Open Journal of Immunology
Vol.06 No.03(2016), Article ID:69776,14 pages
10.4236/oji.2016.63009
Colonization by Superantigen Producing Staphylococcus aureus in Mice Enhances the Capacity to Develop Oral Tolerance
Anna Stern, Erika Lindberg, Forough L. Nowrouzian, Agnes E. Wold, Sofia Östman
Department of Infectious Medicine, Institute of Biomedicine, Sahlgrenska Academy, University of Gothenburg, Gothenburg, Sweden

Copyright © 2016 by authors and Scientific Research Publishing Inc.
This work is licensed under the Creative Commons Attribution International License (CC BY).
http://creativecommons.org/licenses/by/4.0/



Received 30 March 2016; accepted 13 August 2016; published 16 August 2016
ABSTRACT
Microbial stimulation in early childhood may be necessary for proper maturation of the immune system. Infants colonized with Staphylococcus aureus have low risk of developing food allergy. Neonatal exposure to staphylococcal superantigen improves oral tolerance and enhances protection in experimental allergy models. Here, we used three wild-type strains of S. aureus, naturally harboring genes for different superantigens (SElM/SElO alone, or in combination with SEA or TSST-1). We first investigated their in vitro stimulatory capacity of splenocytes from germ-free mice. Secondly, germ-free mice were colonized with the strains and their capacity to develop oral tolerance was tested in a food allergy model. In vitro, S. aureus with only SElM/SElO genes promoted the strongest B-cell stimulation. S. aureus carrying gene for SEA induced the highest proportion of CD4+FoxP3+ T cells. The proportion of regulatory T cells was inversely correlated to B-cell proliferation, indicating suppressive ability of these cells. All strains were equally able to colonize the germ-free gut, initially achieving 1010CFU/g faeces, which decreased to 105 over a period of six weeks. Mice colonized with S. aureus carrying genes for SEA or TSST-1 had improved capacity to develop tolerance compared to germ-free mice. These results suggest that colonization by S. aureus producing superantigens may improve active tolerance to gut allergens.
Keywords:
Oral Tolerance, Superantigen, Staphylococcus aureus

1. Introduction
The newborn infant is born sterile with a relatively immature immune system. Microbes in the environment establish in the gut, leading to development of a complex normal microbiota that is capable of activating the immune system and inducing its maturation [1] - [3] . Immune maturation includes the capacity to develop tolerance towards exogenous inhaled or fed protein antigens, a process called oral tolerance. Thus, germ-free mice are deficient in development of oral tolerance [4] [5] and in tolerogenic processing of a dietary antigen [6] and have less functional regulatory T cells [7] .
Failure of oral tolerance results in allergy, a disease that has increased during the last decades in the Western societies. The cause of this rapid increase is unknown but according to the hygiene hypothesis [8] and many epidemiological studies [8] - [15] , there is a clear correlation between a hygienic lifestyle and the prevalence of allergy. Furthermore, children delivered by cesarean section have an increased risk of developing food allergy [16] [17] and a complex gut microbiota in early infancy protects against later allergy development [18] - [21] .
Staphylococcus aureus is a typical member of the skin microflora, foremost colonizing the anterior nares [22] - [24] . However, S. aureus is common in the gut flora of Swedish infants [25] [26] , perhaps as a result of decreased competition by traditional fecal bacteria whose circulation has decreased, in parallel with improved sanitary conditions [27] . Quite surprisingly, gut colonization with S. aureus was found to protect infants from development of food allergy [28] .
S. aureus strains can produce a range of toxins and to date over 20 different enterotoxins and related toxins have been described, including the staphylococcal enterotoxins (SEs), the staphylococcal enterotoxin-like proteins (SEls) and toxic shock syndrome toxin-1 (TSST-1). The SEs include SEA-E, G-I, R and T and the SEls include SElJ, K-Q, S, U, V and X [29] . These toxins bind to conserved Vβ regions on the T cell receptor and to MHC class II molecules on antigen-presenting cells. This cross-linking leads to massive T-cell activation and the toxins are called superantigens. Approximately 45% of the S. aureus strains that colonize the gut of Swedish infants produce one or more of the superantigens SEA-D or TSST-1. SEC is most common, followed by SEA [25] [26] . In addition, >60% of the S. aureus strains colonizing the gut of Swedish infants carry the egc locus which encodes several superantigens, such as SElM and SElO [30] . Infants colonized by S. aureus producing superantigens were found to have higher levels of serum IgA than infants colonized by enterotoxin-negative strains or not colonized by S. aureus [28] . In a mouse model, mucosal exposure to S. aureus superantigen in the neonatal period improves their ability to develop oral tolerance to an unrelated food protein at adult age. Protection was evident both in a respiratory hypersensitivity model and a food allergy model [31] [32] .
Regulatory T cells (Tregs) are present within the CD4+CD25+ T cell population. They express the transcription factor forkhead box P3 (FoxP3) [33] and can be divided into two major subsets; Tregs that are induced in the periphery from naïve CD4+FoxP3neg T cells and Tregs that are derived from the thymus [34] . Tregs have suppressive function and act by dampening other immune cells either by secretion of suppressive cytokines and/or by contact dependent mechanisms and are essential to preserve self-tolerance and immune homeostasis such as regulating allergic inflammation, as reviewed by Palomares et al. [35] and are known to be induced during development of oral tolerance, as discussed by Pabst and Mowat [36] . Mice that lack Tregs [37] and humans that are deficient in FoxP3 [38] [39] suffer from autoimmune and inflammatory diseases such as allergic disease. In vitro stimulation with S. aureus converts cord CD4+ T cell into suppressive CD4+FoxP3+ regulatory T cells [40] .
Lastly, S. aureus bacterial cells are also potent B cell mitogens, due to their possession of protein A, a surface protein that binds conserved portions of the IgG molecule, as well as a fraction of IgM molecules [41] . Thus, S. aureus has a unique capacity for antigen non-specific activation of antigen-specific cells, a fact that could explain their immunomodulatory potential.
In this study, we have investigated three S. aureus strains, isolated from stool samples of Swedish infants, for their ability to colonize and stimulate the immune system of germ-free mice. All three strains carried the gene cassette egc encoding the SEls M and O. Two of the isolates carried genes for either SEA or TSST-1.
2. Materials and Methods
2.1. Animals
Germ-free (GF) female Swiss-Webster mice, 5 - 10 weeks old, were maintained in flexible film isolators under a strict 12 h light cycle. GF status was verified regularly by anaerobic culturing in addition to PCR for bacterial 16S rDNA. Both GF and conventionally raised (CONV-R) mice were fed autoclaved chow diet (Labdiet) ad libitum. The mice were transferred from the isolators at 5 - 10 weeks-of-age and then maintained during standard housing condition for the course of the experiment. This study was carried out in strict accordance with recommendations from the Swedish board of agriculture and approved by the regional ethical committee (Gothenburg Counties of Västra Götaland and Värmland, Kammarrätteni Göteborg, Box 1531, 401 50GÖTEBORG, Permit No: 229-2010).
