American Journal of Plant Sciences
Vol.5 No.3(2014), Article ID:43053,7 pages DOI:10.4236/ajps.2014.53054
Isolation and Characterization of Novel Microsatellite Markers in Walnut (Juglans regia L.)
1Department of Agronomy and Plant Breeding, Faculty of Agriculture, University of Zabol, Zabol, Iran; 2Agricultural Biotechnology Research Institute of Iran (ABRII), Karaj, Iran; 3Faculty of Agriculture, University of Tehran, Karaj, Iran.
Email: *fatemeh.najafi61@gmail.com
Copyright © 2014 Fatemeh Najafi et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. In accordance of the Creative Commons Attribution License all Copyrights © 2014 are reserved for SCIRP and the owner of the intellectual property Fatemeh Najafi et al. All Copyright © 2014 are guarded by law and by SCIRP as a guardian.
Received November 25th, 2013; revised January 22nd, 2014; accepted February 14th, 2014
KEYWORDS
Isolation; Microsatellite; Juglans regia L.; Polymorphism; SSR Markers
ABSTRACT
Thirteen microsatellite markers were isolated and characterized from Juglans regia L. Successful polymorphism of the 13 primer pairs was observed in various genotypes of J. regia. The number of polymorphic alleles ranged from 2 to 4 (with an average of 4.35). The polymorphic information content values ranged from 0.47 to 0.88 (with an average of 0.69). TC/AG and GAA/CTT class of repeats were the most abundant di-nucleotide and tri-nucleotide repeats, respectively.
1. Introduction
Walnut (Juglans regia) cultivars currently are grown in Iran widely, from Astara to Golidagh with arid climate in north forests. The major producers of this species are USA, China, Turkey and Iran, respectively (www.fao.org). Walnut is probably originated from central Asia Mountains and the main origin of walnut is in minor Asia. Juglans genus includes about 21 species [1]. Walnut (2n = 32) is a monoecious and preferably allogamous species which is mainly cultivated for fruit production.
Cultivation of walnut in Iran has a long history as early as 370 B.C. According to the FAO’s reports, Iran is the third country for producing walnut only after China and America (10% of cultivation area in Iran is allotted to walnut). SSR markers are PCR-based markers.
Microsatellites or Simple Sequence Repeats (SSR) markers are ideal for genetic diversity analysis because of their abundance, high polymorphism content, co-dominance, but there is an urgent need to increase this repertoire for effective deployment of these highly useful markers in conservation genetic studies on pomegranate. But there had been only few studies based on molecular markers to understand the population dynamics of this valuable plant, mainly due to lack of microsatellite markers. Thus, there is an urgent need to increase this repertoire for effective deployment of these highly useful markers in conservation genetic studies on Juglans species.
A number of methods have been used to identify microsatellite markers including a Fast Isolation by AFLP of Sequences Containing repeats (FIASCO) approach [2]. FIASCO is a fast and effective technique to develop SSR markers compared to methods previously used and a major limitation of SSRs in the time and cost required to isolated each locus.
Although, 41 primers have been reported for walnut cultivar in the world, 13 SSRs have been identified in our research. Among the Iranian walnut endemics cultivar Z30 and cultivar Z60 are the most polymorphics. Collection and selection of walnut genotypes was started by Atefi (1983 to 1985). Despite wide range of morphologyical variability in walnut germplasm, only few studies have been performed for molecular markers to determine the population dynamics of this economically important fruit tree [3-7].
Although using of FIASCO in SSR developing is relatively ordinary but this technique was used rarely in some species including fungal pathogens, insects and recently in plants [8-10].
Some studies have assessed walnut plants using different DNA markers including SSR [11]. We have reported the characterization and isolation of gnome-wide SSR marker in walnut. Since Iranian walnuts have been poorly characterized for their diversity, in this study and as a confirmation for our previous report [12], some morphological and SSR markers were applied to survey genetic relationships among walnuts sampled from different regions of Iran along with some foreign cultivars.
In this study, large numbers of samples were analyzed to probe new Short Sequence Repeat (SSR) in walnut. Moreover, SSR fingerprinting of newly found SSRs were compared with other fingerprinting methods such as AFLP, RAPD and ISSR. Microsatellite shows numerous advantages since they are locus specific, co-dominant, highly reproducible and usually highly polymorphic [13].
Margins, column widths, line spacing, and type styles are built-in; examples of the type styles are provided throughout this document and are identified in italic type, within parentheses, following the example. Some components, such as multi-leveled equations, graphics, and tables are not prescribed, although the various table text styles are provided. The formatter will need to create these components, incorporating the applicable criteria that follow.
2. Materials and Method
The 36 genotypes which were used as our material in this study were obtained from the walnut collection maintained in Karaj, Iran, and are part of the walnut breeding program developed at this institution (Table 1). The 36 genotypes which were used as our material in this study were obtained from the walnut collection maintained in Karaj, Iran, and are part of the walnut breeding program developed at this institution.
Totally 35 superior genotypes including 16 from Fars province (10 from Neyriz and 6 from Bavanat), 7 from Chahar-Mahal province (Shahrekord) and 11 from Tehran province (Karaj), one from Azarbayejane-sharghi province (Mianeh) as well as four foreign cultivars (“Vina”, “Serr”, “Franquette” and “Lara”) were used as our material in this study (Table 1).
