 Journal of Biomaterials and Nanobiotechnology, 2012, 3, 487-498 http://dx.doi.org/10.4236/jbnb.2012.324050 Published Online October 2012 (http://www.SciRP.org/journal/jbnb) 487 Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces Li-Chong Xu1*, Christopher A. Siedlecki1,2 1Department of Surgery, Biomedical Engineering Institute, College of Medicine, The Pennsylvania State University, Hershey, USA; 2Department of Bioengineering, Biomedical Engineering Institute, College of Medicine, The Pennsylvania State University, Hershey, USA. Email: *lxx5@psu.edu Received August 11th, 2012; revised September 13th, 2012; accepted September 24th, 2012 ABSTRACT Plasma proteins influence the initial adhesion of bacteria to biomaterials as well as interactions between bacteria and blood platelets on blood-contacting medical devices. In this paper, we study the effects of three human plasma proteins, albumin, fibrinogen (Fg), and fibronectin (Fn), on the adhesion of Staphylococcus epidemidis RP62A to polyurethane biomaterial surfaces, and also address how these three proteins affect bacterial interactions with human platelets on ma- terials. Measurements of bacterial adhesion on polymer surfaces pre-adsorbed with a variety of proteins demonstrate that Fn leads to increased bacterial adhesion, with the order of effectiveness being Fn Fg > albumin. Immuno-AFM (atomic force microscopy) was used to assess the Fn adsorption/activity on surfaces and bacterial cell membranes by looking at molecular scale events. A correlation between molecular scale Fn adsorption and macroscale bacterial adhe- sion was observed, with an increased numbers of Fn-receptor recognition events measured on cell surfaces as compared to Fg-receptor recognition events, suggesting Fn is an important protein in bacterial adhesion. Monoclonal antibodies recognizing either the carboxyl-terminus or amino-terminus of Fn were coupled to AFM probes and used to assess the orientation of Fn adsorbed on a surface, with an increased amount of Fn carboxyl-terminus availability corresponding to higher bacterial adhesion. Interactions between bacteria and platelets were demonstrated with fluorescence and AFM imaging on the polyurethane surfaces, with albumin inhibiting bacteria-platelet interaction and platelet activation, and both Fg and Fn promoting adhesion of bacteria to platelets and apparent platelet activation, resulting in bacteria/platelet aggregation. Keywords: Bacterial Adhesion; Staphylococcus epidermidis; Fibronectin; Bacteria-Platelet Interactions 1. Introduction Bacterial adhesion to biomaterials causing microbial in- fection and poor tissue integration is one of the main problems associated with the use of blood-contacting devices. Bacteria adhere to a material surface, where they proliferate and colonize to form biofilms on implanted devices, eventually leading to a biomaterial associated infection. These infections are extremely difficult to treat by use of antibiotics alone due to the formation of biofilm, which consists of a microbial community en- trapped within a polymer matrix secreted by the adherent microbes, and serves to protect the community from an- timicrobial agents [1,2]. The increase in antibiotic resis- tance of bacterium has also contributed to the increase in infections that are refractory to treatment [3,4]. Thus, surgical removal and replacement of the implanted de- vices is often the only treatment, causing significant mor- bidity and mortality [5,6]. Coagulase-negative staphylococci, particularly Staphy- lococcus epidermidis, rank first among the causative agent of nosocomial infections and represent the most common source of infections associated with the use of implanted medical devices such as intravascular and peritoneal dialysis catheters, prosthetic heart valves or orthopedic prostheses [7]. The defined virulence associ- ated with S. epidermidis is its ability to colonize and form biofilm on biomaterials [8]. Pathogenic bacteria associated with biomaterial-centered infections have the potential to enter the human circulatory system where they can interact with platelets, resulting in platelet acti- vation/aggregation. For example, S. aureus, another im- portant staphylococcal bacterium found in nosocomial infection, leads to platelet activation/aggregation [9-12] and subsequently leads to abnormal blood function such *Corresponding author. Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 488 as blood coagulation and thrombosis [13,14]. While in- teractions of S. aureus and platelets have been inten- sively studied, less information is available on S. epider- mids. Further knowledge of the interactions between S. epidermidis and blood contact devices as well as the in- teractions between bacteria and platelets is crucial in developing effective strategies for preventing biomate- rial-centered infection and its subsequent complications. Bacterial adhesion is the critical step in the pathogene- sis of biomaterial associated infection. Biomaterial sur- face chemistry characteristics have been shown to influ- ence the initial adhesion and aggregation of S. epider- midis on biomaterials [15]. However, when a biomaterial is implanted in contact with blood, plasma proteins can rapidly adsorb onto the material surface to form a “con- ditioning film”. This adsorbed protein layer may mini- mize the effect of biomaterial surface properties on bac- terial adhesion [16,17] so that interactions between the bacteria and the proteins mediate bacterial adhesion. In vitro studies have shown that the presence of serum pro- teins generally suppresses initial bacterial adhesion due to the lack of a specific interaction between albumin and bacteria [18,19] while the effect of plasma proteins fi- brinogen (Fg) and fibronectin (Fn) on bacterial adhesion is inconclusive. It has been reported that Fg or Fn coated substrata enhanced the adhesion of S. epidermidis [20-22] while there were reports also reporting that both proteins inhibited or had no effect on bacterial adhesion [23-25]. The increase in bacterial adhesion related to adsorbed Fg or Fn is regarded to be due to specific ligand/receptor events between plasma proteins and bacterial cell surface proteins known as the microbial surface components re- cognizing adhesive matrix molecules (MSCRAMM) [26-29]. Multiple MSCRAMM have been found on S. epidermidis surface to promote adhesion of bacteria. These molecules include proteins such as SdrG [20,30], SdrF [29,31], and Embp [28,32], which were identified to bind Fg, collagen, and Fn, respectively. It has also been demonstrated that