS-G1D 6 0.68 0.79 0.14 CS-G1D 6 0.76 0.76 0.00 R-G1D 7 0.71 0.73 0.03 NS-G10L 6 0.41 0.48 0.16 CS-G10L 6 0.60 0.61 0.02 R-G10L 8 0.79 0.84 0.06 NS-UaMU50 6 0.76 0.75 −0.01 CS-UaMU50 6 0.72 0.65 −0.11 R-UaMU50 8 0.80 0.82 0.02 NS-UaMU51 6 0.52 0.74 0.30 CS-UaMU51 7 0.73 0.82 0.11 R-UaMU51 7 0.72 0.78 0.08 originated in Slovenia and are therefore part of the west- ern lineage (subclade 1b), whereas Slovakia’s bears be- long to the eastern lineage known as subclade 3a [45]. There are bears of both mtDNA lineages in Romania [5] but the differentiation is not reflected in nuclear loci, perhaps due to male-mediated gene flow and female philopatry [21]. The genetic diversity of brown bears is highest in Romania [46], where the population has never fallen below 800 individuals, and is also high in Slovenia [46] as well as neighbouring Croatia [47]. Our results represent a comparative study of a sub- population which has hitherto received little attention from researchers. However, the brown bear is one of the best-studied mammalian species [13]. In the last two decades there has been a proliferation of genetics studies mapping populations at different geographical scales from regional [7,8,48] to continental [49,50]. Standard- ised procedures have been developed for sampling and analysis in order to facilitate comparisons between stud- ies [46,51,52]. Non-invasive genetic methods, especially appropriate for use with elusive species in small, endan- gered populations or over large areas, are now available to allow identification of individual animals, census populations and monitor migration and gene flow [13]. The potential for further work building on our study is therefore substantial. Knowledge of population size, dis- tribution, social and sexual structure, home range and population trend on the local level as well as migration is crucial for the proper conservation and management of species within and between protected areas. 4. ACKNOWLEDGEMENTS This study was supported by Structural Funds of EU Project of the Agency of the Ministry of Education, Science, Research and Sport of the Slovak Republic, Bratislava, ITMS No. 26110230078. REFERENCES [1] Swenson, J.E., Dahle, B., Gerstl, N. and Zedrosser, A. (2000) Action plan for the conservation of the brown bear in Europe (Ursus arctos)—Convention on the Conserva- tion of European Wildlife and Natural Habitats (Bern Convention). Nature and Environment No. 114, Council of Europe Publishing, Strasbourg. [2] Sommer, R.S. and Benecke, N. (2005) The recolonization of Europe by brown bears Ursus arctos Linnaeus, 1758 after the Last Glacial Maximum. Mammal Review, 35, 156-164. doi:10.1111/j.1365-2907.2005.00063.x [3] Breitenmoser, U. (1998) Large predators in the Alps: The fall and rise of man’s competitors. Biological Conserva- tion, 83, 279-289. doi:10.1016/S0006-3207(97)00084-0 [4] Zedrosser, A., Dahle, B., Swenson, J.E. and Gerstl, N. (2001) Status and management of the brown bear in Europe. Ursus, 12, 9-12. [5] Zachos, F.E., Ottoa, M., Unici, R., Lorenzini and R., Hartl, G.B. (2008) Evidence of a phylogeographic break in the Romanian brown bear (Ursus arctos) population from the Carpathians. Mammalian Biology, 73, 93-101. doi:10.1016/j.mambio.2007.02.007 [6] Randi, E., Gentile, L., Boscagli, G., Huber, D. and Roth, H.U. (1994) Mitochondrial DNA sequence divergence among some west European brown bear (Ursus arctos L.) populations. Lessons for conservation. Heredity, 73, 480- 489. doi:10.1038/hdy.1994.146 [7] Taberlet, P., Camarra, J.J., Griffin, S., Uhrès, E., Hanotte, O., Waits, L.P., Dubois-Paganon, C., Burke, T. and