Paper Menu >>
Journal Menu >>
 Natural Resources, 2010, 1, 95-103 doi:10.4236/nr.2010.12010 Published Online December 2010 (http://www.SciRP.org/journal/nr) Copyright © 2010 SciRes. NR 95 Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Hédia Hannachi1*, Catherine Breton2, Monji Msallem3, Salem Ben El Hadj4, Mohamed El Gazzah1, André Bervillé2 1Faculté des Sciences de Tunis, Département de Biologie, Campus Universitaire, Tunis, Tunisie; 2INRA, UMR1097, Bât. 33, 2 place Viala, F-34060 Montpellier cedex 1, France; 3Institut de l’Olivier, Tunis, Tunisia; 4Institut National Agronomique de Tunisie, Tunis, Mahrajène, Tunisia. Email: hannachi_hedia@yahoo.fr Received October 26th, 2010; revised November 29th, 2010; accepted November 30th, 2010. ABSTRACT The olive is widely cropped in Tunisia where also oleaster trees thrive all around orchards and in natural sites. Little is known on the genetic relationships between the olive crop and oleaster trees in Tunisia. Fifty-two oleaster trees and fifteen cultivars were sampled from Tunisia. SSR genotyping was performed in polyacrylamide gels after fluorescent labeling. We used seven nuclear and two chloroplast SSR markers. AFC analyses showed close genetic relationships between cultivated and oleaster trees. Genetic relationships were also displayed in a dendrogram based on Unweighted Pair Group Method (UPGMA). Five clusters were defined mixing cultivar and oleaster trees suggesting close relation- ship between some cultivar and some oleaster trees. One oleaster is single in a cluster. The chlorotype SSR markers show probably three olive origins. Some cultivars have the CE chlorotype originates from the East of the Mediterra- nean basin, the CCK haplotype originates from Maghreb and the COM chlorotype originates from West Mediterranean. The cultivars were 1) introduced from the East; 2) selected in the West; 3) or selected in the North Africa region. The Tunisian oleaster trees carry eastern and western Mediterranean chlorotype CCK, COM and CE. Keywords: Cultivars, Oleaster, Genetic Relationship, SSR Markers, Haplotype, Origin 1. Introduction Two olive (Olea europaea subsp. europaea var. euro- paea) varieties are distinguished by botanists in the Me- diterranean basin namely var. europaea which is the cul- tivated form and var. sylvestris, the wild olive tree or oleaster. The olive is one of both of the oldest tree crops with the fig tree and it is cultivated for oil and table olives. The olive is the most important oil producing crop in the Mediterranean region. Olive oil has traditionally been used for pharmaceutical, industrial and consumer pur- poses. Tunisia is formerly a major producer of olive oil in North Africa. In Tunisia, about 60 million olive trees are cultivated in third of cultivated areas, most of them represented by two prevalent oil cultivars ‘Chétoui’ and ‘Chemlali’. The rest is represented by several minor cul- tivars [1]. Little is known about the Tunisia oleaster trees and about their genetic relationships with cultivars. Genetic erosion and loss of biodiversity do not seem to be major issues for olive germplasms due to absence of turnover of new genotypes that do not occur as fast as in other woody crops. Moreover, old olive trees survive for a long time once abandoned [2,3]. Morphological traits in the olive do not enable differ- entiation between oleasters and cultivars. Although, sev- eral morphological descriptors show partial differentia- tion of them [4,5]. Recently, molecular markers have been developed in the olive [2,6-11] that enable cultivar differentiation and identification due to their high intra species variability. The use of nuclear microsatellite markers for genetic analysis is well established in the olive [7,12-14]. The  Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 96 principle has been extended to the chloroplast [15,16] and mitochondrial genomes [3,17,18]. The utility of mo- lecular tools for evolutionary studies arises from the in- sensitivity of the genetic markers to environmental fac- tors. Several markers based on DNA amplification tech- nology have been used to look for genetic relationships between the cultivated olive (cultivars) and the oleaster trees as to structure its genetic diversity [3,8,15,16], in- cluding DNA from nucleus, chloroplast (cpDNA) and mitochondria (mtDNA). Simple Sequence Repeats (SSRs) lead to multiallelic fragments and are easily amenable to Polymerase Chain Reaction (PCR) based analysis. Mi- crosatellite markers explore various independent portions of the olive genome and they have been identified in plants’ nuclear and mitochondrial genomes [3,15,19] as well as in the chloroplast genome where they are mono- nucleotide [20]. With SSRs, Restriction Fragment Length Polymorphisms (RFLPs) are complementarily used to define chloroplast and mitochondrial RFLP data, using multiple pair wise combinations of probe and restriction enzyme to recognize distinct genetic patterns, called chlorotypes or mitotypes. Since the organelles are usu- ally passed to offspring from the female parents, cyto- plasm markers (mitochondria and chloroplast) trace ma- ternal lineage only [21]. Many studies have shown the diversity of cultivars using morphological descriptors [1,22], but little atten- tion has been given to the Tunisian oleasters [4,5]. In Tunisia a few studies have been made on cultivars using SSRs markers [23]. In the present study, an analysis of polymorphisms within and among the two olive taxa (cultivar and oleaster trees) was undertaken using nuclear and cytoplasm SSR markers. This will enable the determination of genetic groups or clusters to establish breeding programs that encompasses the genetic diversity of this species. 2. Materials and Methods A total of 15 autochthonous Tunisian cultivars and 52 oleasters were sampled in the north of Tunisia (Table 1), all are presently cropped in wide area. They were subject to genotyping for chlorotypes previously described and developed. We used two markers ccmp5 and ccmp7 re- tained by authors [8,21,22] and seven nuclear microsatel- lite markers, three ssrOeUA-DCA04, 05, 09 [11]; one ssrOe-GAPU 101 [25] and three ssrOe-UD012, 017, 024 [9], chosen as used by Breton et al. [14], (Table 2). 2.1. DNA Amplifications Total DNA preparation was performed using the method described by Besnard et al. [21]. PCR reaction was per- formed in 12.5 µl final volume, containing 40 ng ge- nomic DNA, 0.75 mM MgCl2, 2.5 mM dNTP, 1.25 U Taq polymerase and 0.19 mM M13-Fam. PCR amplify- cation was conducted in a thermal cycler Gradient 96 Robocycler (Stratagene, Germany). The amplification program was 94℃C for 1 min, 52℃ for 1 min, 72℃ for 1 min, followed by 35 cycles at 94℃ for 30 s, at 50℃ for 45 s and at 72℃ for 1 min, with a final elongation cycle at 72℃ for 4 min. Amplification products and ladders were labeled using the tailing method with the Fam fluorochrome. They were separated into 8% polyacrylamide gels enabling reading with a Hitachi scanner system associated with the FMBIO2 software [26]. The method used for chlorotype DNA-RFLP analysis was described by Besnard et al. [21] and Breton [13]. Two restrictions enzyme/probe combinations (HindIII/ atp6 and XbaI/atp6) were used to identify the chlorotype CE, COM and CCK previously determined by Besnard et al. [21]. 2.2. Statistical Analysis Factorial Correspondence Analysis (FCA) was per- formed using GENETIX. Dendrogram was constructed with Unweighted Pair Group Method (UPGMA) algorithm based on Nei’ genetic distances [27] and a neighbour- joining tree was constructed with the nuclear SSR data set using PHYLIP Version 3.5c [28]. 3. Results 3.1. Factorial Correspondence Analysis The plot of FCA coordinates for the first and the second axes, which explain 8.69% and 6.23% of variance, re- spectively and showed continuity in the distribution of oleaster and cultivar trees. However, most of cultivars clustered in the right of the cloud. Thus, we considered two clusters of genotypes (Figure 1). The cultivars clus- ter grouped three oleaster trees. In contrast, most oleaster trees clustered on the left with the cultivar C1 (Sayali). 