SdrG promotes platelet adhe- sion/activation and aggregation [33]. Microscopy analysis and quantification of adherent bacteria are generally used to evaluate bacterial adhesion on material surfaces. This approach reveals cellular and macroscopic scale phenomena of bacterial adhesion un- der a variety of conditions and provides useful informa- tion on the relationships between bacterial adhesion and various experimental parameters. However, the direct measurement of interactions between material surfaces, plasma proteins, and the bacterial cell surface at the mo- lecular scale is particularly important for understanding the mechanisms of bacterial adhesion and pathogenic infection. Atomic force microscopy (AFM) is a powerful tool in studying bacterial adhesion, not only for imaging of bacterial cells under physiological conditions, but also for probing the nano-Newton (or less) interaction forces between bacteria and various substratum surfaces or bio- logical molecules [34]. Méndez-Vilas et al. [35,36] char- acterized the surfaces of slime covered S. epidermidis and the nano-mechanical properties of cell walls, show- ing the importance of cell surface properties to adhesion. Successful coating of bacteria on the AFM probe made it possible to directly measure the molecular interaction forces between bacterium and surfaces, indicating the probability of adhesion. Liu et al. [37] measured the ad- hesion forces between S. epidermidis and self-assembled monolayers surfaces in the presence of proteins and found that molecular adhesion forces between bacteria and Fn were much greater than the forces between bacte- ria and fetal bovine serum. Other investigators [38] measured the time-dependent bacterial adhesion forces of S. epidermidis to hydrophilic and hydrophobic surfaces using a similar approach and found different bond- strengths for staphylococcal adhesion to surfaces with different wettability. In this paper we studied the effects of plasma proteins on adhesion of S. epidermidis as well as bacteria interact- tions with platelets on microphase-separated polyure- thane (PU) biomaterial surfaces, an important material used for blood-contacting devices for over 30 years. Three plasma proteins (albumin, Fg, and Fn) were pre- adsorbed on PU surfaces and bacterial adhesion was measured. AFM was used to detect the molecular-scale Fn adsorption/orientation and to measure the interaction forces between proteins and bacterial cell surfaces to reveal the role of protein in bacterial adhesion. The cor- relation between molecular scale results and macroscale bacterial adhesion yields important information for un- derstanding the mechanisms of bacterial adhesion and biological responses to materials. 2. Material and Methods 2.1. General Phosphate buffered saline (PBS, 0.01 M, pH 7.4, Sigma) was prepared using purified water (18 M) from a Mil- lipore Simplicity 185 system. Human fibrinogen (Fg, 100% clottable) from Calbiochem (La Jolla, CA), human serum albumin (HSA, >99%) and human fibronectin (Fn) from Sigma Inc were used as received. A polyclonal anti-Fn antibody was obtained from Abcam (Cambridge, MA). Monoclonal antibodies (MAb) anti-amino (N-ter- minus) (MAb1936) and anti-carboxy (C-terminus) (MAb- 1935) of human Fn were purchased from Millipore (Bil- lerica, MA). MAB1936 specifically recognizes the N-ter- minal fibrin and heparin binding 29 kDa domain. MAB1935 recognizes the C-terminal domain containing Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 489 the second fibrin binding site [39]. 2.2. Polyurethane Film Preparation A BioSpan® MS/0.4 segmented polyurethane urea (PUU), having 22 wt% hard segments, was obtained from the Polymer Technology Group (Berkeley, CA). PUU films were prepared by a drop of solution casting onto round glass coverslips (15 mm dia, Ted Pella Inc., CA) and dried in a vacuum oven at 65˚C overnight. PUU films on glass coverslip were soaked in purified water over night and equilibrated in PBS for 1 hr before use. 2.3. Bacterial Strain Culture and Adhesion Strain S. epidermidis RP62A (ATCC 35984) was cul- tured in tryptic soy broth (TSB, BD) at 37˚C for 24 hrs and collected by centrifuge at 1360 g for 10 min. The pellet was resuspended in PBS and the concentration of bacteria was measured by a spectrophotometer at 600 nm. PUU films on glass coverslip hydrated in H2O for 24 hr were incubated in protein solutions at the desired con- centration for 15 min in 12-well tissue culture plate (BD). After adsorption of proteins, PUU films were rinsed with PBS three times and incubated in bacterial solution at a concentration of 1 × 108 cfu/ml for 1 hr with shaking at 250 rpm. The samples were rinsed in PBS 3 times and fixed in 2.5% glutaraldehyde for 2 hrs, then the bacteria were stained with Hoechst 33258 (Invitrogen) for 30 mins. Adherent bacteria on PUU films were imaged un- der a fluorescent optical microscopy with magnification of 1000× (Nikon, Eclipse 80i) at six random locations. The DAPI filter was used for the Hoechst 33258 (excita- tion/emission wavelengths of 352/461 nm). Cell numbers were quantified using Image J software (NIH, Bethesda, MD). Adhesion was measured on three replicates of each protein concentration and presented as average ± stan- dard deviation. 2.4. Modification of AFM Probes All AFM experiments were performed using a Multi- mode AFM equipped with a Nanoscope IIIa controller system (Veeco Instruments, Santa Barbara, CA). AFM probes having long-narrow Si3N4 triangular cantilevers (Veeco Instruments, Santa Barbara, CA, nominal k = 0.06 N/m) were modified with anti-Fn antibodies, puri- fied Fn or purified Fg. Probes were treated by glow dis- charge plasma at 100 W power for 30 min and then in- cubated in a 1% (v/v) solution of aminopropyltriethox- ysilane (Gelest Inc., PA) in ethanol for 1 hr to provide reactive amine groups on the tip. After thoroughly rins- ing with Millipore water, the probes were reacted with 10% glutaraldehyde in aqueous solution for 1 hr. The probes were again rinsed with Millipore water and incu- bated in protein solution (~20 µg/ml) for 1 hr. The probes were rinsed with PBS after removal from protein solution and were stored in PBS at 4˚C until use within 2 days. This attachment method has been shown to provide suf- ficient mobility and flexibility for proteins to rotate and orient themselves for binding [40,41]. Multiple probes were prepared together to improve consistency between experiments. The spring constants of cantilevers (all taken from the same wafer) were determined using the thermal tuning method (Nanoscope V6.12r2) using a multimode AFM with a PicoForce attachment and Nan- oscope IIIa control system (Veeco Instruments, Santa Barbara, CA). 