Bou- vet, J. (1997) Noninvasive genetic tracking of the endan- gered Pyrenean brown bear population. Molecular Ecol- ogy, 6, 869-876. doi:10.1111/j.1365-294X.1997.tb00141.x [8] Lorenzini, R., Posillico, M., Lovari, S. and Petrella, A. (2004) Non-invasive genotyping of the endangered Ap- ennine brown bear: A case study not to let one’s hair down. Animal Conservation, 7, 199-209. doi:10.1017/S1367943004001301 [9] Taberlet, P. and Bouvet, J. (1994) Mitochondrial DNA polymorphism, phylogeography, and conservation genet- ics of the brown bear Ursus arctos in Europe. Proceed- ings of the Royal Society B: Biological Sciences, 255, 195-200. Copyright © 2013 SciRes. OPEN ACCESS  J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 181 [10] Kohn, M., Knauer, F., Stoffella, A., Schroder, W. and Paabo, S. (1995) Conservation genetics of the European brown bear—A study using excremental PCR of nuclear and mitochondrial sequences. Molecular Ecology, 4, 95- 103. doi:10.1111/j.1365-294X.1995.tb00196.x [11] Korsten, M., Ho, S.Y.W., Davison, J., Pähn, B., Vulla, E., Roht, M., et al. (2009) Sudden expansion of a single brown bear maternal lineage across northern continental Eurasia after the last ice age: A general demographic model for mammals? Molecular Ecology, 18, 1963-1979. doi:10.1111/j.1365-294X.2009.04163.x [12] Valdiosera, C.E., Garcia, N., Anderung, C., Dalen, L., Cregut-Bonnoure, E., Kahlke, R.-D., Stiller, M., Brand- ström, M., Thomas, M.G., Arsuaga, J.L., Götherström, A. and Barnes, I. (2007) Staying out in the cold: Glacial refugia and mitochondrial DNA phylogeography in an- cient European brown bears. Molecular Ecology, 16, 5140-5148. doi:10.1111/j.1365-294X.2007.03590.x [13] Swenson, J.E., Taberlet, P. and Bellemain, E. (2011) Genetics and conservation of European brown bears Ur- sus arctos. Mammal Review, 2, 87-98. doi:10.1111/j.1365-2907.2010.00179.x [14] Saarma, U., Ho, S.Y.W., Pybus, O.G., Kaljuste, M., Tu- manov, I.L., Kojola, I., Vorobiev, A., Markov, N.I., Saveljev, A.P., Valdmann, H., Lyapunova, E.A., Abra- mov, A.V., Männil, P., Korsten, M., Vulla, E., Pazetnov, S.V., Pazetnov, V.S., Putchkovskiy, S.V. and Rõkov, A.M. (2007) Mitogenetic structure of brown bears (Ursus arctos L.) in northeastern Europe and a new time frame for the formation of European brown bear lineages. Mo- lecular Ecology, 16, 401-413. doi:10.1111/j.1365-294X.2006.03130.x [15] Hartl, G.B. and Hell, P. (1994) Maintenance of high lev- els of allelic variation in spite of a severe bottleneck in population size: The brown bear (Ursus arctos) in the Western Carpathians. Biodiversity and Conservation, 3, 546-554. doi:10.1007/BF00115160 [16] Jamnický J. (1993) The hunt of the brown bear and European wolf in Slovakia a hundred years ago. Folia Venatoria, 23, 221-229. (in Slovak with English Sum- mary) [17] Hell, P. and Slamečka, J. (1999) The bear in the Slovak Carpathians and in the world. PaRPress, Bratislava, 150p. (in Slovak). [18] Sabadoš, K. and Šimiak, M. (1981) Distribution and hunting management of the brown bear (Ursus arctos L.) in Slovakia. Folia Venatoria, 10-11, 15-35. (in Slovak) [19] Rigg, R. and Adamec, M. (2007) Status, ecology and management of the brown bear (Ursus arctos) in Slova- kia. Slovak Wildlife Society, Liptovský Hrádok, 128p. [20] Jakubiec, Z. (2001) The brown bear Ursus arctos L. in the Polish part of the Carpathians. Polska Akademia Nauk, Kraków, 108p. (in Polish with English Summary) [21] Straka, M., Paule, L., Ionescu, O., Štofík, J. and