Advances in Bioscience and Biotechnology
Vol.06 No.02(2015), Article ID:53871,15 pages
10.4236/abb.2015.62008
Protein Folding Mediated by an Intramolecular Chaperone: The Energy Landscape for Unimolecular Pro-Subtilisin E Maturation
Ezhilkani Subbian, Danielle M. Williamson, Ujwal Shinde
Department of Biochemistry and Molecular Biology, Oregon Health & Science University, Portland, USA
Email: shindeu@ohsu.edu
Copyright © 2015 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 21 January 2015; accepted 6 February 2015; published 9 February 2015
ABSTRACT
Efficient and precise assembly of polypeptides into native functional states is critical for normal cellular processes. Understanding how a specific structure is encoded in the polypeptide sequence and what drives the structural progression to the native state is essential to deciphering the folding problem. Several prokaryotic and eukaryotic proteins require their propeptide-domains to function as dedicated intramolecular chaperones (IMCs). In this manuscript, we investigate the elementary steps in the IMC mediated maturation of Subtilisin E, a bacterial serine protease, and a prototype for the eukaryotic proprotein convertases (PCs). Through detailed analyses, we have at- tempted to define the unimolecular folding energy landscape for SbtE to understand how the stabilization of folding intermediates influences the maturation process, an aspect that is difficult to study in eukaryotic PCs. Our studies demonstrate that a rapid hydrophobic collapse precedes acquisition of tertiary structure during the folding of Pro-SbtE and results in formation of a molten- globule like intermediate. Induction of structure within the IMC stabilizes both the molten globule-like folding intermediate and the native state, and appears to expedite initial stages of folding, purely through thermodynamic stabilization of the folded state. While the induced structure does not affect the activation energies in the unimolecular folding reaction, it is detrimental to the autoproteolytic cleavage of the precursor and subsequent release and degradation of the inhibitory IMC-domain since both these stages require some degree of unfolding. Completion of Pro- SbtE maturation results in the formation of a kinetically trapped and extremely stable native state. Hence, our results suggest that the SbtE IMC appears to have evolved to be intrinsically unstructured and to bind with its cognate protease with a specific affinity that is critical for biological regulation.
Keywords:
Propeptide Dependent Folding, Protease Activation, Subtilases, Energetics, Proprotein Convertases

1. Introduction
Protein folding is one of the simplest and most fundamental examples of biological self-assembly [1] . Uncovering the principles of protein folding not only helps us outline the factors that drive this process, but also is the first step in developing rational cures for folding diseases, and designing novel proteins. The broad question of folding is concerned with deciphering how a complex three-dimensional structure is encoded in within a one- dimensional polypeptide sequence, and how the polypeptide attains this native state in a limited time [2] [3] . Extensive studies have led to the new view of folding, which postulates that the native state is attained by a stochastic search of all possible conformations, aided by the stability of sub-structures that are formed [4] [5] . Substantial progress towards this goal has been made for a handful of proteins over the past few years [6] -[8] . However, many of these studies have been based on relatively small proteins that fold with fast kinetics, and in the absence of stable intermediate states [4] [9] . On the other hand, the average biological molecule is large (~300 amino acids), and folds through the formation of stable folding intermediates [10] . Stable intermediates can provide valuable information about the development of the overall fold from a linear polypeptide, and can also help examine whether their existence facilitates the process, or serves as kinetic traps that impede folding [11] -[13] . Characterization of these states, including partially structured folding intermediates, unfolded states and transition states that a protein encounters during the folding process are essential to realize our goal of how proteins fold at an atomic level [6] -[8] [14] .
Subtilisin E (SbtE) is a calcium-dependent extracellular serine protease that is secreted as a precursor (Pro- SbtE) and is an established model for understanding IMC-dependent folding of both prokaryotic proteases as well as eukaryotic orthologues, such as the proprotein convertases (PCs) [15] -[22] . In absence of its 77-residue IMC, SbtE (275-residue) folds to a stable, molten-globule intermediate that is separated from its kinetically- trapped native state by a high-energy barrier [23] [24] . Spontaneous conversion to the native state is slow (T1/2 ~ 1500 yrs), but is catalyzed ~106-fold by the propeptide that functions as a dedicated intramolecular chaperone (IMC) and lowers this high-energy barrier [24] . However, the IMC remains tightly associated with the protease and subsequent degradation of the IMC releases enzymatic activity [25] [26] . Degradation of the IMC enables the uncoupling of folding and unfolding pathways, and stabilizes the native state by increasing the transition state for the unfolding reaction [16] [17] . Degradation of the IMC allows facile regulation of protease activation through exogenous signals such as changes in pH and salt [22] [27] [28] . It is important to note that SbtE is a member of a large family of serine proteases termed subtilases [29] , most members of which fold using IMC-dependent mechanisms [17] [30] [31] . As aforementioned, SbtE is orthologous to the eukaryotic PCs [19] - [21] [32] , which have the significant import on human health and disease [33] . The PCs are likewise calcium dependent endoproteases, and are responsible for mediating a diverse range of regulatory and protective pro- cesses by controlled proteolysis of their substrates [33] -[37] . Each PC catalyzes the maturation of proprotein and prohormone substrates within specific compartments of the secretory pathway of the eukaryotic cell, a task that requires precise spatiotemporal regulation of their own folding and activation. As with SbtE, this control is exerted by the IMCs; misregulation of PC activity has been shown to cause cancer, diabetes, obesity, and heart disease [34] -[37] . Much of our comprehension of the thermodynamics and kinetics of IMC-mediated folding has emerged through detailed analysis of bacterial extracellular proteases [17] [38] . Understanding IMC-mediated folding mechanisms gives not only fundamental insights into the energy landscape of proteins, but also the role of intrinsic chaperones in the overall folding process. Moreover, these studies help elucidate why nature selects between chaperone dependent and independent folding pathways. In this manuscript, we use the unimolecular folding energy landscape of maturation of subtilisin E as a test case for the larger family of IMC-dependent folding mechanisms, defining intermediates and analyzing how stabilization of these intermediates affects the folding landscape.
Studies pertaining to the folding kinetics and thermodynamics of SbtE have been done in a bimolecular reaction by the addition of the IMC in trans [17] . Such studies are limited by the formation of the initial IMC-subti- lisin complex [39] and do not provide information as to why IMC-independent folding is slow, or how the IMC helps to lower the transition barrier to folding. Furthermore, the biological folding reaction is clearly unimolecular. Here, we undertake a detailed characterization of the folding mechanism and energy landscape of unimolecular, IMC-mediated maturation of Pro-SbtE, and establish how properties of the IMC affect this landscape. Our studies demonstrate that a rapid hydrophobic collapse precedes acquisition of tertiary structure during the folding of Pro-SbtE. Furthermore, induction of structure within the IMC stabilizes both the hydrophobic molten- globule intermediate and the native state and appears to expedite initial stages of folding, purely through thermodynamic stabilization of the folded state. It is interesting to note that the induced structure does not affect the activation energies of the folding. However, the induced structure in the unimolecular folding reaction is detrimental to the activation of the precursor, since this stage requires some degree of unfolding. Our results suggest that the SbtE IMC appears to have evolved to be intrinsically unstructured and to bind with its cognate protease with a specific affinity that is critical for biological regulation.