2.5. Protein Adsorption and Orientation Detection An immuno-AFM technique [41] was used to detect the adsorption/orientation of Fn from mixtures with Fg or HSA on PUU film surfaces using a probe modified with anti-Fn antibodies. The orientation of Fn adsorbed on PUU surface was detected by the monocolonal antibodies (MAb) coupled AFM probes. PUU films hydrated in H2O for 24 hr were incubated in protein solutions at the desired concentrations for 15 min, and rinsed with PBS to remove the free proteins. The PUU film was mounted on the AFM stage in an AFM fluid cell filled with PBS. An array of 32 × 32 force curves were collected by force volume image mode with a scanning area of 1 × 1 μm2 at a scan rate of 1Hz and ramp size of 1 μm. As nonspecific controls for these measurements, polymer films were incubated with HSA for 10 min and used for measuring the nonspecific interactions of the antibody with protein. The individual retraction force curves were extracted and analyzed off-line with tools developed with Matlab soft- ware. The maximum debonding forces, defined as rup- ture force, calculated from the distance between the zero deflection value to the point of maximum deflection dur- ing probe separation from the surface for each force curve, was used as the strength of the interactions. The rupture length was calculated from the distance that the tip moves from the zero interaction force during separa- tion to the position where the probe has separated and returned to zero deflection. Both rupture force and rup- ture length can be used to distinguish the specific and non-specific interactions. 2.6. Measurement of Binding Strengths between Proteins and Cell Surface To directly measure the binding strengths between pro- teins and bacterial cell surface, bacteria were nonspecifi- cally attached onto the glass coverslips coated with poly- Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 490 L-lysine for 5 min and rinsed with PBS 3 times to re- move un-attached cells [37]. The cells were kept wet in PBS buffer for AFM measurement. AFM probes modi- fied with Fg or Fn were used to measure the interaction forces between protein-probe and bacterial cell surfaces in PBS. A probe coated with HSA was used as control for non-specific interactions between protein and cell surface. An array of 32 × 32 force curves were collected by AFM force volume mode. Force curve data were ex- tracted from AFM files and analyzed off-line with tools developed using Matlab software, and the maximum de- flection of the cantilever from each retracting curve was used to calculate binding strength. The slope of ap- proaching force curve was used to distinguish the forces curves measured on cell or substrate surfaces. 2.7. Interactions of Bacteria and Platelets Salvaged human platelets with citrate phosphate dextrose anticoagulant were obtained from the Blood Bank at Hershey Medical Center. A previous study showed that salvaged human platelets retain functional activity [42]. The platelets were centrifuged at 200 g for 20 min to remove any remaining red blood cells. The supernatant was centrifuged at 600 g for 20 min to separate platelets into a pellet. The platelet pellet was gently resuspended in 10 ml of PBS, and the platelet count was measured by a hematology analyzer (Sysmex KX-21N, Japan). The PUU films were incubated in 2 ml of PBS solution con- taining bacteria (1 × 108 cfu/ml) and platelets (2.5 × 108/ml) for 1 hr with shaking at 250 rpm on a shaker plate. Plasma proteins (HSA (4 mg/ml), Fg (0.3 mg/ml), and Fn (0.03 mg/ml)) were added into solutions respect- tively in order to study the influences of proteins on bac- teria-platelet interactions. After 1 hr, the adherent plate- lets and bacteria were fixed in 1% paraformaldehyde and 2.5% glutaraldehyde for 1 hr. After washing with PBS, platelets and bacteria on the PUU surface were stained and examined with a fluorescence microscopy. For pla- telet staining, samples were incubated in a primary anti- body solution containing Ab662 anti-human IIb 3 (1.5 µg/ml) in 6% normal donkey serum (Jackson Immu- noResearch) overnight at 4˚C. Following this labeling step, samples were washed with PBS and labeled with a secondary antibody by adding 10 μl/ml of AlexaFluor555 goat anti-mouse antibody IgG (Invitrogen) in 6% normal goat serum in the dark and at room temperature for 1 hr. Samples were rinsed with PBS and incubated in Hoechst 33258 solution for 30 min to stain bacteria. After rinsing in PBS, the sample was mounted under a coverslip with antifade gel (Biomeda) and stored at 4˚C overnight. The adherent platelets and bacteria were examined under fluorescence microscopy with appropriate fluorescence filters. In another experiment, the fixed platelets and bacteria samples on PUU surfaces were washed by pure water and dried in air for AFM imaging. 2.8. Image and Data Analysis The fluorescence images were analyzed by Image J software. Statistical analysis of bacterial adhesion data was performed by ANOVA utilizing the commercial software program GraftPad Instat (version 3.06). p < 0.05 was considered statistically significant. Significant dif- ferences are denoted by symbols (* or #) with one sym- bol denoting p < 0.05, two symbols denoting p < 0.01, and three symbols denoting p < 0.001. 3. Results 3.1. S. epidermidis RP62A Adhesion on PUU Surfaces with Pre-Adsorption of Proteins Bacterial adhesion values for S. epidermidis RP62A on PUU surfaces pre-adsorbed with different proteins and different concentrations are illustrated in Figure 1. For surfaces adsorbed with single proteins, the bacterial ad- hesion was significantly increased when surfaces were pre-adsorbed with Fn (either concentration) compared to either Fg or albumin. HSA appears to impart a small in- hibition of bacterial adhesion, evidenced by the lower adhesion against albumin compared to the control surface without any protein adsorption, although the result is not statistically significant. Fg produced a slight increase in adhesion compared to HSA, but adhesion to Fg was still much lower than Fn. Results showed the general trend of effect of single protein adsorption on bacterial adhesion to be in the order of Fn Fg > HSA. When surfaces were pre-adsorbed with dual component proteins solu- tions (either Fn + Fg or Fn + HSA), the Fg + Fn combi- nation showed greater bacterial adhesion than either the control or Fn + HSA samples, as expected. It is interest- ing to note that increasing the amount of Fg with respect to Fn in solution had no significant effect on bacterial adhesion (perhaps even a small drop in adhesion, though not significant) while increasing Fn concentration sig- nificantly increased the bacterial adhesion (Fn 0.01 + Fg 0.3 mg/ml and Fn 0.03 + Fg 0.3 mg/ml). 