Advances in Microbiology
Vol.05 No.05(2015), Article ID:56305,3 pages
10.4236/aim.2015.55030
Isolation and Identification of Yeasts from Luzhou Flavor Daqu
Xinzhi Cao, Jianming You, Hongmei Ming, Jia Liu, Linsheng Ren
Institute of Food and Biotechnology, Sichuan University of Science & Engineering, Zigong, China
Email: caoxinzhi@163.com
Copyright © 2015 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 5 February 2015; accepted 11 May 2015; published 14 May 2015
ABSTRACT
A yeast strain had been isolated by dilution-plate from the Daqu samples in our study. The strain was identified as a strain of Rhodotorula aurantiaca through observation of its morphological features, micromorphological observation and biolog identification system.
Keywords:
Daqu, Yeast, Separation, Biolog Microbial Identification System

1. Introduction
Liquor is a unique traditional crafts, the Daqu is the essence of wine, the use of starch (carbohydrate) raw material with the Daqu, yeast for the saccharification and fermentation agent, by cooking, saccharification, fermentation, distillation, aging and blending of various types of brewed wine. Wine quality is colorless (or yellow) transparent, aromatic scent of pure, mellow and clean entrance, high alcohol content, having ester compound as the main flavor after the storage of aging. Daqu is the driving force of liquor production, having a very important influence on the Liquor yield and quality. The microorganisms of Daqu, after some time on the song blanks culture medium, warehousing storage, and finally formating the based microbial strains likes bacteria, fungi, yeasts and actinomycetes [1] - [3] . Yeast is one of the main functions Daqu microflora, as these yeast are directly selected from fermentation substrate and nature, they can resistant to high concentrations of alcohol and can evolve in fermentation substrate and nature, they are more stable at high yield, high concentrations of alcohol and genetic performance. In the alcoholic fermentation industry, breeding high-yield, high efficiency advantages of strains can significantly improve the yield and increase the economic benefits of alcohol industry. There are wine yeast alcohol groups and ester production capacity fragrant yeast taxa, strong fermentation efforts such as card’s yeast, producing esters such as abnormal Hansenula. Yeast species and different number has great influence on the yield and style features of wine [4] - [6] , therefore, screening and identification the yeast of Daqu, helping to understand the yeast species and its role in Daqu, having an important guiding significance on improving the quality of Daqu.
Biolog Microbial Identification System was a Microbial Identification System nearly 20 years; the system is mainly microbial utilization of carbon sources by computer analysis and comparison, realizing microbial identification test [7] - [9] . The automation and standardization degree of the system is high, the scope of identify is large, the speed is fast, has become a common technique means of classification and identification of bacteria in international [10] .
2. Materials and Methods
2.1. Materials
2.1.1. Sample Source
The general koji from the market, cryopreservation in the refrigerator at 4˚C.
2.1.2. Medium
Malt extract medium (liquid) Formulation:1kg Dried crushed malt 1 kg, 2 - 3 egg, distilled water 3 L.
Malt extract medium (solid) Formulation: Dried crushed malt 1 kg, 2 - 3 egg, distilled water 3 L, 2% agar.
PDA medium formulations: Potato 200 g, sugar (glucose) 20 g, agar 15 - 20 g, distilled water 1000 ml, PH is nature.
2.1.3. The Main Equipment
Clean Benches (s wc j-1 F), Su net Group Suzhou Aetna air Technology Co.); DG-1 multi-function oven, Shanghai medical equipment repair plant; automatic high-pressure steam sterilizer floor type, Guangzhou South China medical equipment Co., Ltd.; Biolog microbial identification instrument, American Biolog company, micro-imaging system (Japan NIKON company).
2.2. Methods
2.2.1. Isolation of Yeast
1) Enrichment culture of yeast
Take certain amount of Daqu samples (Daqu surface, Daqu heart, Daqu corner), after grinding mix, accurately weighed 10g Daqu samples, placed in 90 mL wort medium oscillation 30 min, the supernatant after allowed to stand. Add 1 - 2 drops of lactic acid and penicillin, 25˚C, culture 48 h.
2) Separation of yeast
Take 1 mL enriched culture medium and placed in test tubes equipped with 9 mL of sterile saline to give 10 - 1 dilution of sample. Then a diluted, made
3) Purified yeast
Suspected yeast colonies were picked on malt extract agar medium and crossed isolated and cultured, continue to pick a single colony, cultured to five generations, microscopic examination, if it is a single form colonies and stored in the fridge. If not pure, then crossed separation, preliminary purebred can be obtained.