3.2. Clustering Olive cultivar and oleaster trees were clustered with the UPGMA method based on the Nei’s similarity coeffi- cient using SSR data (Figure 2). The clustering analysis showed five groups and one single oleaster (O11). Three clusters (CL1, CL2 and CL3) contain cultivars and oleaster trees and two other clusters (CL4 and CL5) con- tain only oleaster trees. Cluster_1 (CL1) aggregated 11 out of 15 cultivars (C1 Sayali, C6 Marsaline, C10 Meski, C30 Rajou, C29 Limi, C26 Tounsi, C14 Gerboui, C20 Besbessi, C28 Zarras, C13 Neb Jmel and C22 Chaïbi) and 12 oleasters from several locations; six of them sampled from natural eco- 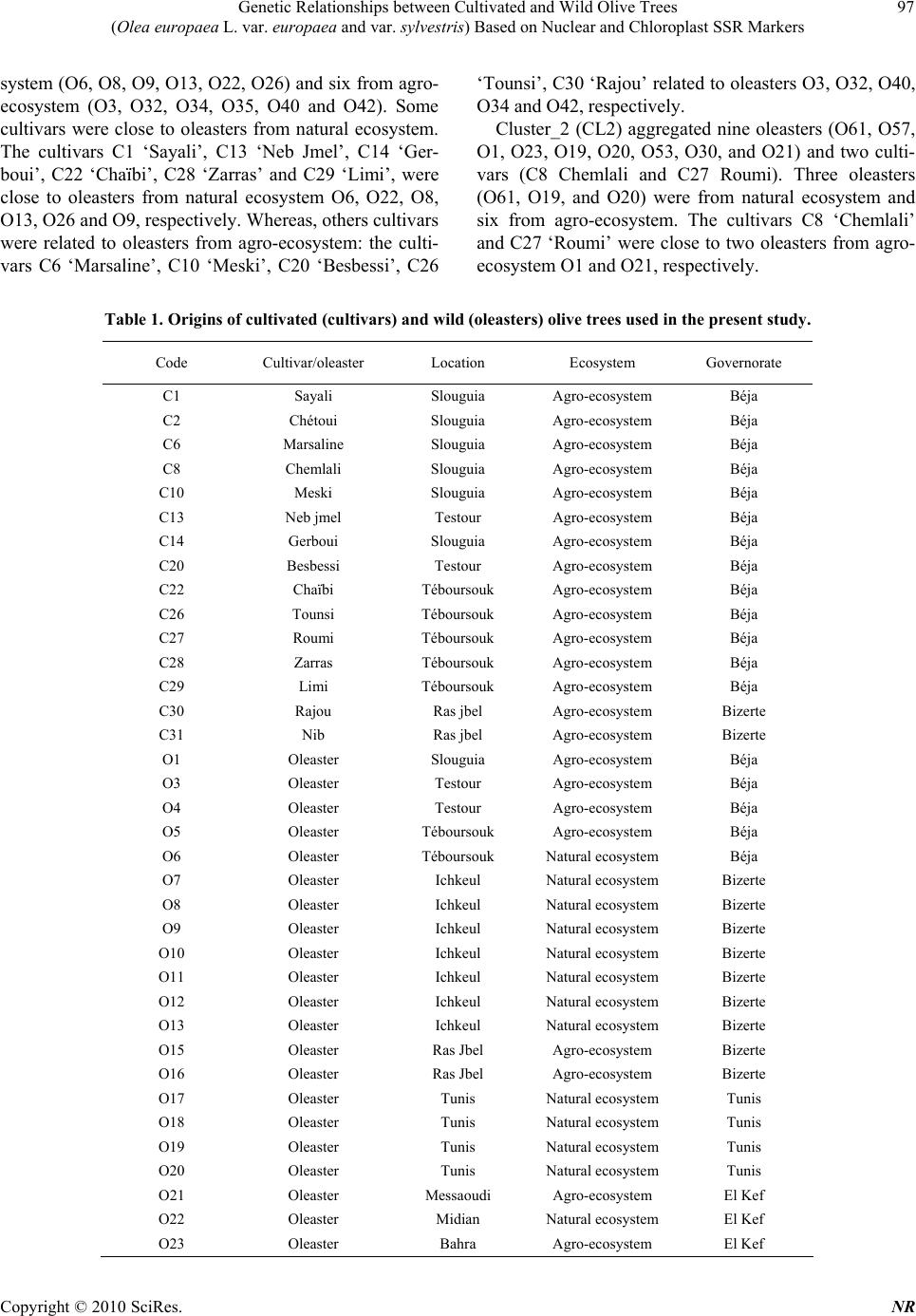 Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 97 system (O6, O8, O9, O13, O22, O26) and six from agro- ecosystem (O3, O32, O34, O35, O40 and O42). Some cultivars were close to oleasters from natural ecosystem. The cultivars C1 ‘Sayali’, C13 ‘Neb Jmel’, C14 ‘Ger- boui’, C22 ‘Chaïbi’, C28 ‘Zarras’ and C29 ‘Limi’, were close to oleasters from natural ecosystem O6, O22, O8, O13, O26 and O9, respectively. Whereas, others cultivars were related to oleasters from agro-ecosystem: the culti- vars C6 ‘Marsaline’, C10 ‘Meski’, C20 ‘Besbessi’, C26 ‘Tounsi’, C30 ‘Rajou’ related to oleasters O3, O32, O40, O34 and O42, respectively. Cluster_2 (CL2) aggregated nine oleasters (O61, O57, O1, O23, O19, O20, O53, O30, and O21) and two culti- vars (C8 Chemlali and C27 Roumi). Three oleasters (O61, O19, and O20) were from natural ecosystem and six from agro-ecosystem. The cultivars C8 ‘Chemlali’ and C27 ‘Roumi’ were close to two oleasters from agro- ecosystem O1 and O21, respectively. Table 1. Origins of cultivated (cultivars) and wild (oleasters) olive trees used in the present study. Code Cultivar/oleaster Location Ecosystem Governorate C1 Sayali Slouguia Agro-ecosystem Béja C2 Chétoui Slouguia Agro-ecosystem Béja C6 Marsaline Slouguia Agro-ecosystem Béja C8 Chemlali Slouguia Agro-ecosystem Béja C10 Meski Slouguia Agro-ecosystem Béja C13 Neb jmel Testour Agro-ecosystem Béja C14 Gerboui Slouguia Agro-ecosystem Béja C20 Besbessi Testour Agro-ecosystem Béja C22 Chaïbi Téboursouk Agro-ecosystem Béja C26 Tounsi Téboursouk Agro-ecosystem Béja C27 Roumi Téboursouk Agro-ecosystem Béja C28 Zarras Téboursouk Agro-ecosystem Béja C29 Limi Téboursouk Agro-ecosystem Béja C30 Rajou Ras jbel Agro-ecosystem Bizerte C31 Nib Ras jbel Agro-ecosystem Bizerte O1 Oleaster Slouguia Agro-ecosystem Béja O3 Oleaster Testour Agro-ecosystem Béja O4 Oleaster Testour Agro-ecosystem Béja O5 Oleaster Téboursouk Agro-ecosystem Béja O6 Oleaster Téboursouk Natural ecosystem Béja O7 Oleaster Ichkeul Natural ecosystem Bizerte O8 Oleaster Ichkeul Natural ecosystem Bizerte O9 Oleaster Ichkeul Natural ecosystem Bizerte O10 Oleaster Ichkeul Natural ecosystem Bizerte O11 Oleaster Ichkeul Natural ecosystem Bizerte O12 Oleaster Ichkeul Natural ecosystem Bizerte O13 Oleaster Ichkeul Natural ecosystem Bizerte O15 Oleaster Ras Jbel Agro-ecosystem Bizerte O16 Oleaster Ras Jbel Agro-ecosystem Bizerte O17 Oleaster Tunis Natural ecosystem Tunis O18 Oleaster Tunis Natural ecosystem Tunis O19 Oleaster Tunis Natural ecosystem Tunis O20 Oleaster Tunis Natural ecosystem Tunis O21 Oleaster Messaoudi Agro-ecosystem El Kef O22 Oleaster Midian Natural ecosystem El Kef O23 Oleaster Bahra Agro-ecosystem El Kef 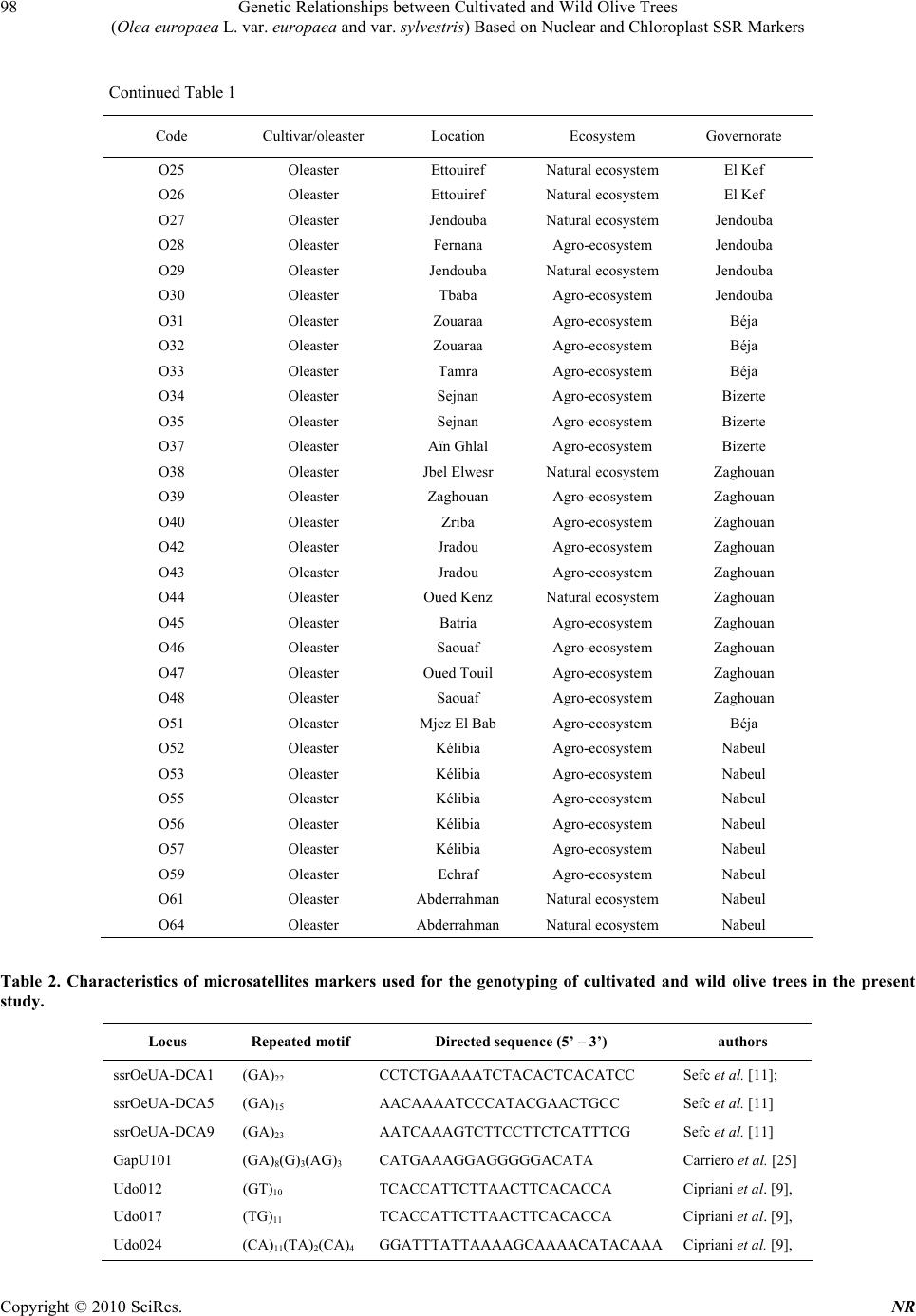 Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 98 Continued Table 1 Code Cultivar/oleaster Location Ecosystem Governorate O25 Oleaster Ettouiref Natural ecosystem El Kef O26 Oleaster Ettouiref Natural ecosystem El Kef O27 Oleaster Jendouba Natural ecosystem Jendouba O28 Oleaster Fernana Agro-ecosystem Jendouba O29 Oleaster Jendouba Natural ecosystem Jendouba O30 Oleaster Tbaba Agro-ecosystem Jendouba O31 Oleaster Zouaraa Agro-ecosystem Béja O32 Oleaster Zouaraa Agro-ecosystem Béja O33 Oleaster Tamra Agro-ecosystem Béja O34 Oleaster Sejnan Agro-ecosystem Bizerte O35 Oleaster Sejnan Agro-ecosystem Bizerte O37 Oleaster Aïn Ghlal Agro-ecosystem Bizerte O38 Oleaster Jbel Elwesr Natural ecosystem Zaghouan O39 Oleaster Zaghouan Agro-ecosystem Zaghouan O40 Oleaster Zriba Agro-ecosystem Zaghouan O42 Oleaster Jradou Agro-ecosystem Zaghouan O43 Oleaster Jradou Agro-ecosystem Zaghouan O44 Oleaster Oued Kenz Natural ecosystem Zaghouan O45 Oleaster Batria Agro-ecosystem Zaghouan O46 Oleaster Saouaf Agro-ecosystem Zaghouan O47 Oleaster Oued Touil Agro-ecosystem Zaghouan O48 Oleaster Saouaf Agro-ecosystem Zaghouan O51 Oleaster Mjez El Bab Agro-ecosystem Béja O52 Oleaster Kélibia Agro-ecosystem Nabeul O53 Oleaster Kélibia Agro-ecosystem Nabeul O55 Oleaster Kélibia Agro-ecosystem Nabeul O56 Oleaster Kélibia Agro-ecosystem Nabeul O57 Oleaster Kélibia Agro-ecosystem