3.2. Fn Adsorption on PUU Surfaces The Fn adsorption on polyurethane surfaces was meas- ured by an immuno-AFM technique. Utilizing an AFM probe modified with polyclonal anti-Fn antibody (pAb), Fn on the polymer surface can be recognized by differen- tiating specific and non-specific interactions between antibody and adsorbed proteins [41]. Either the rupture force or rupture length (stretching of the interaction) Copyright © 2012 SciRes. JBNB 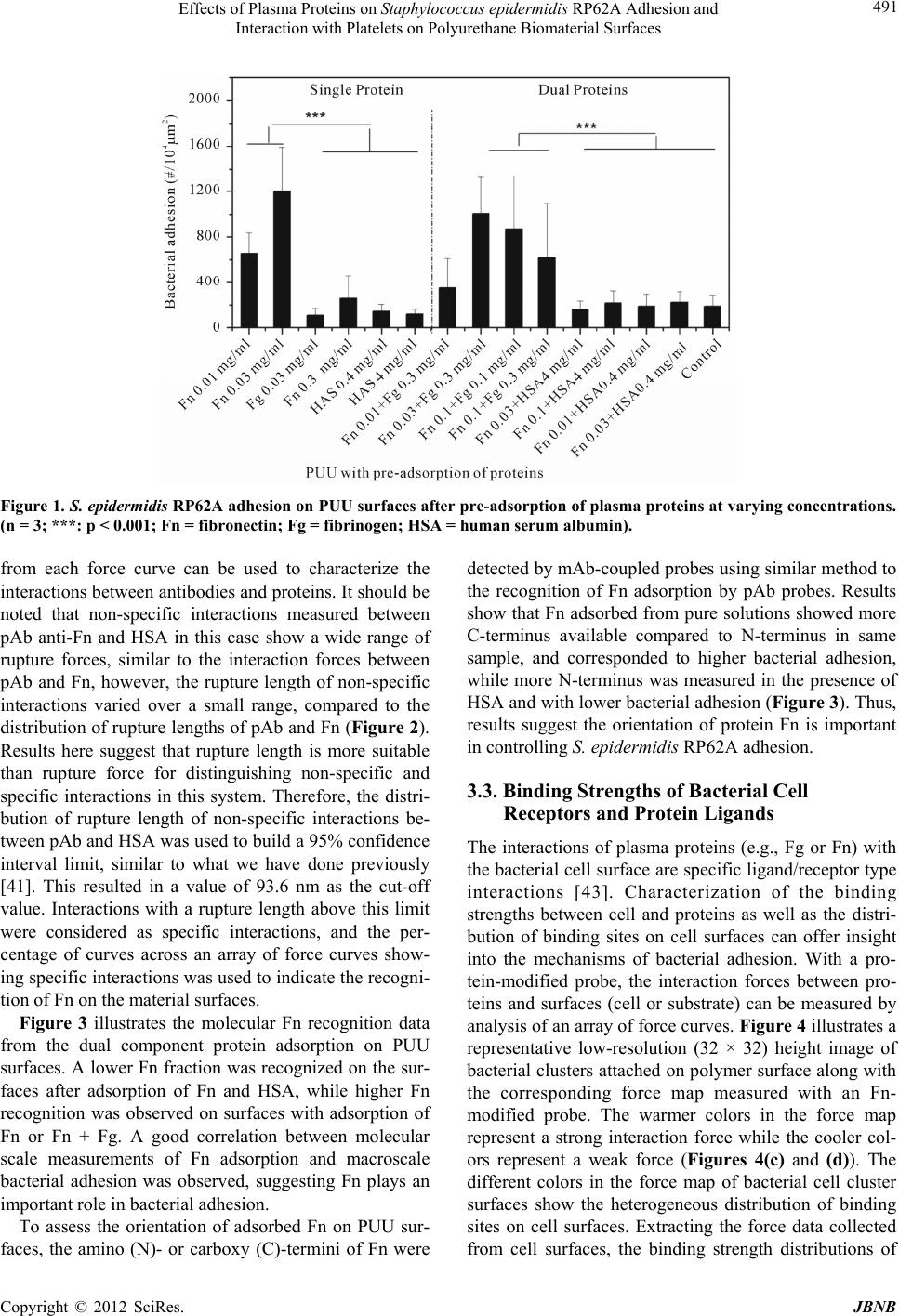 Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces Copyright © 2012 SciRes. JBNB 491 Figure 1. S. epidermidis RP62A adhesion on PUU surfaces after pre-adsorption of plasma proteins at varying concentrations. (n = 3; ***: p < 0.001; Fn = fibronectin; Fg = fibrinogen; HSA = human serum albumin). from each force curve can be used to characterize the interactions between antibodies and proteins. It should be noted that non-specific interactions measured between pAb anti-Fn and HSA in this case show a wide range of rupture forces, similar to the interaction forces between pAb and Fn, however, the rupture length of non-specific interactions varied over a small range, compared to the distribution of rupture lengths of pAb and Fn (Figure 2). Results here suggest that rupture length is more suitable than rupture force for distinguishing non-specific and specific interactions in this system. Therefore, the distri- bution of rupture length of non-specific interactions be- tween pAb and HSA was used to build a 95% confidence interval limit, similar to what we have done previously [41]. This resulted in a value of 93.6 nm as the cut-off value. Interactions with a rupture length above this limit were considered as specific interactions, and the per- centage of curves across an array of force curves show- ing specific interactions was used to indicate the recogni- tion of Fn on the material surfaces. Figure 3 illustrates the molecular Fn recognition data from the dual component protein adsorption on PUU surfaces. A lower Fn fraction was recognized on the sur- faces after adsorption of Fn and HSA, while higher Fn recognition was observed on surfaces with adsorption of Fn or Fn + Fg. A good correlation between molecular scale measurements of Fn adsorption and macroscale bacterial adhesion was observed, suggesting Fn plays an important role in bacterial adhesion. To assess the orientation of adsorbed Fn on PUU sur- faces, the amino (N)- or carboxy (C)-termini of Fn were detected by mAb-coupled probes using similar method to the recognition of Fn adsorption by pAb probes. Results show that Fn adsorbed from pure solutions showed more C-terminus available compared to N-terminus in same sample, and corresponded to higher bacterial adhesion, while more N-terminus was measured in the presence of HSA and with lower bacterial adhesion (Figure 3). Thus, results suggest the orientation of protein Fn is important in controlling S. epidermidis RP62A adhesion. 3.3. Binding Strengths of Bacterial Cell Receptors and Protein Ligands The interactions of plasma proteins (e.g., Fg or Fn) with the bacterial cell surface are specific ligand/receptor type interactions [43]. Characterization of the binding strengths between cell and proteins as well as the distri- bution of binding sites on cell surfaces can offer insight into the mechanisms of bacterial adhesion. With a pro- tein-modified probe, the interaction forces between pro- teins and surfaces (cell or substrate) can be measured by analysis of an array of force curves. Figure 4 illustrates a representative low-resolution (32 × 32) height image of bacterial clusters attached on polymer surface along with the corresponding force map measured with an Fn- modified probe. The warmer colors in the force map represent a strong interaction force while the cooler col- ors represent a weak force (Figures 4(c) and (d)). The different colors in the force map of bacterial cell cluster surfaces show the heterogeneous distribution of binding sites on cell surfaces. Extracting the force data collected from cell surfaces, the bindng strength distributions of i  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 492 (a) (b) (c) (d) Figure 2. Histogram of (a), (b) rupture forces and (c), (d) rupture lengths measured from PUU surfaces pre-adsorbed with HSA and Fn. Figure 3. Correlation between molecular Fn recognition, amino (N)-, carboxy (C)-termini of Fn, and bacterial adhesion on PUU surfaces. Statistical analysis symbol # denotes the comparison of Fn recognition to surfaces adsorbed from pure Fn (0.01 mg/ml) solution, and * denotes comparing N- and C-termini on same sample. Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 493 Figure 4. (a) Height image of bacterial clusters on polymer surface, and (b) corresponding force map measured by Fn-probe (32 × 32). Representative force curves measured on bacterial cell surface were shown as (c) weak interaction force corre- sponding to cool color in (b) and (d) strong force corresponding to warm color in (b) (AFM image size: 5 × 5 µm2). Fn or Fg were calculated and illustrated in Figure 5. To recognize the protein receptors on bacterial cell surfaces, HSA was used as a control and nonspecific interaction forces between HSA and bacterial cell surfaces were measured. The force distribution measured between HSA-probe and bacterial cell surfaces produced a 95% confidence limit at 0.36 nN. Binding strengths over this limit were considered as protein ligand and cell receptor interactions. There are approximately 48% of the meas- urements showing bacteria-Fg interactions while around 69% of the measurements showing an interaction with Fn-probes (Figures 5(a) and (b)). Results suggest more Fn-receptor recognition events than Fg-receptor recogni- tion events on the cell surface. 