2.2.2. Biolog Identification
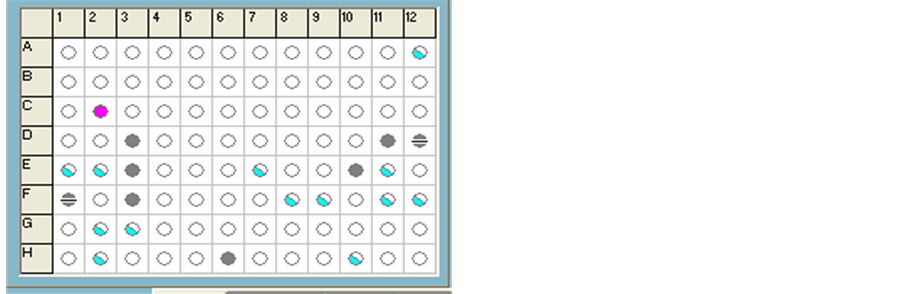
Biolog Microbial Identification System is through the use of 95 kinds of carbon source in a test identification plate. When microbes respire carbon source, will be four azoles redox dye reduction from colorless to purple, resulting in the formation of the microorganism identification plate characteristic response pattern or “fingerprint”, through fiber optics reader to read color change. Computer is through the probability maximum simulation method, the response pattern or “fingerprint” is compared with the database, compare the characteristics of the data related to the target microbes and bacteria database, maximize the match analysis of microorganisms to determine the genus name or species name of the analysis microbiological. Biolog Microbial Identification System consists of turbidity meter reading, reader, software, databases, microporous identification plates.
Turbidity Adjustment: picked good colonies from the BUY culture medium plates with sterile cotton swab, inoculated under aseptic conditions to the turbidity tube. Adjusted to 100% with a blank, raised 47% to 1% with liquid yeast standard pipe. Then inoculated tubes turbidity measurements, turbidity can constantly adjust the size, in order to meet the above criteria haze value
Inoculation board: The turbidity of bacteria have been adjusted is been very carefully inoculated into biolog identification plate by using micro pipette and is placed in an appropriate size tray, and maintain a certain humidity, train 24 h - 72 h.
Read data: open identification system, place the good trained identification plate to the meter reader of colony identification, the computer reads the data, in accordance with the likelihood of 10 given ID name.
Need to consider three parameters on the results identified: the Probability, Similarity, Distance. SIM and DIS are two important parameters, indicate the degree of matching of the test results and the corresponding database data. When DIS < 5.0, SIM > 0.75 is a good match; SIM value closer to 1, the reliability of the test results.
3. The Analysis of the Results
3.1. Isolation and Purification of Yeast
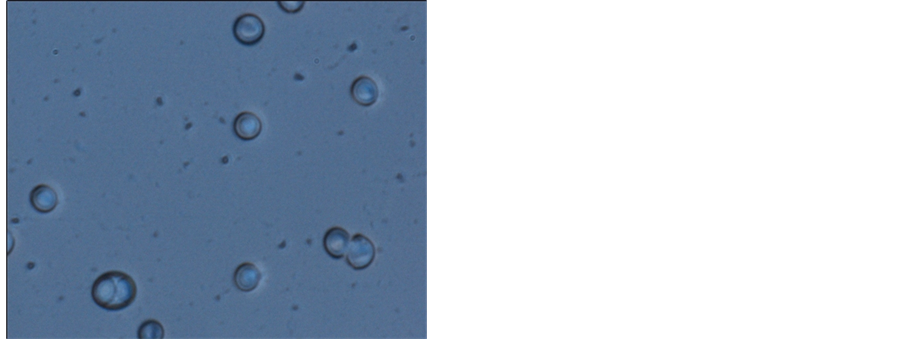
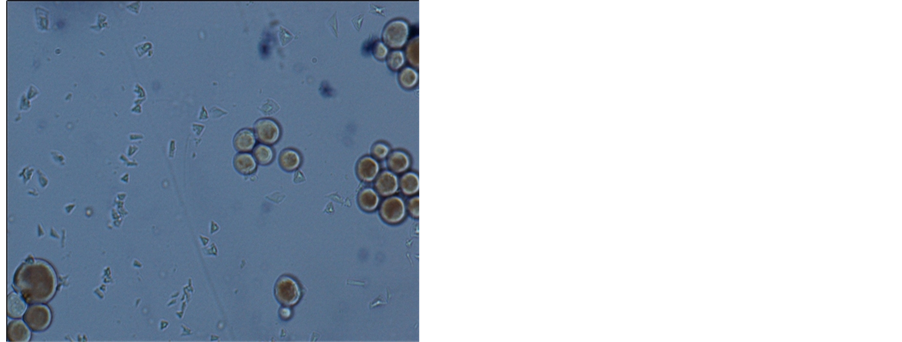
After enrichment culture, purification, filtering obtained a strain of yeast from song samples, the colony surface is smooth, moist, reddish, neat edge, under the microscope cells were round or oval, budding (Figure 1).
