Open Journal of Blood Diseases
Vol.05 No.04(2015), Article ID:62422,11 pages
10.4236/ojbd.2015.54008
Availability of Circulating MicroRNAs as a Biomarker for Early Diagnosis of Diffuse Large B-Cell Lymphoma
Katsushige Inada1*, Yasushi Okoshi1,2,3, Yukiko Cho4, Hitoaki Saito5, Tatsuo Iijima5, Mitsuo Hori1, Hiroshi Kojima2,3,4
1Department of Hematology, Ibaraki Prefectural Central Hospital, Ibaraki, Japan
2Ibaraki Clinical Education and Training Center, University of Tsukuba Hospital, Ibaraki, Japan
3Faculty of Medicine, University of Tsukuba, Ibaraki, Japan
4Department of Medical Oncology, Ibaraki Prefectural Central Hospital, Ibaraki, Japan
5Department of Pathology, Ibaraki Prefectural Central Hospital, Ibaraki, Japan

Copyright © 2015 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 4 December 2015; accepted 27 December 2015; published 30 December 2015
ABSTRACT
Background: MicroRNA (miRNA) regulates post-transcriptional gene expression through binding to complementary sites of target messenger RNA, including that from oncogenes or tumor suppressor genes. This study planned to pursue the possibility that circulating miRNA could be used for the early diagnosis of diffuse large B-cell lymphoma (DLBCL). Materials and Methods: Expression levels of miRNA obtained from serum, exosome-enriched serum, and formalin-fixed paraffin- embedded (FFPE) tissue were evaluated. Samples were collected from patients with newly diagnosed DLBCL (n = 33) or healthy volunteers (n = 22). Based on the results of previous reports, ten miRNAs were selected and expression levels were analyzed by the quantitative real-time PCR. Results: The expression levels of hsa-miR-15a-3p, hsa-miR-21-5p, hsa-miR-181a-5p, and hsa-miR- 210-5p differed significantly between DLBCL patients and controls in serum and/or exosome- enriched serum, but not in FFPE tissue. In contrast, expression levels of hsa-miR-155-5p in FFPE tissue were significantly higher in DLBCL patients, as previously reported. Conclusion: We confirmed that some miRNAs were differentially expressed in serum from DLBCL patients as previously reported. Measurement of these miRNA in exosome-enriched serum did not improve the accuracy in the differential diagnosis of DLBCL. In addition, these miRNAs seem to be produced outside of lymphoma tissue.
Keywords:
DLBCL, miRNA, Serum, Exosome, Formalin-Fixed Paraffin-Embedded Tissue

1. Introduction
MicroRNA (miRNA) is a class of non-coding small RNA (~22 nucleotides) that regulates post-transcriptional gene expression by binding to the complementary sites of target messenger RNA [1] . MiRNAs have important functions in oncogenesis or as tumor suppressors due to their prominent role in regulating cancer pathways. Because of their inherent stability in the blood [2] , circulating miRNAs could be a suitable and convenient biomarker for the early diagnosis of cancer [3] [4] . Recent reports suggested that circulating miRNA is contained within exosomes [2] [5] or complexed with proteins, such as Argonaute2 [6] , which contributes to their stability in the blood.
Diffuse large B-cell lymphoma (DLBCL) is an aggressive tumor that is the most common type of non-Hodg- kin lymphoma. Although the treatment of DLBCL has become increasingly successful, mostly due to the anti- CD20 antibody rituximab, early disease detection and biopsy-proven diagnosis are keys to improving treatment outcome and reducing mortality. DLBCL often occurs in lymph nodes or in Waldeyer’s ring, however, extranodal disease occurs in up to 30% - 40% of cases. The most common site of extranodal involvement is the stomach/gastrointestinal tract [7] [8] , but the disease can arise in virtually any tissue, which sometimes makes biopsy difficult or more invasive than that of superficial lesions. Therefore, sensitive and specific biomarkers for DLBCL will be useful in deciding whether to use an invasive procedure for diagnosis.
To date, elevated levels of some miRNAs, such as miR-15a, miR-21, miR-155, and miR-210, have been found in the serum of patients with DLBCL [9] [10] . Indeed, these miRNAs are potentially useful as biomarkers; however, the sensitivity and specificity should be further increased to improve the ability to distinguish DLBCL from other conditions. To address this issue, we focused on circulating exosome-derived miRNAs and evaluated their expression levels not only in plain serum but also in exosome-enriched serum from patients with newly diagnosed DLBCL. In addition, the expression levels of miRNAs from lymph nodes were also measured to evaluate whether miRNAs showing elevated expression in serum originated in the lymphoma tissue.
2. Materials and Methods
2.1. Study Design
This study was approved by the institutional review board of Ibaraki Prefectural Central Hospital, Japan. We evaluated miRNA expression levels in serum, exosome-enriched serum, and formalin-fixed paraffin-embedded (FFPE) tissue. Based on the results of previous reports, 10 miRNAs (the 5p and 3p strands of hsa-miR-15a, hsa- miR-21, hsa-miR-155, hsa-miR-181a, and hsa-miR-210) were selected as candidate biomarkers (Table 1). The serum samples were collected from patients with DLBCL or from healthy volunteers with consent. All the patients with DLBCL were newly diagnosed at the hospital between March 2012 and August 2013. The FFPE biopsy samples were collected from DLBCL lesions, mainly lymph nodes. FFPE tissues from reactive hyperplastic lymph nodes were used as controls. The small RNA molecules, such as miR-16, miR-39, miR-24, RNU6B, and RNU48, have previously been used to normalize expression of target miRNAs [9] -[13] . In this study, miR-24 was chosen as endogenous control, because it showed proper and relatively stable expression in our preliminary study (data not shown).