2.1. DNA Extract
36 individuals sample of J. regia from different regions of Iran were collected and subsequently the sample’s genomic DNA was extracted using a DNeasy Plant Mini kit (Qiagen, Germany) (Figure 1). 36 individuals sample of J. regia from different regions of Iran were collected and subsequently the sample’s genomic DNA was extracted using a DNeasy Plant Mini kit (Qiagen, Germany) (Figure 1).

Table 1. Genotype code, cultivar/genotype name of walnut and their collection sites.
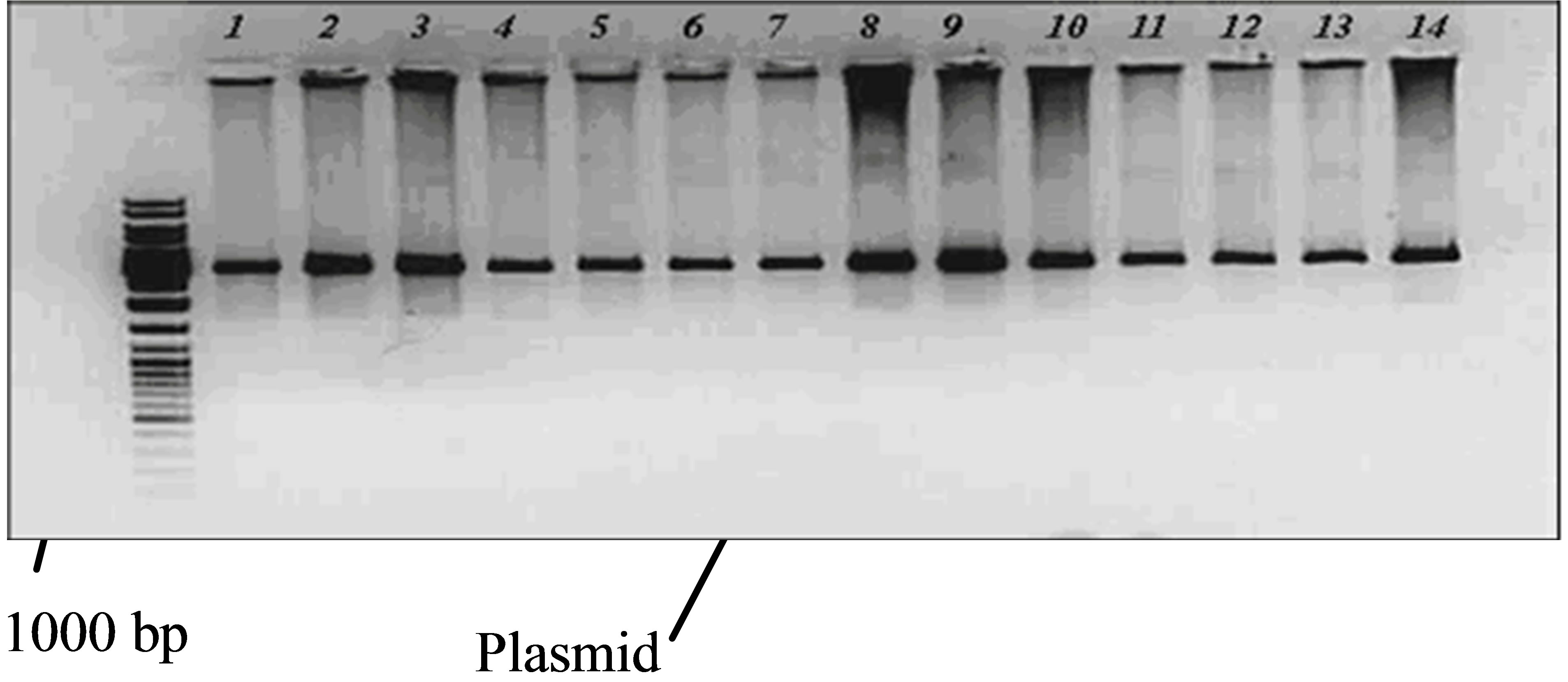
Figure 1. Plasmid extraction. Plasmids were separated by electrophoresis using 2% agarose stained with ethidium bromide. L = commercial 100-bp DNA size standard.
2.2. Isolation of SSR Markers
A SSR-enriched library was constructed from the DNA of an individual from a Walnut cultivar K72 (110296) by using the modified protocol of fast isolation by AFLP of sequences containing repeats (FIASCO) [2]. A total of 250 ng genomic DNAs were first digested with the restriction enzyme MseI (Fermentas, Germany) and then the fragments were ligated to MseI adapters (MWG Biotech). The digestion-ligation fragments were pre-amplified by 20 cycles in a Veriti thermal cycler (Applied biosystems).
The amplified DNA fragments were later hybridized with biotinylated probes (GC)17, (AC)17, (CT)17, (AT)17, (GT)17, (ATT)10, (CTT)10, (AG)17, (ACC)10, (ACT)10, (GTT)10 and (GGT)10 (MWG Biotech) combined with 100 µl of hybridization buffers 4.2 X SSC and 0.07% SDS. The hybridized DNAs were mixed with streptavidin-coated beads (Poromega) and TEN 100 (10 mM Tris-HCL, 1 mM EDTA, 100 mM NaCl, PH = 7.5) then incubated at room temperature for 30 minutes. The biotinylated bead-probe-DNA complex was separated from the hybridization buffer by a BioMag magnetic field (Qiagen, Germany). The collected DNA fragments were amplified for 30 cycles using the MseI primer in the thermal cycler then The PCR fragments ligated into pGEM-T Easy vector (Promega, Germany) and transformed into competent Escherichia coli DH5α following the manufacturer’s instructions. The positive transformants were handpicked by blue/white screening. Sixty one clones with inserts between 400-1000 bp were purified using a plasmid extraction kit (Core-Bio, Sought Korea) and sequenced (Macrogen Sequencing Service, Korea).