2.2. Bacterial Strains
Three different S. aureus strains were used for in vivo colonization of mice (Table 1). The strains were isolated from fecal samples of infants in the ALLERGYFLORA birth-cohort. The culture, isolation and identification procedures have previously been published [27] . All strains carried the enterotoxin-encoding genes selm and selo, which are located on the enterotoxin gene cluster (egc). Two of the strains produced either superantigen SEA or TSST-1, as shown both by production of toxin in vitro (SET-RPLA kit and TST-RPLA; Oxoid, Hampshire, Great Britain) and by the presence of the respective genes by multiplex PCR (unpublished observations). The isolates were further analyzed for carriage of genes and none of them carried genes for enterotoxins SEB-D (seb-d) and SEH (seh); enterotoxin-like SElL (sell), SElR (selr), SElP (selp), SElK (selk) and SElQ (selq); exfoliative toxin A, B and D (eta, etb and etd); epidermal cell differentiation inhibitor toxin A-C (edin); PVL (pvl) and leukotoxin M (lukM). Genes encoding beta-haemolysin (hlB) was only present in the S. aureus strain carrying enterotoxin SElM/SElO and TSST-1.
All strains carried adhesin genes encoding collagen-binding protein (cbp), laminin-binding protein (lbp), elastin binding-protein (ebp), clumping factors (ClfA, ClfB) and fibronectin-binding protein A and B (fnbA, fnbB), whereas genes encoding bone sialoprotein-binding protein (bsp) and fibrinogen-binding protein (fib) were only carried by two strains (Table 1). The bacteria were cultured on Colombia blood agar, harvested, washed and adjusted to109 bacteria/ml. For in vitro use, bacteria were inactivated by exposure to UV-light for 25 min, and inactivation was confirmed by negative viable count.
2.3. Colonization
Germ-free mice were removed from their isolators and given one of three S. aureus strains at approximately 3 ×
Table 1. Expression of virulence genes by Staphylococcus aureus fecal isolates used in vivo and in vitro.
a. Presence or absence of the following toxin genes was assessed by multiplex PCRs. The following virulence genes were not expressed by any of the strains; seb, sec, sed, seh, sell, selr, selp, selk, selq, eta, etb, etd, edin, pvl and lukm; b. agr-accessory gene regulator.
108 colony-forming units (CFU) intragastrically using sterile feeding needles. The mice were subsequently kept under conventional housing conditions with the ability to spontaneously acquire microbes from the environment.
2.4. Collection, Culture and Identification of Samples from the Bowel Flora
Fecal samples were collected prior to colonization, at 48 h as well as 6 and 10 weeks after colonization. The samples were suspended in 1 ml of sterile peptone water, diluted serially, and plated on selective and non-selec- tive media that were incubated under aerobic or anaerobic conditions. The total number of facultative bacteria was quantified on Colombia blood agar plates cultivated aerobically at 37˚C for 24 h. Enterobacteriaceae were quantified on Drigalski agar and speciated using the API20E biotyping system (bioMérieux, Askim, Sweden). Staphylococci were quantified on staphylococcus agar, identified by Gram-stained appearance and a positive catalase reaction. S. aureus was distinguished from coagulase-negative staphylococci by the coagulase reaction. Enterococci were quantified on enterococcosel agar, where they caused esculine hydrolysis and showed typical Gram-stained appearance.
Anaerobes were quantified after growth on Brucella blood agar incubated anaerobically at 37˚C for 48 - 72 h. Each colony type was enumerated separately and checked individually for inability to grow under aerobic conditions and discarded if able to do so. However, scant aerobic growth of gram-positive rods was permitted, as many bifidobacteria and lactobacilli are able to replicate aerobically. Anaerobic strains were speciated by Rapid ID 32A biotyping (bioMérieux). The total counts of anaerobes were quantified on Brucella blood agar, after the numbers of facultative bacteria growing on the plates were subtracted.
2.5. In Vitro Proliferation Assay
To study the effect of S. aureus bacteria on immune cells, in vitro experiments were performed using spleen cells from germ-free mice. The mice were sacrificed and spleens were excised, single cell suspensions were prepared and incubated with NH4Cl buffer (pH 7.3) for 5 min at 37˚C to lyse red blood cells. To measure proliferation of different lymphocyte subsets, splenocytes were stained with CFSE according to the manufacturer’s description (CellTrace™ CFSE Cell Proliferation Kit, Invitrogen, Paisley, UK). The cells were then aliquoted at 1 × 105 cells/well in microtiter plates together with 5 × 108 UV-killed S. aureus/mL (final concentration) and incubated in 5% CO2 at 37˚C. After 4 days of culture, cells were washed and stained for surface markers of B cells (CD19) and T cells (CD4) and for intracellular expression of FoxP3, as described below (flow cytrometry section). Cells were acquired using BD FACSCanto II (BD Biosciences) and analyzed with FlowJo software (Treestarinc, Ashland, OR). Proliferation was measured by examination of CFSE-staining.
2.6. Oral Tolerization and Food Allergy Protocol
Six weeks after monocolonization the mice were fed 5 mg ovalbumin, OVA (grade V, Sigma-Aldrich) in 250 μl PBS. The dose was selected to achieve partial oral tolerance. Non-tolerized controls were fed PBS. Conventionally reared mice were fed 5 mg OVA and included as a positive control for oral tolerance induction.
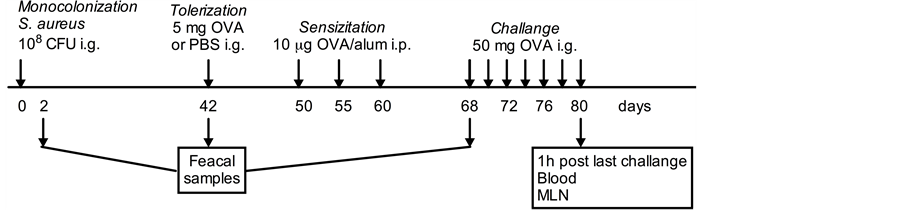
The degree of tolerance to OVA was tested in an OVA-induced food allergy model (Figure 1) [42] . One week after oral tolerization, mice were sensitized three times, 5 days apart, with 10 μg OVA adsorbed onto 2 mg of Al (OH)3 gel (Alum; Sigma-Aldrich) by i.p. injection. Eight days after last sensitization, the animals were challenged intragastrically with 50 mg OVA in 250 μL PBS and this challenge was repeated every other day, six times in total. Mice were deprived of food for 2 h prior to each challenge. Diarrhea was assessed by visually monitoring mice for up to 1 h after each challenge. Mice demonstrating profuse liquid stools were recorded as positive.
One hour after the final OVA challenge, the mice were anesthetized by i.p. injection of a mixture of xylazine (130 mg/kg) and ketamine (670 mg/kg). When the mice were in deep anesthesia, the chest was opened and blood was drawn by heart puncture. The blood was divided into Eppendorf tubes for serum preparation, and EDTA tubes (Sarstedt) for enumeration of eosinophils in the blood. The tubes were centrifuged (3000× g) for 15 min and serum were collected and frozen at −20˚C. Cell pellets in the EDTA tubes were treated with ammonium chloride buffer for lysis of red blood cells. Remaining cells were centrifuged onto glass slides using Cytospin (Thermo scientific, Waltham, MA) and stained with May Grünwald/Giemsa to visualize eosinophils.
Figure 1. Experimental design of food allergy model. Groups of germ-free Swiss-Webster mice (n = 6) were colonized with Staphylococcus aureus (S. aureus) strains carrying genes for different staphylococcal enterotoxins; SElM and SElO alone; S. a (SElM/O) or in combination with SEA; S. a (SEA) or TSST-1; S. a (TSST-1). Fecal samples were collected 48 h and 6 and 10 weeks after colonization to investigate the colonization ability of the different strains. Six weeks after colonization, the mice were fed OVA to induce oral tolerance, or PBS as control. Food allergy was induced one week later by sensitization through three i.p. injections of alum-adsorbed OVA, administrated five days apart, and followed one week later by per oral challenge with OVA every other day for 12 days. The mice were sacrificed 1 h after the last challenge and blood and mesenteric lymph nodes (MLN) were collected for analysis.