2.2. Quantification of Circulating miRNA in Serum
Total RNA was extracted from serum samples using the mirVana PARIS Isolation Kit (Ambion, Austin, TX) in accordance with the manufacturer’s instructions. RNA concentrations were measured using the NanoDrop ND-2000 spectrophotometer (NanoDrop Technologies, Wilmington, DE). To measure miRNA expression levels, TaqMan MicroRNA Assays (Applied Biosystems, Foster City, CA) were used according to the manufacturer’s instructions. Briefly, 10 ng of total RNA was reverse transcribed using the TaqMan MicroRNA Reverse Transcription Kit (Applied Biosystems) in a 15-µL reaction volume for 30 min at 16˚C, 30 min at 42˚C, and 5 min at
Table 1. Selected miRNAs in this study and reported expression levels in previous reports.
*Expression levels of miR-181a influence clinical outcome of R-CHOP chemotherapy in DLBCL. R-CHOP = rituximab and the combination of cyclophosphamide, doxorubicin, vincristine, and prednisone.
85˚C. With the reverse transcription product, the expression level of each miRNA was quantified by real-time PCR in a duplicate manner with the 7500 Fast Dx Real-Time PCR System and the software program SDS v1.4 (Applied Biosystems). The PCR protocol was as follows: incubation for 2 min at 50˚C and 10 min at 95˚C, followed by 40 cycles of 15 s at 95˚C and 1 min at 60˚C. The mean cycle threshold (Ct) value of each miRNA was normalized to the mean Ct of hsa-miR-24-3p, yielding a ΔCT value (ΔCT = CT(miR-target) − CT(miR-24)). Relative expression levels were reported as 2−ΔCT.
2.3. Quantification of miRNA in Exosome-Enriched Serum
To concentrate serum exosomes, the serum sample was centrifuged at 2000 g for 30 min, and combined with 0.2 volume of total exosome isolation (from serum) reagent (Invitrogen, Carlsbad, CA). The sample was incubated for 30 min at 4˚C, and centrifuged for 10 min at 10,000 g. Then, the Total Exosome RNA and Protein Isolation Kit (Invitrogen) was used for extraction of total RNA in accordance with the manufacturer’s instructions, and miRNA expression was measured as described above.
2.4. Quantification of miRNA in FFPE Tissue
In each specimen, total RNA was extracted from 8 FFPE sections of 10-µm thickness. These 8 sections were placed in a tube and deparaffinized using 1 mL of xylene (Wako), and washed with 1 mL of 99.5% ethanol (Wako). After drying, total RNA was extracted with a Recoverall Total Nucleic Acid Isolation Kit (Ambion) in accordance with the manufacturer’s instructions, and miRNA expression was measured as previously described. MiR-24 was used again as an endogenous control based on a previous report [14] . MiR-155, whose upregulation was previously demonstrated in FFPE tissue from DLBCL patients [13] [15] , was used as a positive control.
2.5. Statistical Analysis
Statistical analysis was performed with GraphPad Prism 6.03 (GraphPad Software, San Diego, CA). The Mann- Whitney U test was used to determine statistically significant differences between healthy volunteers and patients with DLBCL. P values < 0.05 were considered statistically significant. Receiver operating characteristic (ROC) curve analysis was performed to determine the areas under the curve (AUCs), sensitivity, and specificity. The point closet to the upper left-hand corner of the ROC curve was chosen as the optimum sensitivity and specificity values.
3. Results
3.1. Patient Characteristics
Serum samples were collected from 33 patients with DLBCL and from 22 healthy volunteers. FFPE samples were from 22 patients with DLBCL and 6 reactive hyperplastic lymph nodes (Table 2).
3.2. Expression of Circulating miRNAs in Serum
The expression of 10 miRNAs in the serum of DLBCL patients (n = 33) was compared with that of healthy vo-
Table 2. Clinical characteristics of patients.
*Reactive hyperplastic lymph nodes. GCB = germinal center B-cell.