Nabeul O59 Oleaster Echraf Agro-ecosystem Nabeul O61 Oleaster Abderrahman Natural ecosystem Nabeul O64 Oleaster Abderrahman Natural ecosystem Nabeul Table 2. Characteristics of microsatellites markers used for the genotyping of cultivated and wild olive trees in the present study. Locus Repeated motif Directed sequence (5’ – 3’) authors ssrOeUA-DCA1 (GA)22 CCTCTGAAAATCTACACTCACATCC Sefc et al. [11]; ssrOeUA-DCA5 (GA)15 AACAAAATCCCATACGAACTGCC Sefc et al. [11] ssrOeUA-DCA9 (GA)23 AATCAAAGTCTTCCTTCTCATTTCG Sefc et al. [11] GapU101 (GA)8(G)3(AG)3 CATGAAAGGAGGGGGACATA Carriero et al. [25] Udo012 (GT)10 TCACCATTCTTAACTTCACACCA Cipriani et al. [9], Udo017 (TG)11 TCACCATTCTTAACTTCACACCA Cipriani et al. [9], Udo024 (CA)11(TA)2(CA)4 GGATTTATTAAAAGCAAAACATACAAACipriani et al. [9], 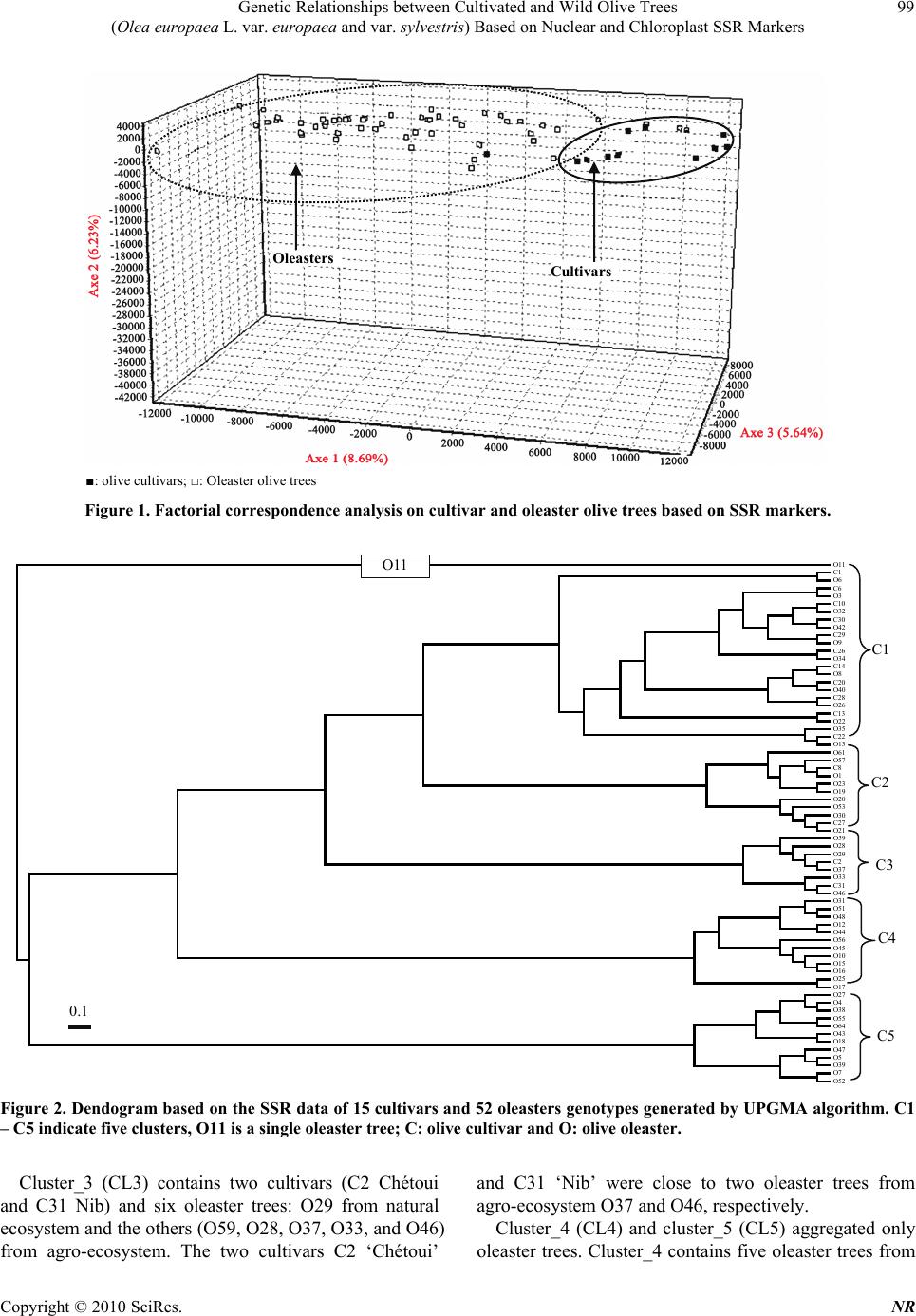 Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 99 Cultivars Oleasters ■: olive cultivars; □: Oleaster olive trees Figure 1. Factorial correspondence analysis on cultivar and oleaster olive trees based on SSR markers. O11 C1 O6 C6 O3 C10 O32 C30 O42 C29 O9 C26 O34 C14 O8 C20 O40 C28 O26 C13 O22 O35 C22 O13 O61 O57 C8 O1 O23 O19 O20 O53 O30 C27 O21 O59 O28 O29 C2 O37 O33 C31 O46 O31 O51 O48 O12 O44 O56 O45 O10 O15 O16 O25 O17 O27 O4 O38 O55 O64 O43 O18 O47 O5 O39 O7 O52 0.1 C1 C2 C3 C4 C5 O11 Figure 2. Dendogram based on the SSR data of 15 cultivars and 52 oleasters genotypes generated by UPGMA algorithm. C1 – C5 indicate five clusters, O11 is a single oleaster tree; C: olive cultivar and O: olive oleaster. Cluster_3 (CL3) contains two cultivars (C2 Chétoui and C31 Nib) and six oleaster trees: O29 from natural ecosystem and the others (O59, O28, O37, O33, and O46) from agro-ecosystem. The two cultivars C2 ‘Chétoui’ and C31 ‘Nib’ were close to two oleaster trees from agro-ecosystem O37 and O46, respectively. Cluster_4 (CL4) and cluster_5 (CL5) aggregated only oleaster trees. Cluster_4 contains five oleaster trees from  Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 100 natural ecosystem (O12, O44, O10, O25, and O17) and seven from agro-ecosystem (O31, O51, O48, O56, O45, O15, and O16). Cluster_5 contains five oleaster trees from natural-ecosystem (O27, O38, O64, O18, and O7) and seven oleaster trees from agro-ecosystem (O4, O55, O43, O47, O5, O39, and O62). However, the oleaster O11 from natural ecosystem is single and represents a cluster by itself. These results showed tight relationships between some oleaster trees and cultivars independently of locations and ecosystem. 3.3. Chloroplast SSR In this study, four chlorotypes CE1, CE2, COM and CCK previously determined in olives were found in cul- tivars and defined the olive origins (Table 3). Oleaster trees from agro-ecosystem and natural sites carry CE1 (6/6), CE2 (0/0), CCK (9/5) and COM (10/7), respect- tively, and the lattes do not reveal significant differences for chlorotype frequencies. Whereas, for olive cultivars six carry CE1, six CE2, one COM and six CCK chloro- type. 4. Discussion We used seven nuclear and two cytoplasmic microsatel- lite markers over 15 cultivars and 52 oleasters that re- vealed several clusters of cultivars, oleaster trees and several chlorotypes. The morphological means showed a continuous variation between cultivated and wild olive trees [4]. In the present study, molecular markers show also continuous variation, but most cultivars clustered together. This genetic structure probably results from the origin of the cultivars and oleaster trees. Besnard et al. [18] and Besnard and Bervillé [15] have shown that the CE1, COM, and CCK chlorotypes are prevalent in oleaster trees from the East (CE1) and the West (COM and CCK). In addition, Breton [13] and Breton et al. [14] have shown that CE2 and COM (COM1 and COM2 are variant of COM) originated in Cyprus and Tunisia where they are prevalent in oleaster trees. Consequently, the deep structure in chlorotypes infers that cultivars carrying CE1 or CE2 have ancestors in oleaster or in cultivars from the East. Whereas, culti- vars carrying COM or CCK have ancestors in oleaster or Table 3. Chlorotypes of cultivated (cultivars) and wild (oleasters) olive trees based on chloroplast SSR markers. (CE1, CE2: East Mediterranean chlorotype; COM, COM2: West Mediterranean chlorotype; CCK: Maghreb chlorotype; C and