Adamec, M. (2012) Microsatellite diversity and structure of Car- pathian brown bears (Ursus arctos): Consequences of human caused fragmentation. Conservation Genetics, 13, 153-164. doi:10.1007/s10592-011-0271-4 [22] Davison, J., Ho, S.Y.W., Bray, S.C., Korsten, M., Tam- meleht, E., Hindrikson, M., Østbye, K., Østbye, E., Lauritzen, S. E., Austin, J., Cooper, A. and Saarma, U. (2011) Late-Quaternary biogeographic scenarios for the brown bear (Ursus arctos), a wild mammal model species. Quaternary Science Reviews, 30, 418-430. doi:10.1016/j.quascirev.2010.11.023 [23] Paule, L., Krajmerová, D., Urban, P. and Adamec, M. (2006) Contribution to genetic diversity of brown bear (Ursus arctos L.) from the Western Carpathians. Výskum a Ochrana Cicavcov na Slovensku, 7, 115-121. (in Slovak with English Abstract) [24] Janík, M., Voskár, J. and Buday, M. (1986) Present dis- tribution of the brown bear (Ursus arctos) in Czechoslo- vakia. Folia Venatoria, 16, 331-352. (in Slovak with English Abstract) [25] Kruckenhauser, L., Rauer, G., Däubl, B. and Haring, E. (2009) Genetic monitoring of a founder population of brown bears (Ursus arctos) in central Austria. Conserva- tion Genetics, 10, 1223-1233. doi:10.1007/s10592-008-9654-6 [26] Brya, J. And Konečný, A. (2003) Fast sex identification in wild mammals using PCR amplification of the SRY gene. Folia Zoologica, 52, 269-274. [27] Bellemain, E. and Taberlet, P. (2004) Improved noninva- sive genotyping method: application to brown bear (Ur- sus arctos) faeces. Molecular Ecology Notes, 4, 519-522. doi:10.1111/j.1471-8286.2004.00711.x [28] Janiga, M., Fečková, M., Korňan, J., Kalaš, M., La- budíková, I., Matiaško, K. and Londa, P. (2006) Prelimi- nary results on genetic tracking of the Brown Bear (Ursus arctos) individuals in the Malá Fatra National Park (Slo- vakia). Oecologia Montana, 15, 24-26. [29] Raymond, M. and Rousset, F. (1995) An exact test for population diferentiation. Evolution, 49, 1283-1286. doi:10.2307/2410454 [30] Weir, B.S. and Cockerham, C.C. (1984) Estimating F- statistics for the analysis of population structure. Evolu- tion, 38, 1358-1370. doi:10.2307/2408641 [31] Robertson, A. and Hill, W.G. (1984) Deviations from Hardy-Weinberg proportions: Sampling variances and use in estimation of inbreeding coefficients. Genetics, 107, 703-718. [32] Guo, S.W. and Thompson, E.A. (1992) Performing the exact test of Hardy-Weinberg proportion for multiple al- leles. Biometrics, 48, 361-372. doi:10.2307/2532296 [33] Goldstein, D.B. and Schlotterer, C. (1999) Microsatellites: Evolution and applications. Oxford University Press, Oxford. [34] Kimura, M. and Crow, J. (1964) The number of alleles that can be maintained in a finite population. Genetics, 49, 725-738. [35] Valdes, A.M., Slatkin, M. and Freimer, N.B. (1993) Al- lele frequencies at microsatellite loci: The stepwise muta- tion model revisited. Genetics, 133, 737-749. [36] Di Rienzo, A., Peterson, A.C., Garza, J.C., Valdes, A.M., Slatkin, M. and Freimer, N.B. (1994) Mutational proc- esses of simple-sequence repeat loci in human popula- tions. Proceedings of the National Academy of Sciences Copyright © 2013 SciRes. OPEN ACCESS  J. Graban et al. / Open Journal of Genetics 3 (2013) 174-182 Copyright © 2013 SciRes. 182 OPEN ACCESS of the United States of America, 91, 3166-3170. doi:10.1073/pnas.91.8.3166 [37] Marshall, T.C., Slate, J., Kruuk, L.E.B. and Pemberton, J.M. (1998) Statistical confidence for likelihood-based paternity inference in natural populations. Molecular Ecology, 7, 639-655. doi:10.1046/j.1365-294x.1998.00374.x [38] Saitou, N. and Nei, M. (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Molecular Biology and Evolution, 4, 406-425. [39] Zedrosser, A., Støen, O.