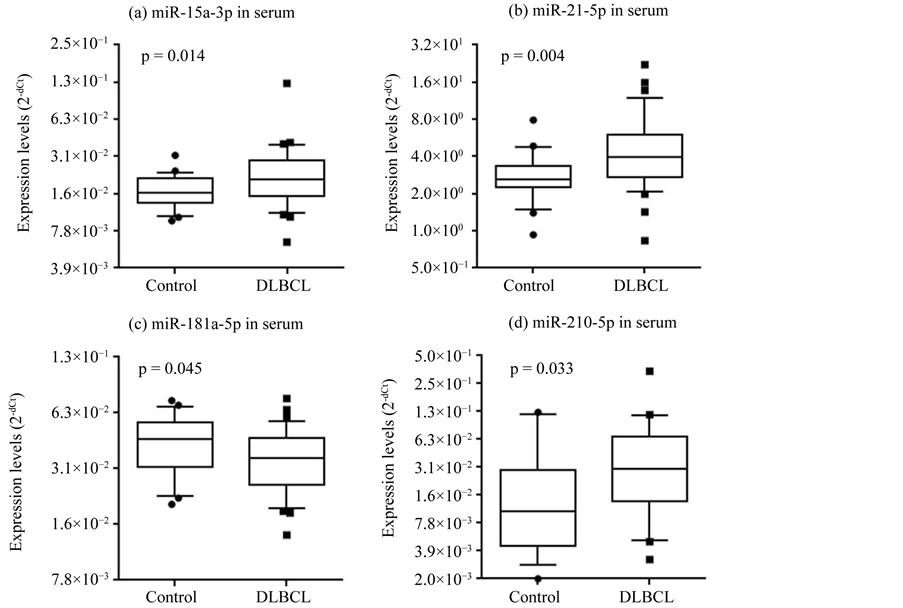
lunteers (n = 22). As described, we chose these miRNAs based on previous reports on DLBCL (Table 1). Among these miRNAs, the expression of 3 miRNAs was significantly increased in DLBCL patients relative to healthy volunteers (hsa-miR-15a-3p, p = 0.014; hsa-miR-21-5p, p = 0.004; hsa-miR-210-5p, p = 0.033. Figure 1(a), Figure 1(b), Figure 1(d)). The expression of hsa-miR-181a-5p was significantly decreased in DLBCL patients relative to healthy volunteers (p = 0.045; Figure 1(c)). The expression of hsa-miR-155-5p and hsa-miR- 210-3p did not differ significantly between the 2 groups (Table 3). In most serum samples, hsa-miR-15a-5p, hsa-miR-21-3p, hsa-miR-155-3p, and hsa-miR-181a-3p were not detected (Table 3).
We next evaluated the value of miRNA as a diagnostic biomarker using ROC curve analysis (Figure 2). The
Figure 1. Expression levels of miRNAs in sera. The expression levels of serum hsa-miR-15a- 3p (a), hsa-miR-21-5p (b), hsa-miR-181a-5p (c), and hsa-miR-210-5p (d) differed significantly between DLBCL patients and healthy volunteers (p < 0.05). Expression levels (log2 scale on the y-axis) were normalized to hsa-miR-24-3p and are shown in box plots. The median is represented as a horizontal line within each box and its lower and upper edges represent the 25 - 75 percentile region. The whiskers represent the 10 - 90 percentile region. Statistically significant differences were determined using the Mann-Whitney U test (one-tailed test).
Table 3. Expression levels and AUC values of the 10 miRNAs in DLBCL patients compared with controls.
The Mann-Whitney U test and ROC curve analysis were used for statistical analysis. n.s. = no significant difference; - = not assessed.
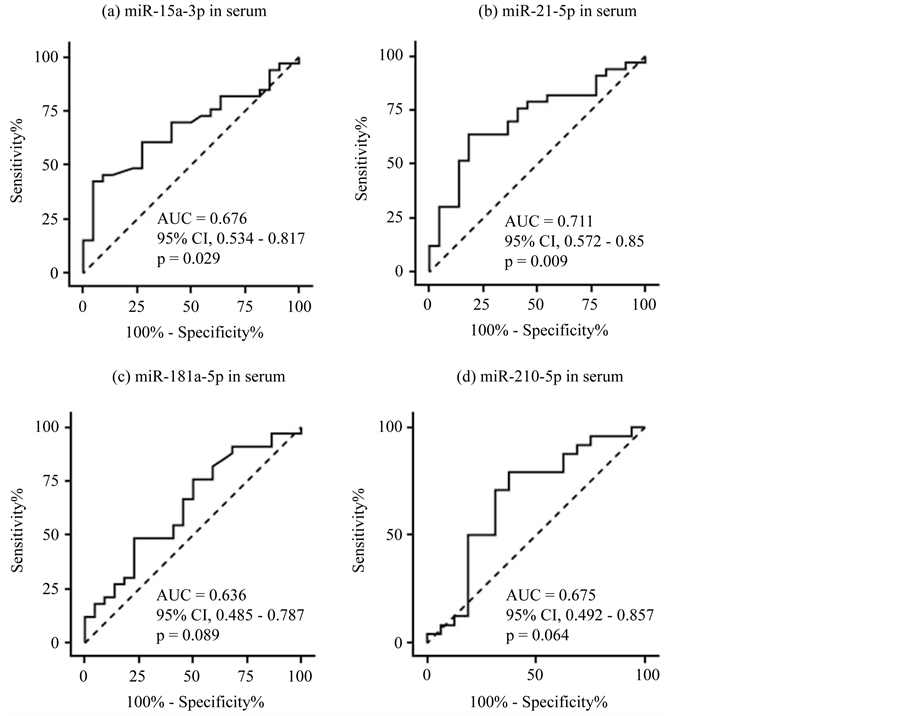
Figure 2. ROC curve analysis for discriminating patients with DLBCL from healthy volunteers based on miRNA expression in serum. 95% CI = 95% confidence interval. (a) hsa-miR-15a-3p; (b) hsa-miR-21-5p; (c) hsa-miR-181a-5p; (d) hsa-miR-210-5p.
