Paper Menu >>
Journal Menu >>
 American Journal of Plant Sciences, 2010, 1, 77-86 doi:10.4236/ajps.2010.12010 Published Online December 2010 (http://www.SciRP.org/journal/ajps) Copyright © 2010 SciRes. AJPS 77 Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Akan Das1, Tapan Kumar Mondal1,2* 1Biotechnology Laboratory, Faculty of Horticulture, Uttar Banga Krishi Viswavidyalaya, Cooch Behar, West Bengal, India; 2National Research Center of DNA Fingerprinting, National Bureau of Plant Genetic Resources, New Delhi, India. Email: *mondaltk@yahoo.com Received November 2nd, 2010; revised November 15th, 2010; accepted November 22nd, 2010. ABSTRACT MicroRNAs (miRNAs) are a class of ~22 nucleotides long non coding RNA molecules which play an important role in gene regulation at the post transcriptional level. The conserved nature of miRNAs provides the basis of new miRNA identification throug h homology search. In an a ttempt to iden tify new conserved miRNAs in tea , previously kno wn plant miRNAs were used for searching their homolog in a tea Expressed Sequence Tags and full length nucleotide sequence database. The sequences showing homolog no more than four mismatches were predicted for their fold back structures and passed through a series of filtration criteria, finally led us to identify 13 conserved miRNAs in tea belonging to 9 miRNA families. A total of 37 potential target genes in Arabidopsis were identified subsequently for 7 miRNA families based on their sequence complementarity which encode transcription factors (8%), enzymes (30%) and transporters (14%) as well as other proteins involved in physiological and metabolic processes (48%). Overall, our findings will accelerate the way for further researches of miRNAs and their functions in tea. Keywords: Camellia sinensis, Computational Identification, Expressed Sequence Tags, microRNA, Targets 1. Introduction MicroRNAs (miRNAs) are short (~22 nt), endogenous non coding RNAs that play an important role in many biological processes [1]. They are generated from long precursor molecules which can fold into hairpin second- dary structures [2]. Mature miRNAs bind to the comple- mentary sites on target mRNAs and repress post tran- scriptional gene expression in both animals and plants [3, 4]. In plants, miRNAs are involved in diverse aspects of plant growth and development such as leaf morphology and polarity, root formation, transition from juvenile to adult vegetative phase and vegetative to flowering phase, flowering time, floral organ identity and reproduction [5, 6]. They are also found to be involved in response to pathogen invasion [7], hormone signaling [8,9], envi- ronmental stress [10,11] and promotion of anti-viral de- fence [12]. Expressed Sequence Tags (ESTs) are complementary DNA (cDNA) sequences, usually 200-500 bp in length that represents the expressed portions of genes. Therefore, ESTs can be used in gene identification, expression pro- filing and polymorphism analysis [13]. The EST se- quencing projects have been enormously successful in the framework of many genome projects. The EST se- quences are being used intensely as a source of informa- tion for the discovery of new genes whose functions can tentatively be deduced from their sequence and verified experimentally. Recently, in silico identification of miRNAs in various plant species have been done by EST analysis [14-17]. The biogenesis of miRNAs suggests that it is possible to find new miRNAs by homology searching of known miRNAs in ESTs. Hence, EST ana- lysis makes it possible for studying conserved miRNAs and their functions in species whose genome sequences have not been well known [10,18,19]. There are both experimental and computational ap- proaches for the investigation of plant miRNAs. The computational approach has been