2.7. Flow Cytometry
Mesenteric lymph nodes were collected and prepared into single cell suspensions and stained for flow cytometric analysis using the following monoclonal antibodies: CD4-APC (clone RM4-5, BD Biosciences), CD19- PerCP-CyTM5.5 (clone 1D3, BD Biosciences) and FoxP3-APC (clone FJK-16s, ebioscience). To examine the expression of cell surface molecules, cells were pre-incubated with FcγR-blocking mAb and then stained with antibodies towards the different surface molecules. For intracellular staining with FoxP3, the cells were fixed and permeabilized with Fix/Perm (eBioscience) for 30 min followed by staining. Cells were acquired using BD FACSCanto II (BD Biosciences) and analyzed with FlowJo software (Treestarinc, Ashland, OR).
2.8. Determination of Total IgE
The concentration of serum IgE was determined by sandwich ELISA. Costar plates were coated with capture antibody (mouse anti-IgE antibody, 1 μg/mL; BD Biosciences). Serum samples were diluted threefold in four steps. Mouse IgE (BD Biosciences) was used as standard, serially diluted twofold in seven steps. Biotinylated anti-mouse IgE (2 μg/mL; BD Biosciences) was used as the detection antibody. The limit of detection was 6 ng/mL.
2.9. Statistical Analysis
Statistical analysis was conducted using nonparametric Mann-Whitney U-test or One-way ANOVA with Tukey’s correction for multiple comparisons. Correlations were analyzed with Pearson’s rank correlation test. P values < 0.05 were considered statistically significant.
3. Results
3.1. UV-Inactivated S. aureus are Potent B Cell Stimulators in Vitro
Natural colonization in the gut by S. aureus during early infancy has been associated with reduced risk of food allergy at 18 months of age, and with signs of immune stimulation, by means of increased levels of IgA and soluble CD14 in serum [28] [43] . Many S. aureus strains can produce enterotoxins with superantigen activity. We here set out to compare the in vitro stimulatory capacity of three isolates of S. aureus differentiating in carriage of superantigen genes. All three strains carried the gene cassette egc encoding the SEls M and O. Two of the isolates carried genes for either SEA or TSST-1, two toxins that are produced by many commensal S. aureus from infants [26] , hereafter referred to as S. aureus (SEA) and S. aureus (TSST-1). The third isolate only carried the egc cassette, referred to as S. aureus (SElM/O). Pathogenic characterization of the bacterial strains, such as virulence factors, can be found in Table 1.
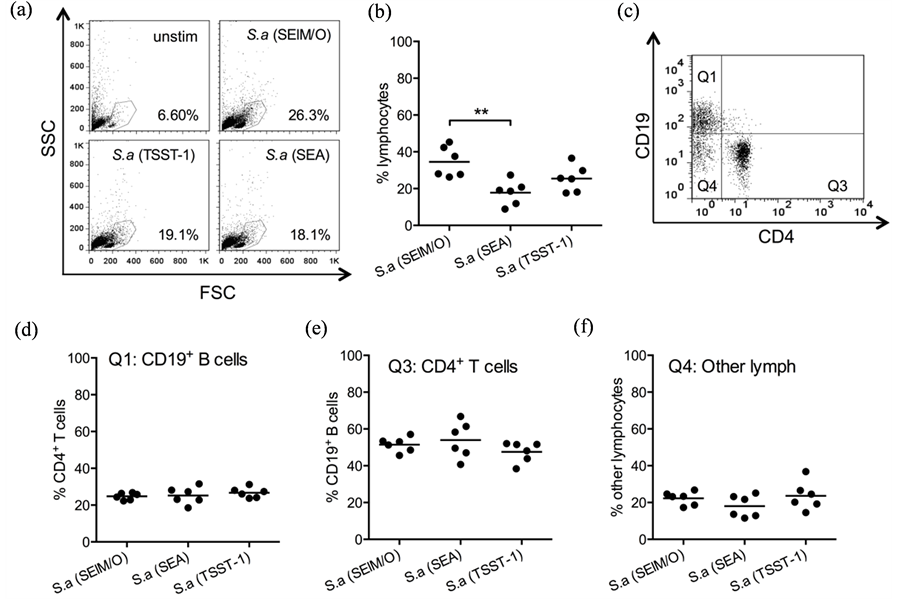
The bacteria were inactivated by UV radiation and washed before use and thereafter unable to produce toxins. As shown in the representative dot plots in Figure 2(a), stimulation of splenocytes by intact S. aureus bacteria
Figure 2. Staphylococcus aureus (S. aureus) carrying genes for SElM and SElO (SElM/O), but not SEA or TSST-1, is the most potent lymphocyte stimulator. Splenocytes from germ-free mice were stained with CFSE and stimulated in vitro with three different intact UV-inactivated S. aureus isolates carrying genes for different superantigens; SElM and SElO alone, S. a (SElM/O), or in combination with SEA, S. a (SEA) or TSST-1, S. a (TSST-1) and analyzed after four days of culture by flow cytometry. a) Representative dot plot showing gating of lymphocytes. b) Proportion of lymphocyte gated cells. c) Representative dot plot showing gating of CD19+ B cells, CD4+ T cells and non-CD4/CD19 lymphocytes so called “other lymphocytes”. (d)-(f) Proportions of indicated lymphocyte population within the lymphocyte gate. **P < 0.01, analyzed with One-way ANOVA with Tukey’s correction for multiple comparisons.
strongly increased the number of lymphocytes compared with unstimulated control cultures. Among the three strains, the S. aureus (SElM/O) yielded the highest proportion of lymphocytes in the culture (Figure 2(b)). Within the lymphocyte gated cells the proportions of T cells, B cells and other lymphocytes (presumably CD8+ T cells and NK cells) were similar in all stimulated cultures (Figures 2(d)-(f), gating strategy in Figure 2(c)).
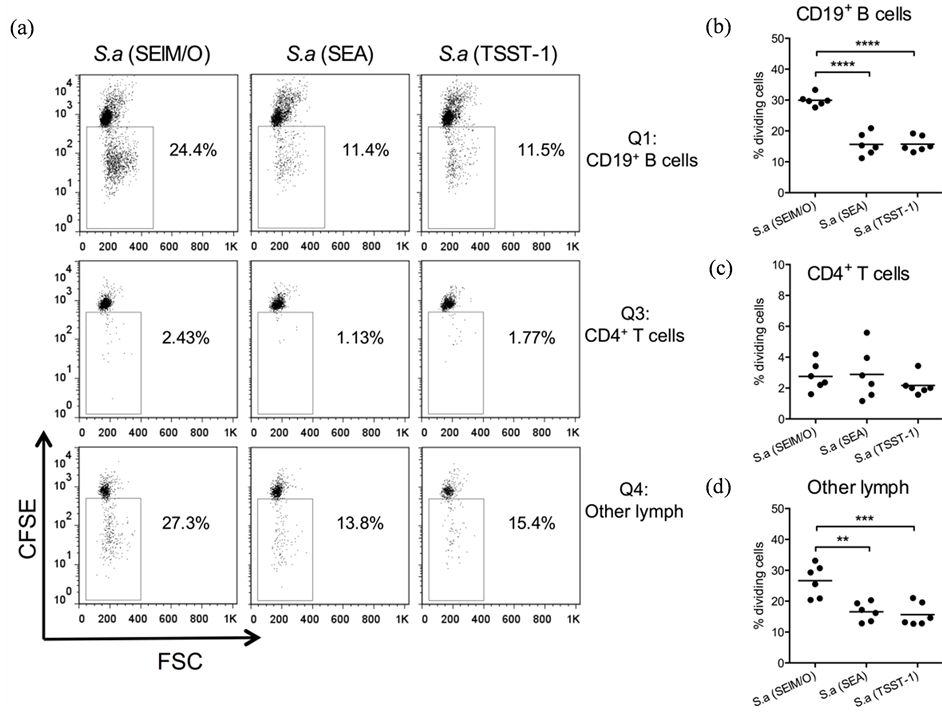
Figure 3 examines which lymphocyte populations that proliferated in response to the different S. aureus isolates, examined by CFSElow/intermediate expression. Representative dot plots are shown in Figure 3(a). Intact but inactivated S. aureus cells were strong stimulators of B cell division and also promoted division of “other lymphocytes” including CD8+ T cells and NK cells, inducing proliferation of 10% - 30% of the B cells and “other lymphocytes”. Interestingly, S. aureus (SElM/O) induced much stronger proliferation of B cells and “other lymphocytes” than did the strains carrying genes for SEA or TSST-1 (Figure 3(b), Figure 3(d)). About 2% - 4% of the T cells underwent proliferation in S. aureus stimulated cultures (Figure 3(c), gating strategy in Figure 2(a), Figure 2(c) with no differences between the three isolates.