AUCs were 0.676 for hsa-miR-15a-3p (57.1% in sensitivity; 72.7% in specificity), 0.711 for hsa-miR-21-5p (60.7% sensitivity; 81.8% specificity), 0.636 for hsa-miR-181a-5p (50% sensitivity; 77.3% specificity) and 0.675 for hsa-miR-210-5p (75% sensitivity; 62.5% specificity). These data suggested that none of the miRNA we evaluated was sensitive or specific enough for use as a diagnostic marker, although expression levels differed significantly between patients and controls.
3.3. Evaluation of miRNA Expression after Exosome Enrichment
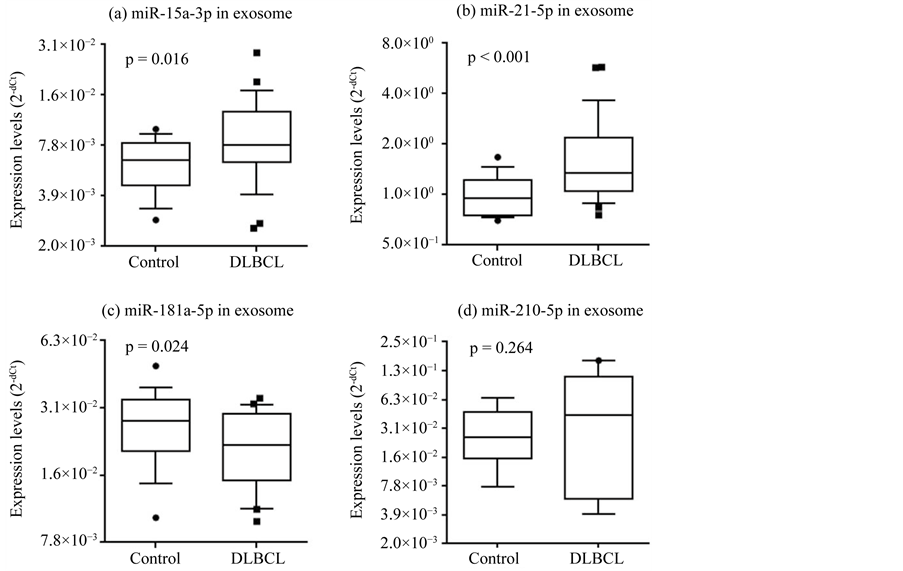
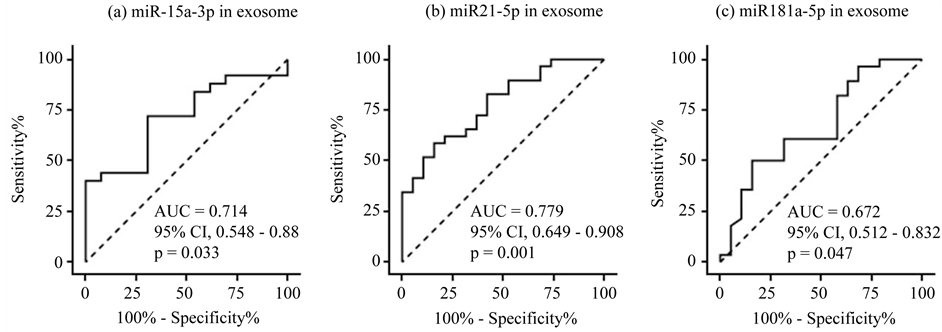
To improve the diagnostic acuity, we enriched exosomes in the sample sera using a commercially available kit because recent studies suggested that some circulating miRNAs are contained within exosomes [2] [5] . The same serum samples used to evaluate miRNA expression were used, but some were not available due to a shortage of storage. Patient characteristics of the final cohort are summarized in Table 2. Expression of the 10 candidate miRNAs was significantly higher in DLBCL patients for hsa-miR-15a-3p and hsa-miR-21-5p (p = 0.016 and p < 0.001, respectively), and significantly lower for hsa-miR-181a-5p (p = 0.024; Figure 3). The AUCs were 0.714 for hsa-miR-15a-3p (72% sensitivity; 69.2% specificity), 0.779 for hsa-miR-21-5p (62.1% sensitivity; 78.9% specificity), and 0.672 for miR-181a-5p (60.7% sensitivity; 68.4% specificity; Figure 4). In conclusion, we could not demonstrate sufficient improvement in diagnostic acuity after exosome enrichment.
3.4. Expression Levels of miRNAs in FFPE Tissue
Finally, we examined whether the miRNAs that showed different expression levels in the plain or exosome-
Figure 3. Expression of hsa-miR-15a-3p (a), hsa-miR-21-5p (b), and hsa-miR-181a-5p (c) differed significantly between DLBCL patients and healthy volunteers (p < 0.05). Expression levels (log2 scale on the y-axis) were normalized to hsa-miR-24-3p. Statistically significant differences were determined using the Mann-Whitney U test (one-tailed test).
Figure 4. ROC curve analysis of miRNA expression in exosome-enriched sera. (a) hsa-miR-15a-3p; (b) hsa-miR-21-5p; (c) hsa-miR-181a-5p.