2. Results
2.1. Maturation Pathway of Pro-SbtE
A complete understanding of the energy landscape of maturation requires 1) determination of the structures of all conformational states along this pathway, including the denatured state, possible intermediates and the native state, 2) a detailed analysis of the thermodynamics and kinetics of these states and, 3) the nature of the transi- tion barriers that separate these states. Our previous studies have established that the unimolecular maturation pathway of Pro-SbtE involves the initial folding of the polypeptide to a stable pre-processed complex (Figure 1,
Figure 1. Schematic representation of Pro-SbtE maturation pathway schematic representation of Pro-SbtE maturation pathway. Maturation of Pro-SbtE can be dissected into the three stages of folding, autoprocessing and, release and degradation. Pro-S221A-SbtE represents an active site mutant that folds to a structured state and is blocked at the stage of folding (uncleaved complex). Pro-S221C-SbtE mutant is able to fold and undergo autoprocessing to result in a structured inhibited complex (Cleaved complex; 1SCJ.pdb) but is unable to release and degrade its IMC. Wildtype Pro-SbtE undergoes complete maturation to yield active protease (1Sbt.pdb).
Stage I), cleavage of the peptide bond between the IMC and the protease (Stage II), and the rate limiting release and degradation of the IMC from the processed complex (Stage III) [26] . Over the years we have isolated specific mutations that enable us to establish the nature of each of the individual stages of this pathway (Figure 1). For example, the active site mutant Pro-S221A-SbtE folds to a structured state (Figure 2) but loses its ability to cleave and degrade its IMC, and represents the maturation intermediate prior to being processed (Stage I) [40] . Similarly Pro-S221C-SbtE variant folds into a structured state (Figure 2) that can auto process its cognate IMC but is unable to subsequently degrade it (Figure 1). This represents the autoprocessed maturation intermediate (Stage II) [41] [42] . Hence, the above active-site variants together with wild type SbtE provide an ideal system to establish the thermodynamics and kinetics of the individual steps of the maturation pathway, and to analyze the implications of the synergy between these different stages.
The structures of mature SbtE (Stage III) and Pro-S221C-SbtE complex (Stage II) have been solved using X- ray crystallography [41] [43] . These structures demonstrate that the conformations of the protease domains are identical in the last two stages of maturation and are unaffected by the active-site substitution. The structure of the uncleaved precursor for pro-SbtE (Stage I) has not been solved. However, the structures of the unautoprocessed, autoprocessed and the mature forms for a related homologue [44] ; establishes that the protease domain in the uncleaved precursor is similar to the wild type protease, but has a distorted 6-residue N-terminus that is covalently attached to the C-terminus of the IMC [45] . Hence, high-resolution structures of the stable maturation intermediates are available. While the thermodynamic properties of the intermediates can be studied at equilibrium, the properties of the high-energy and metastable transition states can be inferred from kinetic studies.
2.2. Hydrophobic Collapse Coincides with Secondary Structure Formation but Precedes Tertiary Structure Acquisition
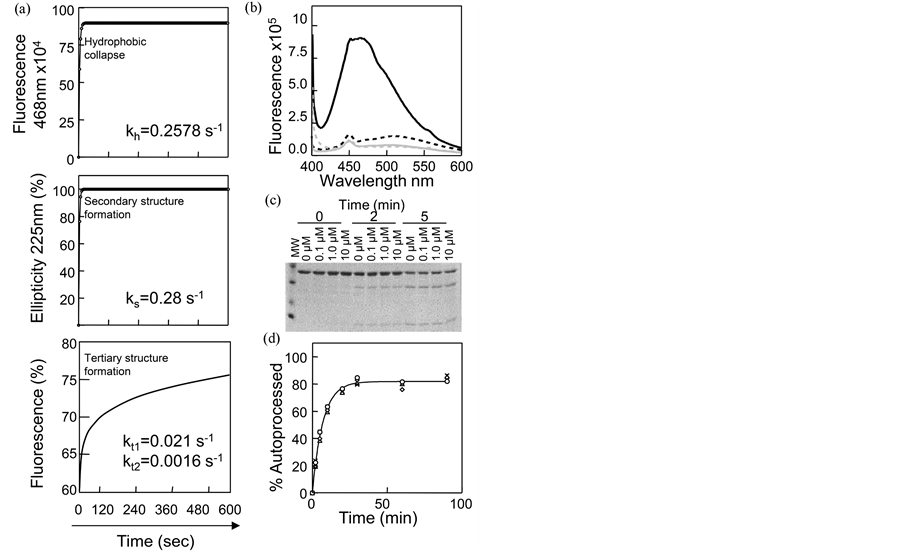
While earlier studies established that the IMC is indispensable to the folding of SbtE, what initiates/drives the Pro-SbtE folding process remains unknown. It is widely accepted that hydrophobicity provides the major driving force for most protein folding reaction. Since the kinetics of the hydrophobic core formation, relative to those for secondary and tertiary structure formation, can provide insights into folding mechanisms [11] , we performed a detailed analysis of the folding kinetics for Pro-SbtE. The formation of a hydrophobic core during folding of a protein can be monitored under both equilibrium and kinetic conditions by using dyes such as 1-anilino 8-napthalene sulphonic acid (ANS). ANS specifically interacts with exposed hydrophobic surfaces on proteins, and upon binding, exhibits an increase in the fluorescence emission intensity. This approach has been successfully utilized to establish the presence of partially structured molten-globule intermediates in several protein- folding models [11] [46] . Figure 3(a) establishes that when denatured Pro-SbtE (0.65 µM) is rapidly diluted (30 fold) into a refolding buffer that contains 6.0 µM ANS, a rapid burst in the fluorescence emission at 458 nm is observed. The fluorescence emission spectra can be fitted to a single exponential rate equation and provides a rate constant (kh = 0.26 s−1) for the formation of the hydrophobic core. It is important to note that in the presence of ANS, only partially folded Pro-SbtE (structure at 600 s) and not its denatured state (6 M GdnHCl), or mature
Figure 2. Secondary structure characterization using circular dichroism (CD) spectroscopy. CD spectra of Pro-SbtE and its active-site mutants: Pro-S221C- SbtE inhibition complex (solid black line), Pro-S221A-SbtE uncleaved complex (dotted grey line), SbtE (dotted black line), isolated IMC (solid grey line) and differential spectra of IMC in complex (brokenline).
SbtE, exhibits significant fluorescence intensity (Figure 3(b)). Hence, the increase in fluorescence reflects the rapid formation of the hydrophobic core of Pro-SbtE (Figure 3(a)).
We next monitored the kinetics of secondary and tertiary structure acquisition of Pro-SbtE. Folding was initiated as described in Methods. Acquisition of secondary structure was monitored as a function of time, using the changes in circular dichroism (CD) ellipticity at 222 nm, while the tertiary structure was monitored using changes in intrinsic tryptophan fluorescence (Figure 3(a)). The kinetics obtained using CD-spectroscopy best fit a single exponential rate equation with a rate constant of 0.35 s−1 at 21˚C. However, the intrinsic fluorescence profiles best fit a double exponential rate equation with rate constants of 0.021 s−1 and 0.0016 s−1 for the fast and slow phases, respectively. The two phases may be consequences of proline isomerization or may reflect the modular folding of the protease domain followed by the relatively slower folding of the IMC domain [17] . Interestingly, the rate constants describing the tertiary structure acquisition are significantly slower than the rate constant for the secondary structure (0.28 s−1) or hydrophobic collapse (0.26 s−1) (Table 1). Hence, these results suggest that a rapid hydrophobic collapse that serves to minimize conformational entropy of Pro-SbtE precedes the relatively slower acquisition of tertiary structure.