Microsatellites were detected and primers were designed using SSR Locator software [14]. PCR amplifications were carried out in a Veriti thermal cycler (Applied Biosystems) in a total volume of 15 μl, which contained 20 ng DNA, 1X PCR buffer, 2 mM MgCl2, 10 mM dNTPs, 10 pmol each primer, and 0.5 Unit Taq DNA polymerase (Fermentas, Germany). The used PCR condition is given as follow: 4 min at 94˚C, followed by 10 touchdown cycles of 30 s at 95˚C, 40 s at 60˚C (with 1˚C decrease per cycle) and 40 s at 72˚C, and 25 cycles of 30 s at 95˚C, 30 s at 50˚C and 40 s at 72˚C, with a final extension step of 7 min at 72˚C. PCR products were denatured by adding 7.5 l formamide loading dye (95% deionized formamide, 10 mM EDTA pH: 8, 0.05% xylene cyanol, 0.05% bromophenol blue), heated for 5 min at 94˚C, then 5:l of denatured preparations were loaded on a pre-warmed (50˚C) 6% polyacrylamide sequencing gel (Bio Rad, Sequi-Gen GT). Permanent record of gels were made using gel scanner (Bio Rad, GS800 calibrated densitometer).
All J. regia individuals were tested for polymorphism. PCR products were visualized by electrophoresis on 6% denaturing polyacrylamide gels followed by silver nitrate staining (Figure 2).
2.3. Data Analysis
The polymorphism information content (PIC) was calculated as described by POPGENE 32 [2] was used to calculate the observed and expected heterozygosities and to evaluate deviation from Hardy-Weinberg equilibrium (HWE) and linkage disequilibrium between pairs of loci. All results were adjusted for multiple simultaneous comparisons using a sequential Bonferroni correction [15]. Polymorphic information content (PIC) was estimated using CERVUS v. 2.0 [1,6,13,15-20]. The number of alleles (na) and the number of effective alleles (ne) were determined. Loci information content was estimated by expected heterozygosity using formula 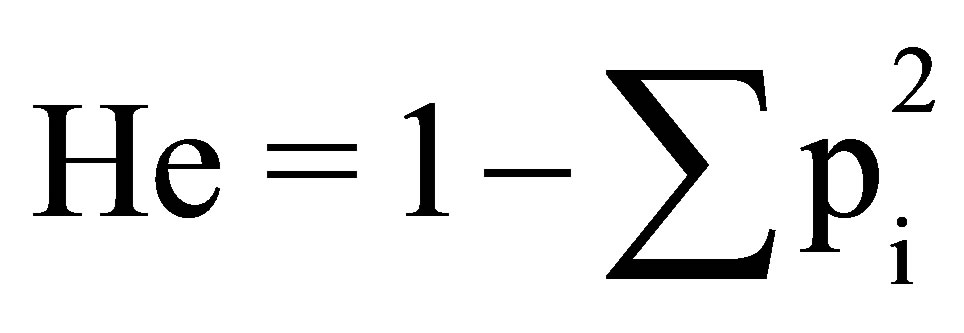 (where pi is the frequency of the ith allele). Discrimination power (DP) was evaluated according to Kloosterman et al. (1993).
(where pi is the frequency of the ith allele). Discrimination power (DP) was evaluated according to Kloosterman et al. (1993).
3. Results
In this study, polymorphic microsatellite markers for walnut were developed from an enriched partial genomic library that was constructed using the fast isolation by AFLP of sequences containing repeats (FIASCO) protocol. The efficiency of this method in this species was approximately 73%, which conformed to the expected percentage of efficiency reported [21]. To the best of our knowledge, polymorphic microsatellite markers have not been reported for J. regia in Iran. These new markers should allow studies of the population structure and genetic diversity of Walnut to be performed in the future.