proved to be faster, affordable and more effective. Most of the miRNAs in miRBase [20] have been contributed through computa- tional approach only. The computational approach has been developed on the principle of looking conserved sequences between different species which can fold into  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 78 hairpin secondary structures [10]. In recent years, a number of programs and bioinformatics tools have been developed and used successfully for the identification and analysis of miRNAs and their targets [21,22]. Tea is one of the most important non-alcoholic bever- age drinks worldwide and gaining further popularity as an important ‘health drink’. Despite the limited genome resources of tea (Camellia sinensis), published EST and full length nucleotide sequences in GenBank (http://www. ncbi.nlm.nih.gov/genbank/) has provided the scope to get more genetic information. In this study, new miRNAs were mined in local tea sequence database for the pur- pose of understanding their roles in regulating growth and development, metabolism and other physiological processes in tea. 2. Methods 2.1. Collection of Reference miRNAs, Full Length Nucleotides and EST Sequences All available plant miRNAs and their fold back sequences were obtained from miRBase (http://www.mirbase.org/) on May, 2010. The homolog miRNAs were eliminated and the rest were defined as reference for searching tea miRNAs. Tea nucleotide and EST sequences (14819 as on May, 2010) were downloaded from NCBI’s nucleo- tide and dbEST database (http://www.ncbi. nlm.nih.gov/). All redundant and poor quality sequences were elimi- nated and created a local nucleotide database. 2.2. Potential miRNAs and Their Precursors The procedure for searching conserved tea miRNA ho- mologues is summarized in Figure 1. The reference se- quences were used as a query for homology search against our local tea nucleotide sequence database at e-value threshold < 0.01 using BLAST + 2.2.22 program [23]. The target sequences with no more than four mismatches were considered for secondary structure prediction using Mfold v 3.2 [24]. The precursor sequences were searched at 50 nucleotides upstream or downstream from the loca- tion of mature miRNAs with an increament of 10 nucleo- tides. While selecting a RNA sequence as a candidate miRNA precursor, following criteria were used accord- ing to Zhang et al. [1] with minor modifications as: 1) a RNA sequence can fold into an appropriate stemloop hairpin secondary structure, 2) a mature miRNA se- quence site in one arm of the hairpin structure, 3) miRNAs Collect all available tea ESTs and genes Exclude redundant and poor quality ESTs Create a local nucleotide database Collect all known plant miRNAs Exclude homolog miRNAs Sequences ≤4 mismatches Total unique hits = 22 Extract pre-miRNAs and predict secondary structures using Mfold Finally select tea miRNAs = 13 Predict target using miRU against Arabidopsis ∆G ≤ -18kcal/mol Bulge sige ≤ 7 Mature miRNAs in the stem regions Figure 1. An overview of different steps involved in miRNA and their target prediction.  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 79 had less than seven mismatches with the opposite miRNA sequence in the other arm, 4) no loop or break in miRNA sequences, 5) predicted secondary structures had higher negative energy MFEs (≤–18 kcal/mol), and iv) 40-70% A + U contents. 