It has been argued that hydrophobic dyes such as ANS may bind and stabilize normally transient folding species and shift their equilibrium towards stable intermediates [47] . To examine the effects of ANS on the maturation pathway, we monitored the kinetics of folding and autoprocessing of Pro-SbtE using quantitative SDS- PAGE (Figure 3(c)). The extent of autoprocessing was measured as a function of time (Figure 3(d)) and establishes that the kinetics remain unaffected in the presence of ~15-fold molar excess of ANS (10 µM). Hence, our results suggest that the hydrophobic dye does not appear to increase stability of the folding intermediate.
Figure 3. Characterization of initial collapse and folding of the precursor: (a) Kinetics of Pro-SbtE folding, monitored by changes in ANS fluorescence (upper), CD ellipticity (middle) and intrinsic fluorescence (lower); (b) Fluorescence emission spectra of ANS: dotted grey line―ANS control; solid black line―Pro-SbtE 600 s after folding initiation; dotted black line―denatured Pro-SbtE; and solid grey line―mature subtilisin; (c) SDS-PAGE of aliquots taken at 0.2 and 5 min after initiation of folding, in presence of denoted amounts of ANS; (d) Extent of autoprocessing, as a function of increasing ANS concentrations, estimated from SDS-PAGE analysis by quantitative gel-scanning densitometry; open circles―0 µM ANS; open diamond―0.1 µM ANS; open triangles―1 µM ANS; inverted triangles―10 µM ANS.
Table 1. Thermodynamic parameters for Stages I, II, and III of SubtE maturation.
2.3. Energetic and Structural Analysis of Stage 1: Pro-Subtilisin Folding
2.3.1. Kinetics of Stage I Analyzed Using the Pro-S221A-SbtE Variant: The Folding Reaction
To obtain insights into the nature of the folding transition state, we next examined the dependence of folding kinetics on the temperature of the reaction. Such an analysis can also yield information about the entropy, enthalpy, heat capacity, flexibility of the polypeptide backbone, as well as the solvation of a protein in its transition state [13] . The kinetics of Pro-S221A-SbtE was monitored as described in Methods, and the temperature dependence of the folding kinetics was established over the range of 0˚C - 25˚C. The Eyring plots thus obtained establish that the rate of folding (kf) is non-linear over the temperature range investigated (Figure 4(a)). Non-linear Eyring plots have been observed in several other proteins. Non-linearity has been attributed to one of several reasons, the most common of which are 1) change in heat capacity, 2) change in the rate-limiting step or 3) the presence of a folding intermediate. To examine possible causes, the kinetics were fitted to Equation (2), to obtain relative changes in the free energy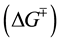 , entropy
, entropy 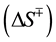 and heat capacity
and heat capacity 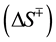 and hence the overall enthalpy
and hence the overall enthalpy 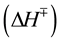 of the transition states. The thermodynamic parameters obtained by best-fits of the Eyring plots are depicted in Table 1 and indicate that the net change in heat capacity between the unfolded and transition state is negligible (~0.1 kcal/mol・K). While a temperature dependent change in the rate-limiting step cannot be overruled, the observed non-linearity in the Eyring plot appears to be most likely due to the presence of a stable folding intermediate (see next section).
of the transition states. The thermodynamic parameters obtained by best-fits of the Eyring plots are depicted in Table 1 and indicate that the net change in heat capacity between the unfolded and transition state is negligible (~0.1 kcal/mol・K). While a temperature dependent change in the rate-limiting step cannot be overruled, the observed non-linearity in the Eyring plot appears to be most likely due to the presence of a stable folding intermediate (see next section).
Previous studies from our laboratory and others have demonstrated that the isolated IMC is an intrinsically unstructured domain [48] [49] . Why the IMC is intrinsically unstructured and how this affects the folding landscape is unknown. Studies establish that the IMC folds into a well-defined α-β structure in the presence of co- solvents such as 10% glycerol and 25% trifluoroethanol [50] . Such concentrations of glycerol do not substantially alter the viscosity of the solution, nor the activity of the protease [51] . Further, the co-solvent induced secondary structure appears to be similar to the structure of the IMC in complex with the protease domain. Introduction of specific mutations can also significantly increase the native-like secondary structure within the isolated IMC domain [52] . The IMC variants with increased structural stability are invariably better chaperones and tighter inhibitors [25] [53] . However, the effects of mutations are cumulative on the entire maturation pathway, and the delineation of their effects on individual stages becomes difficult. Hence, instead of introducing mutations in the IMC, we analyze the folding reaction and the transition state (Stage I), in the presence of 10% glycerol as a co-solvent.
The folding reaction in the presence of glycerol also follows single exponential kinetics with a non-linear dependence upon temperature (Figure 4(a)). The rate of folding is ~2-fold faster in the presence of 10% glycerol, and is consistent with data obtained using stabilizing IMC mutants [25] [53] . The thermodynamic parameters obtained from the fit of the folding kinetics to Equation (2) are depicted in Table 1. The activation energy barriers of the folding reaction in the absence (~18.0 kcal/mol・K) and presence of glycerol (~17.8 kcal/mol・K) suggests that the induced structure does not significantly lower the free energy of the folding transition state. It is however interesting to note that the overall entropy of the transition state is reduced in the presence of glycerol and is compensated by a gain in enthalpy probably due to packing of the side chains (Table 1). This is again consistent with less exposed hydrophobic surface area in the presence of 10% glycerol (data not shown). Hence,
Figure 4. Energetics of Stage I―folding. (a) Temperature dependence of the kinetics of foldingunder normal conditions (filled squares), and in the presence of 10% glycerol (filled triangles); (b) Equilibrium unfolding of folded Pro-S221A-SbtE under normal conditions (filled circles), and in the presence of 10% glycerol (open circles). Deconvoluted constituent fractions under both normal and glycerol conditions are depicted; native (N)―dotted line; intermediate (I)―solidblack line and unfolded (U)―broken line; (c) Temperature dependence of the kinetics of unfolding from the U-state to the I-state under normal conditions (squares), and in the presence of 10% glycerol (triangles).
our results suggest that the induction of structure in the IMC does not significantly lower the activation energy barrier to folding.
2.3.2. Thermodynamic Equilibrium of Stage I Analyzed Using the Pro-S221A-SbtE Variant
To examine thermodynamic stability of the uncleaved precursor, the equilibrium unfolding of Pro-S221A-SbtE was monitored as a function of denaturant concentrations (Figure 4(b)). Under non-denaturing conditions, Pro- S221A-SbtE adopts a well-defined structure as observed using CD (Figure 2). Under increasing concentrations of denaturants Pro-S221A-SbtE unfolds through a partially structured state, which represents a productive folding intermediate (I), as established earlier. This structural transition is completely reversible, and suggests that the folded but un-cleaved precursor adopts a thermodynamically stable conformation. The folding/unfolding profiles best fit a three state transition (Figure 4(b)) and indicate that the complex at Stage I is stabilized by a free energy difference of ~DG0 = −4.6 kcal/mol∙K, relative to the unfolded state. The I-state appears maximally populated at denaturant concentration between ~1.0 to 1.2 M GdnHCl (Figure 4(b)), and has a net free energy of stabilization of ~2.9 kcal/mol∙K, relative to the folded state. The I-state therefore is marginally stable (~1.7 kcal/mol∙K) compared to the unfolded state and exhibits the characteristics of a partially structured molten- globule intermediate (Figure 4(b)). It is important to note that the non-linearity observed in the temperature dependent folding kinetics (Figure 4(a)) may be a consequence of this intermediate.