O: cul- tivars and oleasters codes, respectively, used in UPGMA analysis). Chlorotype Cultivars a Oleastersa Oleastersb CE1 Sayali (C1) Chemlali (C8) Gerboui (C14) Roumi (C27) Zarras (C28) Nib (C31) O32 O37 O45 O48 O56 O57 O6 O9 O18 O22 O44 O64 CE2 Besbessi (C20) COM Neb jmel (C13) O31 O43 O59 O7 COM2 O1 O3 O4 O23 O34 O35 O51 O8 O10 O11 O12 O61 O25 CCK Chétoui (C2) Marsaline (C6) Meski (C10) Chaïbi (C22) Limi (C29) Rajou (C30) O5 O15 O16 O20 O28 O40 O47 O52 O53 O19 O21 O26 O27 O29 a: agro-ecosystem; b: natural ecosystem  Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 101 cultivars from the West. Tunisia offers a peculiar situa- tion due to early colonization by Phoenicians, who have probably introduced cultivars from the East into Carthage colony and their further colonies in the West (Spain, Portugal). We can therefore deduce that oleaster trees carrying CE1 are feral trees (progenies of a cultivar by an oleaster or vice versa) either in the agro-ecosystem or natural sites (Table 3). Gene flow appears responsible for the diffusion of the CE1 chlorotypes, but also for the nuclear markers as shown by Breton et al. [14] using Bayesian methods [29]. From these results, we can deduce that ‘Sayali’ carry- ing CE1 that clustered with oleaster trees is probably of feral origin. ‘Chemlali’, carrying also CE1 but aggre- gated in the FCA intermediate between oleaster trees and cultivars, has probably similar origin. In the Dendrogram ‘Chemlali’ and ‘Roumi’ are in the same clusters with some oleaster trees suggesting that ‘Roumi’ could be a progeny of ‘Chemlali’ with local oleaster trees. ‘Neb Jmel’ characterized by the West chlorotype COM was probably selected in the Maghreb region. Oleasters car- rying western Mediterranean chlorotypes (CCK, COM) may cluster with cultivars carrying the same chlorotypes in different clusters. Consequently, the chlorotypes are not correlated with the clusters (Figure 2). Indeed, the UPGMA clustering revealed that each cluster is independent of the chlorotypes which means that kinship relationships by the chlorotypes have been hindered by gene flow between cultivars and local oleaster trees as well as between oleaster trees and culti- vars. Obviously, we observed events that have occurred through the female side due to the chlorotypes which is maternally inherited in the olive [21]. The same events are likely existing through pollen flow, but too difficult to detect unless using Bayesian methods. However, in this study, the sampled trees are too limited to study per se gene flow events due to the absence of anchor refer- ences for COM and CCK chlorotypes. We suspect that the CCK chlorotype originated from Kabylia [7] where a refuge zone for the olive has been revealed [14], but we do not know whether it has been the only refuge for CCK or if other refuge zones in North Africa or Sicily may have preserved CCK. We also suspected that the COM chlorotypes (COM, COM1, COM2) originate from Tuni- sia where they are prevalent in natural sites and that cor- respond to a refuge zone [13,14], but we cannot exclude that, CCK chlorotypes, were kept in refuge zones from Sicily-Corsica. Unfortunately, oleaster trees from central and south Italy have not been genotyped for the chloro- types [30]. Mixed stands of oleaster trees of natural sites in Tuni- sia display a huge diversity based on the chlorotypes and nuclear polymorphisms in comparison with other oleaster trees in Mediterranean forests. Oleaster trees transformed into new cultivars should be carefully examined since they are the result into crosses not usually done between genotypes from the East and West of the Mediterranean regions. Seed gene flow is locally detected when in a region where cultivars have been introduced and local oleasters do not carry the same cytoplasm [12,15]. Distinction of a crop from its wild relatives is based on several morphological traits and botanists have usually made distinct species of two taxa [31]. For the cultivated olive trees, it has been traditionally carried out by mor- phological, agronomic and chemical traits [1,32-35]. Based on the morphology and molecular markers, it is absolutely impossible to determine whether oleaster trees from natural sites are genuine oleasters or not. The phy- logeography of the oleaster and cultivars