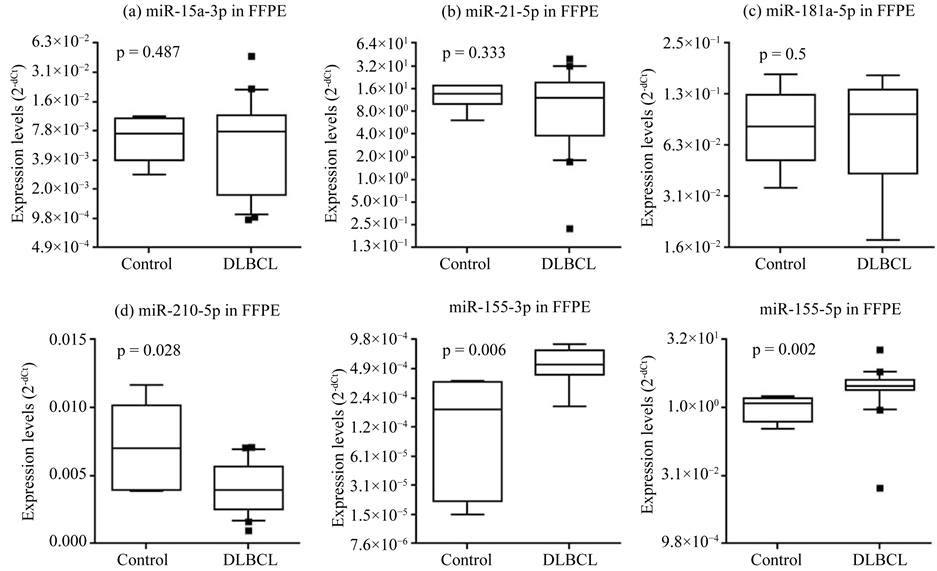
enriched sera originated in lymphoma tissues. MiRNAs were extracted from FFPE biopsy samples of DLBCL patients. These patients were in the same cohort in which their sera and exosome-enriched sera were analyzed, but only biopsy samples containing sufficient miRNA were assessed this time. The patient characteristics of the final cohort are summarized in Table 2. The expression levels of the miRNAs listed in Table 1 were measured while miR-155 was used as the positive control this time. We could not find significantly different from reactive hyperplastic lymph nodes, with the exception of hsa-miR-210-5p, which was significantly decreased in DLBCL patients (p = 0.028; Figure 5).
The miRNA expression results from serum, exosome-enriched serum, and FFPE tissue are summarized in Table 3.
Figure 5. Expression levels of miRNA in FFPE tissue. The expression levels of miRNAs in FFPE tissue were compared between DLBCL and reactive hyperplastic lymph nodes. Only the expression of hsa-miR-210-5p (d) was significantly different (p < 0.05). Hsa-miR-155-3p and hsa-miR-155-5p were used as positive controls. The expression levels of miRNAs (log2 scale on the y-axis) were normalized to hsa-miR-24-3p. Statistically significant differences were determined using the Mann- Whitney U test (one-tailed test).
4. Discussion
It is reported that extracellular miRNA is transported by apoptotic bodies, protein complexes, exosomes, and microvesicles [16] , and exosomal miRNA is reported to play a role in intercellular communication [17] . In this study, we expected that diagnostic accuracy would be improved by exosome enrichment but we could not find it. Of course, there is much room for improving and it is too early to draw firm conclusions. We could try only a single method for the enrichment because of the amount of serum storage. In a recent study, Yoshioka et al. reported that the ExoScreen method, which enables the detection of circulating extracellular vesicles, including exosomes and microvesicles from cancer cells, without the need for a purification step [18] . Diagnostic prediction with circulating miRNAs may be improved by such methods.
We also quantified miRNA expression in DLBCL tissues to investigate the relationship between intracellular and extracellular miRNA levels. We used not frozen but FFPE tissue, as it is easily available and previous studies found a good correlation between miRNA expression in FFPE and frozen cells/tissue samples [14] [19] [20] .
The expression levels of 3 miRNAs (hsa-miR-15a-3p, hsa-miR-21-5p, and hsa-miR-181a-5p) differed significantly between DLBCL patients and healthy volunteers in both plain and exosome-enriched serum (Figure 1, Figure 3, Table 3), suggesting that these miRNAs may be useful as biomarkers. To improve diagnostic sensitivity and specificity, some miRNA pairs such as hsa-miR-15a-3p and hsa-miR-21-5p were evaluated using the cut-off values determined by ROC curves, but no combinations improved the prediction of DLBCL (data not shown). Additionally, the expression of these 3 miRNAs in lymph nodes was not significantly different between DLBCL patients and controls (Figure 5, Table 3), suggesting that the hsa-miR-15a-3p, hsa-miR-21-5p, and hsa- miR-181a-5p in serum are produced outside of lymphoma tissue. If true, we guess that miRNAs which are de-
rived directly from lymphoma cells could be more accurate and sensitive diagnostic biomarkers. Anyway, consistent with our results, Munch-Petersen et al. reported that miR-21 was expressed in stromal cells but not DLBCL tumor cells [21] . A recent study reported that some miRNAs, which is derived from non-cancer cells, had the ability to suppress the growth of cancer cell proliferation, not only in vitro but also in vivo [22] . Thus, these miRNAs in serum might influence lymphoma cell growth.