3.4. Interaction of Platelets and Bacteria in the Presence of Plasma Proteins Interaction of platelets and bacteria was illustrated from the distribution of platelets and bacteria adhered on PUU surfaces. Results show that bacteria adhered on surface and aggregated to form clusters. Although fewer platelets were observed compared to the number of bacteria, most platelets were found to be either entrapped in bacterial aggregates (green arrows in Figure 6) or adherent with bacteria (red arrows in Figure 6), suggesting the forma- tion of platelet-bacteria aggregates. The platelet-bacteria aggregates on the PUU surface were analyzed and counted when plasma proteins were added in bulk solu- tion or proteins were just pre-adsorbed on PUU surfaces (without proteins in bulk solution). Results show that Fn leads to more aggregates than either Fg or albumin in (a) (b) Figure 5. Histogram of binding strengths between bacterial cell surfaces and proteins (a) Fn, and (b) Fg. Figure 6. Fluorescent images of S. epidermidis RP62A bac- teria and platelets interactions on polyurethane surface, (a) platelets and (b) bacteria. Red circles are drawn for com- parison. (Image size: 226 μm × 169 μm). both cases, and more aggregates formed when the protein was present in bulk solution as opposed to just pre-ad- sorbed on the polymer surface (Figure 7). It is interest- ing to note that Fg showed a slight increase in aggrega- tion, not significantly, compared to albumin or control. The interactions of platelets and bacteria were further imaged by AFM. Figure 8 shows the morphology of Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 494 Figure 7. Bacteria-platelet aggregates on PUU surfaces ob- served when proteins were present. platelets and bacteria adhered on PUU surface. Both non-activated (round) and activated (spread) platelets were found on the surface when only platelets were pre- sent in solution (Figure 8(a)), however, platelets were found only activated and spread on the surface when bacteria were present and interacted with platelets. Bac- teria were seen to adhere with activated platelets and form aggregates (Figure 8(b)), suggesting that bacteria increase the platelets activation and aggregation of bacte- ria-platelet. When plasma proteins were present in solu- tion, HSA appeared to decrease the number of bacteria- platelet aggregates on PUU surface (Figure 8(c)) while Fg and Fn increased the formation of aggregates (Fig- ures 8(d) and (e)). Furthermore, non-activated platelets were observed on surface in the case of HSA while all platelets were activated in the cases of Fg and Fn. The images with large magnification show activated platelets either adhered to bacteria or entrapped in bacterial clus- ters (Figure 9). 4. Discussion Bacterial adhesion is the first step in the development of biofilm formation on implanted biomaterials. Factors that influence bacterial adhesion on a polymeric surface in- clude the nature of environment, type of microorganism, and properties of material, and each one of these factors is in turn affected by several other parameters [43]. When a material is implanted, plasma proteins rapidly interact with the surface to form a layer of proteins. The nature of adsorbed proteins is affected by the physico- chemical properties of surface, and in turn moderates the initial bacterial adhesion. S. epidermidis is a predominant bacterial species contributing to cardiovascular implant infection. In this study we measured the adhesion of S. Figure 8. AFM images of platelet and bacteria interactions on PUU surfaces, (a) platelet only, (b) platelet and bacteria without plasma proteins, platelets and bacteria in the pres- ence of (c) HSA, (d) Fg and (e) Fn. The bacteria-platelet aggregates are indicated by circles. (Image size: 50 µm × 50 µm). Figure 9. Aggregates of bacteria-platelets in the presence of Fn showing platelet (a) adherent or (b) entrapped with bac- teria. (Image size: 20 µm × 20 µm). epidermidis and interaction with platelets on polyure- thane biomaterial surfaces in the presence of plasma pro- teins. The molecular scale measurement of protein ad- sorption and interaction forces between protein and cells Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 495 were correlated to bacterial adhesion. Results provided important information to understand the roles of plasma proteins in bacterial adhesion and biological responses to implanted biomaterials. Albumin is the most abundant plasma protein. The ad- dition of HSA to Fn solution decreased the adsorption of Fn and subsequent bacterial adhesion to the PUU sur- faces (Figures 1 and 3). No ligand/receptor binding event was measured between albumin and the cell sur- faces. Furthermore, the presence of albumin appeared to decrease the activation of platelets and interactions with bacteria. Fibrinogen is the most third abundant plasma protein in blood and plays a prominent role in development of surface-induced thrombosis. It serves as a ligand, binding to the platelet integrin receptor IIb 3, leading to platelet immobilization, activation, and aggregation [44,45]. It is also found to promote S. aureus adhesion to material surfaces [17]. The increase in adhesion of S. aureus with Fg was identified to be due to the Fg-binding MSCRA- MM clumping factor on cell surface [27]. However, the presence of Fg shows no significant increase in S. epi- dermidis RP62A adhesion compared to HSA in this study. Lower bacterial adhesion was measured on poly- mer surfaces in the presence of Fg compared to Fn, al- though the solution concentration of Fg (0.3 mg/ml) is 10-times higher than that of Fn (0.03 mg/ml) (Figure 1). Results suggest that S. epidermidis RP62A cell surface has fewer binding sites to Fg than Fn, as evidenced by the molecular scale measurement of protein binding sites on cell surfaces, where approximately 48% of the meas- urements showed bacteria-Fg interactions while ~69% of the measurements showed an interaction with Fn-probes (Figure 5). Fibronectin is one of the main plasma proteins respon- sible for forming a conditioning film on implanted bio- materials. It can bind a variety of extracellular molecules including fibrin, heparin, and collagen, and plays a key role in cell adhesion and proliferation [46,47]. Numerous studies have found that Fn facilitates bacterial adhesion to biomaterials including S. epidermidis [48-50]. In this study, the macroscale measurements of bacterial adhe- sion show higher adhesion of bacteria on surfaces pre- adsorbed with Fn compared to Fg or HSA, and consistent with the amounts of molecular Fn detected by anti-Fn pAb probe on polymer surfaces (Figure 3). Results strongly suggest that Fn plays an important role in adhesion of S. epidermidis to polymer surfaces. Higher adhesion of bacteria on PUU surfaces bearing Fn is believed to be due to the interaction of Fn with MSCRAMM on S. epidermidis RP62A cell surfaces. William et al. identi- fied a giant Fn-binding protein, extracellular matrixbind- ing protein (embp), from S. epidermidis cell surface [28]. Christner et al. further demonstrated that embp mediates binding