trees is due to permanent and recurrent gene flow. Here, we clearly show that oleaster trees carrying the eastern CE1 chloro- type are present in natural sites of Tunisia. In the frame of the hypothesis that CE1 was absent from refuge zones in the west, it should have been introduced from the East into the West 2500 years ago. In this work, it appeared from the dendrogram (Figure 2) that the oleaster and cu-l tivar trees clustered by similarities whatever their chloro- types showing tight genetic relationships. The same re- sults were obtained by Bayesian methods [14]. In Tunisia, many studies have shown the diversity of the Tunisian cultivated olive trees [22,36] but little atten- tion has been given to the Tunisian oleaster trees. Little is known about molecular identification of Tunisian olive. A close genetic proximity between Tunisian oleasters and cultivars was showed by dendrogram based on nu- clear SSR markers (Figure 2). This relationship has al- ready been shown with isozymes [37,38], RAPD and RFLP [3,7,14]. Several molecular studies, including AFLPs, RAPDs, ISSRs, repetitive DNA sequence analysis, chloroplast and mitochondrial DNA polymorphism, have also con- tributed to elucidate the classification of the Olea com- plex and the origin of cultivated olive [2,3,12,15,16, 39-41]. It has been reported that oil composition for oleaster trees were in agreement with the olive oil norms [4]. Here, we show that those oleasters trees are unique to Tunisia. Crossing the oleasters suggest that breeding the olive and could be done in the country where cultivars may have non-equilibrated oil composition. Indeed, to improve the oil quality it should cutting olive oil with others to satisfy European norms. Screening oleaster trees from the agro-ecosystem and natural sites should lead to new genotypes that could be compared for oil  Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 102 composition and yield as it has been done in Australia by Mekuria et al. [42] and Sedgley [43]. These trees are adapted to soil and climate found in Tunisia and therefore they should been screened for their behaviour in the agro-ecosystem to check the yield and quality of the product. 5. Conclusions Cultivars found in Tunisia are of diverse origins based on their chlorotype and nuclear markers. Local genuine oleaster trees are difficult to differentiate from feral trees, and shown to share more or less kinship relationships with autochtonous and introduced cultivars. Those oleaster trees should offer opportunity to screen for new genotypes producing oil with more equilibrated composi- tion than ‘Chemlali’ as an example and acceptable agro- nomic behavior to compete with local cultivars to ensure direct selling of the products to the Europe. REFERENCES [1] A. Trigui and M. Msallem, “Oliviers de Tunisie, Catalogue des Varietes Autochtones & Types Locaux, Identification Varietale & Caracterisation Morpho-Pomologique des Ressources Genetiques Oleicoles de Tunisie,” Vol. 1. (Fr) Ministère de l’Agriculture, IRESA, Institut de l’Olivier, Tunisia, 2002. [2] A. Angiliollo, M. Mencuccini and L. Baldoni, “Olive Genetic Diversity Assessed Using Amplified Fragment Length Polymorphisms,” Theoretical and Applied Genet- ics, Vol. 98, No. 3-4, 1999, pp. 411-421. [3] G. Besnard and A. Bervillé, “Multiple Origins for Medi- terranean Olive (Olea europaea L. subsp. europaea) Ba- sed upon Mitochondrial DNA Polymorphisms,” Comptes Rendus Biologie de l’Académie des Sciences, Vol. 323, No. 2, 2000, pp. 173-181. [4] H. Hannachi, C. Breton, M. Msallem, S. Ben El Hadj, M. El Gazzah and A. Bervillé, “Differences between Native and Introduced Olive Cultivars as Revealed by Morphol- ogy of Drupes, Oil Composition and SSR Polymorphisms: A Case Study in Tunisia,” Scientia Horticultural, Vol. 116, No. 3, 2008, pp. 280-290. [5] H. Hannachi, C. Breton, M. Msallem, S. Ben El Hadj, M. El Gazzah and A. Bervillé, “Are Olive Cultivars Distin- guishable from Oleaster Trees Based on Morphology of Drupes and Pits, Oil Composition and Microsatellite Polymorphisms?” Acta Botanica Gallica, Vol. 155, No. 4, 2008, pp. 531-545. [6] A. Fabbri, J. I. Hormaza and V. S. Polito, “Random Am- plified Polymorphic DNA Analysis of Olive (Olea eu- ropaea L.) Cultivars,” The Journal of the American Soci- ety for