2.3. Prediction of Targets As for tea, since only few gene sequences available, we used Arabidopsis as a reference system for finding the targets of the candidate miRNAs. The predicted tea miRNAs were used as query against the Arabidopsis thaliana DFCI gene index (AGI) release 13 using miRU (http://bioinfo3.noble.org/psRNATarget/) following the criteria as 1) maximum expectation value 3; 2) multiplic- ity of target sites 2; 3) range of central mismatch for translational inhibition 9-11 nucleotide; 4) maximum mismatches at the complementary site ≤4 without any gaps. 2.4. Phylogenetic Analysis Due to the conserved nature of small RNAs, orthologue discovery can be done through bioinformatics analysis. We analysed tea small RNA conservation with their orthologues. A homology search of candidate tea miRNAs was done against all plant miRNAs using NCBI stand- alone BLAST [23] allowing maximum of 3 mismatches and e-value <0.001. The corresponding precursor se- quences of homolog small RNA’s were identified and collected (Appendix). The collected sequences of diverse plant species were aligned with homolog tea miRNA using Clustal W [25]. A query of tea small RNAs against known miRNA families (miRBase, release 15, http://www.mirbase.org/) allowed us to identify 3 previously reported large fami- lies. The precursor sequences of three known family members were selected along with respective precursor sequences of tea (Appendix). Then, the maximum likely- hood trees were constructed for each family based on Tamura-Nei model [26] with default bootstrap values using MEGA 4.0 [27] to illustrate the evolutionary rela- tionships among the members of the family. 2.5. Nomenclature of miRNAs The predicted miRNAs were named in accordance with miRBase [28]. The mature sequences are designated ‘miR’, and the precursor hairpins are labeled as ‘mir’ with the prefix ‘csi’ for C. sinensis, ‘cas’ for C. assamica and ‘cja’ for C. japonica. In the cases where distinct precur- sor sequences have identical miRNAs with different mismatch pattern, they were named as csi-mir-1-a and csi-mir-1-b. 3. Results 3.1. Prediction of miRNAs A total of 14,819 sequences containing 2,023 full length nucleotides and 12,796 ESTs were obtained from Gen- Bank. Out of these, 22 sequences had less than five mis- matches with previously known plant miRNAs. After carefully evaluating the hairpin structures using the crite- ria mentioned in the method, 13 small RNAs were finally identified from different species of Camellia namely sinensis, japonica and assamica. Details of the predicted miRNAs such as source sequences, location in the source sequences, length of precursor sequences and their minimum folding free energies and A+ U content are tabulated below (Table 1). A total of 9 miRNAs were Table 1. Details of the predicted miRNAs in tea. New MiRNAs NS Gene ID Strand SP EPMEMature MiRNAs E Value P L A + U (%) MFE csi-miR 408 EST 206583693 3’ 137 11718/21CUGCACUGCCUCUUCCCUGAG 0.001 336 45.24-120.1 csi-miR1171 EST 171355265 5’ 286 30822/23UGGAGUGGAGUGAAGUGGAGUGG 3E-04 181 56.98-45.97 csi-miR414a EST 206583641 3’ 757 63718/21UCUUCCUCAUCAUCAUCUUCU 0.001 663 57.32-63.18 csi-miR414d EST 284026209 3’ 186 16620/21UCAUCGUCAUCGUCAUCAUCU 0.004 193 61.14-37.72 csi-miR414f EST 212378632 5’ 122 14220/21UCAUCAUCAUCAUCAUCUUCA 6E-05 68 57.35-18.5 cas-miR1122 FL 214011104 5’ 214 23720/24UACUCCCUCCGUCCCAAAAUAAUG 6E-05 294 69.83-91.23 csi-miR414g EST 51453040 3’ 474 45418/21CCUUCCUCAUCAUCAUCGUCC 0.001 70 45.71-25.2 csi-miRf10132-akr EST 51453383 3’ 58 3425/25GCGAGCUUCUCGAAGAUGUCGUUGA 9E-08 200 49.00-69.5 cja-miR2910 FL 1777723 5’ 1262 128221/21UAGUUGGUGGAGCGAUUUGUC 1E-05 301 49.83-91.0 csi-miR2914 FL 34787361 5’ 345 36722/23UAUGGUGGUGACGGGUGACGGAG 5E-06 65 49.23-20.9 cas-miRf10185-akr EST 221071827 3’ 232 21217/21GAAAGGGGAAAACAUUGUAGC 0.004 139 48.92-51.1 cas-miR11590-akr EST 212379609 3’ 113 9417/20UUUUGGUGUGCCUUCAACCU 0.003 75 53.33-23.8 csi-miR414h EST 295345415 3’ 79 5817/21UCAUCCUCAUCAUCGUCAGAA 0.004 644 55.36-86.83 NS = Nucleotide Source, FL = Full-Length, SP = Start Point, EP = End Point, ME = Match Extent, PL = Pre-miRNA Length, MEF = Minimal Free Energy  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 80 predicted