We next examined how inducing structure within the IMC (using 10% glycerol as a co-solvent) affects equilibrium folding/unfolding of Pro-S221A-SbtE. Even under these conditions the complex exhibits a distinct three-state transition (Figure 4(b)). However, stabilities of the intermediate and native states are enhanced by ~2.0 and ~2.6 kcal/mol∙K, respectively. Further, the stable I-state is maximally populated at ~2.25 M GdnHCl (Figure 4(b)). Hence, our data suggests that a structured IMC stabilizes both the folded state and the I-state.
2.3.3. Kinetics of Stage I Analyzed Using the Pro-S221A-SbtE Variant: The Unfolding Reaction
To obtain insights into the transition state separating the I- from the N-state, folded Pro-S221A-SbtE was rapidly diluted into denaturant concentrations wherein the intermediate species are maximally populated. The loss in ellipticity at 222 nm was monitored using CD spectroscopy and a typical unfolding curve is shown in Figure 4(c) (inset). The temperature dependence of the unfolding kinetics from the N-state to I-state follows strong non- linearity, both with and without glycerol (Figure 4(c)). Using Equation (2) the activation free energies, entropies, and activation heat capacities for the unfolding reaction were estimated. The activation enthalpies were then calculated from the free energies and the entropies. It is interesting to note that the rates of unfolding of the precursor to its intermediate state is faster in the presence of glycerol at lower temperatures (Figure 4(c)), and the trend is reversed at higher temperature (above 17˚C). While reasons for this change in dependence are currently unknown, one possible cause may be the presence of additional intermediates that are not resolved using equilibrium unfolding. Alternatively, it is possible that this non-linearity may be due to a change in the rate-deter- mining step of the unfolding reaction.
2.4. Energetic and Structural Analysis of Stage II: Pro-Subtilisin Autoprocessing
Pro-subtilisin processing involves the autoproteolytic cleavage of the peptide bond between the C-terminus of the IMC and the N-terminus of the protease domain and results in a non-covalently associated stoichiometric complex [40] [42] . Auto-cleavage is accompanied by structural rearrangements within the complex, which coincides with the formation of a high-affinity calcium-binding site [42] . To establish the energetics associated with autoprocessing, we measured the temperature dependence of this reaction using Pro-S221C-SbtE variant, both in the presence and absence of glycerol. This variant is ideal for these studies since its rate of folding is similar to Pro-SbtE, while its rates of autoprocessing are approximately 100-fold slower [41] [54] . Hence, unlike Pro-SbtE insignificant levels of autoprocessing are observed during the folding of the Pro-S221C-SbtE variant. Moreover, Pro-S221C-SbtE is unable to degrade its IMC and this prevents trans proteolysis. This allows one to fold the precursor at a fixed temperature (4˚C) and then measure the kinetics of auto-cleavage at any desired temperature using a quantitative SDS-PAGE analysis as described in Methods. Figure 5(a) depicts a representative kinetic
Figure 5. Energetics of Stage II―autoprocessing. (a) Kinetics of autoprocessing estimated from SDS-PAGE (Inset) through quantitative gel-scanning densitometry. Mature protein―open circles; precursor―closed circles; (b) Temperature dependence of autoprocessing kinetics under normal (squares), and glycerol (triangles) conditions; (c) Efficiency of autoprocessing estimated from the relative yield of mature under increasing amounts of denaturant, in the presence (open circles), and absence (filled circles) of glycerol; (d) Equilibrium unfolding of the Pro-S221C-SbtE cleaved complex under normal conditions (upper panel), and in the presence of 10% glycerol (lower panel). The individual fractions obtained from deconvolution of folding curves are Native (N)―dashed grey line; Intermediate (I)―solid grey line and unfolded (U)―broken grey line.
curve for autoprocessing and the rate for this reaction was obtained by fitting the profile to a single exponential equation. The temperature dependence of the rate constant thus obtained provides an Eyring plot for the autoprocessing reaction. The Eyring plot fits equally well to Equation (2) and to a linear function (Equation (3)) and the estimated parameters for the fit are listed in Table 1. It is interesting to note the presence of a “crossover” between the Eyring plots obtained in the presence and absence of glycerol. At this “crossover” temperature the rates of autoprocessing remain unaffected by glycerol, while at lower temperatures the rates of autoprocessing increase. At higher temperatures however, it appears that the increased structural stability of the IMC slows autoprocessing. This mirrors the unfolding behavior of the precursor to its intermediate state (Figure 4(c)). Although reasons for why the rates of unfolding are higher in the presence of glycerol are currently unknown, these results suggest that the kinetics of autoprocessing appear to directly correlate with the kinetics of unfolding of the precursor. Hence, the stabilization of the uncleaved complex appears to be detrimental to autoprocessing. In addition, presence of glycerol in the folding conditions does not lower the activation energy barrier notably as seen from Table 1.
To better understand this behavior we studied the autoprocessing reaction under equilibrium conditions in the presence of varying concentrations of GdnHCl at room temperature. Under non-denaturing conditions, the yield of mature enzyme in the presence of glycerol is lower than that observed in its absence. This suggests that the higher stability (lower unfolding rates) in the presence of glycerol adversely affects autoprocessing. Interestingly, if the stability is decreased by the addition of low concentrations of denaturants (0.75 M GdnHCl) the yield of SbtE is increased. This mirrors the trend observed in the kinetics of unfolding (Figure 4(c)) and autoprocessing (Figure 5(b)). This suggests that partial unfolding maybe essential for efficient autoprocessing. However this is not reflected in the activation energy barriers and requires additional analysis.
Thermodynamic Equilibrium of the Processed Complex
The thermodynamic stability of the autoprocessed complex was established by monitoring the loss in ellipticity as a function of denaturant concentration. It is important to note that this folding/unfolding reaction is reversible due to the presence of the IMC, which can facilitate folding in trans [55] . Analysis of the unfolding curves using Equation (1) establishes the free-energy of unfolding of the autoprocessed complex (Table 2 and Figure 5(b)). The equilibrium unfolding was monitored both in the presence and absence of 10% glycerol. The presence of glycerol significantly increases the stability of the processed complex (DDG = 4.6 kcal/mol/K). Thus, a structured IMC increases the thermodynamic stability of both the uncleaved precursor and the cleaved complex.
2.5. Energetic and Structural Analysis of Stage III: Pro-Subtilisin Activation
The release of the IMC and its subsequent degradation by the active protease represents the rate-determining step to IMC release. Through a careful study, we earlier established the energetics of Pro-SbtE activation [51] . The temperature dependence of the activation kinetics demonstrated that the kinetic barrier of this reaction is ~17.3 ± 0.3 kcal∙mol−1. However, this measured Ea signifies the limiting energy barrier for release and degradation. Since the proteolysis of the IMC is rapid, it should be noted that the energy barrier for release is most likely higher. In the presence of 10% glycerol, the Pro-SbtE inhibition complex is extremely stable, with the protease being inactive till >8 days at 23˚C and >4 weeks at 4˚C. Hence we were unable to determine the kinetics of activation in the presence of glycerol. However, based on the extremely slow activation observed under these conditions, it is reasonable to state that the energy barrier to activation significantly increases in the presence of a structured IMC domain.