Horticultural Science, Vol. 120, No. 3, 1995, pp. 538-542. [7] G. Besnard, P. Baradat, D. Chevalier, A. Tagmount and A. Bervillé, “Genetic Differentiation in the Olive Complex (Olea europaea L.) Revealed by RAPDs and RFLPs in the rRNA Genes,” Genetic Resources and Crop Evolution, Vol. 48, No. 2, 2001, pp. 165-182. [8] V. Bronzini de Caraffa, J. Maury, C. Gambotti, C. Breton, A. Bervillé and J. Giannettini, “Mitochondrial DNA Variation from Western an Eastern Mediterranean,” Theoretical and Applied Genetics, Vol. 104, No. 6-7, 2002, pp. 1209-1216. [9] G. Cipriani, M. T. Marrazzo, R. Marconi, A. Cimato and R. Testolin, “Microsatellite Markers Isolated in Olive (Olea europaea L.) are Suitable for Individual Finger- Printing and Reveal Polymorphism within Ancient Culti- vars,” Theoretical and Applied Genetics, Vol. 104, No. 2- 3, 2002, pp. 223-228. [10] P. Rallo, G. Dorado and A. Martin, “Development of Simple Sequence Repeats (SSR) in Olive Tree (Olea eu- ropaea L.),” Theoretical and Applied Genetics, Vol. 101, No. 5-6, 2000, pp. 984-989. [11] K. M. Sefc, M. S. Lopes, A. D. Mendoc, M. Rodrigues Dos Santos, M. L. da C. Machado and A. da C. Machado, “Identification of Microsatellites Loci in Olive (Olea eu- ropaea) and Their Characterization in Italian and Iberian Trees,” Molecular Ecology, Vol. 9, No. 8, 2000, pp. 1171-1173. [12] C. Breton, G. Besnard and A. Bervillé, “Using Multiple Types of Molecular Markers to Understand Olive Phy- logeography,” In: M. A. Zeder, D. Decker-Walters, D. Bradley, B. Smith, Eds., Documenting Domestication: New Genetic and Archaeological Paradigms, University of California Press, California, 2005. [13] C. Breton, “Reconstruction de l’Histoire De L’Olivier (Olea europaea subsp. europaea) et de son Processus de Domestication en Region Mediterraneenne, Etudies sur des Bases Moleculaires,” Ph.D. Biologie des Populations et Ecologie, Paul Cézanne France, 2006. [14] C. Breton, M. Tersac and A. Bervillé, “SSR Genetic Di- versity in Wild Olive (Oleaster, Olea europaea L.) Sug- gests Several Plio-Pleistocene Refuge Zones in the Medi- terranean Basin and Gene Flow with Olive,” Journal of Biogeography, Vol. 33, No. 11, 2006, pp. 1916-1928. [15] G. Besnard and A. Bervillé, “On Chloroplast DNA Varia- tions in the Olive (Olea europaea L.) Complex: Com- pareson of RFLP and PCR Polymorphisms,” Theoretical and Applied Genetics, Vol. 104, No. 8, 2002, pp. 1157- 1163. [16] G. Besnard, B. Khadari, P. Baradat and A. Bervillé, “Olea europaea (Oleaceae) Phylogeography Based on Chloro- plast DNA Polymorphism,” Theoretical and Applied Ge- netics, Vol. 104, No. 8, 2002, pp. 1353-1361. [17] G. Besnard, P. Baradat, C. Breton, B. Khadari and A. Bervillé, “Olive Domestication from Structure of Oleas- ters and Cultivars Using Nuvlear RAPDs and Mitochon- drial RFLPs,” Genetics Selection Evolution, Vol. 33, Special Issue, 2001, pp. S251-S268. [18] G. Besnard, P. Baradat and A. Bervillé, “Genetic Rela- tionship in the Olive (Ole europaea L.) Reflect Multilocal Selection of Cultivars,” Theoretical and Applied Genetics,  Genetic Relationships between Cultivated and Wild Olive Trees (Olea europaea L. var. europaea and var. sylvestris) Based on Nuclear and Chloroplast SSR Markers Copyright © 2010 SciRes. NR 103 Vol. 102, No. 2-3, 2001, pp. 251-258. [19] C. Sperisen, U. Büchler, F. Gugerli, G. Matyas, T. Gebu- rek and G. G. Vendramin, “Tandem Repeat on Plant Mi- tochondrial Genomes: Application to the Analysis of Population Differentiation in the Conofer Norway Spruce,” Molecular Ecology, Vol. 10, No. 1, 2001, pp. 257-263. [20] W. Powell, M. Morgante, C. Andre, J. W. McNicol, G. C. Machray, J. J. Doyle, S. V. Tingey and J. A. Rafalski, “hypervariable Chloroplast Microsatellites Provide a General Source of Polymorphic DNA Markers for the Chloroplast Genome,” Current Biology, Vol. 5, No. 9, 1995, pp. 1023-1029. [21] G. Besnard, B. Khadari, P. Villemur and A. Bervillé, “A Cytoplasmic Male Sterility in Olive Cultivars (Olea eu- ropaea L.): Phenotypic, Genetic and Molecular Ap- proaches,” Theoretical and Applied Genetics, Vol. 100, No. 7, 2000, pp. 1018-1024. [22] N. Minangoin, “L’olivier en Tunisie,” Direction de l’Agriculture et du Commerce Imprimerie Rapide, Nicolas L., 1901. [23] W. Taamalli, F. Geuna, S. B. Temime, D. Bassi, D. Daoud and M. Zarrouk, “Using