-G., Sæbø, S. and Swenson, J.E. (2007) Should I stay or should I go? Natal dispersal in the brown bear. Animal Behaviour, 74, 369-376. doi:10.1016/j.anbehav.2006.09.015 [40] Fernández, N., Selva, N., Yuste, C., Okarma, H. and Ja- kubiec, Z. (2012) Brown bears at the edge: Modelling habitat constrains at the periphery of the Carpathian population. Biological Conservation, 153, 134-142. doi:10.1016/j.biocon.2012.04.013 [41] Finďo, S., Skuban, M. and Koreň, M. (2007) Brown bear corridors in Slovakia. Carpathian Wildlife Society, Zvo- len. [42] Pérez, T., Naves, J., Vázquez, J.F., Seijas, J., Corao, A., Albornoz, J. and Domínguez, A. (2010) Evidence for im- proved connectivity between Cantabrian brown bear subpopulations. Ursus, 21, 104-108. doi:10.2192/09SC018.1 [43] Waits, L.P., Taberlet, P., Swenson, J.E., Sandegren, F. and Franzén, R. (2000) Nuclear DNA microsatellite analysis of genetic diversity and gene flow in the Scan- dinavian brown bear (Ursus arctos). Molecular Ecology, 9, 421-431. doi:10.1046/j.1365-294x.2000.00892.x [44] Tallmon, D.A., Bellemain, E., Swenson, J. and Taberlet, P. (2004) Genetic monitoring of Scandinavian brown bear effective population size and immigration. Journal of Wildlife Management, 68, 960-965. doi:10.2193/0022-541X(2004)068[0960:GMOSBB]2.0.C O;2 [45] Paunović, M. and Ćirović, D. (2006) Viability increase and recovery of brown bear (Ursus arctos L. 1758) popu- lation in northeastern Serbia—A Feasibility Study. Fac- ulty of Biology. University of Belgrade, Belgrade. [46] Skrbinšek, T., Jelenčič, M., Waits, L.P., Potočnik, H., Kos, I. and Trontelj, P. (2012) Using a reference popu- lation yardstick to calibrate and compare genetic diversity reported in different studies: An example from the brown bear. Heredity, 109, 299-305. [47] Kocijan, I., Galov, A., Ćetković, H., Kusak, J., Gomerčić, T. and Huber, Đ. (2011) Genetic diversity of Dinaric brown bears (Ursus arctos) in Croatia with implications for bear conservation in Europe. Mammalian Biology- Zeitschrift fur Saugetierkunde, 76, 615-621. [48] De Barba, M., Waits, L.P., Garton, E.O., Genovesi, P., Randi, E., Mustoni, A. and Groff, C. (2010) The power of genetic monitoring for studying demography, ecology and genetics of a reintroduced brown bear population. Molecular Ecology, 19, 3938-3951. doi:10.1111/j.1365-294X.2010.04791.x [49] Paetkau, D., Waits, L.P., Clarkson, P., Craighead, L., Vyse, E.R., Ward, R. and Strobeck, C. (1998) Variation in genetic diversity across the range of North American brown bears. Conservation Biology, 12, 418-429. [50] Tammeleht, E., Remm, J., Korsten, M., Davidson, J., Tumanov, I., Saveljev, A., Männil, P., Kojola, I. and Saarma, U. (2010) Genetic structure in large, continuous mammal populations: The example of brown bears in northwestern Eurasia. Molecular Ecolo gy, 19, 5359-5370. doi:10.1111/j.1365-294X.2010.04885.x [51] Aarnes, S.G., Bellemain, E., Eiken, H.G. and Wartainen, I. (2009) Interlaboratory comparison of genetic profiles of brown bears from Sweden (Laboratoire d’Ecologie Alpine) and Norway (Bioforsk Svanhovd). Bioforsk Re- port, 4, 133. [52] Karamanlidis, A.A., De Barba, M., Georgiadis, L., Groff, C., Jelinčić, M., Kocijan, I., Kruckenhauser, L., Rauer, G., Sindičić, M., Skrbinšek, T. and Huber, D. (2009) Com- mon guidelines for the genetic study of brown bears (Ur- sus arctos) in southeastern Europe. LCIE, Athens.
|