3.2. The Frequency of CD4+FoxP3+ T Cells Correlates Negatively with B-Cell Proliferation
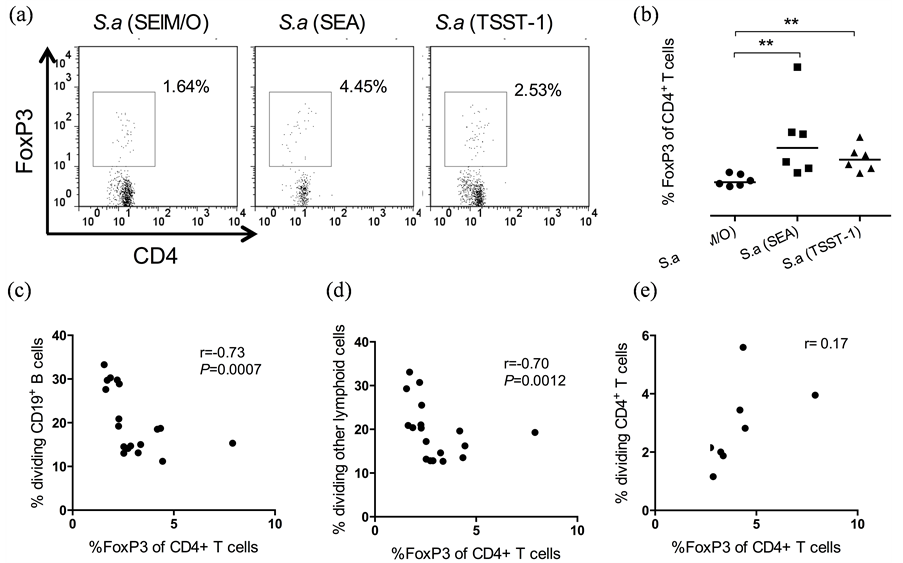
Figure 4 shows intracellular expression of FoxP3 in the stimulated cells, with representative dot plots shown in Figure 4(a). We observed a slight increase in proportion of FoxP3 expressing CD4+ T cells in the cultures stimulated with S. aureus (SEA) and S. aureus (TSST-1) as compared to cultures stimulated with S. aureus (SElM/O) (Figure 4(b), gating strategy in Figure 2(a), Figure 2(c)). The proportion of FoxP3+ T cells among the CD4+ T cells correlated negatively to B-cell proliferation (r = −0.73, P = 0.0007; Figure 4(c)) as well as to proliferation of other lymphoid cells (r = −0.70, P = 0.0012; Figure 4(d)). Proliferation of CD4+ T cells was
Figure 3. Staphylococcus aureus (S. aureus) carrying genes for SElM and SElO (SElM/O), but not SEA or TSST-1, induce the strongest proliferation of B cells and other lymphocytes. Splenocytes from germ-free mice were stained with CFSE, stimulated in vitro with three different intact UV-inactivated S. aureus isolates carrying genes for different superantigens; SElM and SElO alone, S. a (SElM/O), or in combination with SEA, S. a (SEA) or TSST-1, S. a (TSST-1) and analyzed after four days of culture by flow cytometry. a) Representative dot plot showing gating of dividing lymphocyte populations being CFSEintermediate/low. Top panel represent B cells, middle panel T cells and bottom panel non-CD4/CD19 lymphocytes so called “other lymphocytes”. b) Proportion of dividing CD19+ B cells. c) Proportion of dividing CD4+ T cells. d) Proportion of dividing other lymphocytes. **P < 0.01, ***P < 0.001, ****P < 0.0001 analyzed with One-way ANOVA with Tukey’s correction for multiple comparisons.
unrelated to FoxP3 expression in CD4+ T cells (r = 0.17; Figure 4(e)).
3.3. Different S. aureus Strains Colonize the Mouse Gut with Similar Efficiency
To test the in vivo modulatory capacity of the three different S. aureus strains, we colonized germ-free mice with either of the strains, left the mice to acquire bacteria from the environment, and thereafter tested their capacity to actively tolerize a novel protein antigen, ovalbumin (OVA). The experimental layout is shown in Figure 1. Fecal samples were collected prior to colonization and at 48 h, 6 weeks and 10 weeks after colonization. The first fecal culture after colonization showed S. aureus in high numbers, 109 - 1010 CFU/g faeces. This in contrast to conventionally reared (CONV-R) mice that had low counts of S. aureus; 103 CFU/g faeces or less, during the whole experimental period (Figure 5(a)); the competition from a full complex microbiota keeps the population counts low. After 6 weeks, the population counts of S. aureus had decreased to by approximately 4 log units (Figure 5(a)). This occurred in parallel with a less pronounced reduction in facultative bacteria in general (Figure 5(b)), and an increase in obligate anaerobic bacteria (Figure 5(c)). This resembles the situation in human infants where S. aureus population counts decrease from an average of 109 CFU/g to 104 CFU/g over the first year of life [26] in parallel with acquisition of a complex microbiota dominated by obligate anaerobes. The
Figure 4. Staphylococcus aureus (S. aureus) carrying gene for staphylococcal enterotoxin A (SEA) or toxic shock syndrome toxin-1 (TSST-1) induce a slight increase in FoxP3 expression in stimulated CD4+ T cells. Splenocytes from germ-free mice were stained with CFSE, stimulated in vitro with three different intact UV-inactivated S. aureus isolates carrying genes for different staphylococcal enterotoxins; SEM and SEO alone, S. a (SEM/O), or in combination with SEA, S. a (SEA) or TSST-1, S. a (TSST-1) and analyzed after four days of culture by flow cytometry. a) Representative dot plots showing gating strategy of CD4+FoxP3+ T cells. b) Proportion of FoxP3+ T cells among the CD4+ T-cell population. **P < 0.01 analyzed with One-way ANOVA with Tukey’s correction for multiple comparisons. c) Correlation plot between proportions of dividing B cells and FoxP3+ T cells. d) Correlation plot between proportions of dividing other lymphocyte cells and FoxP3+ T cells. e) Correlation plot between proportions of dividing CD4+ T cells and FoxP3+ T cells. Analyzed by Pearson’s rank correlation test.
different strains of S. aureus displayed similar colonization capacity (Figure 5(a)).