Sequence analysis of 61 putative SSR-positive clones showed the presence of one or more unique SSRs in 24 (30%) clones. Out of 24 SSR-positive clones, 14 and 10 SSRs were unique and redundant, respectively. Among the di-nucleotide SSRs, the TC/AG repeat motifs were the most common (61%) followed by AC/TG repeat motifs (36%). In the tri-nucleotide SSRs, TGG/ACC repeat
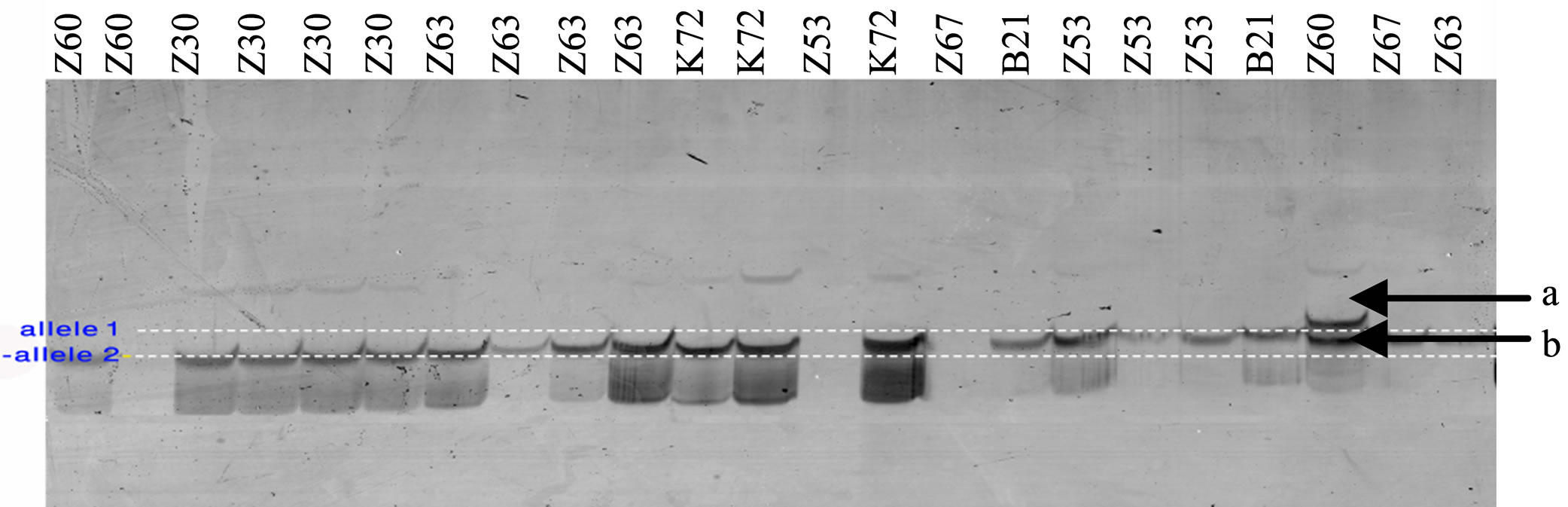

Figure 2. Preliminary genotypic result. Amplification products from J. regia accessions with SSR primers. Fragments were separated by electrophoresis using 6% polyacrylamide gel stained with silver nitrate. Lane L, denotes 100 bp ladder (Invitrogen).The letter b indicates a monomorphic band and the letter a indicates a polymorphic band.
motifs were the most common (69%) followed by GGT/ CCA (6%).
Since the most common di-nucleotide SSRs in this study were TC/AG, thus the (TC/CT) or (GA/AG) probes could be used for further SSR isolation in J. regia. The sequences of 61 putative SSR-positive clones were analyzed and thirteen primer pairs were designed based on 60 unique SSR clones.
Nineteen primer pairs (47%) yielded scorable amplicons and 17 primer pairs showed polymorphism among 36 individuals of J. regia. The polymorphic SSR markers detected 2 to 4 alleles with an average of 4.35 alleles per marker. The highest number of alleles (4) was observed in ABRII/Jr6. The polymorphism information content (PIC) value for these polymorphic markers varied 0.56 (ABRII/Jr6) (Table 2). The most informative marker with highest PIC was ABRII/Jr6. Nei’s genetic diversity (He) ranged 0.63 (ABRII/Jr6).
Among these markers are identified, a number of components will be Monomorph. Of the markers identified so far, 13 markers were used to check the fingerprints of walnut. Also, 14 primers designed by Wost et al. will be used on the population studies of Persian walnut.
The size of amplified fragments of PCR was different in each marker. Amplified fragment lengths were in very close range. Size of the largest fragments was 320 - 270 bp allele with primers 376 WP and the smallest fragment of 155 - 150 bp allele lengths were observed in 8 WP marker. Other primers, amplified pieces that range in size between 376 WP and WGA376 primers (Table 3(.
PCo was obtained by the similarity matrix of Jaccard method and Software NTSYS-PC 2.02 in dealing with molecular data, PCoA is used instead of the PCA. Twodimensional diagram can be seen in Figure below (Figure 3).
4. Discussion
The enrichment efficiency of this study was 30% and it was higher than those obtained in some other SSR-enriched libraries in Juglans [17]. The enrichment efficiency depends on factors such as restriction enzymes used for library construction and the SSR probes used for enrichment. The highest number of isolated microsatellites was di-nucleotides, mostly TC/AG which is consistent with the results obtained by Amy Ross-Davis and Erin et al. and Rodney and colleagues in Juglans which reported perfect TC/AG compound repeats as the frequent di-nucleotides. In the plant genome, (GA)n microsatellites are in high abundance compared to other dinucleotide SSRs and they are well distributed throughout the genome [6].
As observed in this study, most of the SSRs showed a considerable variation in J. regia genotypes. In addition, some of these SSR markers are currently being used in the existing linkage maps of walnut as anchor points and could further facilitate comparison and consolidation of the available maps.
In addition, some of these SSR markers are currently being used in the existing linkage maps of walnut as anchor points and could further facilitate comparison and consolidation of the available maps.
Based on the qualitative and quantitative traits, the 7 walnut genotypes were classified into main cluster (Table 4).
The first sub-group in SSRs markers included seven genotype from Karaj (“Z601”, “Z303”, “Z631”, “Z636”, “Z603”, “Z672”, “Z536”). The second subgroup included two genotypes from Karaj (“Z302”, “Z306”). The third sub-group included five genotypes from Karaj (“Z632”, “Z633”, “Z635”, “Z532”, “K723”). The forth sub-group consisted four genotypes from Karaj (“Z634”, “Z637”, “Z638”, “Z305”). The fifth subgroup included three genotypes from Karaj (“K721”, “K722”, “K725”) and one genotype (“B212”) form Bavanat. Six subgroups included four genotypes from Karaj (“Z534”, “Z602”, “Z537”, “Z671”) and one genotype (“B211”) from Bavanat. The last sub-group included five (“Z531”, “Z307”, “K724”, “K726”, “Z673”). The Z30 and K72 genotypes were nested as discernible groups within the denderograms produced in each marker system. Thus, this region can be considered as a useful and special source of walnut genetic diversity for further breeding studies (Figure 4).