from ESTs whereas 4 were from full length nucleotide sequences. Five of them were located in the direct strand and the rest were in indirect strand. The newly identified precursor miRNAs have minimum folding free energies (mfe) ranging from –186.83 to –18.5 kcal/mol, with an average of about –72.69 kcal/mol and the A + U content were ranges from 45.24 to 69.83% with an average of 53.79%. The length of the precursors ranges from 65 to 663 nt with an average of 248 nt and mature sequences ranges from 20 to 25 nt. The newly predicted two tea miRNA (cja-miR2910, csi-miRf10132-akr) sequences were perfectly (100%) matched with the corresponding homologues of populus and rice, whereas the remaining 11 mature miRNA se- quences differ by 1 to 4 nucleotides from their homo- logues. All the mature miRNAs were found in the stem portion of the hairpin structures (Figure 2) containing less than 7 mismatches in the other arm without break or loop inside the se- quences. It was found that tea miRNA (csi-miR408) has been conserved with diverse plant spe- cies (Figure 3) from monocotyledonous plants such as rice, maize to dicotyledonous plants such as populous.  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 81 Figure 2. Predicted hairpin secondary structures of pre-miRNAs. MiRNAs are highlighted (red color) in the stem portion. Figure 3. Conservation of tea miRNA (csi-miR408) with diverse plant species. Conserved portion is highlighted. Abbreviated names are given in full in appendix.  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 82 3.2. Phylogenetic Analysis The newly identified tea miRNAs belong to 9 miRNA families including three known independent large miRNA families (mir 408, mir414 and mir1122). There are one tea miRNA namely csi-miR 408 and cas-miR1122 in each family of mir-408 and mir-1122, respectively. However, five members of family mir408 were found in tea (csi-miR414a, csi-miR414d, csi-miR414f, csi-miR414g, and csi-miR414h). The comparison of the predicted miRNA precursor sequences with other members in the same family showed that most members could be found to have a high degree of sequence similarity with others. The phylogenetic trees among the members of each fam- ily illustrated the evolutionary relationships of tea miRNAs (Figure 4). 3.3. Target Prediction A total of 37 potential targets were identified for the 7 predicted miRNA families which include 11 miRNAs based on their perfect or nearly perfect complementarity with their target sequences in Arabidopsis (Table 2, Figure 5). For all the miRNAs, single binding site was found in the targets without any gaps in the complement- tary region and expectation value ranges from 0 to 3. These potential miRNA targets were belonged to a num- ber of gene families that involved in different biological functions such as regulation of cell cycle, metal ion transportation, starch metabolic processes etc. There were 8% of genes encoding transcription factors, 30% of (a) (b) (c) Figure 4. Phylogenetic relationships among the miRNA family members of (a) miRNA414 (b) miRNA1122 (c) miRNA408. genes encoding different enzymes and 14% of genes en- coding transporters as well as 48% of genes encoding various proteins of physiological and metabolic proc- esses (Table 2). The miRNA family ‘miR414’ showed the highest 30 numbers of independent target genes fol- lowed by ‘miR408’ family with 2 numbers of target genes. The rest miRNA families were with single target genes in Arabidopsis (Table 2). The ‘miR1171’ and ‘miR1122’ miRNA family members did not bind to any target sequences within our filtration criteria. 