Table 2. Parameters for equilibrium unfolding of Pro-S221A and Pro-S221C SbtE.
The thermodynamic stability of the mature protease relative to the auto processed complex was established by monitoring the extent of binding of the IMC to the mature domain. The Ka for this interaction under folding conditions was 7.34 × 106 M−1 and [51] and the IMC binding with SbtE stabilizes the inhibition complex by ~9.3 kcal/mol・K. Thus, the free-energy diagram for activation favors formation of a thermodynamically stable inhibition complex over the active protease (Figure 6). This established that the native state of the protease is not at its thermodynamically stable conformation but is trapped in a local energy minimum by a high barrier to unfolding.
3. Discussion
3.1. Energy Landscape for Pro-Subtilisin Maturation
A complete study of the folding pathway involves the analysis of the structural acquisition from the unfolded state to the native state of a protein. In this study, we define the folding energy landscape of Pro-SbtE using active site variants. This landscape involves three distinct stages of folding, autoprocessing and activation (Figure 1). Our results establish that folding is initiated by a rapid hydrophobic collapse and occurs through the formation of a stable molten globule-like intermediate. Both the initial folding and the subsequent autoprocessing are thermo- dynamically driven processes. Autoprocessing of the uncleaved precursor is accompanied by the formation of the high-affinity calcium-binding site and significantly increases the thermodynamic stability of the complex [42] . This results in a high enthalpic barrier to autoprocessing. Earlier analysis of Pro-SbtE activation demonstrated that the release of the IMC from the complex is energetically unfavorable. Removal of the IMC through proteolysis results in the formation of a kinetically trapped native state. Taken together, the energetics of Stage I, II and II, establish that the native state is also thermodynamically unstable. A similar native state has been observed with the IMC mediated folding of alpha-lytic protease and appears to offer distinct functional advantages.
3.2. Effect of a Structured IMC on the Folding Energy Landscape
The IMC of SbtE is intrinsically unstructured and folds only in the presence of its cognate protease domain. To understand how this structural instability affects maturation we established the energy landscape in the presence of a structured IMC. Our results demonstrate that the IMC does not significantly contribute to the first transition state to folding and only functions subsequent to the formation of the stable intermediate. Further, inducing structure in the IMC does not significantly affect the kinetic barriers to folding and autoprocessing but enhances
Figure 6. Folding energy landscape for Pro-SbtE maturation. Folding of Pro- SbtE (N) is driven by a hydrophobic collapse that precedes the formation of tertiary structure and proceeds through a distinct structured intermediate (I). Autoprocessing of the IMC results in athermodynamically stable inhibited com- plex. Release and degradation of the IMC drives the transition to a kinetically stable native state M (solid black line). A structured IMC (solid greyline) increases the inherent stability of I- and N-states and stabilizes the folding reaction by ~2.5 kcal/mol. Similarly, the stability of the autoprocessed complex is also significantly enhanced (~4.6 kcal/mol). However the release of the IMC is limited by its structural stability and enhances the barrier to activation.
the thermodynamic stability of the structured states. However such stabilization of the uncleaved complex appears to be detrimental to auto-cleavage and dramatically prolongs activation. While this suggests that autoprocessing of the IMC, similar to its release, may require some structural instability within the IMC, the absence of a significant change in the activation energy barrier necessitates further analysis. While structured native states and thermodynamic stability are an accepted norm for a majority of proteins, the kinetically-determined folding pathway of Pro-SbtE is optimized for a structurally unstable IMC domain (Figure 6). Hence in this case of protein folding interestingly kinetic-stability, its consequent functional advantages, and structural instability appear to have co-evolved [15] [21] .
3.3. Subtilisin E and the Proprotein Convertase Family
The eukaryotic proprotein convertases, as well as the related yeast kexins, are homologous protease subfamilies of the bacterial subtilisins [21] . We are just now beginning to fully grasp the role the PCs have in regulating human health and disease; for example, much attention has been given to PCSK9 in recent years for its role in regulation of LDL, and the role of PC1/3 in nutrient sensing and orexigenic control continues to generate interest in the endocrine field [33] . Regulation of the PCs on a molecular levels remains poorly understood, however we believe that in building a better understanding of the folding and maturation of subtilisin, we can extend these findings to make directed hypotheses regarding these processes in the PCs [33] . The activation pathway of the PCs is largely conserved with reference to that of subtilisin; the IMC, and its autoprocessing, guides folding of the catalytic domain and directs its egress and entry from one compartment of the secretory pathway to the next [20] . The similarities between the maturation pathways of subtilisin and the PCs have been most intensively studied and well described in furin, considered the canonical PC. These studies have indicated a second level of control is required in the PCs; as an evolutionary adaptation to the changing pH of the secretory pathway [19] [22] , we have previously demonstrated that protonation of a conserved histidine (His69 in furin) is necessary and sufficient for the cleavage and release of the IMC, and thus activation of furin. However, while this histidine residue is conserved within the other PCs, each homologue exhibits its own pH specificity of activation, thus leading us to question what additional factors play into this exquisite control mechanism? From our understanding of the activation of subtilisin, we posited that protonation of the pH-sensing histidine induces local conformational changes within the loop of the IMC, driving a destabilization of the core that results in the degradation of the IMC, a hypothesis that we have recently tested and found data to support [20] . Given the results presented here, we can further speculate that differences in the stability of the IMCs of the PCs, modulated by the protonation status of pH-sensing histidine residues, may have been fine-tuned by nature over evolutionary time to allow for the differential activation of the PCs within different compartments of the eukaryotic cell [32] . Only further research can definitively answer these questions, and it is clear that insights from SbtE can be critical in making important advances in the larger arena of protein folding, with implications bridging the gap between basic and clinical science.
4. Conclusion
In the results presented above, we offer a mechanistic and energetic characterization of the unimolecular, IMC- mediated maturation of pro-subtilisin E, with particular interest in how properties of the IMC, a propeptide that is not part of the mature, functional protease, and affects the landscape of subtilisin folding. We propose that there is a rapid hydrophobic collapse of SbtE that drives acquisition of tertiary structure via thermodynamic stabilization of the hydrophobic molten globular intermediate and the native state. From an evolutionary perspective, the need for biological regulation appears to have necessitated the IMC be intrinsically unstructured until binding with its cognate protease. While relatively simple in comparison to the larger family of subtilases, especially eukaryotic subtilases such as the kexins and proprotein convertases, establishing the determinants and landscape of folding in this IMC-dependent system can lend powerful insights into why nature has selected chaperone-dependent versus chaperone-independent folding pathways in specific cases to allow for precise spatial and temporal regulation of protease activity.
5. Materials and Methods
5.1. Circular Dichroism Spectroscopy Measurements
CD measurements were performed on an automated AVIV 215 spectrophotometer maintained at 23˚C and spectra were taken between 190 to 260 nm as described earlier [42] [56] [57] . Protein concentrations were maintained between 0.25 mg to 0.4 mg ml−1. A 1 mm path-length cuvette was used to measure spectra except in case of the IMC, where a 0.5 mm path-length cuvette was used. Each plot represents the average of 3 independent scans.