In this study, hsa-miR-210-5p was overexpressed in plain serum but not in exosome-enriched serum samples from DLBCL patients (Figure 1, Figure 3, Table 3). In addition, the expression levels of hsa-miR-210-5p in FFPE tissues were significantly downregulated in DLBCL patients (Figure 5, Table 3). Taken together, these findings suggest that hsa-miR-210-5p in serum might originate outside of lymphoma tissue and it is transported by structures other than exosomes, such as microvesicles or protein complexes. MiR-210 is involved in regulating the cell cycle, cell survival, differentiation, and DNA repair, as well as the immune response [23] , and has been reported to function as both an oncogene and a tumor suppressor [24] .
As mentioned above, miRNAs evaluated in this study are involved in not only DLBCL but also general cancer biology. MiR-15a, miR-21, miR-181a, and miR-210 were shown to target the anti-apoptotic BCL-2 [23] [25] - [28] , which plays an important role in carcinogenesis, including lymphomagenesis [29] . In addition, miR-21 was reported to promote cell invasion, migration, and growth by suppressing the expression of phosphatase and Tensin Homolog Deleted from Chromosome 10 (PTEN) [30] . MiR-21 was also found to be overexpressed in the serum of patients with breast cancer, ovarian cancer, and non-small cell lung cancer [31] . Alencar et al. reported that high expression of miR-181a in the DLBCL tissue correlates with poor progression-free survival and miR-181a directly downregulates the expression of FOXP1 [12] . However, the reason for the downregulation of hsa-miR-181a-5p in DLBCL serum in our study (Figure 1) remains unclear. Further studies are necessary to determine whether the expression level of miRNAs is useful diagnostic or prognostic biomarker for DLBCL.
5. Conclusion
We confirmed that some miRNAs are differentially expressed in serum from DLBCL patients as previously reported. The miRNAs we evaluated, however, were not sensitive or specific enough for use as a diagnostic marker, and exosome-enriched serum did not improve the accuracy. Finally, these miRNAs seem to be produced outside of lymphoma tissue.
Acknowledgements
We thank the technical staff in the clinical laboratory for collecting samples. This work was partially supported by research grants from Ibaraki Prefecture and Ibaraki Prefectural Central Hospital.
Cite this paper
KatsushigeInada,YasushiOkoshi,11,11,YukikoCho,HitoakiSaito,TatsuoIijima,MitsuoHori,HiroshiKojima,11,11, (2015) Availability of Circulating MicroRNAs as a Biomarker for Early Diagnosis of Diffuse Large B-Cell Lymphoma. Open Journal of Blood Diseases,05,48-58. doi: 10.4236/ojbd.2015.54008
References
- 1. Gregory, R.I. and Shiekhattar, R. (2005) MicroRNA Biogenesis and Cancer. Cancer Research, 65, 3509-3512. http://dx.doi.org/10.1158/0008-5472.CAN-05-0298
- 2. Mitchell, P.S., Parkin, R.K., Kroh, E.M., Fritz, B.R., Wyman, S.K., Pogosova-Agadjanyan, E.L., Peterson, A., Noteboom, J., O’Briant, K.C., Allen, A., Lin, D.W., Urban, N., Drescher, C.W., Knudsen, B.S., Stirewalt, D.L., Gentleman, R., Vessella, R.L., Nelson, P.S., Martin, D.B. and Tewari M. (2008) Circulating MicroRNAs as Stable Blood-Based Markers for Cancer Detection. Proceedings of the National Academy of Sciences of the United States of America, 105, 10513-10518.
http://dx.doi.org/10.1073/pnas.0804549105 - 3. Kosaka, N., Iguchi, H. and Ochiya, T. (2010) Circulating MicroRNA in Body Fluid: A New Potential Biomarker for Cancer Diagnosis and Prognosis. Cancer Science, 101, 2087-2092.
http://dx.doi.org/10.1111/j.1349-7006.2010.01650.x - 4. Sundarbose, K., Kartha, R. and Subramanian, S. (2013) MicroRNAs as Biomarkers in Cancer. Diagnostics, 3, 84-104. http://dx.doi.org/10.3390/diagnostics3010084
- 5. Valadi, H., Ekstrom, K., Bossios, A., Sjostrand, M., Lee, J.J. and Lotvall, J.O. (2007) Exosome-Mediated Transfer of mRNAs and MicroRNAs is a Novel Mechanism of Genetic Exchange between Cells. Nature Cell Biology, 9, 654-659. http://dx.doi.org/10.1038/ncb1596
- 6. Arroyo, J.D., Chevillet J.R., Kroh E.M., Ruf I.K., Pritchard C.C., Gibson D.F., Mitchell P.S., Bennett C.F., Pogosova-Agadjanyan E.L., Stirewalt D.L., Tait J.F. and Tewari M. (2011) Argonaute2 complexes carry a population of circulating MicroRNAs Independent of Vesicles in Human Plasma. Proceedings of the National Academy of Sciences of the United States of America, 108, 5003-5008.