4. Discussion With the availability of sequence resources in public da- tabases, computer based miRNA identification methods have been focused more and more in the recent years due to its advantages of low cost and high efficiency. Se- quence and structure homologies are the main theory behind the computer-based approach for miRNAs pre- diction. At present, four kinds of databases namely ge- nome, GSS, EST and nucleotide are mainly used for plant miRNA mining. Considering the unavailability of genome and genomic survey sequences of tea, both EST and nucleotide databases were mined for miRNA identi- fication. The number and sorts of miRNAs predicted in tea supported the fact that software-based approach is feasible and effective [3,10,14]. The idenfied new miRNAs were belonged to 9 fami- lies where miR414 family has 5 members and the rests have single member in each. This familial distribution of miRNAs was also observed in Arabidopsis, rice and maize 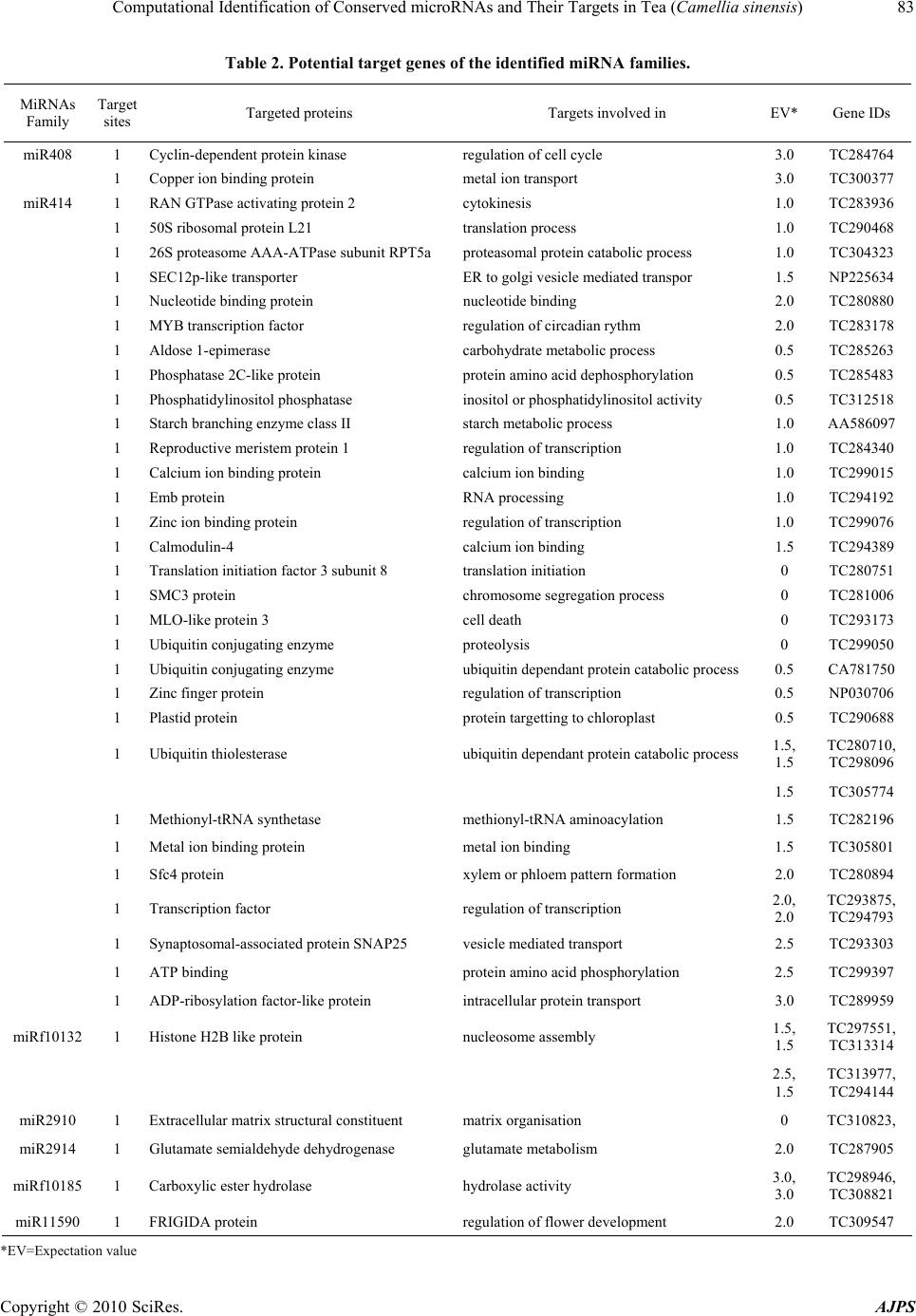 Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 83 Table 2. Potential target genes of the identified miRNA families. MiRNAs Family Target sites Targeted proteins Targets involved in EV* Gene IDs miR408 1 Cyclin-dependent protein kinase regulation of cell cycle 3.0 TC284764 1 Copper ion binding protein metal ion transport 3.0 TC300377 miR414 1 RAN GTPase activating protein 2 cytokinesis 1.0 TC283936 1 50S ribosomal protein L21 translation process 1.0 TC290468 1 26S proteasome AAA-ATPase subunit RPT5a proteasomal protein catabolic process 1.0 TC304323 1 SEC12p-like transporter ER to golgi vesicle mediated transpor 1.5 NP225634 1 Nucleotide binding protein nucleotide binding 2.0 TC280880 1 MYB transcription factor regulation of circadian rythm 2.0 TC283178 1 Aldose 1-epimerase carbohydrate metabolic process 0.5 TC285263 1 Phosphatase 2C-like protein protein amino acid dephosphorylation 0.5 TC285483 1 Phosphatidylinositol phosphatase inositol or phosphatidylinositol activity 0.5 TC312518 1 Starch branching enzyme class II starch metabolic process 1.0 AA586097 1 Reproductive meristem protein 1 regulation of transcription 1.0 TC284340 1 Calcium ion binding protein calcium ion binding 1.0 TC299015 1 Emb protein RNA processing 1.0 TC294192 1 Zinc ion binding protein regulation of transcription 1.0 TC299076 1 Calmodulin-4 calcium ion binding 1.5 TC294389 1 Translation initiation factor 3 subunit 8 translation initiation 0 TC280751 1 SMC3 protein chromosome segregation process 0 TC281006 1 MLO-like protein 3 cell death 0 TC293173 1 Ubiquitin conjugating enzyme proteolysis 0 TC299050 1 Ubiquitin conjugating enzyme ubiquitin dependant protein catabolic process 0.5 CA781750 1 Zinc finger protein regulation of transcription 0.5 NP030706 1 Plastid protein protein targetting to chloroplast 0.5 TC290688 1 Ubiquitin thiolesterase ubiquitin dependant protein catabolic process 1.5, 1.5 TC280710, TC298096 1.5 TC305774 1 Methionyl-tRNA synthetase methionyl-tRNA aminoacylation 1.5 TC282196 1 Metal ion binding protein metal ion binding 1.5 TC305801 1 Sfc4 protein xylem or phloem pattern formation 2.0 TC280894 1 Transcription factor regulation of transcription 2.0, 2.0 TC293875, TC294793 1 Synaptosomal-associated protein SNAP25 vesicle mediated transport 2.5 TC293303 1 ATP binding protein amino acid phosphorylation 2.5 TC299397 1 ADP-ribosylation factor-like protein intracellular protein transport 3.0 TC289959 miRf10132 1 Histone H2B like protein nucleosome assembly 1.5, 1.5 TC297551, TC313314 2.5, 1.5 TC313977, TC294144 miR2910 1 Extracellular matrix structural constituent matrix organisation 0 TC310823, miR2914 1 Glutamate semialdehyde dehydrogenase glutamate metabolism 2.0 TC287905 miRf10185 1 Carboxylic ester hydrolase hydrolase activity 3.0, 3.0 TC298946, TC308821 miR11590 1 FRIGIDA protein regulation of flower development 2.0 TC309547 *EV=Expectation value  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 84 Figure 5. Predicted miRNA targets (black in colour) and