5.2. Kinetics of Folding and Unfolding
Proteins were expressed and purified as described earlier [26] [40] [41] . Folding kinetics were measured by monitoring changes in CD ellipticity, extrinsic fluorescence using ANS and intrinsic tryptophan fluorescence. To monitor the kinetics by CD, Pro-SbtE (in 6 M GdnHCl, pH 4.8) was rapidly diluted into folding buffer (50 mM MES-NaOH, pH 6.5, 0.5 M (NH4)2SO4, 1 mM CaCl2), with or without 10% Glycerol, and changes in CD ellipticity were monitored at 225 nm. To establish the effect of temperature, folding kinetics were monitored as described above, from 4˚C - 25˚C. Changes in fluorescence were measured on a Hi-Tech SF61-DX2 stopped flow fluorimeter using λexcitation of 295 nm and 310 nm band-pass filter, using similar conditions as the CD measurements. To monitor the initial hydrophobic collapse, 5 μM of ANS was included in the folding buffer. The sample was excited at 390 nm and emission was monitored at 468 nm.
To estimate the kinetics of unfolding, fully folded Pro-S221A-SbtE was rapidly diluted into folding buffer containing either 1.1 M GdnHCl or, 2.25 M GdnHCl and 10% glycerol, and the loss in structural content was monitored by CD. The concentrations of denaturant represent conditions at which maximum fractions of intermediate (I) are seen at equilibrium. Folding curves were analyzed using GraphPad Prism and the temperature dependence of the microscopic rate constant (ki) was analyzed according to the equation:
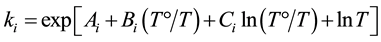 (1)
(1)
where:
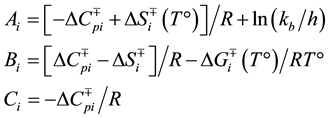
Effect of ANS on folding: Pro-SbtE (200 nM) was folded in the presence of increasing amount of ANS (0 - 10 mM). 20 ml aliquots were taken at different time points after folding initiation, the reaction was stopped by TCA, and samples were analyzed through SDS-PAGE and gel-scanning densitometry.
Equilibrium unfolding: Equilibrium folding-unfolding was monitored (CD at 222 nm) to obtain free-energy differences for each transition as described [14] [16] [24] [58] [59] . The transitions are represented by at least three-states [58] , the native (N), intermediate (I) and unfolded states (U):
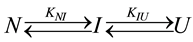 (2)
(2)
The observed ellipticity (Aobs(c)) at any concentration of the denaturant is given by the sum of the contributions from the three states as [59] [60] :
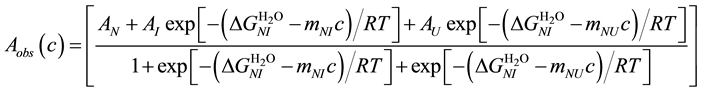 (3)
(3)
where fN (c), fI (c) and fU (c) are the fractions of the three states at a GdnHCl concentration of c (fN + fI+ fU = 1), and AN, AI and AU are the ellipticity values of the N, I and U states, respectively. The fN, fI and fU terms are related to the equilibrium constants, KNI and KNU, of the unfolding transitions from N to I and from N to U, respectively. 

5.3. Kinetics of Activation
20 µl of denatured Pro-SbtE (100 μM in 6 M GdnHCl, pH 4.8) was rapidly mixed in 19,980 μl of the folding buffer at 23˚C with 0.5 mM synthetic substrate (N-succ-AAPF-pNA). After 15 min the stirring was stopped and 200 µl aliquots of the reaction mixture were transferred to a 96-well microplate. Subtilisin activity was monitored by release of p-nitroanilide measured at 405 nm in a microplate reader [18] [26] [42] maintained at the desired temperature. The time of activation was calculated from the X-axis intercept of the “none-to-all” transition curve as described earlier [57] [61] . Ea was calculated from the Arrhenius plots obtained by measuring the activation rate at different temperatures. The activation was also monitored using SDS-PAGE. To analyze the effect of a structured IMC on activation, 10% glycerol was added to the folding buffer and, activation was monitored as discussed above.
Acknowledgements
The authors would like to acknowledge the sources of funding that supported this work, including a National Institutes of Health Individual NRSA Fellowship to DMW (1F30DK096752) and a National Science Foundation CAREER Award to US (MCB0746589).
References
- Wolynes, P.G. (2014) Evolution, Energy Landscapes and the Paradoxes of Protein Folding. Biochimie. http://dx.doi.org/10.1016/j.biochi.2014.12.007
- Alm, E., et al. (2002) Simple Physical Models Connect Theory and Experiment in Protein Folding Kinetics. Journal of Molecular Biology, 322, 463-476. http://dx.doi.org/10.1016/S0022-2836(02)00706-4
- Dobson, C.M. (2004) Principles of Protein Folding, Misfolding and Aggregation. Seminars in Cell Developmental Biology, 15, 3-16. http://dx.doi.org/10.1016/j.semcdb.2003.12.008
- Dobson, C.M. (2003) Protein Folding and Misfolding. Nature, 426, 884-890. http://dx.doi.org/10.1038/nature02261
- Onuchic, J.N. and Wolynes, P.G. (2004) Theory of Protein Folding. Current Opinion in Structural Biology, 14, 70-75. http://dx.doi.org/10.1016/j.sbi.2004.01.009
- Religa, T.L., et al. (2005) Solution Structure of a Protein Denatured State and Folding Intermediate. Nature, 437, 1053- 1056. http://dx.doi.org/10.1038/nature04054
- Bartlett, A.I. and Radford, S.E. (2009) An Expanding Arsenal of Experimental Methods Yields an Explosion of Insights into Protein Folding Mechanisms. Nature Structural Molecular Biology, 16, 582-588. http://dx.doi.org/10.1038/nsmb.1592
- Bartlett, A.I. and Radford, S.E. (2010) Desolvation and Development of Specific Hydrophobic Core Packing during Im7 Folding. Journal of Molecular Biology, 396, 1329-1345. http://dx.doi.org/10.1016/j.jmb.2009.12.048
- Plaxco, K.W., et al. (2000) Topology, Stability, Sequence, and Length: Defining the Determinants of Two-State Protein Folding Kinetics. Biochemistry, 39, 11177-11183. http://dx.doi.org/10.1021/bi000200n
- Kuwajima, K. (1989) The Molten Globule State as a Clue for Understanding the Folding and Cooperativity of Globular-Protein Structure. Proteins, 6, 87-103. http://dx.doi.org/10.1002/prot.340060202
- Shastry, M.C. and Udgaonkar, J.B. (1995) The Folding Mechanism of Barstar: Evidence for Multiple Pathways and Multiple Intermediates. Journal of Molecular Biology, 247, 1013-1027. http://dx.doi.org/10.1006/jmbi.1994.0196
- Nishimura, C., Prytulla, S., Dyson, H.J. and Wright, P.E. (2000) Conservation of Folding Pathways in Evolutionarily Distant Globin Sequences. Nature Structural Biology, 7, 679-686. http://dx.doi.org/10.1038/77985