Bacteria that were spontaneously acquired from the environment were roughly typed. The facultative bacteria foremost included enterococci (average 108 and 107 CFU/g at 6 and 10 weeks respectively; data not shown), which are gram-positive bacteria, while Enterobacteriaceae were not identified, despite that selective media for their isolation were used. Obligate anaerobes acquired spontaneously included clostridia, Prevotella, Bacteroides, lactobacilli and bifidobacteria (all colonizing at >1010 CFU/g faeces; data not shown).
3.4. Colonization with S. aureus Strains Producing SEA or TSST-1 Promotes Oral Tolerance
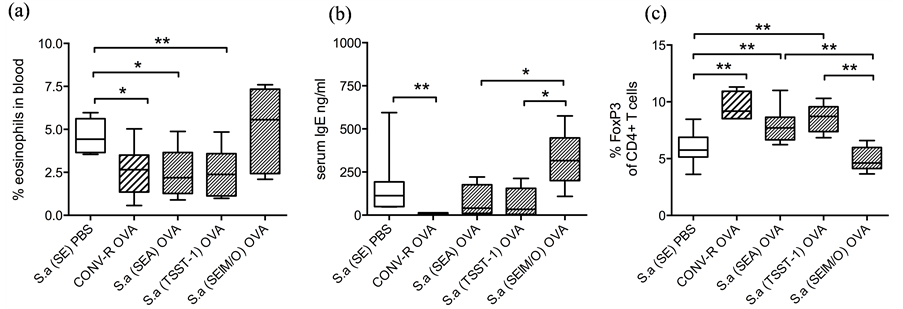
Six weeks after inoculation with S. aureus, the mice were fed OVA in order to induce partial oral tolerance. This was followed by sensitization and challenge in a model of OVA-induced food allergy. For experimental design, see Figure 1. In brief, the mice were immunized with OVA in alum followed by intragastric challenge with OVA which produces hypersensitivity characterized by diarrhea and enhanced numbers of eosinophils in blood in BALB/c mice. After challenge with OVA, we examined the mice for diarrhea. However, in contrast to BALB/c mice that develop profuse diarrhea in this model [32] , none of the Swiss-Webster mice used in the experiment developed diarrhea. Blood eosinophilia is another sign of hypersensitivity in the food allergy model. Non-tolerized S. aureus (SE) colonized mice developed stronger eosinophilia compared to tolerized mice. Tolerized mice colonized with the S. aureus (SElM/O) strain had similar levels of blood eosinophils as non-tolerized mice, while this response was dampened in mice colonized with S. aureus (SEA) or S. aureus (TSST-1) (Figure 6(a)).
Non-tolerized mice exhibited considerably higher levels of IgE in serum than did tolerizedCONV-R mice,
Figure 5. S. aureus strains carrying different superantigen genes colonize the gut in a similar fashion. Germ-free mice (n = 6) were colonized with different wild-type S. aureus strains isolated from Swedish infants. Fecal samples were collected at 48 h (filled bars), 6 weeks (open bars) and 10 weeks (striped bars) after colonization to investigate (a) the colonization ability of the different S. aureus strains as well as the uptake from the environment by estimating population counts of (b) facultative and (c) anaerobic bacteria. Data included are representative of two independent experiments.
Figure 6. Colonization by superantigen producing S. aureus improves oral tolerance. Groups of germ-free mice were colonized with different wild type S. aureus strains (n = 6). Mice were thereafter orally tolerized, sensitized and challenged with OVA in the food allergy model. PBS-fed controls were included as well as OVA-fed conventionally reared (CONV-R) mice. a) The proportion of eosinophils in the blood was estimated by differential counts of blood cells. b) Levels of serum IgE were measured by ELISA. c) The frequency of CD4+ T cells expressing FoxP3 in the mesenteric lymph nodes was estimated by flow cytometry. Data included are representative of two independent experiments. *P < 0.05 and **P < 0.01, analyzed with Mann-Whitney U-test.
which had undetectable levels (Figure 6(b)). Mice colonized by the S. aureus (SElM/O) strain had elevated levels of total IgE, while the mice colonized by the strains carrying genes for the classical superantigens, S. aureus (SEA) or S. aureus (TSST-1), were less elevated (Figure 6(b)).
The proportion of CD4+Foxp3+ cells in mesenteric lymph nodes in mice undergoing the food allergy model was analyzed by flow cytometry. Tolerized CONV-R mice had a higher proportion of FoxP3+ T cells as compared to non-tolerized mice (Figure 6(c)). Also, tolerized S. aureus (SEA) or S. aureus (TSST-1) colonized mice had higher levels of FoxP3+ T cells, compared to the PBS fed controls (Figure 6(c)). The S. aureus (SElM/O) colonized mice had comparable levels of FoxP3+ T cells to the PBS fed control, which in fact was significantly lower than the levels observed in both S. aureus (SEA) and S. aureus (TSST-1) colonized mice (Figure 6(c)).
4. Discussion
We have observed an association between early intestinal colonization with S. aureus and a lower risk of food allergy at 18 months of age [43] . We speculated that exposure to S. aureus bacteria and their enterotoxins during infancy would promote profound immune stimulation acting by improving the maturation of the immune system and consequently reduce the risk of later allergy development. We have investigated the effect of S. aureus superantigens on the neonatal immune system in mice and shown that neonatal oral exposure to SEA improves oral tolerance development, which is crucial to prevent allergic reactions. Here, we sought to investigate the potential of the S. aureus bacteria on immunoregulatory function with respect to the toxins the bacteria could produce, by investigating both the in vitro stimulatory response and the in vivo colonization and immunomodulatory ability of three different S. aureus strains.
We started by investigating the in vitro stimulatory capacity of the S. aureus strains. We observed that all three strains induced adequate stimulation to B cells. However, the strain that only possessed genes encoding the SEl proteins SElM and SElO, and not any SEs, was the strongest B cell stimulator. The basis for this difference is unknown but could relate to e.g. difference in cell wall structure or protein A content. All three strains induced equal stimulation to T cells. Though, we noted a slight increase in FoxP3 expression on T cells stimulated with S. aureus carrying gene for SEA or TSST-1. The expression of FoxP3 on CD4+ T cells was also inversely correlated to B-cell proliferation indicating that the FoxP3-expressing T cells may have suppressive function and act by reducing the stimulatory capacity of the B cells. It was shown in a study by Rabe et al. that stimulation of cord blood cells with S. aureus promoted upregulation of FoxP3 on CD4+ T cells and that these cells demonstrated functional suppressive capacity. However, it was not clear from that study if the strain used for in vitro stimulation expressed any superantigens [40] . It is reasonable to believe that substantial T cell proliferation would be followed by induction of Tregs to dampen the massive immune response. However, in our setting stimulating splenocytes from germ-free mice with UV-killed S. aureus did not induce a profound T cell proliferation but nevertheless the proportion of FoxP3-expressing CD4+ T cells increased, supporting that view.
Taken together, these data imply that S. aureus strains different in their carriage of superantigen genes differentiates in their in vitro stimulatory ability of splenocytes from germ-free mice. Our data indicate that induction of Tregs seen with stimulation of S. aureus with genes for SEA or TSST-1 may account for the reduced B-cell proliferation seen in the cultures. Although, the increase in proportion of FoxP3+ T cells was limited and the observation needs to be further verified in additional experiments.