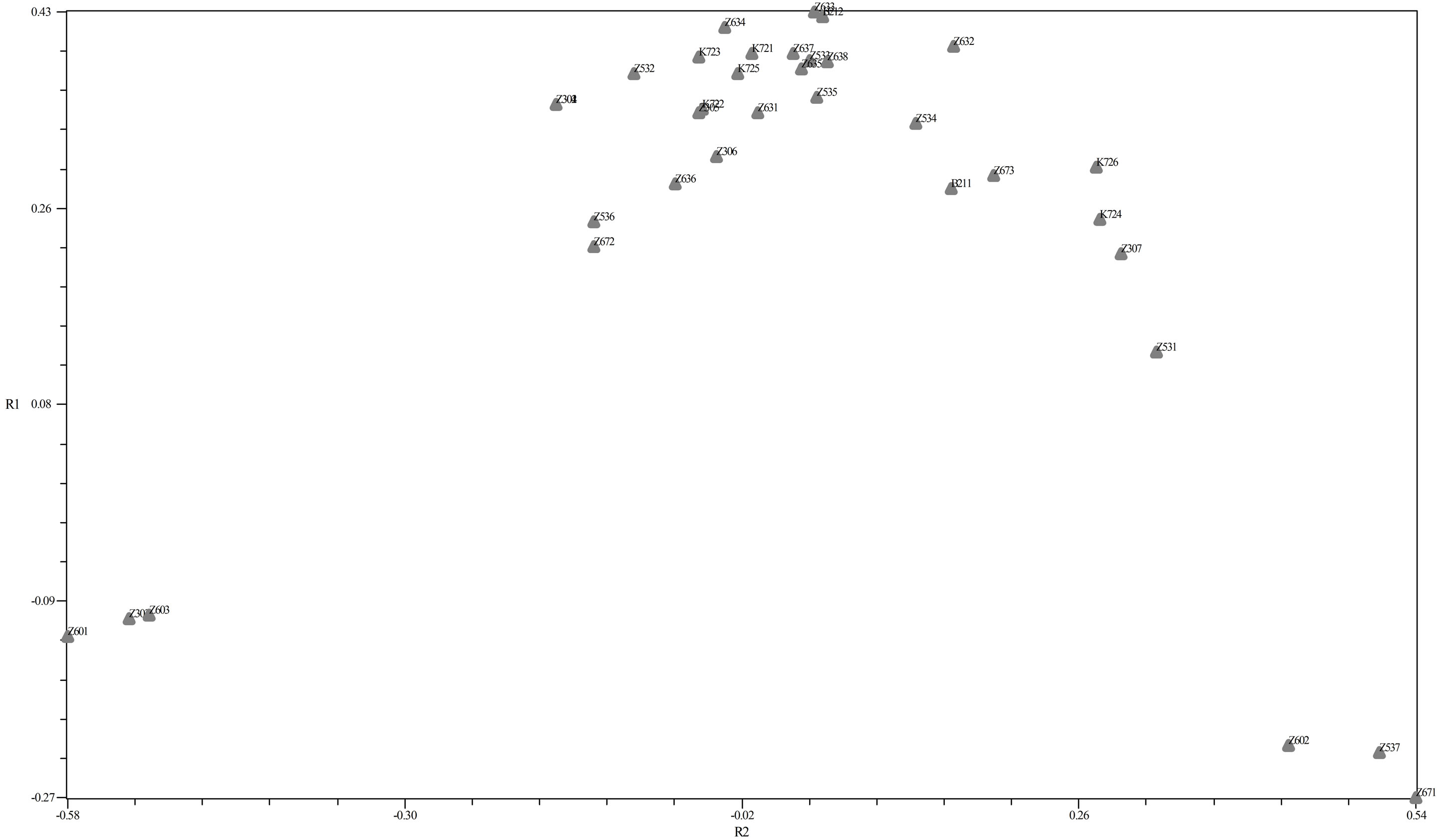
Figure 3. PCo.