http://dx.doi.org/10.1073/pnas.1019055108 - 7. Moller, M.B., Pedersen, N.T. and Christensen, B.E. (2004) Diffuse Large B-Cell Lymphoma: Clinical Implications of Extranodal versus Nodal Presentation—A Population-Based Study of 1575 Cases. British Journal of Haematology, 124, 151-159. http://dx.doi.org/10.1046/j.1365-2141.2003.04749.x
- 8. Lopez-Guillermo, A., Colomo, L., Jimenez, M., Bosch, F., Villamor, N., Arenillas, L., Muntanola, A., Montoto, S., Gine, E., Colomer, D., Bea, S., Campo, E. and Montserrat, E. (2005) Diffuse Large B-Cell Lymphoma: Clinical and Biological Characterization and Outcome According to the Nodal or Extranodal Primary Origin. Journal of Clinical Oncology, 23, 2797-2804. http://dx.doi.org/10.1200/JCO.2005.07.155
- 9. Lawrie, C.H., Gal, S., Dunlop, H.M., Pushkaran, B., Liggins, A.P., Pulford, K., Banham, A.H., Pezzella, F., Boultwood, J., Wainscoat, J.S., Hatton, C.S. and Harris, A.L. (2008) Detection of Elevated Levels of Tumour-Associated MicroRNAs in Serum of Patients with Diffuse Large B-Cell Lymphoma. British Journal of Haematology, 141, 672-675. http://dx.doi.org/10.1111/j.1365-2141.2008.07077.x
- 10. Fang, C., Zhu, D.X., Dong, H.J., Zhou, Z.J., Wang, Y.H., Liu, L., Fan, L., Miao, K.R., Liu, P., Xu, W. and Li, J.Y. (2012) Serum MicroRNAs Are Promising Novel Biomarkers for Diffuse Large B Cell Lymphoma. Annals of Hematology, 91, 553-559. http://dx.doi.org/10.1007/s00277-011-1350-9
- 11. Baraniskin, A., Kuhnhenn, J., Schlegel, U., Chan, A., Deckert, M., Gold, R., Maghnouj, A., Zollner, H., Reinacher-Schick, A., Schmiegel, W., Hahn, S.A. and Schroers, R. (2011) Identification of MicroRNAs in the Cerebrospinal Fluid as Marker for Primary Diffuse Large B-Cell Lymphoma of the Central Nervous System. Blood, 117, 3140-3146. http://dx.doi.org/10.1182/blood-2010-09-308684
- 12. Alencar, A.J., Malumbres, R., Kozloski, G.A., Advani, R., Talreja, N., Chinichian, S., Briones, J., Natkunam, Y., Sehn, L.H., Gascoyne, R.D., Tibshirani, R. and Lossos, I.S. (2011) MicroRNAs Are Independent Predictors of Outcome in Diffuse Large B-Cell Lymphoma Patients Treated with R-Chop. Clinical Cancer Research, 17, 4125-4135. http://dx.doi.org/10.1158/1078-0432.CCR-11-0224
- 13. Zhong, H., Xu, L., Zhong, J.H., Xiao, F., Liu, Q., Huang, H.H. and Chen, F.Y. (2012) Clinical and Prognostic Significance of miR-155 and miR-146a Expression Levels in Formalin-Fixed/Paraffin-Embedded Tissue of Patients with Diffuse Large B-Cell Lymphoma. Experimental and Therapeutic Medicine, 3, 763-770.
- 14. Culpin, R.E., Sieniawski, M., Proctor, S.J., Menon, G. and Mainou-Fowler, T. (2013) MicroRNAs Are Suitable for Assessment as Biomarkers from Formalin-Fixed Paraffin-Embedded Tissue, and miR-24 Represents an Appropriate Reference MicroRNA for Diffuse Large B-Cell Lymphoma Studies. Journal of Clinical Pathology, 66, 249-252. http://dx.doi.org/10.1136/jclinpath-2012-201021
- 15. Lawrie, C.H., Soneji, S., Marafioti, T., Cooper, C.D., Palazzo, S., Paterson, J.C., Cattan, H., Enver, T., Mager, R., Boultwood, J., Wainscoat, J.S. and Hatton, C.S. (2007) MicroRNA Expression Distinguishes between Germinal Center B Cell-Like and Activated B Cell-Like Subtypes of Diffuse Large B Cell Lymphoma. International Journal of Cancer, 121, 1156-1161. http://dx.doi.org/10.1002/ijc.22800
- 16. Xu, L., Yang, B.F. and Ai, J. (2013) MicroRNA Transport: A New Way in Cell Communication. Journal of Cellular Physiology, 228, 1713-1719. http://dx.doi.org/10.1002/jcp.24344
- 17. Kosaka, N., Iguchi, H., Yoshioka, Y., Takeshita, F., Matsuki, Y. and Ochiya, T. (2010) Secretory Mechanisms and Intercellular Transfer of MicroRNAs in Living Cells. The Journal of Biological Chemistry, 285, 17442-17452. http://dx.doi.org/10.1074/jbc.M110.107821
- 18. Yoshioka, Y., Kosaka, N., Konishi, Y., Ohta, H., Okamoto, H., Sonoda, H., Nonaka, R., Yamamoto, H., Ishii, H., Mori, M., Furuta, K., Nakajima, T., Hayashi, H., Sugisaki, H., Higashimoto, H., Kato, T., Takeshita, F. and Ochiya, T. (2014) Ultra-Sensitive Liquid Biopsy of Circulating Extracellular Vesicles Using ExoScreen. Nature Communications, 5, Article No. 3591. http://dx.doi.org/10.1038/ncomms4591
- 19. Xi, Y., Nakajima, G., Gavin, E., Morris, C.G., Kudo, K., Hayashi, K. and Ju, J. (2007) Systematic Analysis of MicroRNA Expression of RNA Extracted from Fresh Frozen and Formalin-Fixed Paraffin-Embedded Samples. RNA, 13, 1668-1674. http://dx.doi.org/10.1261/rna.642907
- 20. Li, J., Smyth, P., Flavin, R., Cahill, S., Denning, K., Aherne, S., Guenther, S.M., O’Leary, J.J. and Sheils, O. (2007) Comparison of miRNA Expression Patterns Using Total RNA Extracted from Matched Samples of Formalin-Fixed Paraffin-Embedded (FFPE) Cells and Snap Frozen Cells. BMC Biotechnology, 7, 36.