of S. epidermidis to solid phase attached Fn, constituting the first step of biofilm formation on condi- tioned surfaces. Embp is also a multifunctional cell sur- face protein that mediates attachment to host extracellu- lar matrix, biofilm accumulation and escape from pha- gocytosis, promoting biomaterial-associated infections [32]. Although the Fn-binding proteins on S. epidermidis RP62A cell surface were not identified in this study, such proteins are expected to be present on cell surface, as evidenced by the larger binding strengths and increased binding events measured on cell surfaces by Fn-probe (Figure 5(a)). The binding between Fn and cell surface can be considered as a ligand/receptor interaction. Bus- tanji et al. measured the energy landscape of this bind- ing/unbinding process through dynamic force spectros- copy under different loading rates, and revealed the mo- lecular mechanism of Fn in bacterial adhesion [51]. The orientation of Fn influences bacterial adhesion. Fn is a dimer of two similar polypeptides linked by disulfide bonds at the carboxyl terminus, possessing several func- tional domains that bind to a variety of extracellular molecules such as heparin and collagen [52]. There are two particularly important relevant binding sites for S. epidermidis, which are located at the N-terminus and the C-terminus of Fn [39]. The immuno-AFM measurements show that C-terminus is much more available than N-terminus of Fn molecules when adsorbed from pure Fn solution. The presence of albumin appears to influence the orientation of Fn, with more N-terminus available although the total amount of Fn adsorbed appears low. Bacteria in blood can interact with platelets. This in- teraction appears mediated by plasma proteins. Albumin appears to inhibit the bacteria-platelet interaction and activation/aggregation of platelets, while Fg and Fn promote the interactions of bacteria and platelets along with platelet activation, leading to bacteria/platelet ag- gregation (Figure 8). Interactions between bacteria and platelets are characterized as the binding of bacteria to platelets either directly through a bacterial surface pro- tein or indirectly by a plasma bridging molecule that links bacteria and platelet receptors [53]. S. epidermidis induced platelet activation and aggregation of bacteria and platelets in the absence of plasma proteins, showing the direct mechanism mostly involved (Figure 8(b)), while more bacteria-platelet aggregates were observed in the presence of Fg and Fn, suggesting that both mecha- nisms may be involved in interaction of bacteria and platelets (Figures 8(d) and (e)). More aggregates of bac- teria-platelet were measured on PUU surface when Fn or Fg were added in bulk solution compared to the case of proteins only pre-adsorbed on surface (Figure 7). This suggests that Fg or Fn may serve as linker in interaction Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 496 of bacteria and platelets. Fn leads to more bacteria adhering to platelets and forming more aggregates than Fg does. This may be due to the different functions of MSCRAMM involved in bacteria-platelet interactions. Different MSCRAMM on S. aureus cell surface have been identified including clump- ing factors (Clf) and Fn binding proteins (FnBP), and they were shown to promote bacterial adhesion to and activation of platelets [11,43]. Both Clf and FnBP bind Fg, allowing an interaction with platelet GPIIb/IIIa, leading to platelet adhesion. However, less information on the interactions of platelets and S. epidermidis is available. Recently Brennan et al. reported that Fg-bind- ing serine-aspartate repeat protein G (SdrG) from S. epi- dermidis supports adhesion to platelets and aggregation through both a direct interaction with platelet integrin receptor IIb3, and an indirect mechanism by a bridge of Fg [33], however, the roles of other MSCRAMM, e.g., the giant Fn-binding protein (embp) from S. epidermidis cell surfaces, have been identified in bacterial adhesion and biofilm formation, but without report of interactions of bacteria-platelet. 5. Conclusion Bacterial adhesion to polyurethane biomaterial surfaces as well as interactions with platelets is complex and can be mediated by plasma proteins. This study demonstrated the roles of plasma proteins (albumin, fibrinogen, and fibronectin) in the adhesion of bacteria, S. epidermidis RP62A, on polyurethane biomaterial surfaces. Results show that Fn leads to increased bacterial adhesion, with the order of effectiveness being Fn Fg > albumin. A correlation between molecular scale Fn adsorption and macroscale bacterial adhesion was observed, with in- creased numbers of Fn-receptor recognition events meas- ured on cell surfaces as compared to Fg-receptor recog- nition events, suggesting Fn is an important protein in bacterial adhesion. Interactions between bacteria and platelets induced platelet activation and bacteria-platelet aggregation. Albumin inhibited bacteria-platelet interact- tions and platelet activation, while both Fg and Fn appear to serve as a linker, promote the adhesion of bacteria to platelets and platelet activation, resulting in bacteria- platelet aggregation. 6. Acknowledgements Financial support was from a Department of Surgery Feasibility Grant at the Penn State College of Medicine. REFERENCES [1] J. W. Costerton, P. S. Stewart and E. P. Greenberg, “Bac- terial Biofilms: A Common Cause of Persistent Infec- tions,” Science, Vol. 284, No. 5418, 1999, pp. 1318-1322. doi:10.1126/science.284.5418.1318 [2] J. D. Bryers, “Medical Biofilms,” Biotechnology and Bioengineering, Vol. 100, No. 1, 2008, pp. 1-18. doi:10.1002/bit.21838 [3] F. D. Lowy, “Antimicrobial Resistance: The Example of Staphylococcus aureus,” The Journal of Clinical Investi- gation, Vol. 111, No. 9, 2003, pp. 1265-1273. [4] A. R. Costa, M. Henriques, R. Oliveira and J. Azeredo “The Role of Polysaccharide Intercellular Adhesin (PIA) in Staphylococcus epidermidis Adhesion to Host Tissues and Subsequent Antibiotic Tolerance,” European Journal of Clinical Microbiology & Infectious Diseases. Vol. 28, No. 6, 2009, pp. 623-629. doi:10.1007/s10096-008-0684-2 [5] R. Gutiérrez-González, G. R. Boto and Á. Pérez-Zamar- rón, “Cerebrospinal Fluid Diversion Devices and Infec- tion: A Comprehensive Review,” European Journal of Clinical Microbiology & Infectious Diseases, Vol. 31, No. 6, 2012, pp. 889-897. doi:10.1007/s10096-011-1420-x [6] B. Johnson, I. Starks, G. Bancroft and P. J. Roberts, “The Effect of Care Bundle Development on Surgical Site In- fection after Hemiarthroplasty: An 8-Year Review,” Jour- nal