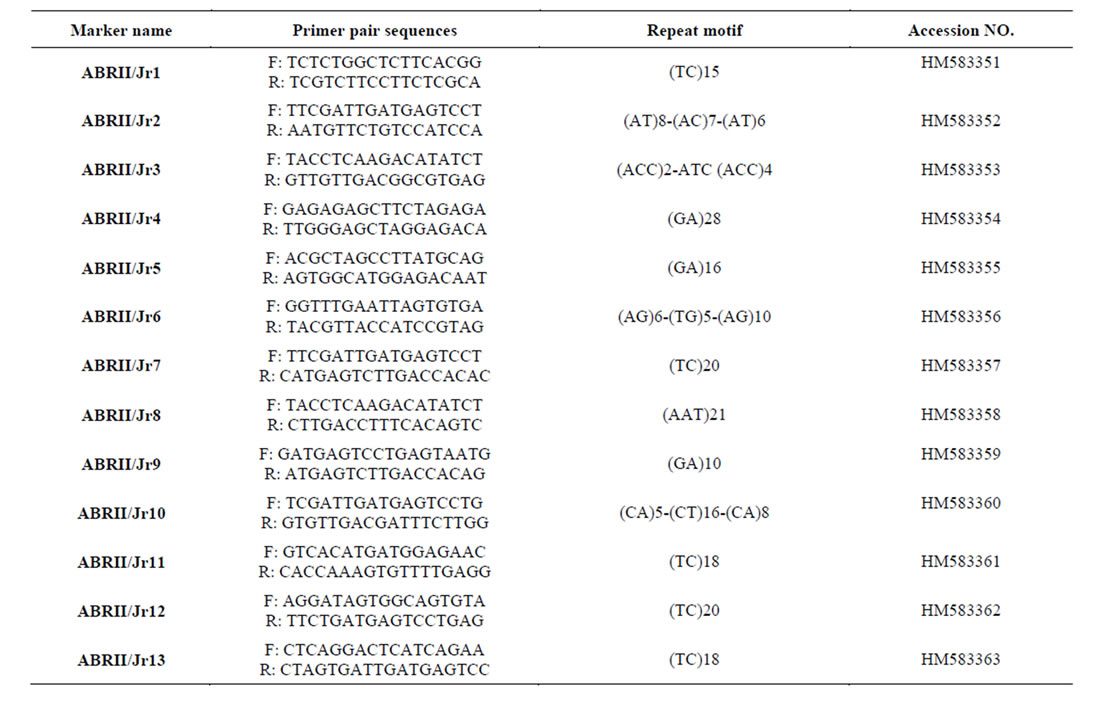
Table 2. Characteristics of 13 microsatellite markers for Juglans. Primer sequences for microsatellite loci that amplify in Juglans rejia.
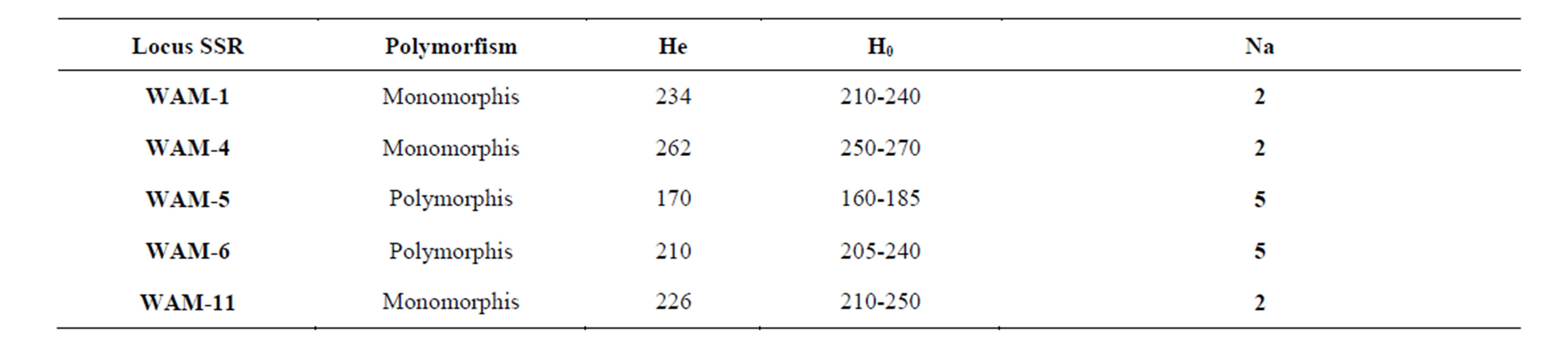
Table 3. Walnut SSRs loci used to study 35 walnut genotypes (Na: number of alleles, Ne: number of effective alleles, Ho: observed heterozygosity, He: expected heterozygosity, DP: Discrimination power).
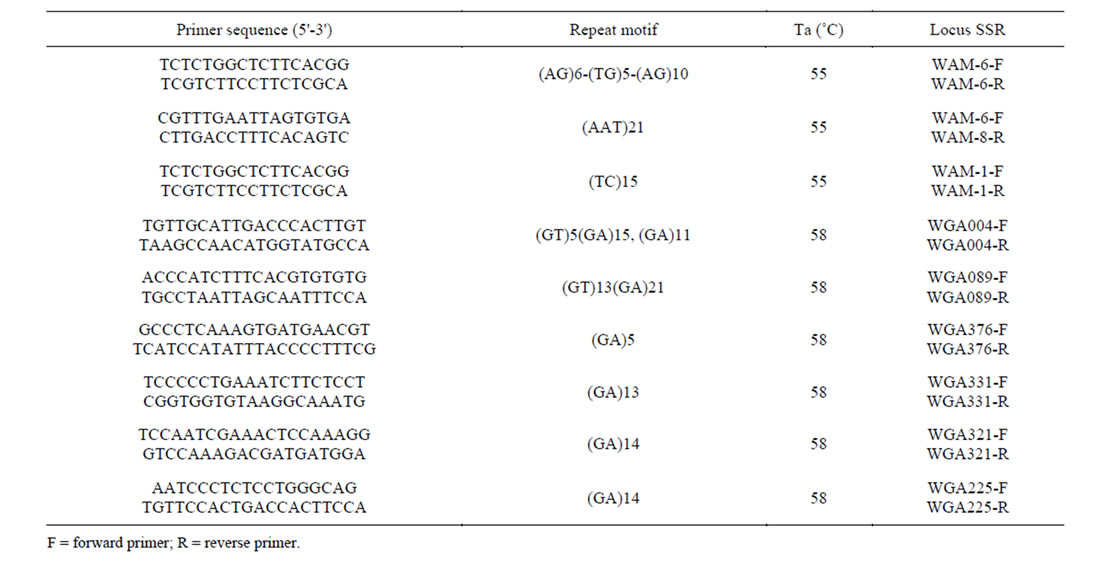
Table 4. The highest number of alleles was seen in 5, 6WP and 376WP primers with 5 polymorphic alleles. The lowest number of alleles was occured in 331WP primers with 1 polymorphic allele.