Microsatellite Markers to Characterise the Main Tunisian Olive Cultivars ‘Chem- lali’ and ‘Chétoui’,” Journal of Horticultural Science and Biotechnology, Vol. 82, No. 1, 2007, pp. 25-28. [24] B. Khadari, C. Breton, N. Moutier, J. P. Roger, G. Besnard and A. Bervillé, “The Use of Molecular Markers for Germplasm Management in a French Olive Collec- tion,” Theoretical and Applied Genetics, Vol. 106, No. 3, 2003, pp. 521-529. [25] F. Carriero, G. Fontanazza, F. Cellini and G. Giorio, “Identification of Simple Sequence Repeats SSRs (GAPU) in Olive Olea europaea L,” Theoretical and Applied Ge- netics, Vol. 104, No. 2-3, 2002, pp. 301-307. [26] W. S. Oetting, H. K. Lee, D. J. Flanders, G. L. Wiesner, T. A. Sellers and R. A. King, “Linkage Analysis with Multi- plexed Short Tandem Repeat Polymorphisms Using In- frared Fluorescence and M13 Tailed Primers,” Genomics, Vol. 30, No. 3, 1995, pp. 450-458. [27] M. Nei, “Genetic Distance between Populations,” The American Naturalist, Vol. 106, No. 949, 1972, pp. 283- 292. [28] J. Felsenstein, “PHYLIP (Phylogeny Inference Package),” Version 3.5c. Department of Genetics, University of Washington, Seattle, 1993. [29] C. Breton, C. Pinatel, F. Médail, F. Bonhomme and A. Bervillé, “Comparison between Classical and Bayesian Methods to Investigate the History of Olive Cultivars Using SSR – Polymorphisms,” Plant Science, Vol. 175, No. 4, 2008, pp. 524-532. [30] L. Baldoni, N. Tosti, C. Ricciolini, A. Belaj, S. Arcioni, S, G. Pannelli, M. A. Germana, M. Mulas and A. Porceddu, “Genetic Structure of Wild and Cultivated Olives in the Central Mediterranean Basin,” Annals of Botany, Vol. 98, No. 5, 2006, pp. 935-942. [31] J. Gressel, “Crop Ferality and Volunteerism,” Taylor and Francis Boca Raton, 2005. [32] J. Ruby, “Recherche Morphologique et Biologique sur l’Olivier et sur ses Variétés Cultivées en France,” Annales des Sciences Naturelles Botanique, 1918, pp. 1-286. [33] G. Valdeyron and P. Crossa-Raynaud, “Les Fruits de Tunisie,” Annales de Services Botanique et Agronomique de Tunisie, 1950, pp. 23-44. [34] A. Koutsaftakis, F. Kotsifakis, E. Stefanoudaki and A. Cert, “Etude Triennale sur les Variations de Plusieurs Caractéristiques Chimiques et de Divers Composants Mineurs des Huiles d’olive Vierges Obtenues à Partir d’Olives Cueillies à Différents Degrés de Maturité,” Olivae, Vol. 80, 2000, pp. 22-27. [35] H. Hannachi, M. Msallem, M. El Gazzah and S. Ben Elhadj, “Etude de la Variabilité Pomologique des Olives et de la Composition en Acides Gras des Huiles de 15 Variétés d’Olivier Tunisiens (Olea europaea L.),” Revue des Régions Arides, Vol. 17, No. 1, 2006, pp. 43-64. [36] H. Camps-Fabrer, “L’Olivier 1ère Partie, L’Olivier et l’Huile dans l’Afrique Romaine,” Gouvernement Général de l’Algérie, Direction de l’Intérieur et des Beaux Arts, Service des Antiquités, Imp. Off., Alger, 1953. [37] R. Lumaret, N. Ouazzani, H. Michaud and P. Villemur, “Cultivated Olive and Oleaster, Two Very Closely Con- nected Partners of the Same Species (Olea europaea). Evidence from Enzyme Polymorphism,” Bocconea, Vol. 7, 1997, pp. 39-42. [38] R. Lumaret and N. Ouazzani, “Ancient Wild Olives in Mediterranean Forests,” Nature, Vol. 413, No. 6857, 2001, p. 700. [39] M. Amane, N. Ouazzani, R. Lumaret and C. Debain, “Chloroplast-DNA Variation in the Wild and Cultivated Olives (Olea europae L.) of Morocco,” Euphytica, Vol. 116, No. 1. 2000, pp. 59-64. [40] J. Hess, J. W. Kadereit and P. Vargas, “The Colonization History of L. in Macaronesia Based on Internal Tran- scribed Spacer 1 (ITS-1) Sequences, Randomly Ampli- fied Polymorphic DNAs (RAPD) and Intersimple Se- quence Repeats (ISSR),” Molecular Ecology, Vol. 9, No 7, 2000, pp. 857-868. [41] A. Contento, M. Ceccarelli, M. T. Gelati, F. Maggini, L. Baldoni and P. G. Cionini, “Diversity of Olea Genotypes and the Origin of Cultivated Olives,” Theoretical and Ap- plied Genetics, Vol. 104, No. 8, 2002, pp. 1229-1238. [42] G. T. Mekuria, G. Collins and M. Sedgley, “Genetic Di- versity within an Isolated Olive Olea Europaea L Popula- tion in Relation to Feral Spread,” Scientia Horticulturae, Vol. 94, No. 1, 2002, pp. 91-105. [43] M. Sedgley, “Wild Olive Selection for Quality Oil Pro- duction,” A Report for the Rural Industries Research and Development Corporation, RIRDC Publication No 04/ 101, 2004. |