- Matagne, A., Jamin, M., Chung, E.W., Robinson, C.V., Radford, S.E. and Dobson, C.M. (2000) Thermal Unfolding of an Intermediate Is Associated with Non-Arrhenius Kinetics in the Folding of Hen Lysozyme. Journal of Molecular Biology, 297, 193-210. http://dx.doi.org/10.1006/jmbi.2000.3540
- Raschke, T.M. and Marqusee, S. (1997) The Kinetic Folding Intermediate of Ribonuclease H Resembles the Acid Molten Globule and Partially Unfolded Molecules Detected under Native Conditions. Nature Structural Biology, 4, 298-304. http://dx.doi.org/10.1038/nsb0497-298
- Shinde, U. and Inouye, M. (2000) Intramolecular Chaperones: Polypeptide Extensions That Modulate Protein Folding. Seminars in Cell & Developmental Biology, 11, 35-44. http://dx.doi.org/10.1006/scdb.1999.0349
- Eder, J., Rheinnecker, M. and Fersht, A.R. (1993) Folding of Subtilisin BPN': Role of the Pro-Sequence. Journal of Molecular Biology, 233, 293-304. http://dx.doi.org/10.1006/jmbi.1993.1507
- Bryan, P.N. (2002) Prodomains and Protein Folding Catalysis. Chemical Reviews, 102, 4805-4816. http://dx.doi.org/10.1021/cr010190b
- Yabuta, Y., Subbian, E., Oiry, C. and Shinde, U. (2003) Folding Pathway Mediated by an Intramolecular Chaperone. A Functional Peptide Chaperone Designed Using Sequence Databases. Journal of Biological Chemistry, 278, 15246- 15251. http://dx.doi.org/10.1074/jbc.M212003200
- Williamson, D.M., Elferich, J., Ramakrishnan, P., Thomas, G. and Shinde, U. (2013) The Mechanism by Which a Propeptide-Encoded pH Sensor Regulates Spatiotemporal Activation of Furin. The Journal of Biological Chemistry, 288, 19154-19165. http://dx.doi.org/10.1074/jbc.M112.442681
- Dillon, S.L., Williamson, D.M., Elferich, J., Radler, D., Joshi, R., Thomas, G. and Shinde, U. (2012) Propeptides Are Sufficient to Regulate Organelle-Specific pH-Dependent Activation of Furin and Proprotein Convertase 1/3. Journal of Molecular Biology, 423, 47-62. http://dx.doi.org/10.1016/j.jmb.2012.06.023
- Shinde, U. and Thomas, G. (2011) Insights from Bacterial Subtilases into the Mechanisms of Intramolecular Chaperone-Mediated Activation of Furin. Methods in Molecular Biology, 768, 59-106. http://dx.doi.org/10.1007/978-1-61779-204-5_4
- Feliciangeli, S.F., Thomas, L., Scott, G.K., Subbian, E., Hung, C.-H., Molloy, S.S., et al. (2006) Identification of a pH Sensor in the Furin Propeptide That Regulates Enzyme Activation. Journal of Biological Chemistry, 281, 16108-16116. http://dx.doi.org/10.1074/jbc.M600760200
- Ikemura, H., Takagi, H. and Inouye, M. (1987) Requirement of Pro-Sequence for the Production of Active Subtilisin E in Escherichia coli. The Journal of Biological Chemistry, 262, 7859-7864.
- Eder, J., Rheinnecker, M. and Fersht, A.R. (1993) Folding of Subtilisin BPN': Characterization of a Folding Intermediate. Biochemistry, 32, 18-26. http://dx.doi.org/10.1021/bi00052a004
- Li, Y., Hu, Z., Jordan, F. and Inouye, M. (1995) Functional Analysis of the Propeptide of Subtilisin E as an Intramolecular Chaperone for Protein Folding. Refolding and Inhibitory Abilities of Propeptide Mutants. The Journal of Biological Chemistry, 270, 25127-25132. http://dx.doi.org/10.1074/jbc.270.42.25127
- Yabuta, Y., Takagi, H., Inouye, M. and Shinde, U. (2001) Folding Pathway Mediated by an Intramolecular Chaperone: Propeptide Release Modulates Activation Precision of Pro-Subtilisin. The Journal of Biological Chemistry, 276, 44427-44434. http://dx.doi.org/10.1074/jbc.M107573200
- Subbian, E., Yabuta, Y. and Shinde, U.P. (2005) Folding Pathway Mediated by an Intramolecular Chaperone: Intrinsically Unstructured Propeptide Modulates Stochastic Activation of Subtilisin. Journal of Molecular Biology, 347, 367- 383. http://dx.doi.org/10.1016/j.jmb.2005.01.028
- Subbian, E., Yabuta, Y. and Shinde, U. (2004) Positive Selection Dictates the Choice between Kinetic and Thermodynamic Protein Folding and Stability in Subtilases. Biochemistry, 43, 14348-14360. http://dx.doi.org/10.1021/bi048397x
- Siezen, R.J. and Leunissen, J.A. (1997) Subtilases: The Superfamily of Subtilisin-Like Serine Proteases. Protein Sci- ence, 6, 501-523. http://dx.doi.org/10.1002/pro.5560060301
- Shinde, U. and Inouye, M. (1993) Intramolecular Chaperones and Protein Folding. Trends in Biochemical Sciences, 18, 442-446. http://dx.doi.org/10.1016/0968-0004(93)90146-E
- Anderson, E.D., Molloy, S.S., Jean, F., Fei, H., Shimamura, S. and Thomas, G. (2002) The Ordered and Compartment- Specific Autoproteolytic Removal of the Furin Intramolecular Chaperone Is Required for Enzyme Activation. Journal of Biological Chemistry, 277, 12879-12890. http://dx.doi.org/10.1074/jbc.M108740200
- Elferich, J., Williamson, D.M., Krishnamoorthy, B. and Shinde, U. (2013) Propeptides of Eukaryotic Proteases Encode Histidines to Exploit Organelle pH for Regulation. The FASEB Journal, 27, 2939-2945. http://dx.doi.org/10.1096/fj.12-226886
- Seidah, N.G. and Prat, A. (2012) The Biology and Therapeutic Targeting of the Proprotein Convertases. Nature Reviews Drug Discovery, 11, 367-383. http://dx.doi.org/10.1038/nrd3699
- Khatib, A.M., Siegfried, G., Prat, A., Luis, J., Chrétien, M., Metrakos, P. and Seidah, N.G. (2001) Inhibition of Proprotein Convertases Is Associated with Loss of Growth and Tumorigenicity of HT-29 Human Colon Carcinoma Cells: Importance of Insulin-Like Growth Factor-1 (IGF-1) Receptor Processing in IGF-1-Mediated Functions. Journal of Biological Chemistry, 276, 30686-30693. http://dx.doi.org/10.1074/jbc.M101725200
- Seidah, N.G., Khatib, A.M. and Prat, A. (2006) The Proprotein Convertases and Their Implication in Sterol and/or Lipid Metabolism. Biological Chemistry, 387, 871-877. http://dx.doi.org/10.1515/BC.2006.110
- Seidah, N.G., Mayer, G., Zaid, A., Rousselet, E., Nassoury, N., Poirier, S., et al. (2008) The Activation and Physiological Functions of the Proprotein Convertases. The International Journal of Biochemistry & Cell Biology, 40, 1111-1125. http://dx.doi.org/10.1016/j.biocel.2008.01.030
- Seidah, N.G. and Prat, A. (2002) Precursor Convertases in the Secretory Pathway, Cytosol and Extracellular Milieu. Essays in Biochemistry, 38, 79-94.