http://dx.doi.org/10.1186/1472-6750-7-36 - 21. Munch-Petersen, H.D., Ralfkiaer, U., Sjo, L.D., Hother, C., Asmar, F., Nielsen, B.S., Brown, P., Ralfkiaer, E. and Gronbaek, K. (2015) Differential Expression of miR-155 and miR-21 in Tumor and Stroma Cells in Diffuse Large B-Cell Lymphoma. Applied Immunohistochemistry & Molecular Morphology, 23, 188-195.
http://dx.doi.org/10.1097/PAI.0000000000000073 - 22. Kosaka, N., Iguchi, H., Yoshioka, Y., Hagiwara, K., Takeshita, F. and Ochiya, T. (2012) Competitive Interactions of Cancer Cells and Normal Cells via Secretory MicroRNAs. The Journal of Biological Chemistry, 287, 1397-1405. http://dx.doi.org/10.1074/jbc.M111.288662
- 23. Qin, Q., Wei, F.R. and Li, B.S. (2014) Multiple Functions of Hypoxia-Regulated miR-210 in Cancer. Journal of Experimental & Clinical Cancer Research, 33, 50. http://dx.doi.org/10.1186/1756-9966-33-50
- 24. Volinia, S., Calin, G.A., Liu, C.G., Ambs, S., Cimmino, A., Petrocca, F., Visone, R., Iorio, M., Roldo, C., Ferracin, M., Prueitt, R.L., Yanaihara, N., Lanza, G., Scarpa, A., Vecchione, A., Negrini, M., Harris, C.C. and Croce, C.M. (2006) A MicroRNA Expression Signature of Human Solid Tumors Defines Cancer Gene Targets. Proceedings of the National Academy of Sciences of the United States of America, 103, 2257-2261. http://dx.doi.org/10.1073/pnas.0510565103
- 25. Aqeilan, R.I., Calin, G.A. and Croce, C.M. (2010) MiR-15a and miR-16-1 in Cancer: Discovery, Function and Future Perspectives. Cell Death and Differentiation, 17, 215-220.
http://dx.doi.org/10.1038/cdd.2009.69 - 26. Li, H., Hui, L. and Xu, W. (2012) MiR-181a Sensitizes a Multidrug-Resistant Leukemia Cell Line K562/A02 to Daunorubicin by Targeting BCL-2. Acta Biochimica et Biophysica Sinica, 44, 269-277.
http://dx.doi.org/10.1093/abbs/gmr128 - 27. Neilson, J.R., Zheng, G.X., Burge, C.B. and Sharp, P.A. (2007) Dynamic Regulation of miRNA Expression in Ordered Stages of Cellular Development. Genes & Development, 21, 578-589.
http://dx.doi.org/10.1101/gad.1522907 - 28. Wickramasinghe, N.S., Manavalan, T.T., Dougherty, S.M., Riggs, K.A., Li, Y. and Klinge, C.M. (2009) Estradiol Downregulates miR-21 Expression and Increases miR-21 Target Gene Expression in MCF-7 Breast Cancer Cells. Nucleic Acids Research, 37, 2584-2595. http://dx.doi.org/10.1093/nar/gkp117
- 29. Kelly, P.N. and Strasser, A. (2011) The Role of BCL-2 and Its Pro-Survival Relatives in Tumourigenesis and Cancer Therapy. Cell Death and Differentiation, 18, 1414-1424.
http://dx.doi.org/10.1038/cdd.2011.17 - 30. Meng, F., Henson, R., Wehbe-Janek, H., Ghoshal, K., Jacob, S.T. and Patel, T. (2007) MicroRNA-21 Regulates Expression of the PTEN Tumor Suppressor Gene in Human Hepatocellular Cancer. Gastroenterology, 133, 647-658. http://dx.doi.org/10.1053/j.gastro.2007.05.022
- 31. Weiland, M., Gao, X.H., Zhou, L. and Mi, Q.S. (2012) Small RNAs Have a Large Impact: Circulating MicroRNAs as Biomarkers for Human Diseases. RNA Biology, 9, 850-859.
http://dx.doi.org/10.4161/rna.20378
NOTES
*Corresponding author.