of Trauma and Acute Care Surgery, Vol. 72, No. 5, 2012, pp. 1375-1379. [7] R. F. Padera, Infection in Ventricular Assist Devices: The Role of Biofilm,” Cardiovascular Pathology, Vol. 15, No. 5, 2006, pp. 264-270. doi:10.1016/j.carpath.2006.04.008 [8] P. D. Fey and M. E. Olson, “Current Concepts in Biofilm Formation of Staphylococcus epidermidis,” Future Mi- crobiology, Vol. 5, No. 6, 2010, pp. 917-933. doi:10.2217/fmb.10.56 [9] S. W. Kerrigan, N. Clarke, A. Loughman, G. Meade, T. J. Foster and D. Cox, “Molecular Basis for Staphylococcus aureus-Mediated Platelet Aggregate Formation under Ar- terial Shear in Vitro,” Arteriosclerosis Thrombosis and Vascular Biology, Vol. 28, No. 2, 2008, pp. 335-340. doi:10.1161/ATVBAHA.107.152058 [10] L. Obrien, S. W. Kerrigan, G. Kaw, M. Hogan, J. Penades, D. Litt, et al., “Multiple Mechanisms for the Activation of Human Platelet Aggregation by Staphylococcus aureus: Roles for the Clumping Factors ClfA and ClfB, the Ser- ine-Aspartate Repeat Protein Sdre and Protein A,” Mo- lecular Microbiology, Vol. 44, No. 4, 2002, pp. 1033-1044. doi:10.1046/j.1365-2958.2002.02935.x [11] J. R. Fitzgerald, A. Loughman, F. Keane, M. Brennan, M. Knobel, J. Higgins, et al., “Fibronectin-Binding Proteins of Staphylococcus aureus Mediate Activation of Human Platelets via Fibrinogen and Fibronectin Bridges to In- tegrin GPIIb/IIIa and IgG Binding to the FcyRlla Recep- tor,” Molecular Microbiology, Vol. 59, No. 1, 2006, pp. 212-230. doi:10.1111/j.1365-2958.2005.04922.x [12] D. Cox, S. W. Kerrigan and S. P. Watson, “Platelets and the Innate Immune System: Mechanisms of Bacterial- Induced Platelet Activation,” Journal of Thrombosis and Haemostasis, Vol. 9, No. 6, 2011, pp. 1097-1107. Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces 497 doi:10.1111/j.1538-7836.2011.04264.x [13] M. A. Johnson and J. M. Ross, “Staphylococcal Presence Alters Thrombus Formation under Physiological Shear Conditions in Whole Blood Studies,” Annals of Biomedi- cal Engineering, Vol. 36, No. 3, 2008, pp. 349-355. doi:10.1007/s10439-007-9434-3 [14] O. Shannon, E. Hertzen, A. Norrby-Teglund, M. Mörge- lin1, U. Sjöbring and L. Björck, “Severe Streptococcal Infection Is Associated with M Protein-Induced Platelet Activation and Thrombus Formation,” Molecular Micro- biology, Vol. 65, No. 5, 2007, pp. 1147-1157. doi:10.1111/j.1365-2958.2007.05841.x [15] E. E. MacKintosh, J. D. Patel, R. E. Marchant and J. M. Anderson, “Effects of Biomaterial Surface Chemistry on the Adhesion and Biofilm Formation of Staphylococcus epidermidis in Vitro,” Journal of Biomedical Materials Research Part A, Vol. 78A, No. 4, 2006, pp. 836-842. doi:10.1002/jbm.a.30905 [16] K. Vacheethasanee, J. S. Temenoff, J. M. Higashi, A. Gary, J. M. Anderson, R. Bayston, et al., “Bacterial Sur- face Properties of Clinically Isolated Staphylococcus epi- dermidis Strains Determine Adhesion on Polyethylene,” Journal of Biomedical Materials Research, Vol. 42, No. 3, 1998, pp. 425-432. doi:10.1002/(SICI)1097-4636(19981205)42:3<425::AID- JBM12>3.0.CO;2-F [17] V. A. Tegoulia and S. L. Cooper, “Staphylococcus aureus Adhesion to Self-Assembled Monolayers: Effect of Sur- face Chemistry and Fibrinogen Presence,” Colloids and Surfaces B: Biointerfaces, Vol. 24, No. 3-4, 2002, pp. 217-228. doi:10.1016/S0927-7765(01)00240-5 [18] R. Ardehali, L. Shi, J. Janatova, S. F. Mohammad and G. L. Burns, “The Inhibitory Activity of Serum to Prevent Bacterial Adhesion Is Mainly Due to Apo-Transferrin,” Journal of Biomedical Materials Research Part A, Vol. 66A, No. 1, 2003, pp. 21-28. doi:10.1002/jbm.a.10493 [19] J. D. Patel, M. Ebert, R. Ward and J. M. Anderson, “S. epidermidis Biofilm Formation: Effects of Biomaterial Surface Chemistry and Serum Proteins,” Journal of Bio- medical Materials Research Part A, Vol. 80A, No. 3, 2007, pp. 742-751. doi:10.1002/jbm.a.31103 [20] O. Hartford, L. O’Brien, K. Schofield, J. Wells and T. J. Foster, “The Fbe (SdrG) Protein of Staphylococcus epi- dermidis HB Promotes Bacterial Adherence to Fibrino- gen,” Microbiolo gy-SGM, Vol. 147, No. 9, 2001, pp. 2545- 2552. [21] M. Hussain, C. Heilmann, G. Peters and M. Herrmann, “Teichoic Acid Enhances Adhesion of Staphylococcus epidermidis to Immobilized Fibronectin,” Microbial Pa- thogenesis, Vol. 31, No. 6, 2001, pp. 261-270. doi:10.1006/mpat.2001.0469 [22] A. C. Schroeder, J. M. Schmidbauer, A. Sobke, B. Seitz, K. W. Ruprecht and M. Herrmann, “Influence of Fi- bronectin on the Adherence of Staphylococcus epider- midis to Coated and Uncoated Intralocular Lenses,” Jour- nal of Cataract and Refractive Surgery, Vol. 34, No. 3, 2008, pp. 497-504. doi:10.1016/j.jcrs.2007.10.042 [23] W. M. Dunne and E. M. Burd, “Fibronectin and Prote- olytic Fragments of Fibronectin Interfere with the Adhe- sion of Staphylococcus epidermidis to Plastic,” Journal of Applied Microbiology, Vol. 74, No. 4, 1993, pp. 411-416. doi:10.1111/j.1365-2672.1993.tb05147.x [24] S. Galliani, M. Viot, A. Cremieux and P. Vanderauwera, “Early Adhesion of Bacteremic Strains of Staphylococcus Epidermidis to Polystyrene-Influence of Hydrophobicity, Slime Production, Plasma Albumin, Fibrinogen, and Fi- bronectin,” Journal of Laboratory and Clinical Medicine, Vol. 123, No. 5, 1994, pp. 685-692. [25] J. C. Linnes, K. Mikhova and J. D. Bryers, “Adhesion of Staphylococcus epidermidis to Biomaterials Is Inhibited by Fibronectin and Albumin,” Journal of Biomedical Ma- terials Research Part A, Vol. 100A, No. 8, 2012, pp. 1990- 1997. doi:10.1002/jbm.a.34036 [26] E. R. Wann, S. Gurusiddappa and M. Hook, “The Fibro- nectin-Binding MSCRAMM FnbpA of Staphylococcus aureus is a Bifunctional Protein That Also Binds to Fi- brinogen,” Journal of Biological Chemistry, Vol. 275, No. 18, 2000, pp. 13863-13871. doi:10.1074/jbc.275.18.13863 [27] O. M. Hartford, E. R. Wann, M. Hook and T. J. Foster, “Identification of Residues in the Staphylococcu s aureus Fi- brinogen-Binding MSCRAMM Clumping Factor A (ClfA) That Are Important for Ligand Binding,” Journal of Bio- logical Chemistry, Vol. 276, No. 4, 2001, pp. 2466-2473. doi:10.1074/jbc.M007979200 [28] R. J. Williams, B. Henderson, L. J. Sharp and S. P. Nair, “Identification of a Fibronectin-Binding Protein from Staphylococcus epidermidis,” Infe ction and Immunity, Vol. 70, No. 12, 2002, pp. 6805-6810. doi:10.1128/IAI.70.12.6805-6810.2002 [29] C. Arrecubieta, F. A. Toba, M. Von Bayern, H. Akashi, M. C. Deng, Y. Naka, et al., “SdrF, a Staphylococcus epi- dermidis Surface Protein, Contributes to the Initiation of Ventricular Assist Device Driveline-Related Infections,” Plos Pathogens, Vol. 5, No. 5, 