- Jaswal, S.S., Sohl, J.L., Davis, J.H. and Agard, D.A. (2002) Energetic Landscape of Alpha-Lytic Protease Optimizes Longevity through Kinetic Stability. Nature, 415, 343-346. http://dx.doi.org/10.1038/415343a
- Bryan, P., Alexander, P., Strausberg, S., Schwarz, F., Lan, W., Gilliland, G. and Gallagher, D.T. (1992) Energetics of Folding Subtilisin BPN'. Biochemistry, 31, 4937-4945. http://dx.doi.org/10.1021/bi00136a003
- Shinde, U. and Inouye, M. (1995) Folding Pathway Mediated by an Intramolecular Chaperone: Characterization of the Structural Changes in Pro-Subtilisin E Coincident with Autoprocessing. Journal of Molecular Biology, 252, 25-30. http://dx.doi.org/10.1006/jmbi.1995.0472
- Jain, S.C., Shinde, U., Li, Y.Y., Inouye, M. and Berman, H.M. (1998) The Crystal Structure of an Autoprocessed Ser221Cys-Subtilisin E-Propeptide Complex at 2.0 Å Resolution. Journal of Molecular Biology, 284, 137-144. http://dx.doi.org/10.1006/jmbi.1998.2161
- Yabuta, Y., Subbian, E., Takagi, H., Shinde, U. and Inouye, M. (2002) Folding Pathway Mediated by an Intramolecular Chaperone: Dissecting Conformational Changes Coincident with Autoprocessing and the Role of Ca2+ in Subtilisin Maturation. Journal of Biochemistry, 131, 31-37. http://dx.doi.org/10.1093/oxfordjournals.jbchem.a003074
- Chu, N.M., Chao, Y. and Bi, R.C. (1995) The 2 Å Crystal Structure of Subtilisin E with PMSF Inhibitor. Protein Engineering, Design and Selection, 8, 211-215. http://dx.doi.org/10.1093/protein/8.3.211
- Comellas-Bigler, M., Maskos, K., Huber, R., Oyama, H., Oda, K. and Bode, W. (2004) 1.2 Å Crystal Structure of the Serine Carboxyl Proteinase Pro-Kumamolisin. Structure of an Intact Pro-Subtilase. Structure, 12, 1313-1323. http://dx.doi.org/10.1016/j.str.2004.04.013
- Shinde, U. and Inouye, M. (1995) Folding Mediated by an Intramolecular Chaperone: Autoprocessing Pathway of the Precursor Resolved via a Substrate Assisted Catalysis Mechanism. Journal of Molecular Biology, 247, 390-395. http://dx.doi.org/10.1006/jmbi.1994.0147
- Engelhard, M. and Evans, P.A. (1995) Kinetics of Interaction of Partially Folded Proteins with a Hydrophobic Dye: Evidence That Molten Globule Character Is Maximal in Early Folding Intermediates. Protein Science, 4, 1553-1562. http://dx.doi.org/10.1002/pro.5560040813
- Ali, V., Prakash, K., Kulkarni, S., Ahmad, A., Madhusudan, K.P. and Bhakuni, V. (1999) 8-Anilino-1-Naphthalene Sulfonic Acid (ANS) Induces Folding of Acid Unfolded Cytochrome c to Molten Globule State as a Result of Electrostatic Interactions. Biochemistry, 38, 13635-13642. http://dx.doi.org/10.1021/bi9907835
- Marie-Claire, C., Yabuta, Y., Suefuji, K., Matsuzawa, H. and Shinde, U. (2001) Folding Pathway Mediated by an Intramolecular Chaperone: The Structural and Functional Characterization of the Aqualysin I Propeptide. Journal of Molecular Biology, 305, 151-165. http://dx.doi.org/10.1006/jmbi.2000.4233
- Buevich, A.V., Shinde, U.P., Inouye, M. and Baum, J. (2001) Backbone Dynamics of the Natively Unfolded Pro-Pep- tide of Subtilisin by Heteronuclear NMR Relaxation Studies. Journal of Biomolecular NMR, 20, 233-249. http://dx.doi.org/10.1023/A:1011243116136
- Shinde, U., Li, Y., Chatterjee, S. and Inouye, M. (1993) Folding Pathway Mediated by an Intramolecular Chaperone. Proceedings of the National Academy of Sciences of the United States of America, 90, 6924-6928. http://dx.doi.org/10.1073/pnas.90.15.6924
- Subbian, E., Yabuta, Y. and Shinde, U. (2005) Folding Pathway Mediated by an Intramolecular Chaparone: Intrinsically Unstructured Propeptide Modulates Stochastic Activation of Subtilisin. Journal of Molecular Biology, 347, 367- 383. http://dx.doi.org/10.1016/j.jmb.2005.01.028
- Ruvinov, S., Wang, L., Ruan, B., Almog, O., Gilliland, G.L., Eisenstein, E. and Bryan, P.N. (1997) Engineering the Independent Folding of the Subtilisin BPN' Prodomain: Analysis of Two-State Folding versus Protein Stability. Biochemistry, 36, 10414-10421. http://dx.doi.org/10.1021/bi9703958
- Wang, L., Ruan, B., Ruvinov, S. and Bryan, P.N. (1998) Engineering the Independent Folding of the Subtilisin BPN' Pro-Domain: Correlation of Pro-Domain Stability with the Rate of Subtilisin Folding. Biochemistry, 37, 3165-3171. http://dx.doi.org/10.1021/bi972741r
- Li, Y. and Inouye, M. (1996) The Mechanism of Autoprocessing of the Propeptide of Prosubtilisin E: Intramolecular or Intermolecular Event? Journal of Molecular Biology, 262, 591-594. http://dx.doi.org/10.1006/jmbi.1996.0537
- Zhu, X.L., Ohta, Y., Jordan, F. and Inouye, M. (1989) Pro-Sequence of Subtilisin Can Guide the Refolding of Denatured Subtilisin in an Intermolecular Process. Nature, 339, 483-484. http://dx.doi.org/10.1038/339483a0
- Shinde, U.P., Liu, J.J. and Inouye, M. (1997) Protein Memory through Altered Folding Mediated by Intramolecular Chaperones. Nature, 389, 520-522. http://dx.doi.org/10.1038/39097
- Fu, X., Inouye, M. and Shinde, U. (2000) Folding Pathway Mediated by an Intramolecular Chaperone. The Inhibitory and Chaperone Functions of the Subtilisin Propeptide Are Not Obligatorily Linked. The Journal of Biological Chemistry, 275, 16871-16878. http://dx.doi.org/10.1074/jbc.275.22.16871
- Fujiwara, K., Arai, M., Shimizu, A., Ikeguchi, M., Kuwajima, K. and Sugai, S. (1999) Folding-Unfolding Equilibrium and Kinetics of Equine Beta-Lactoglobulin: Equivalence between the Equilibrium Molten Globule State and a Burst- Phase Folding Intermediate. Biochemistry, 38, 4455-4463. http://dx.doi.org/10.1021/bi982683p
- Takei, J., Chu, R.A. and Bai, Y. (2000) Absence of Stable Intermediates on the Folding Pathway of Barnase. Proceedings of the National Academy of Sciences of the United States of America, 97, 10796-10801. http://dx.doi.org/10.1073/pnas.190265797
- Mizuguchi, M., Arai, M., Ke, Y., Nitta, K. and Kuwajima, K. (1998) Equilibrium and Kinetics of the Folding of Equine Lysozyme Studied by Circular Dichroism Spectroscopy. Journal of Molecular Biology, 283, 265-277. http://dx.doi.org/10.1006/jmbi.1998.2100
- Inouye, M., Fu, X. and Shinde, U. (2001) Substrate-Induced Activation of a Trapped IMC-Mediated Protein Folding Intermediate. Nature Structural Biology, 8, 321-325. http://dx.doi.org/10.1038/86194