Gut colonization by S. aureus is more common today in Swedish infants, which probably is a result of disappearance of “classical” gut commensals, such as E. coli [26] [27] [44] . The colonization model used in this study bears some resemblance to the mode of colonization of infants delivered by cesarean section, who are not exposed to their mother’s vaginal or fecal flora and acquire their strains from other environmental sources. The mice picked up enterococci as well as anaerobic bacteria (clostridia, Prevotella, Bacteroides, lactobacilli and bifidobacteria) from the environment but did not acquire E. coli, as this bacterium is confined to the fecal microflora and its establishment is delayed in caesarean delivered infants [45] . In parallel with acquisition of an anaerobic flora, S. aureus population counts decreased from 109 to 105 CFU/g, which is similar to what is seen in Western infants in whom S. aureus levels reach 107 on average by one week of age in colonized infants, 105 by 6 months of age and 104 by 12 months of age [25] - [27] [46] . This confirms that S. aureus is less fit for competition with an increasingly complex anaerobic flora than, for example, enterococci which were able to maintain population levels of >107 CFU/g. It is therefore likely that S. aureus strains mainly exert their effects early after colonization, when their population levels are high and antibodies to the superantigens have not yet been produced.
In the Swedish infants, colonization with S. aureus strains carrying the genes encoding for SElM/O were associated to higher population counts in the infant stools and was thus suggested to be a virulence factor promoting gut colonization. In the same study, S. aureus carrying gene for SEA were associated to lower population counts in infant stools and suggested to be a less potent gut colonizer [30] . However, here we observed no difference in the colonization ability between the S. aureus strains used. The strain having the gene for SEA colonized equally well as the other two strains. This might be due to the complete lack of competition from other microbes in the germ-free mice giving all S. aureus strains a unique colonization possibility. Also, the milieu in the animal facilities is relatively clean and the colonized S. aureus strains may experience less competition from other microbes as compared to human infants. Thus, our observed difference in the outcome in the food allergy model cannot be explained by differences in colonization ability between the S. aureus strains used in our model. Germ-free mice are known to have a poor ability for oral tolerance induction [5] [6] [47] , which goes in line with a poorly developed immune system, especially in the gut. Conventionalization of germ-free mice restores maturation of the immune system and oral tolerance development [48] . Mono-colonization, with selected bacteria is in general less potent for immune maturation but one exception is the spore-forming segmented filamentous bacterium (SFB) which has been shown in mono-colonization experiments to have a unique immunomodulatory ability [49] . By the eye, germ-free mice have few payers’ patches, enlargement of the caecum and increased mucus production in the intestine. Here, colonization with S. aureus, followed by conventionalization was fairly potent in restoring the phenotype of conventional mice. At the termination of the experiment we observed that the S. aureus colonized mice had comparable number of Payer’s patches, reduced size of the caecum (although not normalized) and less mucus in the intestine (data not shown), which indicates normalization from a germ-free to a conventional state. However, when we studied the ability for tolerance induction we found differences between the S. aureus colonized mice. We have previously shown that neonatal treatment of mice with commercially purified SEA leads to improved oral tolerance demonstrated as improved protection in an airway allergy model [31] as well as in a food allergy model [32] . Here, we demonstrate that colonization by S. aureus strains carrying genes for SEA or TSST-1 had some improvement in oral tolerance induction, as measured in the food allergy model. This was in contrast to mice colonized with a S. aureus strain only carrying genes for SElM and SElO. These mice were equally allergic as the PBS fed controls. These results suggest that in vivo exposure to certain superantigens might promote maturation of immunoregulatory mechanisms that favor oral tolerance development. The slight improvement in oral tolerance observed in germ-free mice colonized with S. aureus carrying genes for SEA or TSST-1 was accompanied by an increased proportion of CD4+ T cells in the mesenteric lymph nodes expressing the regulatory T cell marker FoxP3. Furthermore, blood eosinophilia was less pronounced after OVA challenge in animals that had been colonized by the strains having SEA or TSST-1 genes compared to animals colonized with strain having only SElM/O genes. This implies that colonization with S. aureus with the ability to produce SEA or TSST-1 augmented the capacity to induce oral tolerance to OVA. Colonization with S. aureus carrying the gene for SEA was most efficient regarding this improvement. SEA has been proposed to be the most potent superantigen [50] and this might be related to its ability to cross-link two MHC class II molecules and, hence, more potent cell activation [51] [52] . Tolerance feeding was performed six weeks after initial colonization, when S. aureus counts were quite low suggesting that colonization resulted in long-term modulation of immune function.
5. Conclusion
Colonization of germ-free mice with commensal S. aureus derived from the gut microbiota of infants gave rise to immune modulation that in many aspects resembled that was seen in newborn human infants colonized with these strains. The effects on the immune system are also similar to these seen in animals neonatally given purified toxin, without the bacteria. These observations suggest that S. aureus colonizing the gut indeed elaborates toxins in quantities sufficient to afford immune activation. The uniquely strong T-cell activating capacity of S. aureus enterotoxins may provide the developing immune system with the adequate activation and maturation signals that may be lacking in infants with a poorly developed intestinal microflora.
Acknowledgements
We would like to thank Professor Fredrik Bäckhed at the University of Gothenburg for providing us with the germ-free mice. This work was funded by the Swedish Research Council, the Medical Faculty of Gothenburg (LUA) and the Torsten and Ragnar Söderberg foundation.
Cite this paper
Anna Stern,Erika Lindberg,Forough L. Nowrouzian,Agnes E. Wold,Sofia Östman, (2016) Colonization by Superantigen Producing Staphylococcus aureus in Mice Enhances the Capacity to Develop Oral Tolerance. Open Journal of Immunology,06,79-92. doi: 10.4236/oji.2016.63009
References
- 1. Hesselmar, B., Aberg, N., Aberg, B., Eriksson, B. and Bjorksten, B. (1999) Does Early Exposure to Cat or Dog Protect Against Later Allergy Development? Clinical & Experimental Allergy, 29, 611-617.
http://dx.doi.org/10.1046/j.1365-2222.1999.00534.x - 2. Kramer, U., Heinrich, J., Wjst, M. and Wichmann, H.E. (1999) Age of Entry to Day Nursery and Allergy in Later Childhood. The Lancet, 353, 450-454.
http://dx.doi.org/10.1016/S0140-6736(98)06329-6 - 3. Braun-Fahrlander, C., Gassner, M., Grize, L., Neu, U., Sennhauser, F.H., Varonier, H.S., et al. (1999) Prevalence of Hay Fever and Allergic Sensitization in Farmer’s Children and Their Peers Living in the Same Rural Community. SCARPOL Team. Swiss Study on Childhood Allergy and Respiratory Symptoms with Respect to Air Pollution. Clinical & Experimental Allergy, 29, 28-34.
http://dx.doi.org/10.1046/j.1365-2222.1999.00479.x - 4. Eggesbo, M., Botten, G., Stigum, H., Nafstad, P. and Magnus, P. (2003) Is Delivery by Cesarean Section a Risk Factor for Food Allergy? Journal of Allergy and Clinical Immunology, 112, 420-426.
http://dx.doi.org/10.1067/mai.2003.1610 - 5. Eggesbo, M., Botten, G., Stigum, H., Samuelsen, S.O., Brunekreef, B. and Magnus, P. (2005) Cesarean Delivery and Cow Milk Allergy/Intolerance. Allergy, 60, 1172-1173.
http://dx.doi.org/10.1111/j.1398-9995.2005.00857.x - 6. Wang, M., Karlsson, C., Olsson, C., Adlerberth, I., Wold, A.E., Strachan, D.P., et al. (2008) Reduced Diversity in the Early Fecal Microbiota of Infants with Atopic Eczema. Journal of Allergy and Clinical Immunology, 121, 129-134.