their complementary sites with miRNAs (red in colour). [29]. This may be an indicative of dominant nature of miR414 family in miRNA-mediated gene regulation in tea. The miRNAs were found diverse in nature such as location of mature miRNA sequences and length of pre- cursor sequences. The average length of precursor se- quence was 248 nucleotides; however a majority of them (62%) have 65-200 nucleotides. This finding is similar to other plants where the length of precursors varied in con- trast to consistent miRNA length of animal miRNAs (70-80 nt) [30,31]. In tea miRNAs, diversity was also observed within the members of same family which was also found in maize [1]. The identified precursor miRNAs were fold into hairpin secondary structures us- ing minimum free energies, with an average –72.69 kcal/mol which was lower than the values of Arabidopsis thaliana precursor miRNAs and much lower than the folding free energies of tRNA (–27.5 kcal /mol) and rRNA (–33 kcal/ mol) [32]. Out of 13 newly identified miRNAs, ten were from ESTs. There are several reports on miRNA identification from ESTs in various plant species [1,33]. The source sequences of miRNAs show a link between miRNAs and their tissues, organs, developmental stages or expression to which it belongs. On that basis, it was recognized that csi-miR414f, cas-miRf10185 and cas-miR11590-akr might be expressed in root and the rests were in leaf tissues. Moreover, csi-miR1171 and csi-miR414d were found in leaf tissue under the stress of winter dormancy and pest infestation, respectively. Three miRNAs namely cas- miR1122, cja-miR2910 and csi-miR2914 were identified from the full length nucleotides of RNA polymerase second largest subunit (intron 23) and 18S ribosomal subunit, respectively. Plant miRNAs are highly con- served among distantly related plant species, both in terms of primary and mature miRNAs [4]. This finding is also supported by our results, the identified miRNA was  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 85 found conserved in diverse plant species from mono- cotyledonous to dicotyledonous plants. These results suggested that different miRNAs might have evolved at different rates not only within the same plant species, but also in different ones. The miRNA target gene identification is an important step for understanding the role of miRNAs in gene regu- latory networks. Our prediction of target genes for the tea miRNAs revealed that more than one gene was regulated by individual miRNA. This result was similar to the re- cent findings in other plant species [1,10] which sug- gested that miRNA research should be focused on net- works rather than individual connections between miRNA and strongly predicted targets. MiRNAs may directly target transcription factors which affect plant growth and development, and also specific genes which control me- tabolism [4]. In this study, we identified a total of 37 potential targets for the 7 identified miRNA families in tea. The identified target genes appeared to be associated with diverse biological functions. There were genes en- coding transcription factors such as MYB, translation intiation factor such as TIF3, important proteosome de- grading pathway enzyme such as ubiquitin conjugating enzyme, different ion transporters such as copper ion binding protein, carbohydrate metabolism related enzyme such as aldose 1-epimerase, glutamate metabolism related enzymes such Glutamate semialdehyde dehydrogenase, important protein for nucleosome assembly such as his- tone as well as ribosomal proteins. In an earlier report, it was found as 20% transcription factors and 53% proteins related to diverse physiological processes, however their investigation was limited to only four miRNAs [34]. Overall, these findings made us clear that tea miRNAs