2009, p. e1000411. [30] S. L. Davis, S. Gurusiddappa, K. W. McCrea, S. Perkins and M. Hook, “SdrG, a Fibrinogen-Binding Bacterial Adhesin of the Microbial Surface Components Recog- nizing Adhesive Matrix Molecules Subfamily from Sta- phylococcus epidermidis, Targets the Thrombin Cleavage Site in the B Beta Chain,” Journal of Biological Chemis- try, Vol. 276, No. 30, 2001, pp. 27799-27805. doi:10.1074/jbc.M103873200 [31] C. Arrecubieta, M. H. Lee, A. Macey, T. J. Foster and F. D. Lowy, “SdrF, a Staphylococcus epidermidis Surface Protein, Binds Type I Collagen,” Journal of Biological Chemistry, Vol. 282, No. 26, 2007, pp. 18767-18776. doi:10.1074/jbc.M610940200 [32] M. Christner, G. C. Franke, N. N. Schommer, U. Wendt, K. Wegert, P. Pehle, et al., “The Giant Extracellular Ma- trix-Binding Protein of Staphylococcus epidermidis Me- diates Biofilm Accumulation and Attachment to Fi- bronectin,” Molecular Microbiology, Vol. 75, No. 1, 2010, pp. 187-207. doi:10.1111/j.1365-2958.2009.06981.x [33] M. P. Brennan, A. Loughman, M. Devocelle, S. Arasu, A. J. Chubb, T. J. Foster, et al., “Elucidating the Role of Copyright © 2012 SciRes. JBNB  Effects of Plasma Proteins on Staphylococcus epidermidis RP62A Adhesion and Interaction with Platelets on Polyurethane Biomaterial Surfaces Copyright © 2012 SciRes. JBNB 498 Staphylococcus epidermidis Serine-Aspartate Repeat Pro- tein G in Platelet Activation,” Journal of Thrombosis and Haemostasis, Vol. 7, No. 8, 2009, pp. 1364-1372. doi:10.1111/j.1538-7836.2009.03495.x [34] M. Katsikogianni and Y. F. Missirlis, “Concise Review of Mechanisms of Bacterial Adhesion to Biomaterials and of Techniques Used in Estimating Bacteria-Material Interac- tions,” European Cells & Materials Journal, Vol. 8, 2004, pp. 37-57. [35] A. Mendez-Vilas, A. M. Gallardo-Moreno and M. L. Gonzalez-Martin, “Nano-Mechanical Exploration of the Surface and Sub-Surface of Hydrated Cells of Staphylo- coccus epidermidis,” Antonie Van Leeuwenhoek Interna- tional Journal of General and Molecular Microbiology, Vol. 89, No. 3, 2006, pp. 373-386. doi:10.1007/s10482-005-9041-y [36] A. Méndez-Vilasa, A. M. Gallardo-Morenoa, M. L. Gon- zález-Martı́na, R. Calzado-Monteroa, M. J. Nuevoa, J. M. Bruquea, et al., “Surface Characterisation of Two Strains of Staphylococcus epidermidis with Different Slime- Production by AFM,” Applied Surface Science, Vol. 238, No. 1-4, 2004, pp. 18-23. doi:10.1016/j.apsusc.2004.05.183 [37] Y. Liu, J. Strauss and T. A. Camesano, “Adhesion Forces between Staphylococcus epidermidis and Surfaces Bear- ing Self-Assembled Monolayers in the Presence of Model Proteins,” Biomaterials, Vol. 29, No. 33, 2008, pp. 4374- 4382. doi:10.1016/j.biomaterials.2008.07.044 [38] N. P. Boks, H. J. Busscher, H. C. van der Mei and W. Norde, “Bond-Strengthening in Staphylococcal Adhesion to Hydrophilic and Hydrophobic Surfaces Using Atomic Force Microscopy,” Langmuir, Vol. 24, No. 22, 2008, pp. 12990-12994. doi:10.1021/la801824c [39] R. A. Jarvis and J. D. Bryers, “Effects of Controlled Fi- bronectin Surface Orientation on Subsequent Staphylo- coccus epidermidis Adhesion,” Journal of Biomedical Materials Resear ch Part A, Vol. 75A, No. 1, 2005, pp. 41- 55. doi:10.1002/jbm.a.30404 [40] P. B. Chowdhury and P. F. Luckham, “Probing Recogni- tion Process between an Antibody and an Antigen Using Atomic Force Microscopy,” Colloids and Surfaces A: Physicochemical and Engineering Aspects, Vol. 143, No. 1, 1998, pp. 53-57. doi:10.1016/S0927-7757(98)00408-7 [41] A. Agnihotri and C. A. Siedlecki, “Adhesion Mode Ato- mic Force Microscopy Study of Dual Component Protein Films,” Ultramicroscopy, Vol. 102, No. 4, 2005, pp. 257- 268. doi:10.1016/j.ultramic.2004.10.006 [42] P. Soman, Z. Rice and C. A. Siedlecki, “Measuring the Time-Dependent Functional Activity of Adsorbed Fi- brinogen by Atomic Force Microscopy,” Langmuir, Vol. 24, No. 16, 2008, pp. 8801-8806. doi:10.1021/la801227e [43] D. Pavithra and M. Doble, “Biofilm Formation, Bacterial Adhesion and Host Response on Polymeric Implants Is- sues and Prevention, Biomedical Materials, Vol. 3, No. 3, 2008, pp. 1-13. [44] A. Chiumiento, S. Lamponi and R. Barbucci, “Role of Fibrinogen Conformation in Platelet Activation,” Biomac- romolecules, Vol. 8, No. 2, 2007, pp. 523-531. doi:10.1021/bm060664m [45] I. Lee and R. E. Marchant, “Molecular Interaction Studies of Hemostasis: Fibrinogen Ligand-Human Platelet Re- ceptor Interactions,” Ultramicroscopy, Vol. 97, No. 1-4, 2003, pp. 341-352. doi:10.1016/S0304-3991(03)00059-7 [46] A. J. Garcia, M. D. Vega and D. Boettiger, “Modulation of Cell Proliferation and Differentiation through Sub- strate-Dependent Changes in Fibronectin Conformation,” Molecular Biology of the Cell, Vol. 10, No. 3, 1999, pp. 785-798. [47] U. Schwarz-Linek, M. Hook and J. R. Potts, “The Mo- lecular Basis of Fibronectin-Mediated Bacterial Adher- ence to Host Cells,” Molecular Microbiology, Vol. 52, No. 3, 2004, pp. 631-641. doi:10.1111/j.1365-2958.2004.04027.x [48] S. Kerdudou, M. W. Laschke, B. Sinha, K. T. Preissner, M. D. Menger and M. Herrmann, “Fibronectin Binding Proteins Contribute to the Adherence of Staphylococcus Aureus to Intact Endothelium in Vivo,” Thrombosis and Haemostasis, Vol. 96, No. 2, 2006, pp. 183-189. [49] K. H. Simpson, M. G. Bowden, M. Hook and B. Anvari, “Measurement of Adhesive Forces between S. epider- midis and Fibronectin-Coated Surfaces Using Optical Tweezers,” Lasers in Surgery and Medicine, Vol. 31, No. 1, 2002, pp. 45-52. doi:10.1002/lsm.10070 [50] C. P. Xu, N. P. Boks, J. de Vries, H. J. Kaper, W. Norde, H. J. Busscher, et al., “Staphylococcus aur eus-Fibronectin Interactions with and without Fibronectin-Binding Pro- teins and Their Role in Adhesion and Desorption,” Ap- plied and Environmental Microbiology, Vol. 74, No. 24, 2008, pp. 7522-7528. doi:10.1128/AEM.00948-08 [51] Y. Bustanji, C. R. Arciola, M. Conti, E. Mandello, L. Montanaro and B. Samori, “Dynamics of the Interaction between a Fibronectin Molecule and a Living Bacterium under Mechanical Force,” Proceedings of the National Academy of Sciences of the United States of America, Vol. 100, No. 23, 2003, pp. 13292-13297. doi:10.1073/pnas.1735343100 [52] B. Henderson, S. Nair, J. Pallas and M. A. Williams, “Fi- bronectin: A Multidomain Host Adhesin Targeted by Bacterial Fibronectin-Binding Proteins,” FEMS Microbi- ology Reviews, Vol. 35, No. 1, 2011, pp. 147-200. doi:10.1111/j.1574-6976.2010.00243.x [53] J. R. Fitzgerald, T. J. Foster and D. Cox, “The Interaction of Bacterial Pathogens with Platelets,” Nature Reviews Microbiology, Vol. 4, No. 6, 2006, pp. 445-457. doi:10.1038/nrmicro1425
|