3.2. Identification Results of Biolog Microbial Identification System
The purified strain was inoculated yeast identification and plate on the train reading of Biolog Microbial Identification System after the 48 h to reader data, the results shown below:
As can be seen from the above Figure 2, only α-D-glucose showed positive, indicating that the strain can
Figure 1. Strain B microscopic observation (100×), left for flooding piece, right for glycogen staining.
Figure 2. Carbon utilization of strain B Biolog identification plate.
utilize α-D-glucose. After culturing the strain B 48h, three important parameters were obtained in the system, SIM value = 0.748; DIT = 3.35 < 5.0; PROB value = 0. 96, relatively close to 1, so the database has a good match, in ID address bar shows a best match name, Rhodotorula aurantiaca B (refer to orange red yeast B).
4. Conclusion
By doing the separation of Daqu in yeast, purified culture, and observing colony morphology, cell morphology, using Biolog microbial identification system, identified as the orange-yellow-red yeast strain B (Rhodotorula aurantiaca B). Biolog Microbial Identification System has become the international classification and identification of yeasts multiphase common techniques. Lee Yun [10] , who reported the use of Biolog system successfully, identified a strain of Saccharomyces cerevisiae producing a good performance. This article will introduce Biolog identification system into the identification of yeasts Daqu, favor further study of microbial distribution in the Daqu.
Acknowledgements
The authors thank the Sichuan University of Science & Engineering for the basic laboratory and laboratory equipment.
References
- Ao, Z.H., Shan, X.H., Shen, C.H., Zhang, L., Wang, X.J., Shen, C.P., et al. (2011) Luzhou Daqu Industry Development in Our Country. Sichuan University of Science & Engineering, Zigong.
- Yao, W.C., Tang, Y.M. and Ren, D.Q. (2005) Study on the Differences of Microbes in the Different Layers of Guojiao Daqu. Sichuan University of Science & Engineering, Zigong.
- Sun, J.F. (2005) Application and Cultivation of Effective Microzyme in Daqu. Shandong Taishan Miguel Sources Group Co., Ltd., Shandong.
- Zhang, L., Shi, S. and Zhang, X.W. (2010) Isolation and Identification of Yeast in Liquor Daqu. Sichuan University, Chengdu.
- Wu, S.W., Zhang, Z.G. and Li, X.H. (2011) Okuma Microbial Research and Development Status and Development Prospects in Liquor Production. Four Wine Co., Ltd., Jiangxi.
- Yao, S., Cheng, C. and Li, J.X. (2006) 1Biolog Microbial Identification System―Study of Production and Research Experience of Filamentous Fungi Identification Procedures. China Research Institute of Food and Fermentation Industry, Beijing.
- Zhang, C.H. (1994) The Application of 1Biolog Bacterial Automatic Identification System. The Ministry of Agriculture Plant Quarantinr Laboratory, Beijing.
- Liao, Y.H., Ren, W.Y., Wu, S.L. and Shen, H. (2010) Koji Mucor Two Separate and Biolog Microbial Identification System Analysis. Beijing Industry and Commerce University, Beijing.
- Li, J.X., Cheng, C., Yao, S. and Hu, H.R. (2006) Biolog Microbial Identification System―Study of Yeast Identification Procedures. Chinese Food Fermentation Industry Research Institute, Beijing.
- Li, Y., Sheng, H. and Zhao, R.H. (2005) The Application of Biolog Microbial Identification System in Species Identification, Beijing.