targeted both trancription factors as well as specific genes. This findings of miRNAs in tea will pave the way for understanding the function and processing of tea small RNAs in future. Moreover, it shows a path for the pre- diction and analysis of miRNAs to those species whose genomes are not available through bioinformatics tools. REFERENCES [1] B. Zhang, X. Pan and A. T. Anderson, “Identification of 188 Conserved Maize microRNAs and Their Targets”, FEBS Letters, Vol. 580, No. 15, June 2006, pp. 3753-3762. [2] X. Zhou, J. Ruan, G. Wang and W. Zhang, “Characteriza- tion and Identification of MicroRNA Core Promoters in Four Model Species,” PLoS Computational Biology, Vol. 3, No. 3, January 2007, p. e37. [3] J. Singh and J. Nagaraju, “In silico Prediction and Char- acterization of microRNAs from Red Flour Beetle (Tri- bolium castaneum),” Insect Molecular Biology, Vol. 17, No. 4, August 2008, pp. 427-436. [4] B. Zhang, X. Pan, G. P. Cobb and T. A. Anderson, “Plant microRNA: A Small Regulatory Molecule with Big Im- pact,” Developmental Biology, Vol. 289, No. 1, January 2006, pp. 3-16. [5] C. A. Mallory and H. Vaucheret, “Functions of mi- croRNAs and Related Small RNAs in Plants,” Nature Genetics, Vol. 38, May 2006, pp. S31-S36. [6] R. Sunkar, V. Chinnusamy, J. Zhu and K. J. Zhu, “Small RNAs as Big Players in Plant Abiotic Stress Responses and Nutrient Deprivation,” Trends in Plant Science, Vol. 12, No. 7, July 2007, pp. 301-309. [7] B. Zhang, X. Pan, C. H. Cannon, G. P. Cobb and A. T. Anderson, “Conservation and Divergence of Plant mi- croRNA Genes,” Plant Journal, Vol. 46, No. 2, April 2006, pp. 243-259. [8] A. N. Eckardt, “MicroRNAs Regulate Auxin Homeosta- sis and Plant Development,” The Plant Cell, Vol. 17, May 2005, pp. 1335-1338. [9] S. H. Guo, Q. Xie, F. J. Fei and H. N. Chua, “MicroRNA Directs mRNA Cleavage of the Transcription Factor NAC1 to Downregulate Auxin Signals for Arabidopsis Lateral Root Development,” The Plant Cell, Vol. 17, May 2005, pp. 1376-1386. [10] M. W. Jones-Rhoades and P. D. Bartel, “Computtional Identification of Plant microRNAs and Their Targets, In- cluding a Stress-Induced miRNA,” Molecular Cell, Vol. 14, June 2004, pp. 787-799. [11] R. Sunkar and J. K. Zhu, “Novel and Stress-Regulated microRNAs and Other Small RNAs from Arabidopsis”, The Plant Cell, Vol. 16, No. 8, August 2004, pp. 2001- 2019. [12] Y. Lu, Q. Gan, X. Chi and S. Qin, “Roles of microRNA in Plant Defense and Virus Offense Interaction,” Plant Cell Reports, Vol. 27, No. 10, October 2008, pp. 1571- 1579. [13] R. Alba, Z. Fei, P. Payton, Y. Liu, L. S. Moore, P. Debbie, J. Cohn and M. D’Ascenzo, “ESTs, cDNA Microarrays, and Gene Expression Profiling: Tools for Dissecting Plant Physiology and Development,” The Plant Journal, Vol. 39, No. 5, September 2004, pp. 697-714. [14] B. H. Zhang, X. P. Pan, Q. L. Wang, G. P. Cobb and T. A. Anderson, “Identification and Characterization of New Plant microRNAs Using EST Analysis,” Cell Research, Vol. 15, No. 5, May 2005, pp. 336-360. [15] Q. Guo, A. Xiang, Q. Yang and Z. Y. Yang, “Bioinfor- matic Identification of microRNAs and Their Target Genes from Solanum tuberosum Expressed Sequence Tags,” Chinese Science Bulletin, Vol. 52, No. 17, Sep- tember 2007, pp. 2380-2389. [16] M. N. Nasaruddin, K. Harikrishna, Y. R. Othman, S. L. Hoon and A. J. Harikrishna, “Computational Prediction of microRNAs from Oil Palm (Elaeis guineensis Jacq.) Expressed Sequence Tags,” Asian Pacific Journal of Mo- lecular Biology and Biotechnology, Vol. 15, No. 3, Oc- tober 2007, pp. 107-113. [17] W. Jin, N. Li, B. Zhang, F. Wu, W. Li, A. Guo and Z. Deng, “Identification and Verification of microRNA in  Computational Identification of Conserved microRNAs and Their Targets in Tea (Camellia sinensis) Copyright © 2010 SciRes. AJPS 86 Wheat (Triticum aestivum),” Journal