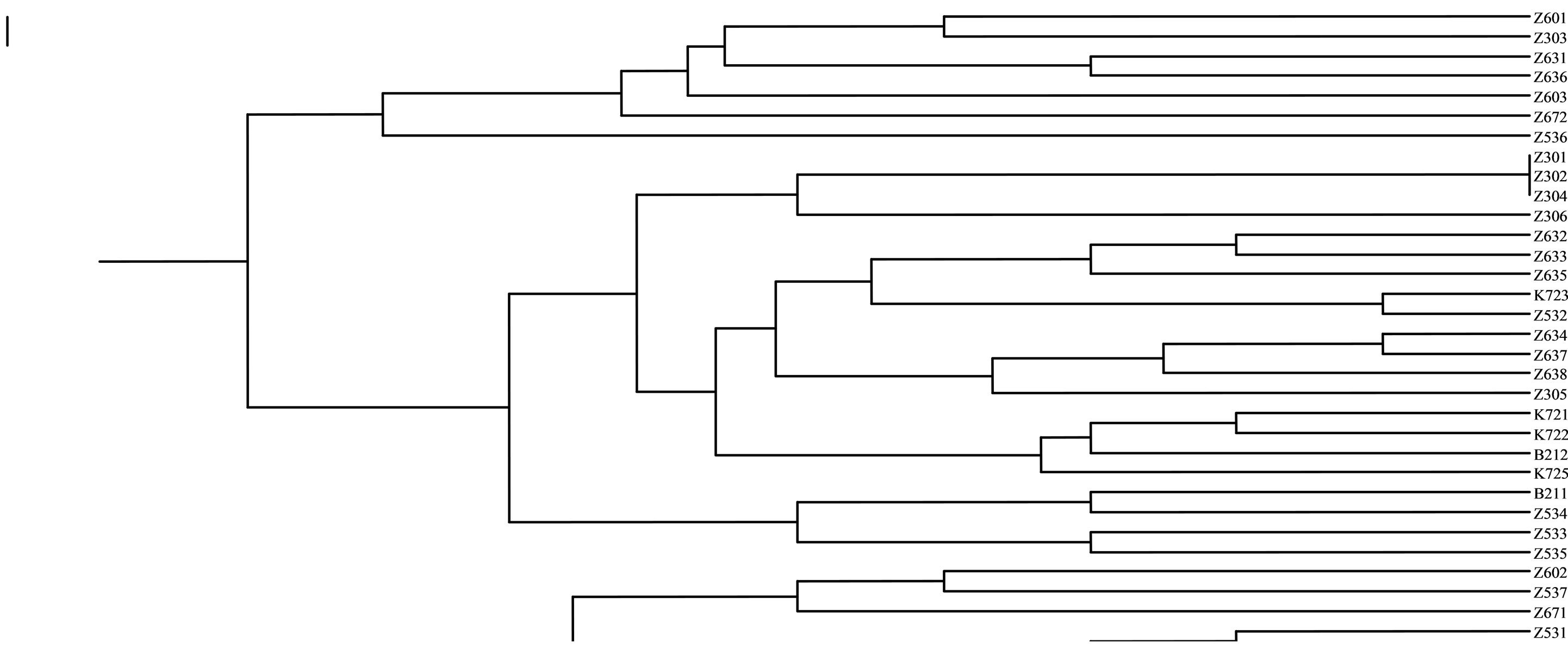
Figure 4. Preliminary genotypic result. Amplification products from J. regia accessions with SSR primers.
Acknowledgements
This research was funded by ABRII (No. 20131400- 0001850485002). Primers were obtained from MWG biotech companies.
REFERENCES
- G. McGranahan and C. Leslie, “Walnuts (Juglans),” In: J. N. Moore and J. R. Ballington, Eds., Genetic Resources of Temperate Fruit and Nut Crops, Acta Horticulturae, Vol. 290, No. 2, 1990, pp. 907-974.
- L. Zane, L. Bargelloni and T. Patarnello, “Strategies for Microsatellite Isolation: A Review,” Molecular Ecology, Vol. 11, No. 1, 2002, pp. 1-16. http://dx.doi.org/10.1046/j.0962-1083.2001.01418.x
- S. Bayazit, K. Kazan, V. Gulbitti, C. Evik, H. Ayanoglu and A. Ergul, “AFLP Analysis of Genetic Diversity in Low Chill Requiring Walnut (Juglans regia L.) Genotypes from Hatay, Turkey,” Scientia Horticulturae, Vol. 111, No. 4, 2007, pp. 394-398. http://dx.doi.org/10.1016/j.scienta.2006.11.006
- G. S. Dangl, K. Woeste, M. K. Aradhya, A. Koehmstedt, C. Simon, D. Potter, C. A. Leslie and G. McGranahan, “Characterization of 14 Microsatellite Markers for Genetic Analysis and Cultivar Identification of Walnut,” Journal of American Society for Horticultural Science, Vol. 130, No. 3, 2005, pp. 348-354.