http://dx.doi.org/10.1016/j.jaci.2007.09.011 - 7. Bisgaard, H., Li, N., Bonnelykke, K., Chawes, B.L., Skov, T., Paludan-Muller, G., et al. (2011) Reduced Diversity of the Intestinal Microbiota during Infancy Is Associated with Increased Risk of Allergic Disease at School Age. Journal of Allergy and Clinical Immunology, 128, 646-652.e5.
http://dx.doi.org/10.1016/j.jaci.2011.04.060 - 8. Abrahamsson, T.R., Jakobsson, H.E., Andersson, A.F., Bjorksten, B., Engstrand, L. and Jenmalm, M.C. (2012) Low Diversity of the Gut Microbiota in Infants with Atopic Eczema. Journal of Allergy and Clinical Immunology, 129, 434-440.e2.
http://dx.doi.org/10.1016/j.jaci.2011.10.025 - 9. Ismail, I.H., Oppedisano, F., Joseph, S. J., Boyle, R.J., Licciardi, P.V., R, Robins-Browne, R.M. and Tang, M.L. (2012) Reduced Gut Microbial Diversity in Early Life Is Associated with Later Development of Eczema but Not Atopy in High-Risk Infants. Pediatric Allergy and Immunology, 23, 674-681.
http://dx.doi.org/10.1111/j.1399-3038.2012.01328.x - 10. Kluytmans, J., van Belkum, A. and Verbrugh, H. (1997) Nasal Carriage of Staphylococcus aureus: Epidemiology, Underlying Mechanisms, and Associated Risks. Clinical Microbiology Reviews, 10, 505-520.
- 11. Nouwen, J.L., van Belkum, A. and Verbrugh, H.A. (2001) Determinants of Staphylococcus aureus Nasal Carriage. Netherlands Journal of Medicine, 59, 126-133.
http://dx.doi.org/10.1016/S0300-2977(01)00150-4 - 12. Vandenbergh, M.F. and Verbrugh, H.A. (1999) Carriage of Staphylococcus aureus: Epidemiology and Clinical Relevance. Journal of Laboratory and Clinical Medicine, 133, 525-534.
http://dx.doi.org/10.1016/S0022-2143(99)90181-6 - 13. Lindberg, E., Adlerberth, I., Hesselmar, B., Saalman, R., Strannegard, I.L., Aberg, N. and Wold, A.E. (2004) High Rate of Transfer of Staphylococcus aureus from Parental Skin to Infant Gut Flora. Journal of Clinical Microbiology, 42, 530-534.
http://dx.doi.org/10.1128/JCM.42.2.530-534.2004 - 14. Lindberg, E., Nowrouzian, F., Adlerberth, I. and Wold, A.E. (2000) Long-Time Persistence of Superantigen-Producing Staphylococcus aureus Strains in the Intestinal Microflora of Healthy Infants. Pediatric Research, 48, 741-747.
http://dx.doi.org/10.1203/00006450-200012000-00007 - 15. Adlerberth, I., Lindberg, E., Aberg, N., Hesselmar, B., Saalman, R., Strannegard, I.L. and Wold, A.E. (2006) Reduced Enterobacterial and Increased Staphylococcal Colonization of the Infantile Bowel: An Effect of Hygienic Lifestyle? Pediatric Research, 59, 96-101.
http://dx.doi.org/10.1203/01.pdr.0000191137.12774.b2 - 16. Lundell, A.C., Hesselmar, B., Nordstrom, I., Saalman, R., Karlsson, H., Lindberg, E., et al. (2009) High Circulating Immunoglobulin a Levels in Infants Are Associated with Intestinal Toxigenic Staphylococcus aureus and a Lower Frequency of Eczema. Clinical & Experimental Allergy, 39, 662-670.
http://dx.doi.org/10.1111/j.1365-2222.2008.03176.x - 17. Xu, S.X. and McCormick, J.K. (2012) Staphylococcal Superantigens in Colonization and Disease. Front Cell Infect Microbiol, 2, 52.
http://dx.doi.org/10.3389/fcimb.2012.00052 - 18. Nowrouzian, F.L., Dauwalder, O., Meugnier, H., Bes, M., Etienne, J., Vandenesch, F., et al. (2011) Adhesin and Superantigen Genes and the Capacity of Staphylococcus aureus to Colonize the Infantile Gut. Journal of Infectious Diseases, 204, 714-721.
http://dx.doi.org/10.1093/infdis/jir388 - 19. Lonnqvist, A., Ostman, S., Almqvist, N., Hultkrantz, S., Telemo, E., Wold, A.E. and Rask, C. (2009) Neonatal Exposure to Staphylococcal Superantigen Improves Induction of Oral Tolerance in a Mouse Model of Airway Allergy. European Journal of Immunology, 39, 447-456.
http://dx.doi.org/10.1002/eji.200838418 - 20. Stern, A., Wold, A.E. and Ostman, S. (2013) Neonatal Mucosal Immune Stimulation by Microbial Superantigen Improves the Tolerogenic Capacity of CD103+ Dendritic Cells. PLoS ONE, 8, e75594.
http://dx.doi.org/10.1371/journal.pone.0075594 - 21. Hori, S., Nomura, T. and Sakaguchi, S. (2003) Control of Regulatory T Cell Development by the Transcription Factor Foxp3. Science, 299, 1057-1061.
http://dx.doi.org/10.1126/science.1079490 - 22. Curotto de Lafaille, M.A. and Lafaille, J.J. (2009) Natural and Adaptive Foxp3+ Regulatory T Cells: More of the Same or a Division of Labor? Immunity, 30, 626-635.
http://dx.doi.org/10.1016/j.immuni.2009.05.002 - 23. Palomares, O., Yaman, G., Azkur, A.K., Akkoc, T., Akdis, M. and Akdis, C.A. (2010) Role of Treg in Immune Regulation of Allergic Diseases. European Journal of Immunology, 40, 1232-1240.
http://dx.doi.org/10.1002/eji.200940045 - 24. Pabst, O. and Mowat, A.M. (2012) Oral Tolerance to Food Protein. Mucosal Immunology, 5, 232-239.
http://dx.doi.org/10.1038/mi.2012.4 - 25. Sakaguchi, S., Sakaguchi, N., Asano, M., Itoh, M. and Toda, M. (1995) Immunologic Self-Tolerance Maintained by Activated T Cells Expressing IL-2 Receptor Alpha-Chains (CD25). Breakdown of a Single Mechanism of Self-Tolerance Causes Various Autoimmune Diseases. Journal of Immunology, 155, 1151-1164.