of Plant Research, Vol. 121, No. 3, March 2008, pp. 351-355. [18] B. Billoud, D. R. Paepe, D. Baulcombe and M. Boccara, “Identification of New Small Non-Coding RNAs from Tobacco and Arabidopsis,” Biochimie, Vol. 87, No. 9, September 2005, pp. 905-910. [19] R. Sunkar and G. Jagadeeswaran, “In silico Identification of Conserved microRNAs in Large Number of Diverse Plant Species,” BMC Plant Biology, Vol. 8, No. 37, 2008, p. 13. [20] S. Griffiths-Jones, H. K. Saini, D. S. Van and A. J. En- right, “MiRBase: Tools for microRNA Genomics,” Nu- cleic Acids Research, Vol. 36, November 2008, pp. D154- D158. [21] N. Rajewsky and D. N. Socci, “Computational Identifica- tion of microRNA Targets,” Developmental Biology, Vol. 267, No. 2, March 2004, pp. 529-535. [22] L. Zhang, M. J. Chia, S. Kumari, C. J. Stein, Z. Liu, A. Narechania, A. C. Maher, K. Guill, D. M. McMullen and D. Ware, “A Genome-Wide Characterization of Mi- croRNA Genes in Maize,” PLoS Genetics, Vol. 5, No. 11, November 2009, pp. 5e1000716. [23] S. F. Altschul, T. L. Madden, A. A. Schäffer, J. Zhang, Z. Zhang, W. Miller and D. J. Lipman, “Gapped BLAST and PSI-BLAST: A New Generation of Protein Database Search Programs,” Nucleic Acids Research, Vol. 25, No. 17, September 1997, pp. 3389-3402. [24] M. Zuker, “Mfold Web Server for Nucleic Acid Folding and Hybridization Prediction,” Nucleic Acids Research, Vol. 31, No. 13, July 2003, pp. 3406-3415. [25] J. D. Thompson, D. G. Higgins and T. J. Gibson, “CLUSTAL W: Improving the Sensitivity of Progressive Multiple Sequence Alignments through Sequence Weight- ing, Position Specific Gap Penalties and Weight Matrix Choice,” Nucleic Acids Research, Vol. 22, No. 22, No- vember 1994, pp. 4673-4680. [26] K. Tamura and M. Nei, “Estimation of the Number of Nucleotide Substitutions in the Control Region of Mito- chondrial DNA in Humans and Chimpanzees,” Molecular Biology and Evolution, Vol. 10, No. 3, May 1993, pp. 512-526. [27] K. Tamura, J. Dudley, M. Nei and S. Kumar, “MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0.,” Molecular Biology and Evolution, Vol. 24, No. 8, August 2007, pp. 1596-1599. [28] S. Griffiths-Jones, R. J. Grocock, D. S. Van, A. Bateman and A. J. Enright, “miRBase: microRNA Sequences, Tar- gets and Gene Nomenclature,” Nucleic Acids Research, Vol. 34, November 2006, pp. D140-D144. [29] E. Bonnet, J. Wuyts, P. Rouze and Y. V. de Peer, “Evi- dence that microRNA Precursors, unlike Other Non-Coding RNAs, Have Lower Folding Free Energies than Random Sequences,” Bioinformatics, Vol. 20, No. 17, November 2004, pp. 2911-2917. [30] P. D. Bartel, “MicroRNAs: Genomics, Biogenesis, Mechanism and Function,” Cell, Vol. 116, No. 2, January 2004, pp. 281-297. [31] V. Ambros, “The Functions of Animal microRNAs,” Na- ture, Vol. 431, September 2004, pp. 350-355. [32] M. Y. Khan-Barozai, M. Irfan, R. Yousaf, I. Ali, U. Qaisar, A. Maqbool, M. Zahoor, B. Rashid, T. Hussnain and S. Riazuddin, “Identification of microRNA in Cot- ton,” Plant Physiology and Biochemistry, Vol. 46, No. 8, August 2008, pp. 739-751. [33] C. X. Qiu, F. L. Xie, Y. Y. Zhu, K. Guo, S. Q. Huang, L. Nie and Z. M. Yang, “Computational Identification of microRNAs and Their Targets in Gossypium hirsutum Expressed Sequence Tags,” Gene, Vol. 395, No. 1, June 2007, pp. 49-61. [34] G. R. Prabu and A. K. A. Mandal, “Computational Identi- fication of miRNAs and Their Target Genes from Ex- pressed Sequence Tags of Tea (Camellia sinensis),” Ge- nomics Proteomics and Bioinformatics, Vol. 8, No. 2, June 2010, pp. 113-121. Abbreviations used: BLAST, basic local alignment search tool; dbEST, database of expressed sequence tags; ESTs, expressed sequence tags; mRNA, messenger RNA; miRNA, microRNA; mfe, minimum free energy; nt, nu- cleotide; NCBI, national center for biotechnological in- formation |