- I. Foroni, R. Rao, K. Woeste and M. Gattitelli, “Characterisation of Juglans regia L. with SSR Markers and Evaluation of Genetic Relationships among Cultivars and the ‘Sorrento’ Landrace,” The Journal of Horticultural Science and Biotechnology, Vol. 80, No. 1, 2005, pp. 49-53.
- P. Pollegioni, K. Woeste, A. Major, G. Scarascia Mugnozza and M. E. Malvolti, “Characterization of Juglans nigra L., Juglans regia L. and Juglans x Intermedia (Carr) by SSR Markers: A Case Study in Italy,” Silvae Genetica, Vol. 58, No. 1-2, 2009, pp. 68-78.
- F. C. Yeh and T. J. B. Boyle, “Population Genetic Analysis of Codominant and Dominant Markers and Quantitative Traits,” Belgium Journal of Botany, Vol. 129, 1997, pp. 157-163.
- M. N. Cortinas, I. Barnes, B. D. Wingfield and M. J. Wingfield, “Polymorphic Microsatellite Markers for the Eucalyptus Fungal Pathogen Colletogloeopsis zuluensis,” Molecular Ecology Notes, Vol. 6, No. 3, 2006 pp. 780- 783. http://dx.doi.org/10.1111/j.1471-8286.2006.01342.x
- A. Forlani, B. Crestanello, S. Mantovani, B. Livoreil, L. Zane, G. Bertorelle and L. Congiu, “Identification and Characterization of Microsatellite Markers in Hermann’s Tortoise (Testudo hermanni, Testudinidae),” Molecular Ecology Notes, Vol. 5, No. 2, 2005, pp. 228-230. http://dx.doi.org/10.1111/j.1471-8286.2005.00890.x
- A. Gallini, L. Zane and P. M. Bisol, “Islolation and Characterization of Microsatellites in Zosterisessor ophiocephalus (Perciformes, Gobiidae),” Molecular Ecology Notes, Vol. 5, No. 1, 2005 pp. 24-26. http://dx.doi.org/10.1111/j.1471-8286.2004.00817.x
- H. W. Huang, Z. P. Cheng, Z. H. Zhang and Y. Wang, “History of Cultivation and Trends in China,” In: D. R. Layne, Ed., The Peach: Botany, Production and Uses (Chapter 2), CABI, Wallingford, Oxfordshire, 2008, pp. 37-60.
- R. Fatahi, A. Ebrahimi and Z. Zamani, “Characterization of Some Iranians and Foreign Walnut Genotypes Using Morphological Traits and RAPDs Markers,” Horticulture, Environment, and Biotechnology, Vol. 51, No. 1, 2010, pp. 51-60.
- W. Powell, G. C. Machray and J. Provan, “Polymorphism Revealed by Simple Sequence Repeats,” Trends in Plant Science, Vol. 1, No. 7, 1996, pp. 215-222. http://dx.doi.org/10.1016/1360-1385(96)86898-1
- L. C. Da Maia, D. A. Palmieri, V. Q. de Souza, M. M. Kopp, F. I. de Carvalho and A. Costa de Oliveira, “SSR Locator: Tool for Simple Sequence Repeat Discovery Integrated with Primer Design and PCR Simulation,” International Journal of Plant Genomics, Vol. 2008, 2008, p. 2696.
- W. R. Rice, “Analyzing Tables of Statistical Tests,” Evolution, Vol. 43, No. 1, 1989, pp. 223-225. http://dx.doi.org/10.2307/2409177
- T. C. Marshall, J. Slate, L. E. B. Kruuk and J. M. Pemberton, “Statistical Confidence for Likelihood-Based Paternity Inference in Natural Population,” Molecular Ecology, Vol. 7, No. 5, 1998, pp. 639-655. http://dx.doi.org/10.1046/j.1365-294x.1998.00374.x
- P. Pollegioni, K. Woeste, G. Scarascia Mugnozza and M. E. Malvolti, “Retrospective Identification of Hybridogenic Walnut Plants by SSR Fingerprinting and Parentage Analysis,” Molecular Breeding, Vol. 24, No. 4, 2009, pp. 321-335. http://dx.doi.org/10.1007/s11032-009-9294-7
- A. Ross-Davis and K. E. Woeste, “Microsatellite Markers for Juglans cinerea L. and Their Utility in Other Juglandaceae Species,” Conservation Genetics, Vol. 9, No. 2, 2008, pp. 465-469. http://dx.doi.org/10.1007/s10592-007-9337-8
- J. Slate, T. C. Marshall and J. M. Pemberton, “A Retrospective Assessment of the Accuracy of the Paternity Inference Program CERVUS,” Molecular Ecology, Vol. 9, No. 6, 2000, pp. 801-808. http://dx.doi.org/10.1046/j.1365-294x.2000.00930.x
- E. R. Victory, J. C. Glaubitz, O. E. Rhodes Jr. and K. E. Woeste, “Genetic Homogeneity in Juglans nigra (JUGLANDACEAE) at Nuclear Microsatellites,” American Journal of Botany, Vol. 93, No. 1, 2006, pp. 118-126. http://dx.doi.org/10.3732/ajb.93.1.118
- L. Hibrand-Saint Oyant, L. Crespel, S. Rajapakse, L. Zhang and F. Foucher, “Genetic Linkage Maps of Rose Constructed with New Microsatellite Markers and Locating QTL Controlling Flowering Traits,” Tree Genetics and Genomes, Vol. 4, No. 1, 2008, pp. 11-23. http://dx.doi.org/10.1007/s11295-007-0084-2
NOTES
*Corresponding author.