- 26. Chatila, T.A., Blaeser, F., Ho, N., Lederman, H.M., Voulgaropoulos, C., Helms, C. and Bowcock, A.M. (2000) JM2, Encoding a Fork Head-Related Protein, Is Mutated in X-Linked Autoimmunity-Allergic Disregulation Syndrome. Journal of Clinical Investigation, 106, R75-R81.
http://dx.doi.org/10.1172/jci11679 - 27. Wildin, R.S., Smyk-Pearson, S. and Filipovich, A.H. (2002) Clinical and Molecular Features of the Immunodysregulation, Polyendocrinopathy, Enteropathy, X Linked (IPEX) Syndrome. Journal of Medical Genetics, 39, 537-545.
http://dx.doi.org/10.1136/jmg.39.8.537 - 28. Rabe, H., Nordstrom, I., Andersson, K., Lundell, A.C. and Rudin, A. (2014) Staphylococcus aureus Convert Neonatal Conventional CD4+ T Cells into FOXP3+ CD25+ CD127low T Cells via the PD-1/PD-L1 Axis. Immunology, 141, 467-481.
http://dx.doi.org/10.1111/imm.12209 - 29. Boyle, M.D.P. and Reis, K.J. (1987) Bacterial Fc Receptors. Nature Biotechnology, 5, 697-703.
http://dx.doi.org/10.1038/nbt0787-697 - 30. Brandt, E.B., Strait, R.T., Hershko, D., Wang, Q., Muntel, E.E., Scribner, T.A., et al. (2003) Mast Cells Are Required for Experimental Oral Allergen-Induced Diarrhea. Journal of Clinical Investigation, 112, 1666-1677.
http://dx.doi.org/10.1172/JCI19785 - 31. Lundell, A.C., Adlerberth, I., Lindberg, E., Karlsson, H., Ekberg, S., Aberg, N., et al. (2007) Increased Levels of Circulating Soluble CD14 but Not CD83 in Infants Are Associated with Early Intestinal Colonization with Staphylococcus aureus. Clinical & Experimental Allergy, 37, 62-71.
http://dx.doi.org/10.1111/j.1365-2222.2006.02625.x - 32. Nowrouzian, F., Hesselmar, B., Saalman, R., Strannegard, I.L., Aberg, N., Wold, A.E. and Adlerberth, I. (2003) Escherichia Coli in Infants’ Intestinal Microflora: Colonization Rate, Strain Turnover, and Virulence Gene Carriage. Pediatric Research, 54, 8-14.
http://dx.doi.org/10.1203/01.PDR.0000069843.20655.EE - 33. Matricardi, P.M. (1997) Infections Preventing Atopy: Facts and New Questions. Allergy, 52, 879-882.
http://dx.doi.org/10.1111/j.1398-9995.1997.tb01246.x - 34. Strachan, D.P. (2000) Family Size, Infection and Atopy: The First Decade of the “Hygiene Hypothesis”. Thorax, 55, S2-S10.
http://dx.doi.org/10.1136/thorax.55.suppl_1.S2 - 35. Matricardi, P.M. and Bonini, S. (2000) High Microbial Turnover Rate Preventing Atopy: A Solution to Inconsistencies Impinging on the Hygiene Hypothesis? Clinical & Experimental Allergy, 30, 1506-1510.
http://dx.doi.org/10.1046/j.1365-2222.2000.00994.x - 36. Hjern, A., Rasmussen, F. and Hedlin, G. (1999) Age at Adoption, Ethnicity and Atopic Disorder: A Study of Internationally Adopted Young Men in Sweden. Pediatric Allergy and Immunology, 10, 101-106.
http://dx.doi.org/10.1034/j.1399-3038.1999.00023.x - 37. Strachan, D.P. (1989) Hay Fever, Hygiene, and Household Size. BMJ, 299, 1259-1260.
http://dx.doi.org/10.1136/bmj.299.6710.1259 - 38. Ostman, S., Rask, C., Wold, A.E., Hultkrantz, S. and Telemo, E. (2006) Impaired Regulatory T Cell Function in Germ-Free Mice. European Journal of Immunology, 36, 2336-2346.
http://dx.doi.org/10.1002/eji.200535244 - 39. Rask, C., Evertsson, S., Telemo, E. and Wold, A.E. (2005) A Full Flora, But Not Monocolonization by Escherichia coli or Lactobacilli, Supports Tolerogenic Processing of a Fed Antigen. Scandinavian Journal of Immunology, 61, 529-535.
http://dx.doi.org/10.1111/j.1365-3083.2005.01598.x - 40. Moreau, M.C. and Gaboriau-Routhiau, V. (1996) The Absence of Gut Flora, the Doses of Antigen Ingested and Aging Affect the Long-Term Peripheral Tolerance Induced by Ovalbumin Feeding in Mice. Research in Immunology, 147, 49-59.
http://dx.doi.org/10.1016/0923-2494(96)81548-3 - 41. Moreau, M.C. and Corthier, G. (1988) Effect of the Gastrointestinal Microflora on Induction and Maintenance of Oral Tolerance to Ovalbumin in C3H/HeJ Mice. Infection and Immunity, 56, 2766-2768.
- 42. Crabbe, P.A., Nash, D.R., Bazin, H., Eyssen H. and Heremans, J.F. (1970) Immunohistochemical Observations on Lymphoid Tissues from Conventional and Germ-Free Mice. Laboratory Investigation, 22, 448-457.
- 43. Crabbe, P.A., Bazin, H., Eyssen, H. and Heremans, J.F. (1968) The Normal Microbial Flora as a Major Stimulus for Proliferation of Plasma Cells Synthesizing IgA in the Gut. The Germ-Free Intestinal Tract. International Archives of Allergy and Immunology, 34, 362-375.
http://dx.doi.org/10.1159/000230130 - 44. Carter, P.B. and Pollard, M. (1971) Host Responses to “Normal” Microbial Flora in Germ-Free Mice. Journal of the Reticuloendothelial Society, 9, 580-587.
- 45. Adlerberth, I. and Wold, A.E. (2009) Establishment of the Gut Microbiota in Western Infants. Acta Paediatrica, 98, 229-238.
http://dx.doi.org/10.1111/j.1651-2227.2008.01060.x - 46. Adlerberth, I., Strachan, D.P., Matricardi, P.M., Ahrne, S., Orfei, L., Aberg, N., et al. (2007) Gut Microbiota and Development of Atopic Eczema in 3 European Birth Cohorts. Journal of Allergy and Clinical Immunology, 120, 343-350.
http://dx.doi.org/10.1016/j.jaci.2007.05.018 - 47. Moreau, M.C., Coste, M., Gaboriau, V. and Dubuquoy, C. (1995) Oral Tolerance to Ovalbumin in Mice: Effect of Some Parameters on the Induction and Persistence of the Suppression of Systemic IgE and IgG Antibody Responses. Advances in Experimental Medicine and Biology, 371B, 1229-1234.
- 48. Shroff, K.E. and Cebra, J.J. (1995) Development of Mucosal Humoral Immune Responses in Germ-Free (GF) Mice. Advances in Experimental Medicine and Biology, 371A, 441-446.
http://dx.doi.org/10.1007/978-1-4615-1941-6_92 - 49. Cebra,J.J., Periwal, S.B., Lee, G., Lee, F. and Shroff, K.E. (1998) Development and Maintenance of the Gut-Associated Lymphoid Tissue (GALT): The Roles of Enteric Bacteria and Viruses. Developmental Immunology, 6, 13-18.
http://dx.doi.org/10.1155/1998/68382 - 50. Mollick, J.A., Chintagumpala, M., Cook, R.G. and Rich, R.R. (1991) Staphylococcal Exotoxin Activation of T Cells. Role of Exotoxin-MHC Class II Binding Affinity and Class II Isotype. Journal of Immunology, 146, 463-468.
- 51. Al-Daccak, R., Mehindate, K., Damdoumi, F., Etongue-Mayer, P., Nilsson, H., Antonsson, P., Sundstrom, M., Dohlsten, M., Sekaly, R.P. and Mourad, W. (1998) Staphylococcal Enterotoxin D Is a Promiscuous Superantigen Offering Multiple Modes of Interactions with the MHC Class II Receptors. Journal of Immunology, 160, 225-232.
- 52. Tiedemann, R.E. and Fraser, J.D. (1996) Cross-Linking of MHC Class II Molecules by Staphylococcal Enterotoxin A Is Essential for Antigen-Presenting Cell and T Cell Activation. Journal of Immunology, 157, 3958-3966.